Universal Molecular Triggers of Stress Responses in Cyanobacterium Synechocystis
Abstract
1. Introduction
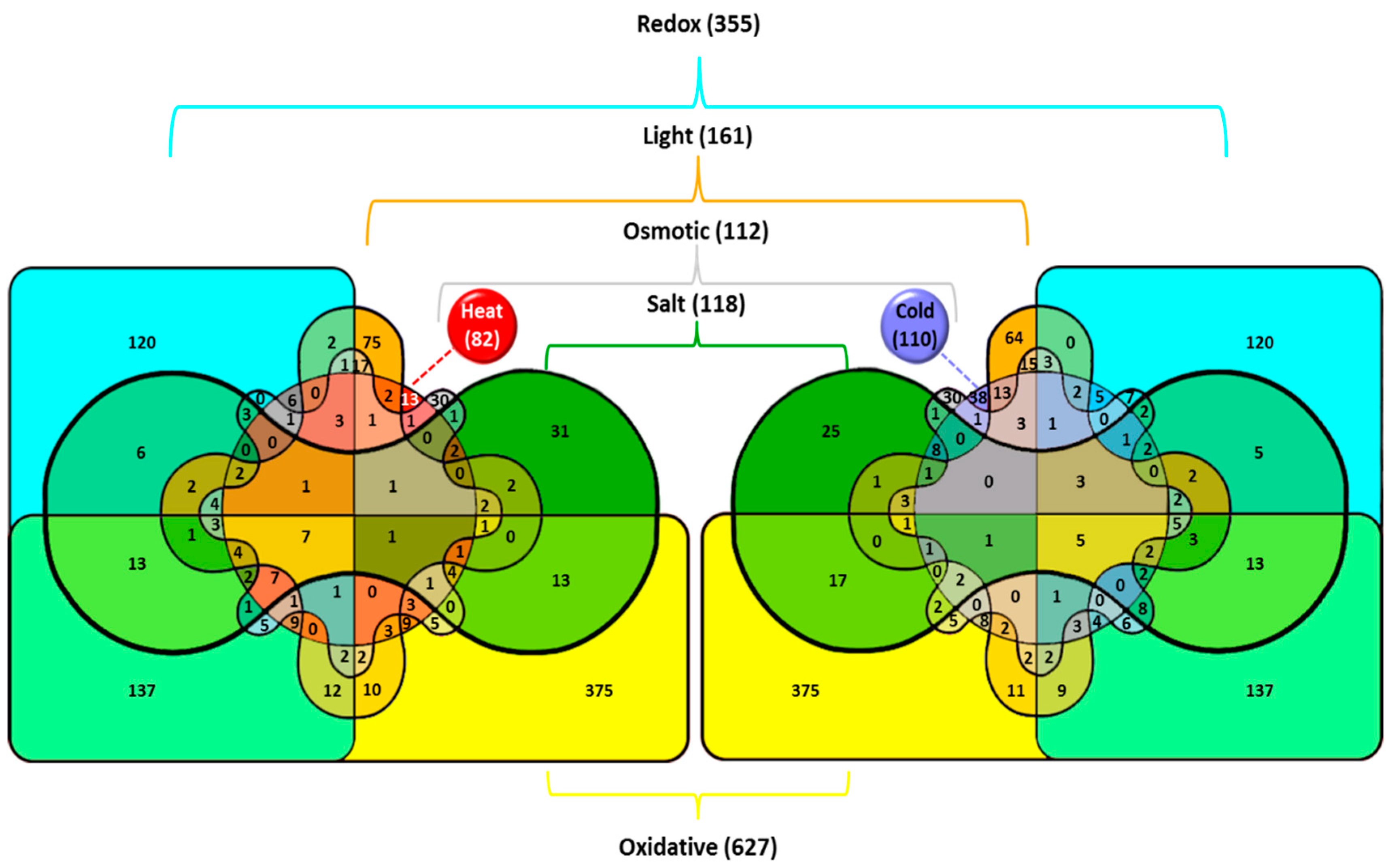
2. Systemic Analysis of Cyanobacterial Stress Transcriptomes
2.1. Genes Induced by Heat Stress
2.2. Genes Induced by Strong Light and Ultraviolet-B (UV-B)
2.3. Genes Induced by Salt and Hyperosmotic Stress
2.4. Genes Induced by Cold Shock and Other Stressors
2.5. Redox Regulation and Oxidative Stress
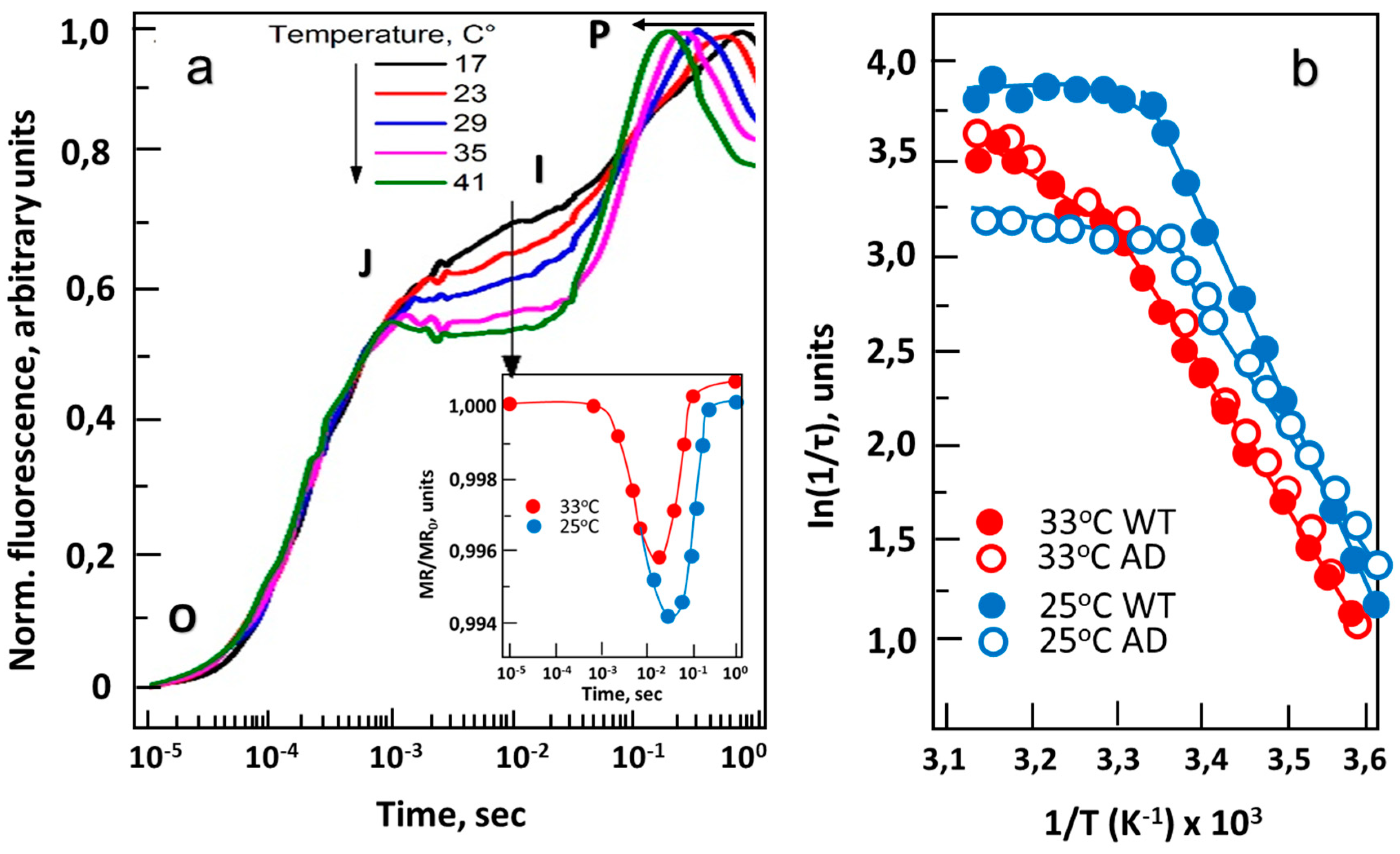
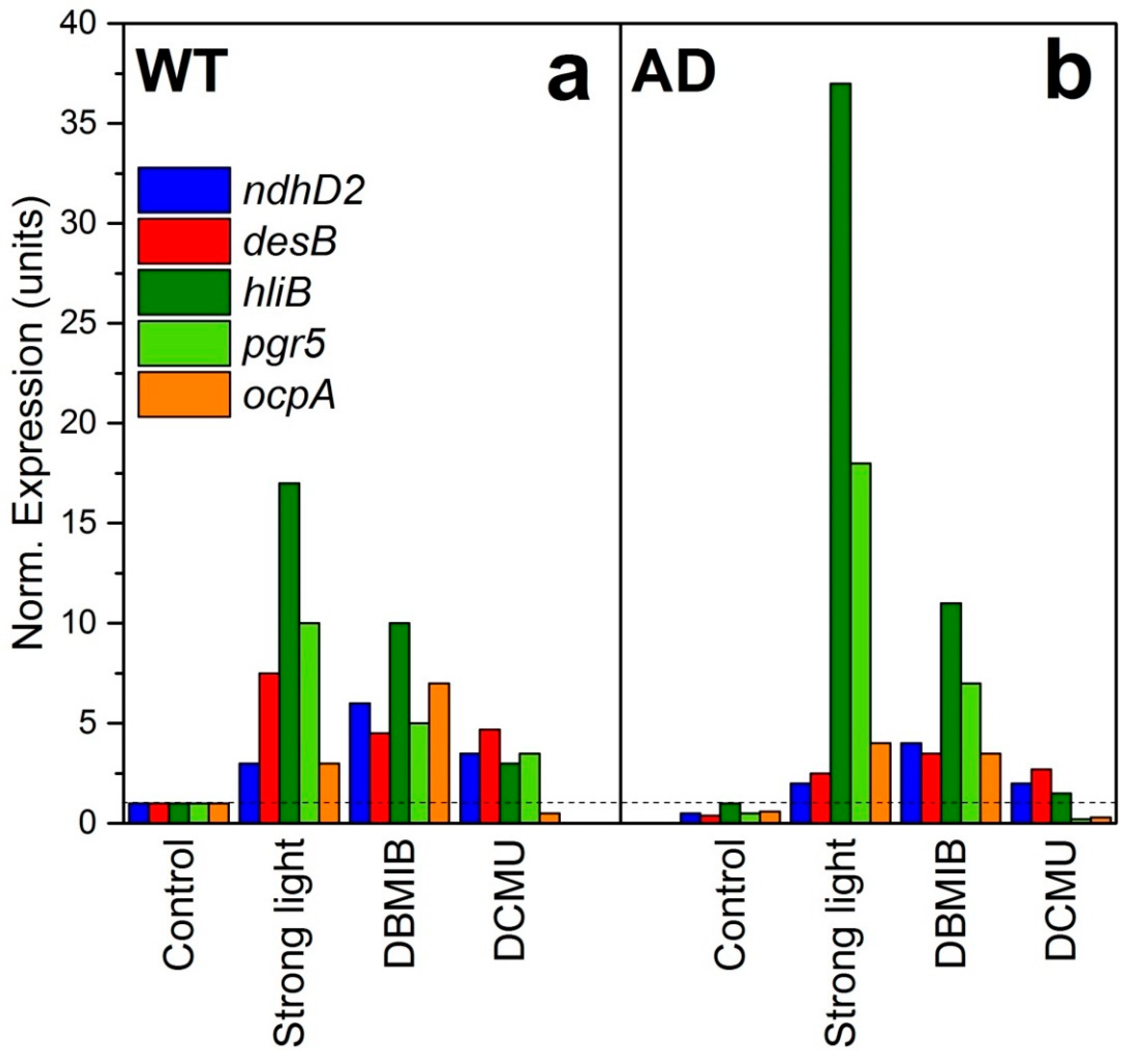
3. The Redox State of Plastoquinone (PQ) Depends on Membrane Fluidity
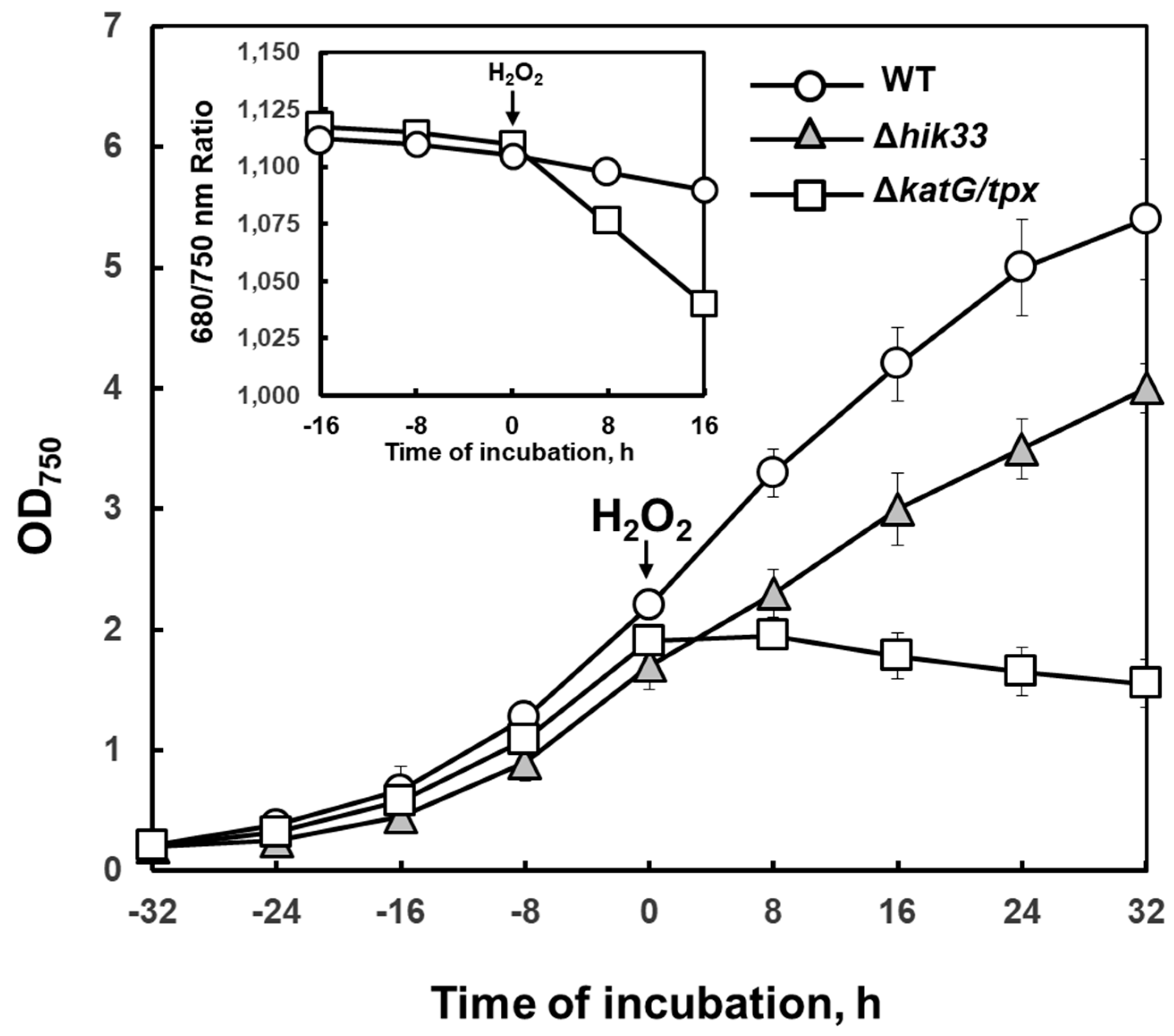
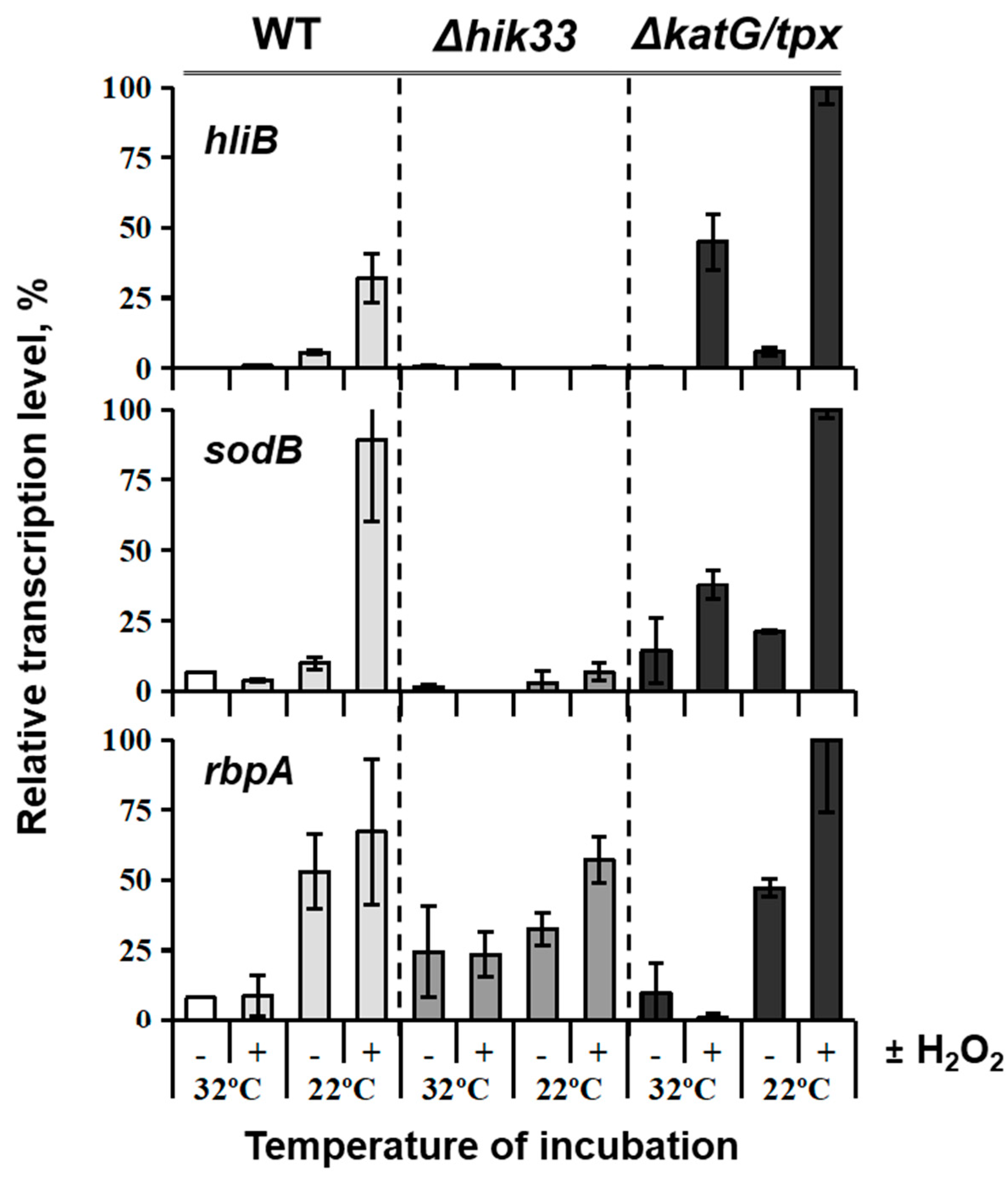
4. H2O2 is Involved in Regulation of Cold Stress Responses
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Los, D.A.; Zorina, A.; Sinetova, M.; Kryazhov, S.; Mironov, K.; Zinchenko, V.V. Stress sensors and signal transducers in cyanobacteria. Sensors 2010, 10, 2386–2415. [Google Scholar] [CrossRef] [PubMed]
- Bachin, D.; Nazarenko, L.V.; Mironov, K.S.; Pisareva, T.; Allakhverdiev, S.I.; Los, D.A. Mechanosensitive ion channel MscL controls ionic fluxes during cold and heat stress in Synechocystis. FEMS Microbiol. Lett. 2015, 362, fnv090. [Google Scholar] [CrossRef] [PubMed]
- Los, D.A.; Mironov, K.S.; Allakhverdiev, S.I. Regulatory role of membrane fluidity in gene expression and physiological functions. Photosynth. Res. 2013, 116, 489–509. [Google Scholar] [CrossRef] [PubMed]
- Červený, J.; Sinetova, M.A.; Zavřel, T.; Los, D.A. Mechanisms of high temperature resistance of Synechocystis sp. PCC 6803: An impact of histidine kinase 34. Life 2015, 5, 676–699. [Google Scholar] [CrossRef] [PubMed]
- Sinetova, M.A.; Los, D.A. Systemic analysis of transcriptomics of Synechocystis: Common stress genes and their universal triggers. Mol. BioSyst. 2016, 12, 3254–3258. [Google Scholar] [CrossRef]
- Sinetova, M.A.; Los, D.A. Lessons from cyanobacterial transcriptomics: Universal genes and triggers of stress responses. Mol. Biol. (Moscow) 2016, 50, 685–694. [Google Scholar] [CrossRef]
- Murata, N.; Los, D.A. Histidine kinase Hik33 is an important participant in cold signal transduction in cyanobacteria. Physiol. Plant. 2006, 126, 17–27. [Google Scholar] [CrossRef]
- Kahlon, S.; Beeri, K.; Ohkawa, H.; Hihara, Y.; Murik, O.; Suzuki, I.; Ogawa, T.; Kaplan, A. A putative sensor kinase, Hik31, is involved in the response of Synechocystis sp. strain PCC 6803 to the presence of glucose. Microbiology 2006, 152, 647–655. [Google Scholar] [CrossRef]
- Nagarajan, S.; Sherman, D.M.; Shawhttp, I.; Sherman, L.A. Functions of the duplicated hik31 operons in central metabolism and responses to light, dark, and carbon sources in Synechocystis sp. strain PCC 6803. J. Bacteriol. 2012, 194, 448–459. [Google Scholar] [CrossRef]
- Nagarajan, S.; Srivastava, S.; Sherman, L.A. Essential role of the plasmid hik31 operon in regulating central metabolism in the dark in Synechocystis sp. PCC 6803. Mol. Microbiol. 2014, 91, 79–97. [Google Scholar] [CrossRef] [PubMed]
- Giner-Lamia, J.; López-Maury, L.; Reyes, J.C.; Florencio, F.J. The CopRS two-component system is responsible for resistance to copper in the cyanobacterium Synechocystis sp. PCC 6803. Plant Physiol. 2012, 159, 1806–1818. [Google Scholar] [CrossRef]
- Giner-Lamia, J.; López-Maury, L.; Florencio, F.J. Global transcriptional profiles of the copper responses in the cyanobacterium Synechocystis sp. PCC 6803. PLoS ONE 2014, 9, e108912. [Google Scholar] [CrossRef]
- Suzuki, I.; Kanesaki, Y.; Hayashi, H.; Hall, J.J.; Simon, W.J.; Slabas, A.R.; Murata, N. The histidine kinase Hik34 is involved in thermotolerance by regulating the expression of heat shock genes in Synechocystis. Plant Physiol. 2005, 138, 1409–1421. [Google Scholar] [CrossRef]
- Kanesaki, Y.; Yamamoto, H.; Paithoonrangsarid, K.; Shumskaya, M.; Suzuki, I.; Hayashi, H.; Murata, N. Histidine kinases play important roles in the perception and signal transduction of hydrogen peroxide in the cyanobacterium, Synechocystis sp. PCC 6803. Plant J. 2007, 49, 313–324. [Google Scholar] [CrossRef]
- Inaba, M.; Suzuki, I.; Szalontai, B.; Kanesaki, Y.; Los, D.A.; Hayashi, H.; Murata, N. Genetically-engineered rigidification of membrane lipids enhances the cold inducibility of gene expression in Synechocystis. J. Biol. Chem. 2003, 278, 12191–12198. [Google Scholar] [CrossRef]
- Prakash, J.S.S.; Sinetova, M.; Kupriyanova, E.; Zorina, A.; Suzuki, I.; Murata, N.; Los, D.A. DNA supercoiling regulates the stress-inducible expression of genes in the cyanobacterium. Mol. BioSyst. 2009, 5, 1904–1912. [Google Scholar] [CrossRef]
- Imamura, S.; Asayama, M.; Takahashi, H.; Tanaka, K.; Takahashi, H.; Shirai, M. Antagonistic dark/light-induced SigB/SigD, group 2 sigma factors, expression through redox potential and their roles in cyanobacteria. FEBS Lett. 2003, 554, 357–362. [Google Scholar] [CrossRef]
- Tuominen, I.; Pollari, M.; Tyystjärvi, E.; Tyystjärvi, T. The SigB sigma factor mediates high-temperature responses in the cyanobacterium Synechocystis sp. PCC 6803. FEBS Lett. 2006, 580, 319–323. [Google Scholar] [CrossRef]
- Shumskaya, M.A.; Paithoonrangsarid, K.; Kanesaki, Y.; Los, D.A.; Zinchenko, V.V.; Tanticharoen, M.; Suzuki, I.; Murata, N. Identical Hik-Rre systems are involved in perception and transduction of salt signals and hyperosmotic signals but regulate the expression of individual genes to different extents in Synechocystis. J. Biol. Chem. 2005, 280, 21531–21538. [Google Scholar] [CrossRef]
- Pollari, M.; Gunnelius, L.; Tuominen, I.; Ruotsalainen, V.; Tyystjarvi, E.; Salminen, T.; Tyystjarvi, T. Characterization of single and double inactivation strains reveals new physiological roles for group 2 sigma factors in the cyanobacterium Synechocystis sp. PCC 6803. Plant Physiol. 2008, 147, 1994–2005. [Google Scholar] [CrossRef]
- Paithoonrangsarid, K.; Shoumskaya, M.A.; Kanesaki, Y.; Satoh, S.; Tabata, S.; Los, D.A.; Zinchenko, V.V.; Hayashi, H.; Tanticharoen, M.; Suzuki, I.; et al. Five histidine kinases perceive osmotic stress and regulate distinct sets of genes in Synechocystis. J. Biol. Chem. 2004, 279, 53078–53086. [Google Scholar] [CrossRef]
- Marin, K.; Kanesaki, Y.; Los, D.A.; Murata, N.; Suzuki, I.; Hagemann, M. Gene expression profiling reflects physiological processes in salt acclimation of Synechocystis sp. strain PCC 6803. Plant Physiol. 2004, 136, 3290–3300. [Google Scholar] [CrossRef]
- Hagemann, M. Molecular biology of cyanobacterial salt acclimation. FEMS Microbiol Rev. 2011, 35, 87–123. [Google Scholar] [CrossRef]
- Hihara, Y.; Kamei, A.; Kanehisa, M.; Kaplan, A.; Ikeuchi, M. DNA microarray analysis of cyanobacterial gene expression during acclimation to high light. Plant Cell 2001, 13, 793–806. [Google Scholar] [CrossRef]
- Huang, L.; McCluskey, M.P.; Ni, H.; LaRossa, R.A. Global gene expression profiles of the cyanobacterium Synechocystis sp. strain PCC 6803 in response to irradiation with UV-B and white light. J. Bacteriol. 2003, 184, 6845–6858. [Google Scholar] [CrossRef] [PubMed]
- Hihara, Y.; Sonoike, K.; Kanehisa, M.; Ikeuchi, M. DNA microarray analysis of redox-responsive genes in the genome of the cyanobacterium Synechocystis sp. strain PCC 6803. J. Bacteriol. 2003, 185, 1719–1725. [Google Scholar] [CrossRef]
- Ohta, H.; Shibata, Y.; Haseyama, Y.; Yoshino, Y.; Suzuki, T.; Kagasawa, T.; Kamei, A.; Ikeuchi, M.; Enami, I. Identification of genes expressed in response to acid stress in Synechocystis sp. PCC 6803 using DNA microarrays. Photosynth. Res. 2005, 84, 225–230. [Google Scholar] [CrossRef]
- Uchiyama, J.; Asakura, R.; Kimura, M.; Moriyama, A.; Tahara, H.; Kobayashi, Y.; Kubo, Y.; Yoshihara, T.; Ohta, H. Slr0967 and Sll0939 induced by the SphR response regulator in Synechocystis sp. PCC 6803 are essential for growth under acid stress conditions. Biochim. Biophys. Acta 2012, 1817, 1270–1276. [Google Scholar] [CrossRef] [PubMed]
- Hsiao, H.Y.; He, Q.; Van Waasbergen, L.G.; Grossman, A.R. Control of photosynthetic and high-light-responsive genes by the histidine kinase DspA: Negative and positive regulation and interactions between signal transduction pathways. J. Bacteriol. 2004, 186, 3882–3888. [Google Scholar] [CrossRef]
- Shapiguzov, A.; Lyukevich, A.A.; Allakhverdiev, S.I.; Sergeyenko, T.V.; Suzuki, I.; Murata, N.; Los, D.A. Osmotic shrinkage of cells of Synechocystis sp. PCC 6803 by water efflux via aquaporins regulates osmostress-inducible gene expression. Microbiology 2005, 151, 447–455. [Google Scholar] [CrossRef] [PubMed]
- Sinetova, M.A.; Mironov, K.S.; Mustardy, L.; Shapiguzov, A.; Bachin, D.; Allakhverdiev, S.I.; Los, D.A. Aquaporin-deficient mutant of Synechocystis is sensitive to salt and high-light stress. J. Photochem. Photobiol. B. Biol. 2015, 152, 377–382. [Google Scholar] [CrossRef]
- Kanesaki, Y.; Suzuki, I.; Allakhverdiev, S.I.; Mikami, K.; Murata, N. Salt stress and hyperosmotic stress regulate the expression of different sets of genes in Synechocystis sp. PCC 6803. Biochem. Biophys. Res. Commun. 2002, 290, 339–348. [Google Scholar] [CrossRef]
- Suzuki, I.; Kanesaki, Y.; Mikami, K.; Kanehisa, M.; Murata, N. Cold-regulated genes under control of the cold sensor Hik33 in Synechocystis. Mol. Microbiol. 2001, 40, 235–244. [Google Scholar] [CrossRef]
- Sinetova, M.A.; Los, D.A. New insights in cyanobacterial cold stress responses: Genes, sensors, and molecular triggers. Biochim. Biophys. Acta 2016, 1860, 2391–2403. [Google Scholar] [CrossRef] [PubMed]
- Takala, H.; Björling, A.; Berntsson, O.; Lehtivuori, H.; Niebling, S.; Hoernke, M.; Kosheleva, I.; Henning, R.; Menzel, A.; Ihalainen, J.A.; et al. Signal amplification and transduction in phytochrome photosensors. Nature 2014, 509, 245–248. [Google Scholar] [CrossRef]
- Wang, H.L.; Postier, B.L.; Burnap, R.L. Alterations in global 5 patterns of gene expression in Synechocystis sp. PCC 6803 in response to 6 inorganic carbon limitation and the inactivation of ndhR, a LysR family regulator. J. Biol. Chem. 2004, 279, 5739–5751. [Google Scholar] [CrossRef] [PubMed]
- Steczkiewicz, K.; Muszewska, A.; Knizewski, L.; Rychlewski, L.; Ginalski1, K. Sequence, structure and functional diversity of PD-(D/E)XK phosphodiesterase superfamily. Nucleic Acids Res. 2012, 40, 7016–7045. [Google Scholar] [CrossRef]
- Li, H.; Singh, A.K.; McIntyre, L.M.; Sherman, L.A. Differential gene expression in response to hydrogen peroxide and the putative PerR regulon of Synechocystis sp. strain PCC 6803. J. Bacteriol. 2004, 186, 3331–3345. [Google Scholar] [CrossRef] [PubMed]
- Peters, J.M.; Mooney, R.A.; Grass, J.A.; Jessen, E.D.; Tran, F.; Landick, R. Rho and NusG suppress pervasive antisense transcription in Escherichia coli. Genes Dev. 2012, 26, 2621–2633. [Google Scholar] [CrossRef]
- Karzai, A.W.; Susskind, M.M.; Sauer, R.T. SmpB, a unique RNA-binding protein essential for the peptide-tagging activity of SsrA (tmRNA). EMBO J. 1999, 18, 3793–3799. [Google Scholar] [CrossRef]
- Sireesha, K.; Radharani, B.; Krishna, P.S.; Sreedhar, N.; Subramanyam, R.; Mohanty, P.; Prakash, J.S. RNA helicase, CrhR is indispensable for the energy redistribution and the regulation of photosystem stoichiometry at low temperature in Synechocystis sp. PCC6803. Biochim. Biophys. Acta 2012, 1817, 1525–1536. [Google Scholar] [CrossRef] [PubMed]
- Georg, J.; Rosana, A.R.R.; Chamot, D.; Migur, A.; Hess, W.R.; Owttrim, G.W. Inactivation of the RNA helicase CrhR impacts a specific subset of the transcriptome in the cyanobacterium Synechocystis sp. PCC 6803. RNA Biol. 2019, 24, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Los, D.A.; Murata, N. Structure and expression of fatty acid desaturases. Biochim. Biophys. Acta 1998, 1394, 3–15. [Google Scholar] [CrossRef]
- Los, D.A.; Ray, M.K.; Murata, N. Differences in the control of the temperature-dependent expression of four genes for desaturases in Synechocystis sp. PCC 6803. Mol. Microbiol. 1997, 25, 1167–1175. [Google Scholar] [CrossRef]
- Tang, Q.; Tan, X.; Xu, X. Effects of a type-II RNA-binding protein on fatty acid composition in Synechocystis sp. PCC 6803. Chin. Sci. Bull. 2010, 55, 2416–2421. [Google Scholar] [CrossRef]
- He, Q.; Dolganov, N.; Bjorkman, O.; Grossman, A.R. The high light-inducible polypeptides in Synechocystis PCC6803. Expression and function in high light. J. Biol. Chem. 2001, 276, 306–314. [Google Scholar] [CrossRef]
- Mironov, K.S.; Sidorov, R.A.; Trofimova, M.S.; Bedbenov, V.S.; Tsydendambaev, V.D.; Allakhverdiev, S.I.; Los, D.A. Light-dependent cold-induced fatty acid unsaturation, changes in membrane fluidity, and alterations in gene expression in Synechocystis. Biochim. Biophys. Acta 2012, 1817, 1352–1359. [Google Scholar] [CrossRef] [PubMed]
- Chidgey, J.W.; Linhartová, M.; Komenda, J.; Jackson, P.J.; Dickman, M.J.; Canniffe, D.P.; Koník, P.; Pilný, J.; Hunter, C.N.; Sobotka, R. A cyanobacterial chlorophyll synthase-HliD complex associates with the Ycf39 protein and the YidC/Alb3 insertase. Plant Cell 2014, 26, 1267–1279. [Google Scholar] [CrossRef]
- Kufryk, G.; Hernandez-Prieto, M.A.; Kieselbach, T.; Miranda, H.; Vermaas, W.; Funk, C. Association of small CAB-like proteins (SCPs) of Synechocystis sp. PCC 6803 with photosystem II. Photosynth. Res. 2008, 95, 135–145. [Google Scholar] [CrossRef]
- Nakamura, K.; Hihara, Y. Photon flux density-dependent gene expression in Synechocystis sp. PCC 6803 is regulated by a small redox-responsive, LuxR-type regulator. J. Biol. Chem. 2006, 281, 36758–36766. [Google Scholar] [CrossRef]
- Horiuchi, M.; Nakamura, K.; Kojima, K.; Nishiyama, Y.; Hatakeyama, W.; Hisabori, T.; Hihara, Y. The PedR transcriptional regulator interacts with thioredoxin to connect photosynthesis with gene expression in cyanobacteria. Biochem. J. 2010, 431, 135–140. [Google Scholar] [CrossRef]
- Cameron, J.C.; Pakrasi, H.B. Essential role of glutathione in acclimation to environmental and redox perturbations in the cyanobacterium Synechocystis sp. PCC 6803. Plant Physiol. 2010, 154, 1672–1685. [Google Scholar] [CrossRef]
- Schmitt, F.-J.; Renger, G.; Friedrich, T.; Kreslavski, V.D.; Zharmukhamedov, S.K.; Los, D.A.; Kuznetsov, V.V.; Allakhverdiev, S.I. Reactive oxygen species: Re-evaluation of generation, monitoring and role in stress-signaling in phototrophic organisms. Biochim. Biophys. Acta 2014, 1837, 835–848. [Google Scholar] [CrossRef] [PubMed]
- Singh, A.K.; Li, H.; Sherman, L.A. Microarray analysis and redox control of gene expression in the cyanobacterium Synechocystis sp. PCC 6803. Physiol. Plant. 2004, 120, 27–35. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.W.; Helmann, J.D. The PerR transcription factor senses H2O2 by metal-catalysed histidine oxidation. Nature 2006, 440, 363–367. [Google Scholar] [CrossRef]
- Saita, E.; Albanesi, D.; de Mendoza, D. Sensing membrane thickness: Lessons learned from cold stress. Biochim. Biophys. Acta 2016, 1861, 837–846. [Google Scholar]
- Mironov, K.S.; Sidorov, R.A.; Kreslavski, V.D.; Bedbenov, V.S.; Tsydendambaev, V.D.; Los, D.A. Cold-induced gene expression and ω3 fatty acid unsaturation is controlled by red light in Synechocystis. J. Photochem. Photobiol. B Biol. 2014, 137, 84–88. [Google Scholar] [CrossRef]
- Guskov, A.; Kern, J.; Gabdulkhakov, A.; Broser, M.; Zouni, A.; Saenger, W. Cyanobacterial photosystem II at 2.9-A resolution and the role of quinones, lipids, channels and chloride. Nat. Struct. Mol. Biol. 2009, 16, 334–342. [Google Scholar] [CrossRef]
- Maksimov, E.G.; Mironov, K.S.; Trofimova, M.S.; Nechaeva, N.L.; Todorenko, D.A.; Klementiev, K.E.; Tsoraev, G.V.; Tyutyaev, E.V.; Zorina, A.A.; Feduraev, P.V.; et al. Membrane fluidity controls redox-regulated cold stress responses in cyanobacteria. Photosynth. Res. 2017, 133, 215–223. [Google Scholar]
- Tasaka, Y.; Gombos, Z.; Nishiyama, Y.; Mohanty, P.; Ohba, T.; Ohki, K.; Murata, N. Targeted mutagenesis of acyl-lipid desaturases in Synechocystis: Evidence for the important roles of polyunsaturated membrane lipids in growth, respiration and photosynthesis. EMBO J. 1996, 15, 6416–6425. [Google Scholar] [CrossRef]
- Allahverdiyeva, Y.; Mustila, H.; Ermakova, M.; Bersanini, L.; Richaud, P.; Ajlani, G.; Battchikova, N.; Cournac, L.; Aro, E.M. Flavodiiron proteins Flv1 and Flv3 enable cyanobacterial growth and photosynthesis under fluctuating light. Proc. Natl. Acad. Sci. USA 2013, 110, 4111–4116. [Google Scholar] [CrossRef]
- Mironov, K.S.; Shumskaya, M.; Sidorov, R.A.; Trofimova, M.S.; Los, D.A. Membrane physical state and stress regulation in Synechocystis: Fluidizing alcohols repress fatty acid desaturation. Plant J. 2018, 96, 1007–1017. [Google Scholar] [CrossRef]
- Vigh, L.; Los, D.A.; Horváth, I.; Murata, N. The primary signal in the biological perception of temperature: Pd-catalyzed hydrogenation of membrane lipids stimulated the expression of the desA gene in Synechocystis PCC6803. Proc. Natl. Acad. Sci. USA 1993, 90, 9090–9094. [Google Scholar] [CrossRef]
- Horváth, I.; Glatz, A.; Nakamoto, H.; Mishkind, M.L.; Munnik, T.; Saidi, Y.; Goloubinoff, P.; Harwood, J.L.; Vigh, L. Heat shock response in photosynthetic organisms: Membrane and lipid connections. Prog Lipid Res. 2012, 51, 208–220. [Google Scholar] [CrossRef]
- Yamamoto, H.; Miyake, C.; Dietz, K.J.; Tomizawa, K.; Murata, N.; Yokota, A. Thioredoxin peroxidase in the cyanobacterium Synechocystis sp. PCC 6803. FEBS Lett. 1999, 447, 269–273. [Google Scholar] [CrossRef]
- Fedurayev, P.V.; Mironov, K.S.; Gabrielyan, D.A.; Bedbenov, V.S.; Zorina, A.A.; Shumskaya, M.; Los, D.A. Hydrogen peroxide participates in perception and transduction of cold stress signal in Synechocystis. Plant Cell Physiol. 2018, 59, 1255–1264. [Google Scholar] [CrossRef]
- Kujirai, J.; Nanba, S.; Kadowaki, T.; Oka, Y.; Nishiyama, Y.; Hayashi, Y.; Arai, M.; Hihara, Y. Interaction of the GntR-family transcription factor Sll1961 with thioredoxin in the cyanobacterium Synechocystis sp. PCC 6803. Sci. Rep. 2018, 8, 6666. [Google Scholar]
- Hakkila, K.; Valev, D.; Antal, T.; Tyystjävi, E.; Tyystjävi, T. Group 2 sigma factors are central regulators of oxidative stress acclimation in cyanobacteria. Plant Cell Physiol. 2019, 60, 436–447. [Google Scholar] [CrossRef]
- Marinho, H.S.; Real, C.; Cyrne, L.; Soares, H.; Antunes, F. Hydrogen peroxide sensing, signaling and regulation of transcription factors. Redox Biol. 2014, 2, 535–562. [Google Scholar] [CrossRef]
- Kojima, K.; Oshita, M.; Nanjo, Y.; Kasai, K.; Tozawa, Y.; Hayashi, H.; Nishiyama, Y. Oxidation of elongation factor G inhibits the synthesis of the D1 protein of photosystem II. Mol. Microbiol. 2007, 65, 936–947. [Google Scholar] [CrossRef]
- Yutthanasirikul, R.; Nagano, T.; Jimbo, H.; Hihara, Y.; Kanamori, T.; Ueda, T.; Haruyama, T.; Konno, H.; Yoshida, K.; Hisabori, T.; et al. Oxidation of a cysteine residue in elongation factor EF-Tu reversibly inhibits translation in the cyanobacterium Synechocystis sp. PCC 6803. J. Biol. Chem. 2016, 291, 5860–5870. [Google Scholar] [CrossRef] [PubMed]
- Khursigara, C.M.; Wu, X.; Zhang, P.; Lefman, J.; Subramaniam, S. Role of HAMP domains in chemotaxis signaling by bacterial chemoreceptors. Proc. Natl. Acad. Sci. USA 2008, 105, 16555–16560. [Google Scholar] [CrossRef]
- Sporer, A.J.; Kahl, L.J.; Price-Whelan, A.; Dietrich, L.E.P. Redox-based regulation of bacterial development and behavior. Annu. Rev. Biochem. 2017, 86, 777–797. [Google Scholar] [CrossRef]
| Abiotic Stressors | Redox | ROS | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ORF | Gene | Protein | Function | Heat | Salt | Osmo | Light | UV-B | pH | DBMIB | DCMU | H2O2 |
| slr1285 | hik34 | Hik34 | Sensor histidine kinase Hik34 | + | + | + | + | + | + | + | ||
| sll0306 | sigB | SigB | Sigma factor B of RNA polymerase | + | + | + | + | + | + | + | ||
| slr2075 | groES | GroES | Heat-shock protein (HSP) 10 kDa co-chaperonin | + | + | + | + | + | + | |||
| slr2076 | groEL1 | GroEL1 | HSP 60 kDa chaperonin 1 | + | + | + | + | + | + | |||
| sll0416 | groEL2 | GroEL2 | HSP 60 kDa chaperonin 2 | + | + | + | + | |||||
| sll1514 | hspA | HspA | HSP 17 kDa | + | + | + | + | + | + | + | + | |
| sll0170 | dnaK2 | DnaK2 | HSP 70 kDa | + | + | + | + | + | + | + | + | |
| slr0093 | dnaJ | DnaJ | HSP 40 kDa | + | + | + | + | + | + | + | ||
| sll0430 | htpG | HtpG | HSP 90 kDa | + | + | + | + | + | + | |||
| slr1641 | clpB1 | ClpB1 | HSP 100 kDa chaperone | + | + | + | + | + | + | + | ||
| slr1204 | htrA | HtrA | Serine protease | + | + | + | + | + | + | + | ||
| slr0008 | ctpA | CtpA | C-terminal processing protease | + | + | + | + | + | ||||
| sll1621 | sll1621 | PrxA | Peroxiredoxin | + | + | + | + | + | + | |||
| slr0095 | smtA | SmtA | S-adenosyl methionine methyltransferase | + | + | + | + | + | ||||
| slr1674 | slr1674 | Slr1674 | Thermoprotector protein of PS II | + | + | + | + | + | + | + | ||
| slr1963 | ocpA | OCP | Orange carotenoid-binding protein | + | + | + | + | + | ||||
| slr1512 | sbtA | SbtA | Na-dependent bicarnonate transporter | + | + | + | ||||||
| slr1516 | sodB | SodB | Superoxide dismutase | + | + | + | + | + | + | |||
| sll0528 | sll0528 | Sll0528 | Site-2-protease | + | + | + | + | + | + | + | + | |
| slr1675 | hypA1 | HypA1 | Hydrogenase maturation factor HypA1 | + | + | + | + | + | + | + | + | |
| ssl3044 | ssl3044 | Fdx | Ferredoxin | + | + | + | + | + | + | + | + | |
| slr1687 | nblB1 | NblB1 | Phycobilisome degradation protein | + | + | + | + | + | + | + | + | + |
| sll0939 | sll0939 | Sll0939 | Low pH resistance protein | + | + | + | + | + | + | + | + | |
| slr0967 | slr0967 | Slr0967 | Low pH resistance protein | + | + | + | + | + | + | + | + | |
| slr1686 | slr1686 | Unknown function | + | + | + | + | + | |||||
| slr1603 | slr1603 | Unknown function | + | + | + | + | + | + | ||||
| sll1853 | sll1853 | Unknown function | + | + | + | |||||||
| sll0846 | sll0846 | Unknown function | + | + | + | + | + | + | + | + | ||
| sll1884 | sll1884 | Unknown function | + | + | + | + | + | |||||
| sll1009* | frpC* | FrpC | Membrane Fe-regulated protein | + | ||||||||
| sll1501* | cbiA* | CbiA | Cobalamine biosynthesis protein | + | ||||||||
| sll0441* | sll0441* | Unknown function | + | |||||||||
| sll1892* | sll1892* | Unknown function | + | |||||||||
| slr0670* | slr0670* | Unknown function | + | |||||||||
| sll0982* | sll0982* | Unknown function | + | |||||||||
| slr1127* | slr1127* | Unknown function | + | |||||||||
| ORF | Gene | Protein | Function | Abiotic stressor | Redox | ROS | ||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Cold | Salt | Osmo | Light | UV-B | pH | DBMIB | DCMU | H2O2 | ||||
| sll0790 | hik31 | Hik31 | Two-component sensor histidine kinase | + | + | |||||||
| slr1594 | rre5 | Rre5 | Response regulator | + | + | + | ||||||
| slr0083 | crhR | CrhR | RNA helicase | + | + | + | + | + | ||||
| sll0517 | rbpA1 | RbpA1 | RNA binding protein A1 | + | + | |||||||
| sll0533 | tig | Tig | Ribosome trigger factor | + | + | + | ||||||
| sll1096 | rpsL | Rps12 | 30S ribosomal protein S12 | + | + | + | ||||||
| slr0082 | rimO | RimO | Methyltransferase of ribosomal protein S12 | + | + | + | + | + | ||||
| slr1105 | typA | TypA | GTP-binding protein TypA/BipA | + | + | + | + | + | ||||
| slr1512 | sbtA | SbtA1 | Na-dependent bicarbonate transporter | + | + | + | ||||||
| sll1541 | syc2 | Syc2 | Carotenoid oxygenase | + | + | + | + | + | + | |||
| slr1291 | ndhD2 | NdhD2 | NADH dehydrogenase subunit 4 | + | + | + | + | + | ||||
| ssl2542 | hliA | HliA | High light inducible protein | + | + | + | + | + | + | + | ||
| ssr2595 | hliB | HliB | High light inducible protein | + | + | + | + | + | + | + | + | |
| slr1544 | lilA | LilA | Light-harvesting protein LilA | + | + | + | + | + | + | + | + | + |
| slr1687 | nblB | NblB | Phycocyanobilin lyase NblB | + | + | + | + | + | + | + | + | + |
| sll1742 | nusG | NusG | Transcription antitermination protein | + | + | + | + | + | ||||
| sll1818 | rpoA | RpoA | RNA polymerase alpha subunit σ70 | + | + | + | ||||||
| sll20121 | sigD1 | SigD1 | RNA polymerase sigma factor SigD | + | + | + | + | + | + | + | + | |
| ssl30441 | ssl30441 | Fdx1 | Ferredoxin | + | + | + | + | + | + | + | + | |
| slr1350 | desA | DesA | Δ12 fatty acid desaturase | + | + | |||||||
| sll1441 | desB | DesB | ω3 fatty acid desaturase | + | + | |||||||
| slr1992 | gpx2 | Gpx2 | Hydroperoxy fatty acid reductase | + | + | + | ||||||
| sll1483 | sll1483 | Salt-induced periplasmic protein | + | + | + | + | + | + | + | + | + | |
| sll0157 | sll0157 | Zn-dependent hydrolase | + | + | + | + | + | |||||
| sll1863 | sll1863 | Unknown protein | + | + | + | + | ||||||
| sll1862 | sll1862 | Unknown protein | + | + | + | + | ||||||
| sll1853 | sll1853 | Unknown protein | + | + | + | |||||||
| slr0551 | slr0551 | Unknown protein | + | + | + | + | + | + | ||||
| slr0959 | slr0959 | Unknown protein | + | + | + | + | ||||||
| slr1686 | slr1686 | Unknown protein | + | + | + | + | + | |||||
| slr1687 | slr1687 | Unknown protein | + | + | + | + | + | |||||
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mironov, K.S.; Sinetova, M.A.; Shumskaya, M.; Los, D.A. Universal Molecular Triggers of Stress Responses in Cyanobacterium Synechocystis. Life 2019, 9, 67. https://doi.org/10.3390/life9030067
Mironov KS, Sinetova MA, Shumskaya M, Los DA. Universal Molecular Triggers of Stress Responses in Cyanobacterium Synechocystis. Life. 2019; 9(3):67. https://doi.org/10.3390/life9030067
Chicago/Turabian StyleMironov, Kirill S., Maria A. Sinetova, Maria Shumskaya, and Dmitry A. Los. 2019. "Universal Molecular Triggers of Stress Responses in Cyanobacterium Synechocystis" Life 9, no. 3: 67. https://doi.org/10.3390/life9030067
APA StyleMironov, K. S., Sinetova, M. A., Shumskaya, M., & Los, D. A. (2019). Universal Molecular Triggers of Stress Responses in Cyanobacterium Synechocystis. Life, 9(3), 67. https://doi.org/10.3390/life9030067






