Clustering and Kernel Density Estimation for Assessment of Measurable Residual Disease by Flow Cytometry
Abstract
1. Introduction
2. Experimental Section
2.1. Study Material
2.2. Study Methods
- (a)
- From a diagnostic AML sample, clustering is used on MFC data to obtain cell clusters.
- (b)
- Suspicious cell cluster(s) are then identified on the basis of their aberrant immunophenotypic profile. These are usually CD45 low, with low SSC [13], and are often CD34-positive and form in most cases a unique cluster of >10% of total cells. However, given the important heterogeneity of AML, a suspicious cell cluster should always be identified by experienced cytometrists on the basis of their scientific knowledge of the disease.
- (c)
- Once suspicious cell clusters are identified, “Cloud(s)” are created. A “Cloud” is created by the Boolean intersection of the contour gates of 45 bi-parametric plots (each parameter vs. each parameter, using logicle transformation for fluorescence parameters and linear transformation for FSC and SSC).
- (d)
- Once a Cloud is created at diagnosis, its Abnormality Ratio (AR) can be calculated. The AR is calculated as follows: AR = ((Cloud cells/total cells) of patient sample)/((Cloud cells/total cells) of control group sample). Note: If # Cloud cells = 0, use 1.
- (e)
- The Cloud with the highest AR, formed by a cell cluster of at least 1 × 104 cells (arbitrary value) at diagnosis will define a “Leukemic Cloud” or “L-Cloud”. Of note, if multiple Clouds of at least 5 × 103 cells have an AR of >1000 (arbitrary value), all should be considered as L-Clouds. The Cloud with the highest AR and at least 1 × 104 cells will be considered the “the major L-Cloud”. MRD assessment at follow-up will be done through the AR calculation at follow-up of L-Clouds.
2.2.1. Endpoint 1: Theoretical Evaluation of the AR/L-Cloud Concept
Global Evaluation of the AR/L-Cloud Concept
- Obtaining AML cells
- Simulation of MRD samples
- Evaluation procedure
- L-Clouds were created using the AML-FCS files.
- Performance characteristics (specificity and sensitivity) of the different L-Clouds were calculated using the AML-FCS files and the NORM-FCS file.
- The AR was calculated for each of the 36 simulated follow-up data and compared to the expected theoretical results for each patient.
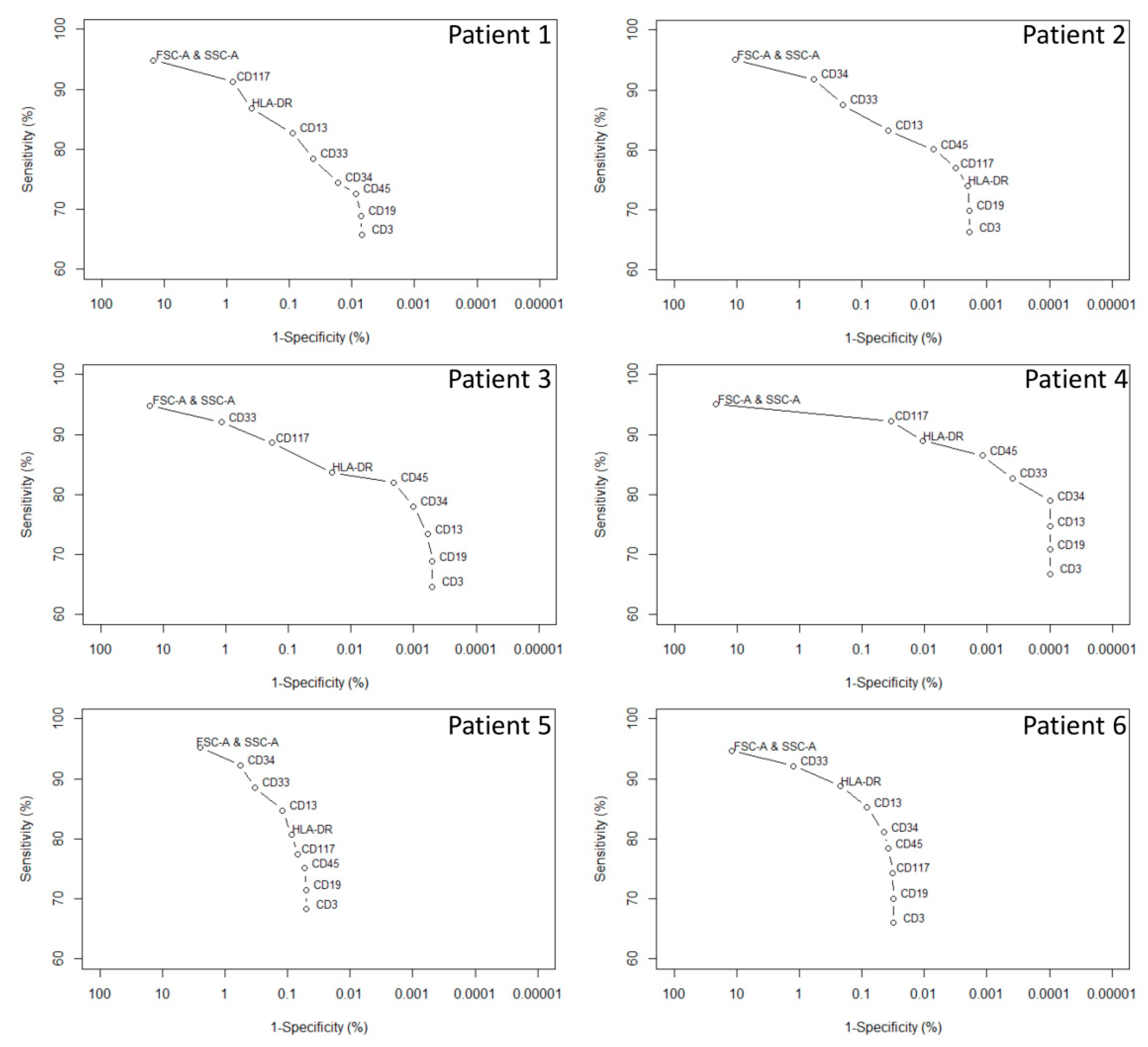
Evaluation of the Influence of Each Measured Parameter on the Intrinsic Performance of the L-Cloud
- Evaluation procedure
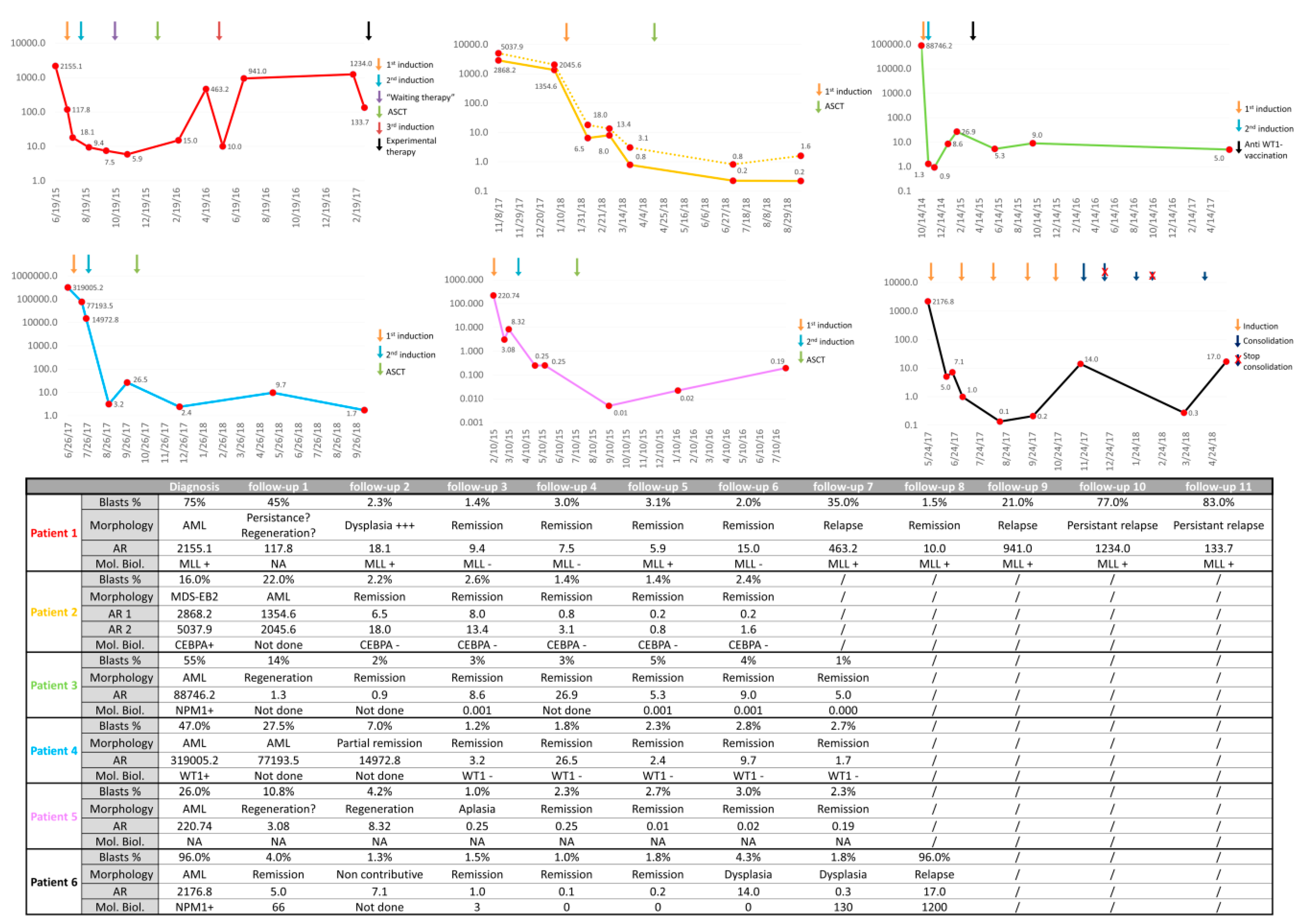
2.2.2. Endpoint 2: Clinical Evaluation of the AR/L-Cloud Concept
3. Results
4. Discussion
4.1. Limitations of This Study
4.2. Perspectives of This Study
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Adan, A.; Alizada, G.; Kiraz, Y.; Baran, Y.; Nalbant, A. Flow cytometry: Basic principles and applications. Crit. Rev. Biotechnol. 2016, 37, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Da Costa, E.S.; Pedreira, C.E.; Barrena, S.; Lécrevisse, Q.; Flores-Montero, J.; Quijano, S.; Almeida, J.; Macias, M.D.C.G.; Böttcher, S.; Van Dongen, J.J.; et al. Automated pattern-guided principal component analysis vs expert-based immunophenotypic classification of B-cell chronic lymphoproliferative disorders: A step forward in the standardization of clinical immunophenotyping. Leukemia 2010, 24, 1927–1933. [Google Scholar] [CrossRef] [PubMed]
- Kalina, T.; Flores-Montero, J.; Van Der Velden, V.H.J.; Martín-Ayuso, M.; Böttcher, S.; Ritgen, M.; Almeida, J.; Lhermitte, L.; Asnafi, V.; Mendonça, A.; et al. EuroFlow standardization of flow cytometer instrument settings and immunophenotyping protocols. Leukemia 2012, 26, 1986–2010. [Google Scholar] [CrossRef]
- Johansson, U.; Bloxham, D.; Couzens, S.; Jesson, J.; Morilla, R.; Erber, W.; Macey, M. British Committee for Standards in Haematology Guidelines on the use of multicolour flow cytometry in the diagnosis of haematological neoplasms. Br. J. Haematol. 2014, 165, 455–488. [Google Scholar] [CrossRef] [PubMed]
- Lacombe, F.; Dupont, B.; Lechevalier, N.; Vial, J.-P.; Pigneux, A.; Bene, M.C. New Concepts of Flow Cytometry Analysis in Oncohematology: Application to Diagnosis and Follow up (Minimal Residual Disease) in AML, ALL and MDS. Blood 2017, 130 (Suppl. 1). [Google Scholar]
- Dohner, H.; Weisdorf, D.J.; Bloomfield, C.D. Acute Myeloid Leukemia. N. Engl. J. Med. 2015, 373, 1136–1152. [Google Scholar] [CrossRef]
- Döhner, H.; Estey, E.; Grimwade, D.; Amadori, S.; Appelbaum, F.R.; Büchner, T.; Dombret, H.; Ebert, B.L.; Fenaux, P.; Larson, R.A.; et al. Diagnosis and management of AML in adults: 2017 ELN recommendations from an international expert panel. Blood 2017, 129, 424–447. [Google Scholar] [CrossRef]
- Chen, X.; Wood, B.L. Monitoring minimal residual disease in acute leukemia: Technical challenges and interpretive complexities. Blood Rev. 2017, 31, 63–75. [Google Scholar] [CrossRef]
- Hourigan, C.; Gale, R.P.; Gormley, N.J.; Ossenkoppele, G.J.; Walter, R.B. Measurable residual disease testing in acute myeloid leukaemia. Leukemia 2017, 31, 1482–1490. [Google Scholar] [CrossRef]
- Ossenkoppele, G.; Schuurhuis, G.J. MRD in AML: Does it already guide therapy decision-making? Hematology 2016, 2016, 356–365. [Google Scholar] [CrossRef]
- Schuurhuis, G.J.; Heuser, M.; Freeman, S.; Béné, M.C.; Lococo, F.; Cloos, J.; Grimwade, D.; Haferlach, T.; Hills, R.K.; Hourigan, C.; et al. Minimal/measurable residual disease in AML: A consensus document from the European LeukemiaNet MRD Working Party. Blood 2018, 131, 1275–1291. [Google Scholar] [CrossRef] [PubMed]
- Paietta, E. Consensus on MRD in AML? Blood 2018, 131, 1265–1266. [Google Scholar] [CrossRef] [PubMed]
- Arnoulet, C.; Béné, M.C.; Durrieu, F.; Feuillard, J.; Fossat, C.; Husson, B.; Jouault, H.; Maynadié, M.; Lacombe, F. Four- and five-color flow cytometry analysis of leukocyte differentiation pathways in normal bone marrow: A reference document based on a systematic approach by the GTLLF and GEIL. Cytom. Part B Clin. Cytom. 2009, 9999. [Google Scholar] [CrossRef] [PubMed]
- Feller, N.; Van Der Velden, V.H.J.; A Brooimans, R.; Boeckx, N.; Preijers, F.; Kelder, A.; De Greef, I.; Westra, G.; Marvelde, J.G.T.; Aerts, P.; et al. Defining consensus leukemia-associated immunophenotypes for detection of minimal residual disease in acute myeloid leukemia in a multicenter setting. Blood Cancer J. 2013, 3, e129. [Google Scholar] [CrossRef]
- Lacombe, F.; Dupont, B.; Lechevalier, N.; Vial, J.P.; Béné, M.C. Innovation in Flow Cytometry Analysis: A New Paradigm Delineating Normal or Diseased Bone Marrow Subsets Through Machine Learning. HemaSphere 2019, 3, e173. [Google Scholar] [CrossRef]
- Mosna, F.; Capelli, D.; Gottardi, M. Minimal Residual Disease in Acute Myeloid Leukemia: Still a Work in Progress? J. Clin. Med. 2017, 6, 57. [Google Scholar] [CrossRef]
- Böttcher, S.; Van Der Velden, V.H.J.; Villamor, N.; Ritgen, M.; Flores-Montero, J.; Escobar, H.M.; Kalina, T.; Brüggemann, M.; Grigore, G.; Martin-Ayuso, M.; et al. Lot-to-lot stability of antibody reagents for flow cytometry. J. Immunol. Methods 2017, 112294. [Google Scholar] [CrossRef] [PubMed]
- Panteghini, M. Implementation of standardization in clinical practice: Not always an easy task. Clin. Chem. Lab. Med. 2012, 50. [Google Scholar] [CrossRef]
- Langebrake, C.; Brinkmann, I.; Teigler-Schlegel, A.; Creutzig, U.; Griesinger, F.; Puhlmann, U.; Reinhardt, D. Immunophenotypic differences between diagnosis and relapse in childhood AML: Implications for MRD monitoring. Cytom. Part B: Clin. Cytom. 2004, 63, 1–9. [Google Scholar] [CrossRef]
- Zeijlemaker, W.; Gratama, J.W.; Schuurhuis, G.J. Tumor heterogeneity makes AML a “moving target” for detection of residual disease. Cytom. Part B Clin Cytom. 2014, 86, 3–14. [Google Scholar] [CrossRef]
- Duetz, C.; Bachas, C.; Westers, T.M.; Van De Loosdrecht, A.A. Computational analysis of flow cytometry data in hematological malignancies. Curr. Opin. Oncol. 2020, 32, 162–169. [Google Scholar] [CrossRef] [PubMed]
- Pedreira, C.; Da Costa, E.S.; Lecrevise, Q.; Grigore, G.; Fluxa, R.; Verde, J.; Hernandez, J.; Van Dongen, J.J.; Orfao, A. From big flow cytometry datasets to smart diagnostic strategies: The EuroFlow approach. J. Immunol. Methods 2019, 475, 112631. [Google Scholar] [CrossRef] [PubMed]
- Saeys, Y.; Van Gassen, S.; Lambrecht, B.N. Computational flow cytometry: Helping to make sense of high-dimensional immunology data. Nat. Rev. Immunol. 2016, 16, 449–462. [Google Scholar] [CrossRef] [PubMed]


| Patient 1 | Patient 2 | Patient 3 | Patient 4 | Patient 5 | Patient 6 | |
|---|---|---|---|---|---|---|
| Age at diagnosis | 47 years | 66 years | 65 years | 59 years | 62 years | 71 years |
| Sex | Male | Male | Male | Female | Male | Female |
| WBC/µL at diagnosis | <100,000 | <100,000 | <100,000 | <100,000 | <100,000 | >100,000 |
| WHO classification (2016) | AML–NOS (Acute myelomonocytic leukemia) | AML with MDS-related changes | NPM1-mutated AML | AML–NOS (AML with maturation) | AML with MDS-related changes | Therapy-related AML |
| 2017 ELN risk classification | Adverse | Adverse | Favorable | Adverse | Intermediate | Favorable |
| Initial chemotherapy | aracytin–idarubicin–lenalidomide | aracytin–daunorubicin–selinexor | aracytin–daunorubicin | aracytin–idarubicin | aracytin–idarubicin | cytarabine–daunoubicin–midostaurin |
| Consolidation therapy | ASCT | ASCT | Anti-WT1 vaccination | ASCT | ASCT | Aracytin–midostaurin |
| Cytogenetic at diagnosis | t(6;11), t(11,14), partial tetrazomy of 11q | Hyperdiploidy–trisomy 8 | Normal karyotype | Hyperdiploidy–t(2;12), trisomy 4 | Normal karyotype | Normal karyotype |
| Mutations at diagnosis | FLT3 and partial MLL tandem duplication | CEBPA–ASLX1–STAG2 | NPM1–WT1 | WT1–ASLX1–GATA2 | / | NPM1–FLT3 |
| Patient 1 | Patient 2 | Patient 3 | Patient 4 | Patient 5 | Patient 6 | ||
|---|---|---|---|---|---|---|---|
| L-Cloud sensitivity ‡ | 65.75% | 66.25% | 64.63% | 66.76% | 68.28% | 65.99% | |
| L-Cloud specificity § | 99.9930% | 99.9981% | 99.9995% | 99.9999% | 99.9502% | 99.9690% | |
| 1–(L-Cloud specificity) | 0.0070% | 0.0019% | 0.0005% | 0.0001% | 0.0498% | 0.0310% | |
| Number of cells retrieved into the L-Cloud * | Control group | 70 | 19 | 5 | 1 | 498 | 310 |
| 0.5% MRD simulation | 3234 | 3228 | 3791 | 3567 | 3948 | 4037 | |
| 0.1% MRD simulation | 707 | 680 | 773 | 721 | 1185 | 1066 | |
| 0.05% MRD simulation | 391 | 345 | 385 | 361 | 841 | 685 | |
| 0.01% MRD simulation | 122 | 85 | 81 | 83 | 566 | 373 | |
| 0.005% MRD simulation | 101 | 48 | 41 | 35 | 531 | 344 | |
| 0.001% MRD simulation | 78 | 26 | 12 | 6 | 506 | 317 | |
| Abnormality Ratio (AR) † | 0.5% MRD simulation | 45.97 | 169.05 | 754.43 | 3549.25 | 7.89 | 12.96 |
| 0.1% MRD simulation | 10.09 | 35.75 | 154.45 | 720.28 | 2.38 | 3.44 | |
| 0.05% MRD simulation | 5.58 | 18.15 | 76.96 | 360.82 | 1.69 | 2.21 | |
| 0.01% MRD simulation | 1.74 | 4.47 | 16.20 | 82.99 | 1.14 | 1.20 | |
| 0.005% MRD simulation | 1.44 | 2.53 | 8.20 | 35.00 | 1.07 | 1.11 | |
| 0.001% MRD simulation | 1.11 | 1.37 | 2.40 | 6.00 | 1.02 | 1.02 |
| Flow Cytometry MRD Approach | LAIP | DfN | AR/L-Cloud |
|---|---|---|---|
| Type of approach | Diagnostic-based | Reference-based | Diagnostic-based and reference-based |
| Foundations | Expert knowledge of malignancy (LAIP knowledge) | Knowledge of normality | Unsupervised data analysis (clustering) and comparison to reference samples $ |
| Subjectivity | Manual gating | Methodology (unknown *) | Cloud modeling (clustering and contour gating algorithms) |
| Standardization | Impossible | Depends upon methodology | Possible $ |
| Automation | Impossible | Depends upon methodology | Possible $ |
| Reference samples | Preferable | Required | Required |
| Diagnostic sample | Required | Not required | Required |
| Data analysis tools | Basics | Depends upon methodology | Advanced |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jacqmin, H.; Chatelain, B.; Louveaux, Q.; Jacqmin, P.; Dogné, J.-M.; Graux, C.; Mullier, F. Clustering and Kernel Density Estimation for Assessment of Measurable Residual Disease by Flow Cytometry. Diagnostics 2020, 10, 317. https://doi.org/10.3390/diagnostics10050317
Jacqmin H, Chatelain B, Louveaux Q, Jacqmin P, Dogné J-M, Graux C, Mullier F. Clustering and Kernel Density Estimation for Assessment of Measurable Residual Disease by Flow Cytometry. Diagnostics. 2020; 10(5):317. https://doi.org/10.3390/diagnostics10050317
Chicago/Turabian StyleJacqmin, Hugues, Bernard Chatelain, Quentin Louveaux, Philippe Jacqmin, Jean-Michel Dogné, Carlos Graux, and François Mullier. 2020. "Clustering and Kernel Density Estimation for Assessment of Measurable Residual Disease by Flow Cytometry" Diagnostics 10, no. 5: 317. https://doi.org/10.3390/diagnostics10050317
APA StyleJacqmin, H., Chatelain, B., Louveaux, Q., Jacqmin, P., Dogné, J.-M., Graux, C., & Mullier, F. (2020). Clustering and Kernel Density Estimation for Assessment of Measurable Residual Disease by Flow Cytometry. Diagnostics, 10(5), 317. https://doi.org/10.3390/diagnostics10050317





