Liquid Biopsy as Novel Tool in Precision Medicine: Origins, Properties, Identification and Clinical Perspective of Cancer’s Biomarkers
Abstract
1. Introduction
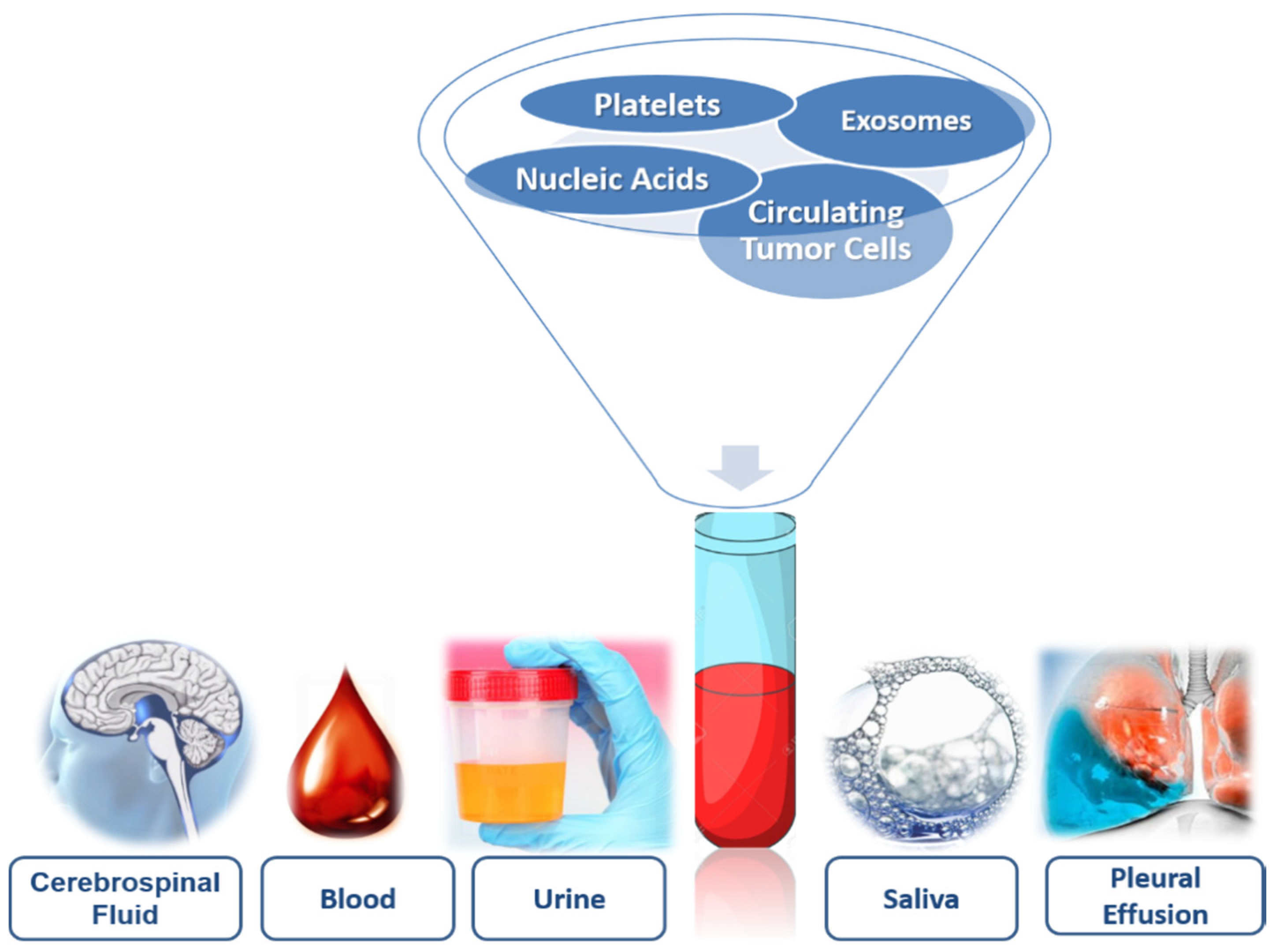
2. Biofluids: Storage of Biomarkers
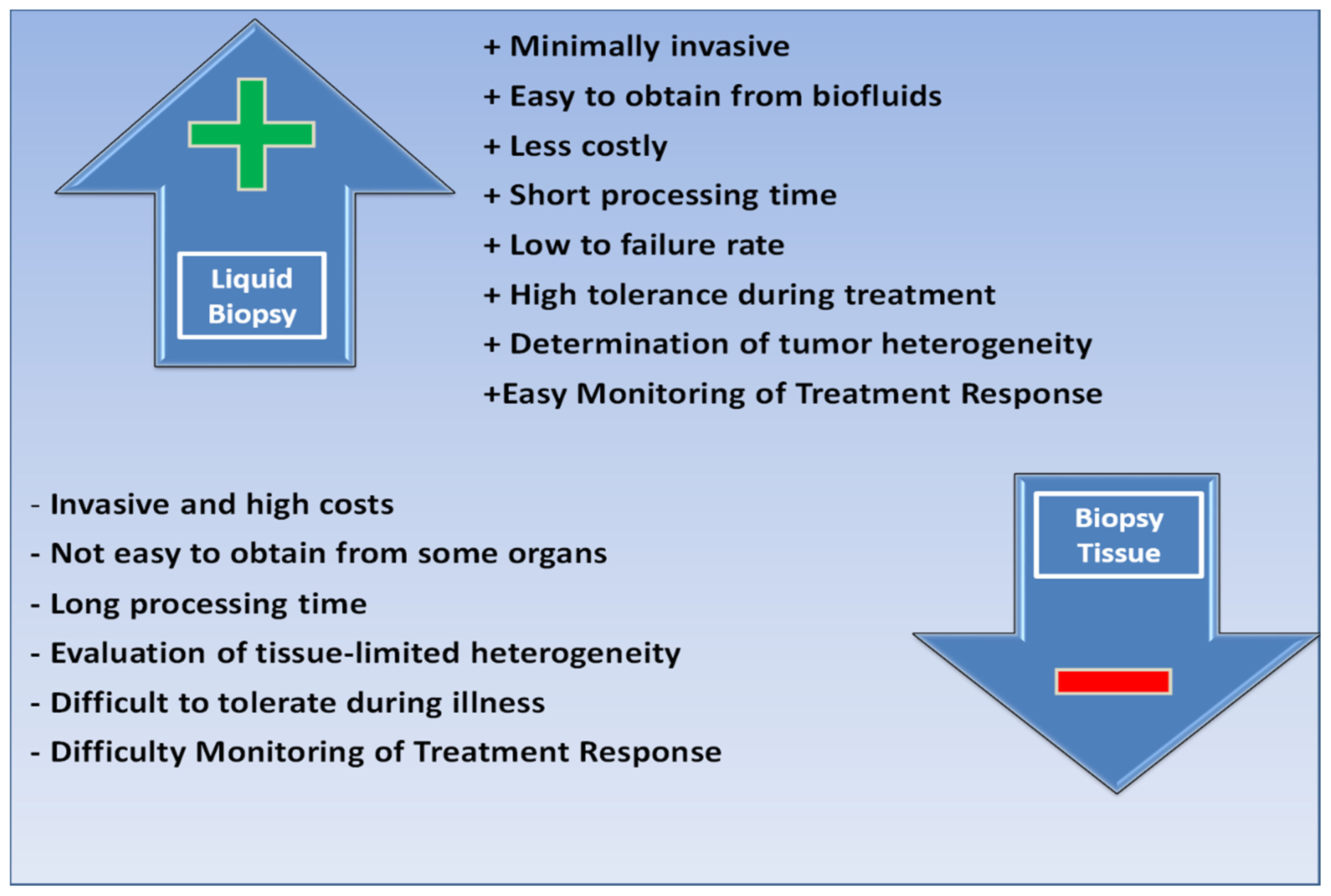
3. Comparison of Tissue Biopsy and Liquid Biopsy
4. Liquid Biopsy Components
4.1. Circulating Tumor Cells (CTCs)
4.1.1. General Characteristics
4.1.2. Origin: Epithelial to Mesenchymal Transition (EMT)
4.1.3. Methods: Isolation, Identification and Culture
4.1.4. Potential Application of Circulating Tumor Cells (CTCs) in Medical Oncology
4.2. Cell-Free Circulating DNA (cfDNA)
4.2.1. Circulating Tumor DNA (ctDNA)
4.2.2. Methodologies for Detection of ctDNA
4.2.3. Potential Application of ctDNA in Medical Oncology
4.3. Cell-Free Circulating Free RNA (cfRNA)
4.3.1. Circulating Tumor Messenger RNA
4.3.2. Circulating Tumor MicroRNAs
4.4. Exosomes
Potential Application of Tumor-Derived Exosomes in Medical Oncology
4.5. Microvesicles
4.6. Platelet
5. Applications of Liquid Biopsy’s Biomarkers
6. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Venesio, T.; Siravegna, G.; Bardelli, A.; Sapino, A. Liquid biopsies for monitoring temporal genomic heterogeneity in breast and colon cancers. Pathobiology 2018, 85, 146–154. [Google Scholar] [CrossRef] [PubMed]
- Hanahan, D.; Weinberg, R.A. Hallmarks of cancer: The next generation. Cell 2011, 144, 646–674. [Google Scholar] [CrossRef] [PubMed]
- Jameson, J.L.; Longo, D.L. Precision medicine—Personalized, problematic, and promising. N. Engl. J. Med. 2015, 372, 2229–2234. [Google Scholar] [CrossRef] [PubMed]
- Imamura, T.; Komatsu, S.; Ichikawa, D.; Kawaguchi, T.; Miyamae, M.; Okajima, W.; Ohashi, T.; Arita, T.; Konishi, H.; Shiozaki, A.; et al. Liquid biopsy in patients with pancreatic cancer: Circulating tumor cells and cell-free nucleic acids. World J. Gastroenterol. 2016, 22, 5627. [Google Scholar] [CrossRef]
- Bardelli, A.; Pantel, K. Liquid biopsies, what we do not know (yet). Cancer Cell 2017, 31, 172–179. [Google Scholar] [CrossRef]
- Pantel, K.; Alix-Panabières, C. Circulating tumour cells in cancer patients: Challenges and perspectives. Trends Mol. Med. 2010, 16, 398–406. [Google Scholar] [CrossRef]
- Siravegna, G.; Marsoni, S.; Siena, S.; Bardelli, A. Integrating liquid biopsies into the management of cancer. Nat. Rev. Clin. Oncol. 2017, 14, 531–548. [Google Scholar] [CrossRef]
- Tan, G.H.; Nason, G.; Ajib, K.; Woon, D.T.S.; Herrera-Caceres, J.; Alhunaidi, O.; Perlis, N. marter screening for prostate cancer. World J. Urol. 2019, 37, 991–999. [Google Scholar] [CrossRef]
- Gahan, P.; Stroun, M. The biology of circulating nucleic acids in plasma and serum (CNAPS). Extracell. Nucleic Acids 2010, 1, 167–189. [Google Scholar]
- Porto-Mascarenhas, E.C.; Assad, D.X.; Chardin, H.; Gozal, D.; Canto, G.D.L.; Acevedo, A.C.; Guerra, E.N.S. Salivary biomarkers in the diagnosis of breast cancer: A review. Crit. Rev. Oncol. Hematol. 2017, 110, 62–73. [Google Scholar] [CrossRef]
- Lawrence, H.P. Salivary markers of systemic disease: Noninvasive diagnosis of disease and monitoring of general health. J. Can. Dent. Assoc. 2002, 68, 170–174. [Google Scholar] [PubMed]
- Zhang, L.; Farrell, J.J.; Zhou, H.; Elashoff, D.; Akin, D.; Park, N.H.; Chia, D.; Wong, D.T. Salivary transcriptomic biomarkers for detection of resectable pancreatic cancer. Gastroenterology 2010, 138, 949–957. [Google Scholar] [CrossRef] [PubMed]
- Gao, S.; Chen, L.-Y.; Wang, P.; Liu, L.-M.; Chen, Z. MicroRNA expression in salivary supernatant of patients with pancreatic cancer and its relationship with ZHENG. Biomed Res. Int. 2014, 756347. [Google Scholar] [CrossRef] [PubMed]
- Humeau, M.; Vignolle-Vidoni, A.; Sicard, F.; Martins, F.; Bournet, B.; Buscail, L.; Torrisani, J.; Cordelier, P. Salivary microRNA in pancreatic cancer patients. PLoS ONE 2015, 10, e0130996. [Google Scholar] [CrossRef] [PubMed]
- Lau, C.; Kim, Y.; Chia, D.; Spielmann, N.; Eibl, G.; Elashoff, D.; Wei, F.; Lin, Y.-L.; Moro, A.; Grogan, T.; et al. Role of pancreatic cancer-derived exosomes in salivary biomarker development. J. Biol. Chem. 2013, 288, 26888–26897. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Yang, T.; Lin, J. Spectral analysis of human saliva for detection of lung cancer using surface-enhanced Raman spectroscopy. J. Biomed. Opt. 2012, 17, 037003. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Xiao, H.; Zhou, H.; Santiago, S.; Lee, J.M.; Garon, E.B.; Yang, J.; Brinkmann, O.; Yan, X.; Akin, D.; et al. Development of transcriptomic biomarker signature in human saliva to detect lung cancer. Cell. Mol. Life Sci. 2012, 69, 3341–3350. [Google Scholar] [CrossRef]
- Wu, Z.-Z.; Wang, J.-G.; Zhang, X.-L. Diagnostic model of saliva protein finger print analysis of patients with gastric cancer. World J. Gastroenterol. 2009, 15, 865–870. [Google Scholar] [CrossRef]
- Wang, X.; Kaczor-Urbanowicz, K.E.; Wong, D.T. Salivary biomarkers in cancer detection. Med. Oncol. 2017, 34, 7. [Google Scholar] [CrossRef]
- Buyse, M.; Sargent, D.J.; Grothey, A.; Matheson, A.; De Gramont, A. Biomarkers and surrogate end points—The challenge of statistical validation. Nat. Rev. Clin. Oncol. 2010, 7, 309–317. [Google Scholar] [CrossRef]
- Heitzer, E.; Haque, I.S.; Roberts, C.E.; Speicher, M.R. Current and future perspectives of liquid biopsies in genomics-driven oncology. Nat. Rev. Genet. 2018, 20, 71–88. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Chang, S.; Li, G.; Sun, Y. Application of liquid biopsy in precision medicine: Opportunities and challenges. Front. Med. 2017, 11, 522–527. [Google Scholar] [CrossRef] [PubMed]
- Maltoni, R.; Gallerani, G.; Fici, P.; Rocca, A.; Fabbri, F. CTCs in early breast cancer: A path worth taking. Cancer Lett. 2016, 376, 205–210. [Google Scholar] [CrossRef] [PubMed]
- Barriere, G.; Fici, P.; Gallerani, G.; Fabbri, F.; Zoli, W.; Rigaud, M. Circulating tumor cells and epithelial, mesenchymal and stemness markers: Characterization of cell subpopulations. Ann. Transl. Med. 2014, 2, 109. [Google Scholar]
- Nagrath, S.; Sequist, L.V.; Maheswaran, S.; Bell, D.W.; Irimia, D.; Ulkus, L.; Smith, M.R.; Kwak, E.L.; Digumarthy, S.; Muzikansky, A.; et al. Isolation of rare circulating tumour cells in cancer patients by microchip technology. Nature 2007, 450, 1235–1239. [Google Scholar] [CrossRef]
- Lim, M.; Kim, C.-J.; Sunkara, V.; Kim, M.-H.; Cho, Y.-K. Liquid biopsy in lung cancer: Clinical applications of circulating biomarkers (CTCs and ctDNA). Micromachines (Basel) 2018, 9, E100. [Google Scholar] [CrossRef]
- Das, V.; Bhattacharya, S.; Chikkaputtaiah, C.; Hazra, S.; Pal, M. The basics of epithelial–mesenchymal transition (EMT): A study from a structure, dynamics, and functional perspective. J. Cell. Physiol. 2019, 234, 14535–14555. [Google Scholar] [CrossRef]
- Lourenço, A.R.; Coffer, P.J. SOX4: Joining the master regulators of epithelial-to-mesenchymal transition? Trends Cancer 2017, 3, 571–582. [Google Scholar] [CrossRef]
- Kalluri, R.; Weinberg, R.A. The basics of epithelial-mesenchymal transition. J. Clin. Investig. 2010, 120, 1420–1428. [Google Scholar] [CrossRef]
- Tiwari, N.; Gheldof, A.; Tatari, M.; Christofori, G. EMT as the ultimate survival mechanism of cancer cells. Semin Cancer Biol. 2012, 22, 194–207. [Google Scholar] [CrossRef]
- Armstrong, A.J.; Marengo, M.S.; Oltean, S.; Kemeny, G.; Bitting, R.L.; Turnbull, J.D.; Herold, C.I.; Marcom, P.K.; George, D.; Garcia-Blanco, M. Circulating tumor cells from patients with advanced prostate and breast cancer display both epithelial and mesenchymal markers. Mol. Cancer Res. 2011, 9, 997–1007. [Google Scholar] [CrossRef] [PubMed]
- Maltoni, R.; Casadio, V.; Ravaioli, S.; Foca, F.; Tumedei, M.M.; Salvi, S.; Martignano, F.; Calistri, D.; Rocca, A.; Schirone, A.; et al. Cell-free DNA detected by “liquid biopsy” as a potential prognostic biomarker in early breast cancer. Oncotarget 2017, 8, 16642–16649. [Google Scholar] [CrossRef] [PubMed]
- Adams, D.L.; Alpaugh, R.K.; Tsai, S.; Tang, C.-M.; Stefansson, S. Multi-Phenotypic subtyping of circulating tumor cells using sequential fluorescent quenching and restaining. Sci. Rep. 2016, 6, 33488. [Google Scholar] [CrossRef] [PubMed]
- Swennenhuis, J.F.; Tibbe, A.G.; Levink, R.; Sipkema, R.C.; Terstappen, L.W. Characterization of circulating tumor cells by fluorescence in situ hybridization. Cytom. A 2009, 75, 520–527. [Google Scholar] [CrossRef]
- Alix-Panabières, C.; Pantel, K. Circulating tumor cells: Liquid biopsy of cancer. Clin. Chem. 2013, 59, 110–118. [Google Scholar] [CrossRef]
- Olivier, C.G.; Carballido, J.R. Circulating tumor cells: Isolation, quantification, and relevance of their translation into clinical practice. Actas Urol. Esp. 2010, 34, 3–5. [Google Scholar] [CrossRef]
- Pantel, K.; Speicher, M. The biology of circulating tumor cells. Oncogene 2016, 35, 1216–1224. [Google Scholar] [CrossRef]
- Rossi, G.; Mu, Z.; Rademaker, A.W.; Austin, L.K.; Strickland, K.S.; Costa, R.L.B.; Nagy, R.J.; Zagonel, V.; Taxter, T.J.; Behdad, A.; et al. Cell-free DNA and circulating tumor cells: Comprehensive liquid biopsy analysis in advanced breast cancer. Clin. Cancer Res. 2018, 24, 560–568. [Google Scholar] [CrossRef]
- Wang, L.; Balasubramanian, P.; Chen, A.P.; Kummar, S.; Evrard, Y.A.; Kinders, R.J. Promise and limits of the CellSearch platform for evaluating pharmacodynamics in circulating tumor cells. Semin. Oncol. 2016, 43, 464–475. [Google Scholar] [CrossRef]
- Cieślikowski, W.A.; Budna-Tukan, J.; Świerczewska, M.; Ida, A.; Hrab, M.; Jankowiak, A.; Mazel, M.; Nowicki, M.; Milecki, P.; Pantel, K.; et al. Circulating Tumor Cells as a Marker of Disseminated Disease in Patients with Newly Diagnosed High-Risk Prostate Cancer. Cancers 2020, 12, 160. [Google Scholar] [CrossRef]
- Duan, G.-C.; Zhang, X.-P.; Wang, H.-E.; Wang, Z.-K.; Zhang, H.; Yu, L.; Xue, W.-F.; Xin, Z.-F.; Hu, Z.-H.; Zhao, Q.-T. Circulating Tumor Cells as a Screening and Diagnostic Marker for Early-Stage Non-Small Cell Lung Cancer. Onco Targets Ther. 2020, 13, 1931–1939. [Google Scholar] [CrossRef] [PubMed]
- Dizdar, L.; Fluegen, G.; van Dalum, G.; Honisch, E.; Neves, R.P.; Niederacher, D.; Neubauer, H.; Fehm, T.; Rehders, A.; Krieg, A.; et al. Detection of circulating tumor cells in colorectal cancer patients using the GILUPI CellCollector: Results from a prospective, single-center study. Mol. Oncol. 2019, 13, 1548–1558. [Google Scholar] [CrossRef] [PubMed]
- Pantel, K.; Alix-Panabieres, C. Functional studies on viable circulating tumor cells. Clin. Chem. 2016, 62, 328–334. [Google Scholar] [CrossRef] [PubMed]
- Alix-Panabières, C. EPISPOT assay: Detection of viable DTCs/CTCs in solid tumor patients. Recent Results Cancer Res. 2012, 195, 69–76. [Google Scholar] [PubMed]
- Franken, A.; Driemel, C.; Behrens, B.; Meier-Stiegen, F.; Endris, V.; Stenzinger, A.; Niederacher, D.; Fischer, J.C.; Stoecklein, N.H.; Ruckhaeberle, E.; et al. Label-free enrichment and molecular characterization of viable circulating tumor cells from diagnostic leukapheresis products. Clin. Chem. 2019, 65, 549–558. [Google Scholar] [CrossRef]
- Kaigorodova, E.V.; Savelieva, O.E.; Tashireva, L.A.; Tarabanovskaya, N.A.; Simolina, E.I.; Denisov, E.V.; Slonimskaya, E.M.; Choynzonov, E.L.; Perelmuter, V.M. Heterogeneity of circulating tumor cells in neoadjuvant chemotherapy of breast cancer. Molecules 2018, 23, 727. [Google Scholar] [CrossRef]
- Gold, B.; Cankovic, M.; Furtado, L.V.; Meier, F.; Gocke, C.D. Do circulating tumor cells, exosomes, and circulating tumor nucleic acids have clinical utility? A report of the association for molecular pathology. J. Mol. Diagn. 2015, 17, 209–224. [Google Scholar] [CrossRef]
- Lv, Q.-B.; Fu, X.; Jin, H.-M.; Xu, H.-C.; Huang, Z.-Y.; Xu, H.-Z.; Chi, Y.-Z.; Wu, A.-M. The relationship between weight change and risk of hip fracture: Meta-analysis of prospective studies. Sci. Rep. 2015, 5, 16030. [Google Scholar] [CrossRef]
- Joosse, S.A.; Gorges, T.M.; Pantel, K. Biology, detection, and clinical implications of circulating tumor cells. EMBO Mol. Med. 2015, 7, 1–11. [Google Scholar] [CrossRef]
- Sorber, L.; Zwaenepoel, K.; Deschoolmeester, V.; Van Schil, P.; Van Meerbeeck, J.; Lardon, F.; Rolfo, C.; Pauwels, P. Circulating cell-free nucleic acids and platelets as a liquid biopsy in the provision of personalized therapy for lung cancer patients. Lung Cancer 2017, 107, 100–107. [Google Scholar] [CrossRef]
- Elazezy, M.; Joosse, S.A. Techniques of using circulating tumor DNA as a liquid biopsy component in cancer management. Comput. Struct. Biotechnol. J. 2018, 16, 370–378. [Google Scholar] [CrossRef] [PubMed]
- Alix-Panabières, C.; Pantel, K. Clinical applications of circulating tumor cells and circulating tumor DNA as liquid biopsy. Cancer Discov. 2016, 6, 479–491. [Google Scholar] [CrossRef] [PubMed]
- Cheng, F.; Su, L.; Qian, C. Circulating tumor DNA: A promising biomarker in the liquid biopsy of cancer. Oncotarget 2016, 7, 48832–48841. [Google Scholar] [CrossRef] [PubMed]
- García-Olmo, D.C.; Domínguez, C.; García-Arranz, M.; Anker, P.; Stroun, M.; García-Verdugo, J.M.; García-Olmo, D. Cell-free nucleic acids circulating in the plasma of colorectal cancer patients induce the oncogenic transformation of susceptible cultured cells. Cancer Res. 2010, 70, 560–567. [Google Scholar] [CrossRef] [PubMed]
- Openshaw, M.R.; Page, K.; Fernandez-Garcia, D.; Guttery, D.; Shaw, J.A. The role of ctDNA detection and the potential of the liquid biopsy for breast cancer monitoring. Expert Rev. Mol. Diagn. 2016, 16, 751–755. [Google Scholar] [CrossRef] [PubMed]
- Forshew, T.; Murtaza, M.; Parkinson, C.; Gale, D.; Tsui, D.W.; Kaper, F.; Dawson, S.-J.; Piskorz, A.M.; Jimenez-Linan, M.; Bentley, D.; et al. Noninvasive identification and monitoring of cancer mutations by targeted deep sequencing of plasma DNA. Sci. Transl. Med. 2012, 4, 136ra68. [Google Scholar] [CrossRef]
- Czeiger, D.; Shaked, G.; Eini, H.; Vered, I.; Belochitski, O.; Avriel, A.; Ariad, S.; Douvdevani, A. Measurement of circulating cell-free DNA levels by a new simple fluorescent test in patients with primary colorectal cancer. Am. J. Clin. Pathol. 2011, 135, 264–270. [Google Scholar] [CrossRef]
- Beaver, J.A.; Jelovac, D.; Balukrishna, S.; Cochran, R.L.; Croessmann, S.; Zabransky, D.J.; Wong, H.Y.; Toro, P.V.; Cidado, J.; Blair, B.G.; et al. Detection of cancer DNA in plasma of patients with early-stage breast cancer. Clin. Cancer Res. 2014, 20, 2643–2650. [Google Scholar] [CrossRef]
- Otsuji, K.; Sasaki, T.; Tanaka, A.; Kunita, A.; Ikemura, M.; Matsusaka, K.; Tada, K.; Fukayama, M.; Seto, Y. Use of droplet digital PCR for quantitative and automatic analysis of the HER2 status in breast cancer patients. Breast Cancer Res. Treat. 2017, 162, 11–18. [Google Scholar] [CrossRef]
- Escors, D.; Gato-Cañas, M.; Zuazo, M.; Arasanz, H.; García-Granda, M.J.; Vera, R.; Kochan, G. The intracellular signalosome of PD-L1 in cancer cells. Signal Transduct. Target Ther. 2018, 3, 26. [Google Scholar] [CrossRef]
- Ishiba, T.; Hoffmann, A.-C.; Usher, J.; Elshimali, Y.; Sturdevant, T.; Dang, M.; Jaimes, Y.C.C.; Tyagi, R.; Gonzalez, R.; Grino, M.; et al. Frequencies and expression levels of programmed death ligand 1 (PD-L1) in circulating tumor RNA (ctRNA) in various cancer types. Biochem. Biophys. Res. Commun. 2018, 500, 621–625. [Google Scholar] [CrossRef] [PubMed]
- Junqueira-Neto, S.; Batista, I.A.; Costa, J.L.; Melo, S.A. Liquid Biopsy beyond Circulating Tumor Cells and Cell-Free DNA. Acta Cytol. 2019, 63, 479–488. [Google Scholar] [CrossRef] [PubMed]
- Leong, S.P.; Ballesteros-Merino, C.; Jensen, S.M.; Marwitz, S.; Bifulco, C.; Fox, B.A.; Skoberne, M. Novel frontiers in detecting cancer metastasis. Clin. Exp. Metastasis 2018, 35, 403–412. [Google Scholar] [CrossRef] [PubMed]
- Chen, W.; Cai, F.; Zhang, B.; Barekati, Z.; Zhong, X.Y. The level of circulating miRNA-10b and miRNA-373 in detecting lymph node metastasis of breast cancer: Potential biomarkers. Tumor Biol. 2013, 34, 455–462. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Mao, F.; Shen, T.; Luo, Q.; Ding, Z.; Qian, L.; Huang, J. Plasma miR-145, miR-20a, miR-21 and miR-223 as novel biomarkers for screening early-stage non-small cell lung cancer. Oncol. Lett. 2017, 13, 669–676. [Google Scholar] [CrossRef] [PubMed]
- Thakur, B.K.; Zhang, H.; Becker, A.; Matei, I.; Huang, Y.; Costa-Silva, B.; Zheng, Y.; Hoshino, A.; Brazier, H.; Xiang, J.; et al. Double-stranded DNA in exosomes: A novel biomarker in cancer detection. Cell Res. 2014, 24, 766–769. [Google Scholar] [CrossRef]
- Jia, Y.; Chen, Y.; Wang, Q.; Jayasinghe, U.; Luo, X.; Wei, Q.; Wang, J.; Xiong, H.; Chen, C.; Xu, B.; et al. Exosome: Emerging biomarker in breast cancer. Oncotarget 2017, 8, 41717–41733. [Google Scholar] [CrossRef]
- Yoshioka, Y.; Kosaka, N.; Konishi, Y.; Ohta, H.; Okamoto, H.; Sonoda, H.; Nonaka, R.; Yamamoto, H.; Ishii, H.; Mori, M.; et al. Ultra-sensitive liquid biopsy of circulating extracellular vesicles using ExoScreen. Nat. Commun. 2014, 5, 3591. [Google Scholar] [CrossRef]
- Vader, P.; Breakefield, X.O.; Wood, M.J. Extracellular vesicles: Emerging targets for cancer therapy. Trends Mol. Med. 2014, 20, 385–393. [Google Scholar] [CrossRef]
- Joosse, S.A.; Pantel, K. Tumor-educated platelets as liquid biopsy in cancer patients. Cancer Cell 2015, 28, 552–554. [Google Scholar] [CrossRef]
- Gandara, D.R.; Paul, S.M.; Kowanetz, M.; Schleifman, E.; Zou, W.; Li, Y.; Rittmeyer, A.; Fehrenbacher, L.; Otto, G.; Malboeuf, C.; et al. Blood-based tumor mutational burden as a predictor of clinical benefit in non-small-cell lung cancer patients treated with atezolizumab. Nat. Med. 2018, 24, 1441–1448. [Google Scholar] [CrossRef] [PubMed]
- Vidal, J.; Muinelo, L.; Dalmases, A.; Jones, F.; Edelstein, D.; Iglesias, M.; Orrillo, M.; Abalo, A.; Rodríguez, C.; Brozos, E.; et al. Plasma ctDNA RAS mutation analysis for the diagnosis and treatment monitoring of metastatic colorectal cancer patients. Ann. Oncol. 2017, 28, 1325–1332. [Google Scholar] [CrossRef] [PubMed]
- García, J.L.; Lozano, R.; Misiewicz-Krzeminska, I.; Fernández-Mateos, J.; Krzeminski, P.; Alfonso, S.; Marcos, R.A.; García, A.; Herrero, M.; Olmos, D.; et al. A novel capillary nano-immunoassay for assessing androgen receptor splice variant 7 in plasma. Correlation with CD133 antigen expression in circulating tumor cells. A pilot study in prostate cancer patients. Clin. Transl. Oncol. 2017, 19, 1350–1357. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Ascierto, P.A.; Minor, D.; Ribas, A.; Lebbe, C.; O’Hagan, A.; Arya, N.; Guckert, M.; Schadendorf, D.; Kefford, R.F.; Grob, J.J.; et al. Phase II trial (BREAK-2) of the BRAF inhibitor dabrafenib (GSK2118436) in patients with metastatic melanoma. J. Clin. Oncol. 2013, 31, 3205–3211. [Google Scholar] [CrossRef]
- Santiago-Walker, A.; Gagnon, R.; Mazumdar, J.; Casey, M.; Long, G.V.; Schadendorf, D.; Flaherty, K.; Kefford, R.; Hauschild, A.; Hwu, P.; et al. Correlation of BRAF mutation status in circulating-free DNA and tumor and association with clinical outcome across four BRAFi and MEKi clinical trials. Clin. Cancer Res. 2016, 22, 567–574. [Google Scholar] [CrossRef]
- Tantiwetrueangdet, A.; Panvichian, R.; Wongwaisayawan, S.; Sueangoen, N.; Lertsithichai, P. Droplet digital PCR using HER2/EIF2C1 ratio for detection of HER2 amplification in breast cancer tissues. Med. Oncol. 2018, 35, 149. [Google Scholar] [CrossRef]
- Cohen, J.D.; Javed, A.A.; Thoburn, C.; Wong, F.; Tie, J.; Gibbs, P.; Schmidt, C.M.; Yip-Schneider, M.T.; Allen, P.J.; Schattner, M.; et al. Combined circulating tumor DNA and protein biomarker-based liquid biopsy for the earlier detection of pancreatic cancers. Proc. Natl. Acad. Sci. USA 2017, 114, 10202–10207. [Google Scholar] [CrossRef]
| Elements | CTCs | ctDNA/ctRNA | Exosomes |
|---|---|---|---|
| Potential to assess tumor spatial and temporal heterogeneity | No | Yes | No |
| Detection of somatic mutations, indels, alteration of the number of copies and gene fusions | Yes | Yes | Si |
| Analysis of mRNA, miRNA, IncRNA and gene variants | Yes | Yes | Yes |
| Evaluation of methylation patterns | Yes | Yes | Yes |
| Methylation Studies | Yes | No | Yes |
| Expression Studies | Yes | No | Si |
| Cell morphology and functional studies ex vivo | Yes | No | No |
| Signal co-location demonstration | Yes | No | No |
| Proteomic analysis | Yes | No | Yes |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Fernández-Lázaro, D.; García Hernández, J.L.; García, A.C.; Córdova Martínez, A.; Mielgo-Ayuso, J.; Cruz-Hernández, J.J. Liquid Biopsy as Novel Tool in Precision Medicine: Origins, Properties, Identification and Clinical Perspective of Cancer’s Biomarkers. Diagnostics 2020, 10, 215. https://doi.org/10.3390/diagnostics10040215
Fernández-Lázaro D, García Hernández JL, García AC, Córdova Martínez A, Mielgo-Ayuso J, Cruz-Hernández JJ. Liquid Biopsy as Novel Tool in Precision Medicine: Origins, Properties, Identification and Clinical Perspective of Cancer’s Biomarkers. Diagnostics. 2020; 10(4):215. https://doi.org/10.3390/diagnostics10040215
Chicago/Turabian StyleFernández-Lázaro, Diego, Juan Luis García Hernández, Alberto Caballero García, Alfredo Córdova Martínez, Juan Mielgo-Ayuso, and Juan Jesús Cruz-Hernández. 2020. "Liquid Biopsy as Novel Tool in Precision Medicine: Origins, Properties, Identification and Clinical Perspective of Cancer’s Biomarkers" Diagnostics 10, no. 4: 215. https://doi.org/10.3390/diagnostics10040215
APA StyleFernández-Lázaro, D., García Hernández, J. L., García, A. C., Córdova Martínez, A., Mielgo-Ayuso, J., & Cruz-Hernández, J. J. (2020). Liquid Biopsy as Novel Tool in Precision Medicine: Origins, Properties, Identification and Clinical Perspective of Cancer’s Biomarkers. Diagnostics, 10(4), 215. https://doi.org/10.3390/diagnostics10040215









