The Role of Orthogonality in Genetic Code Expansion
Abstract
1. Introduction
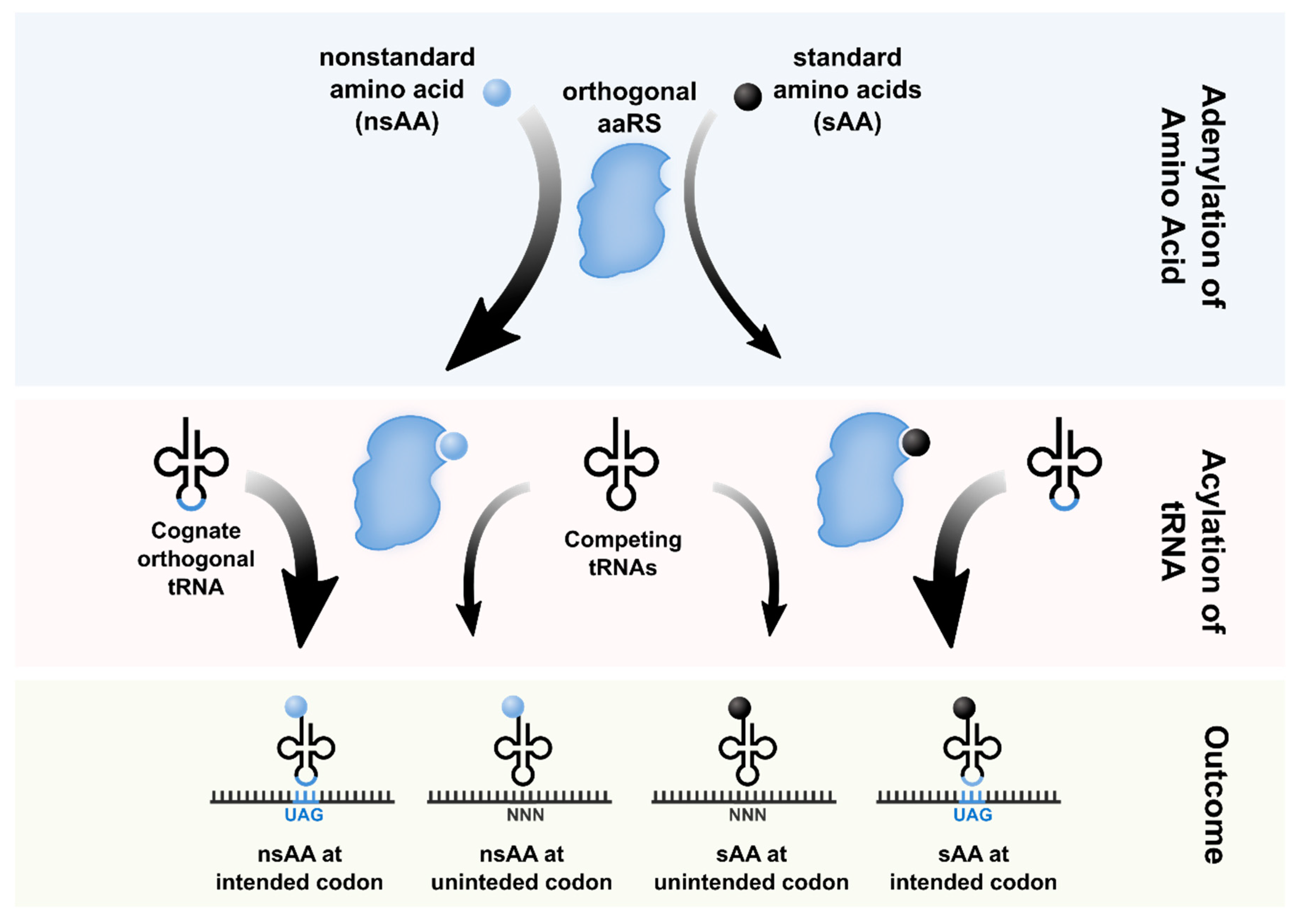
2. Orthogonal Translation Systems
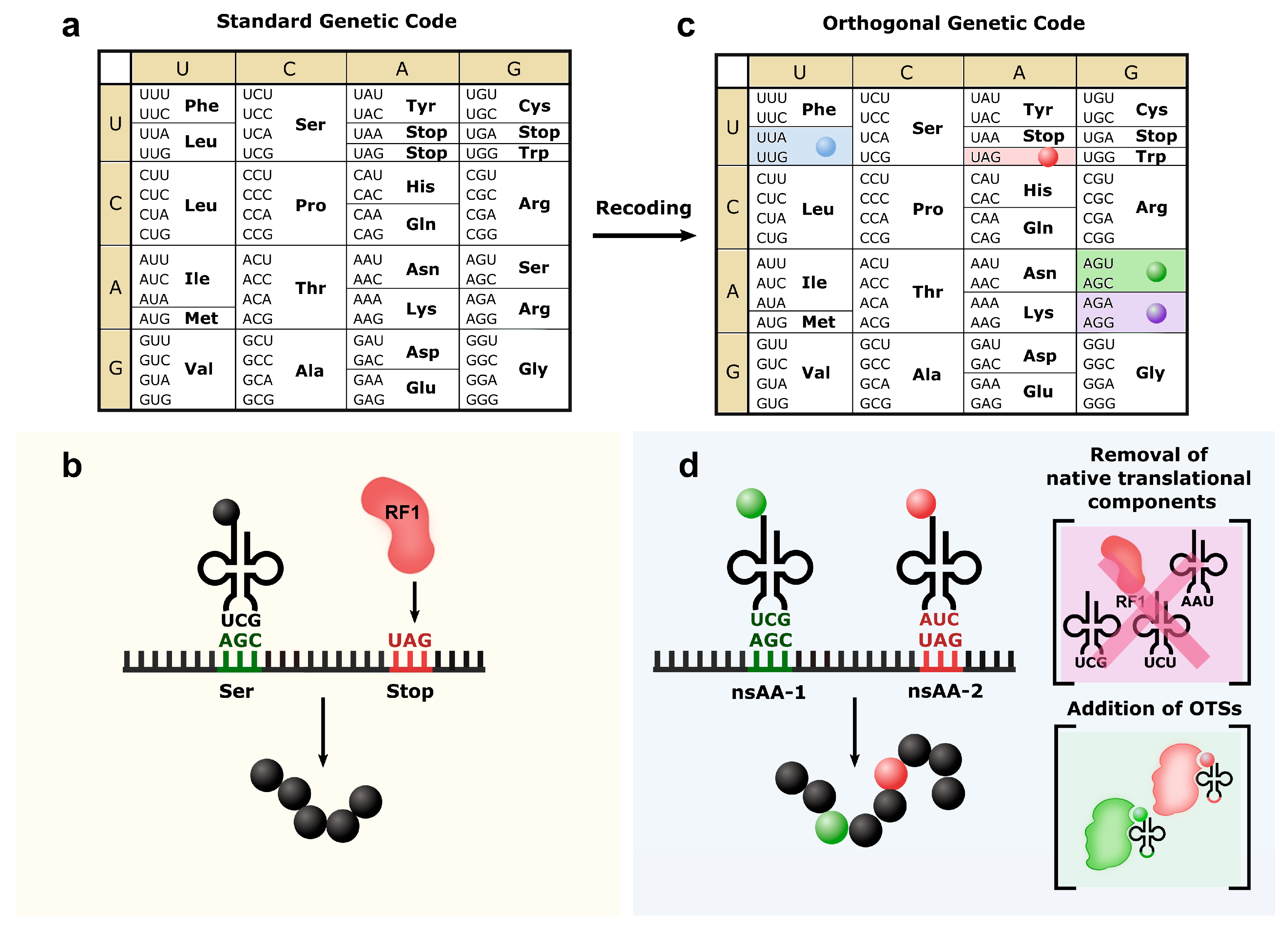
3. Orthogonal Genetic Codes
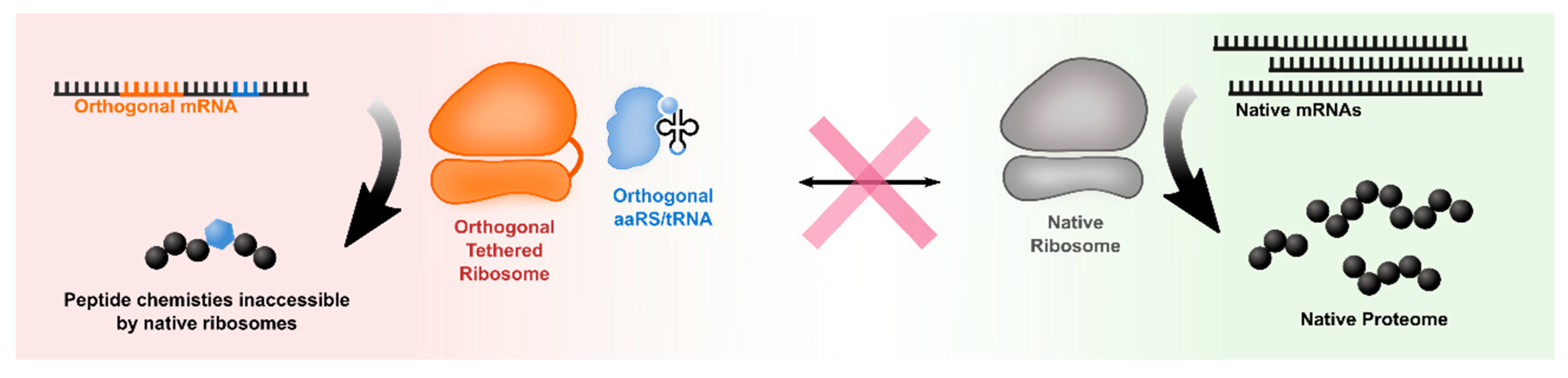
4. Orthogonal Ribosomes
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Bonitz, S.G.; Berlani, R.; Coruzzi, G.; Li, M.; Macino, G.; Nobrega, F.G.; Nobrega, M.P.; Thalenfeld, B.E.; Tzagoloff, A. Codon recognition rules in yeast mitochondria. Proc. Natl. Acad. Sci. USA 1980, 77, 3167–3170. [Google Scholar] [CrossRef]
- Fearnley, I.M.; Walker, J.E. Initiation codons in mammalian mitochondria: Differences in genetic code in the organelle. Biochemistry 1987, 26, 8247–8251. [Google Scholar] [CrossRef] [PubMed]
- Kano, A.; Andachi, Y.; Ohama, T.; Osawa, S. Novel anticodon composition of transfer rnas in micrococcus luteus, a bacterium with a high genomic g+c content: Correlation with codon usage. J. Mol. Biol. 1991, 221, 387–401. [Google Scholar] [CrossRef]
- Low, S.C.; Berry, M.J. Knowing when not to stop: Selenocysteine incorporation in eukaryotes. Trends Biochem. Sci. 1996, 21, 203–208. [Google Scholar] [CrossRef]
- Srinivasan, G.; James, C.M.; Krzycki, J.A. Pyrrolysine encoded by uag in archaea: Charging of a uag-decoding specialized trna. Science 2002, 296, 1459–1462. [Google Scholar] [CrossRef] [PubMed]
- Craigen, W.J.; Caskey, C.T. Expression of peptide chain release factor 2 requires high-efficiency frameshift. Nature 1986, 322, 273–275. [Google Scholar] [CrossRef] [PubMed]
- Arranz-Gibert, P.; Vanderschuren, K.; Isaacs, F.J. Next-generation genetic code expansion. Curr. Opin. Chem. Biol. 2018, 46, 203–211. [Google Scholar] [CrossRef]
- Nessen, M.A.; Kramer, G.; Back, J.; Baskin, J.M.; Smeenk, L.E.; de Koning, L.J.; van Maarseveen, J.H.; de Jong, L.; Bertozzi, C.R.; Hiemstra, H.; et al. Selective enrichment of azide-containing peptides from complex mixtures. J. Proteome. Res. 2009, 8, 3702–3711. [Google Scholar] [CrossRef][Green Version]
- Noren, C.J.; Anthony-Cahill, S.J.; Griffith, M.C.; Schultz, P.G. A general method for site-specific incorporation of unnatural amino acids into proteins. Science 1989, 244, 182–188. [Google Scholar] [CrossRef]
- Melnikov, S.V.; Söll, D. Aminoacyl-trna synthetases and trnas for an expanded genetic code: What makes them orthogonal? Int. J. Mol. Sci. 2019, 20, 1929. [Google Scholar] [CrossRef]
- Wang, L.; Magliery, T.J.; Liu, D.R.; Schultz, P.G. A new functional suppressor trna/aminoacyl−trna synthetase pair for the in vivo incorporation of unnatural amino acids into proteins. J. Am. Chem. Soc. 2000, 122, 5010–5011. [Google Scholar] [CrossRef]
- Xue, H.; Shen, W.; Giege, R.; Wong, J.T. Identity elements of trna(trp). Identification and evolutionary conservation. J. Biol. Chem. 1993, 268, 9316–9322. [Google Scholar]
- Wang, L.; Schultz, P.G. A general approach for the generation of orthogonal trnas. Chem. Biol. 2001, 8, 883–890. [Google Scholar] [CrossRef]
- Italia, J.S.; Addy, P.S.; Erickson, S.B.; Peeler, J.C.; Weerapana, E.; Chatterjee, A. Mutually orthogonal nonsense-suppression systems and conjugation chemistries for precise protein labeling at up to three distinct sites. J. Am. Chem. Soc. 2019, 141, 6204–6212. [Google Scholar] [CrossRef]
- Willis, J.C.W.; Chin, J.W. Mutually orthogonal pyrrolysyl-trna synthetase/trna pairs. Nat. Chem. 2018, 10, 831–837. [Google Scholar] [CrossRef]
- Monk, J.W.; Leonard, S.P.; Brown, C.W.; Hammerling, M.J.; Mortensen, C.; Gutierrez, A.E.; Shin, N.Y.; Watkins, E.; Mishler, D.M.; Barrick, J.E. Rapid and inexpensive evaluation of nonstandard amino acid incorporation in escherichia coli. ACS Synth. Biol. 2017, 6, 45–54. [Google Scholar] [CrossRef]
- Amiram, M.; Haimovich, A.D.; Fan, C.; Wang, Y.-S.; Aerni, H.-R.; Ntai, I.; Moonan, D.W.; Ma, N.J.; Rovner, A.J.; Hong, S.H.; et al. Evolution of translation machinery in recoded bacteria enables multi-site incorporation of nonstandard amino acids. Nat. Biotechnol. 2015, 33, 1272. [Google Scholar] [CrossRef]
- Goodman, H.M.; Abelson, J.; Landy, A.; Brenner, S.; Smith, J.D. Amber suppression: A nucleotide change in the anticodon of a tyrosine transfer rna. Nature 1968, 217, 1019–1024. [Google Scholar] [CrossRef]
- Knott, G.J.; Doudna, J.A. Crispr-cas guides the future of genetic engineering. Science 2018, 361, 866–869. [Google Scholar] [CrossRef]
- Hsu, P.D.; Lander, E.S.; Zhang, F. Development and applications of crispr-cas9 for genome engineering. Cell 2014, 157, 1262–1278. [Google Scholar] [CrossRef]
- Wang, H.H.; Isaacs, F.J.; Carr, P.A.; Sun, Z.Z.; Xu, G.; Forest, C.R.; Church, G.M. Programming cells by multiplex genome engineering and accelerated evolution. Nature 2009, 460, 894–898. [Google Scholar] [CrossRef]
- Ma, N.J.; Moonan, D.W.; Isaacs, F.J. Precise manipulation of bacterial chromosomes by conjugative assembly genome engineering. Nat. Protoc. 2014, 9, 2285–2300. [Google Scholar] [CrossRef]
- Haimovich, A.D.; Muir, P.; Isaacs, F.J. Genomes by design. Nat. Rev. Genet. 2015, 16, 501–516. [Google Scholar] [CrossRef]
- Isaacs, F.J.; Carr, P.A.; Wang, H.H.; Lajoie, M.J.; Sterling, B.; Kraal, L.; Tolonen, A.C.; Gianoulis, T.A.; Goodman, D.B.; Reppas, N.B.; et al. Precise manipulation of chromosomes in vivo enables genome-wide codon replacement. Science 2011, 333, 348–353. [Google Scholar] [CrossRef]
- Lajoie, M.J.; Rovner, A.J.; Goodman, D.B.; Aerni, H.-R.; Haimovich, A.D.; Kuznetsov, G.; Mercer, J.A.; Wang, H.H.; Carr, P.A.; Mosberg, J.A.; et al. Genomically recoded organisms expand biological functions. Science 2013, 342, 357–360. [Google Scholar] [CrossRef]
- Mukai, T.; Hayashi, A.; Iraha, F.; Sato, A.; Ohtake, K.; Yokoyama, S.; Sakamoto, K. Codon reassignment in the escherichia coli genetic code. Nucleic Acids Res. 2010, 38, 8188–8195. [Google Scholar] [CrossRef]
- Heinemann, I.U.; Rovner, A.J.; Aerni, H.R.; Rogulina, S.; Cheng, L.; Olds, W.; Fischer, J.T.; Söll, D.; Isaacs, F.J.; Rinehart, J. Enhanced phosphoserine insertion during escherichia coli protein synthesis via partial uag codon reassignment and release factor 1 deletion. FEBS Lett. 2012, 586, 3716–3722. [Google Scholar] [CrossRef]
- Johnson, D.B.F.; Xu, J.; Shen, Z.; Takimoto, J.K.; Schultz, M.D.; Schmitz, R.J.; Xiang, Z.; Ecker, J.R.; Briggs, S.P.; Wang, L. Rf1 knockout allows ribosomal incorporation of unnatural amino acids at multiple sites. Nat. Chem. Biol. 2011, 7, 779–786. [Google Scholar] [CrossRef]
- Fredens, J.; Wang, K.; de la Torre, D.; Funke, L.F.H.; Robertson, W.E.; Christova, Y.; Chia, T.; Schmied, W.H.; Dunkelmann, D.L.; Beránek, V.; et al. Total synthesis of escherichia coli with a recoded genome. Nature 2019, 569, 514–518. [Google Scholar] [CrossRef]
- Lajoie, M.J.; Kosuri, S.; Mosberg, J.A.; Gregg, C.J.; Zhang, D.; Church, G.M. Probing the limits of genetic recoding in essential genes. Science 2013, 342, 361–363. [Google Scholar] [CrossRef]
- Lau, Y.H.; Stirling, F.; Kuo, J.; Karrenbelt, M.A.; Chan, Y.A.; Riesselman, A.; Horton, C.A.; Schäfer, E.; Lips, D.; Weinstock, M.T.; et al. Large-scale recoding of a bacterial genome by iterative recombineering of synthetic DNA. Nucleic Acids Res. 2017, 45, 6971–6980. [Google Scholar] [CrossRef]
- Napolitano, M.G.; Landon, M.; Gregg, C.J.; Lajoie, M.J.; Govindarajan, L.; Mosberg, J.A.; Kuznetsov, G.; Goodman, D.B.; Vargas-Rodriguez, O.; Isaacs, F.J.; et al. Emergent rules for codon choice elucidated by editing rare arginine codons in <em>escherichia coli</em>. Proc. Natl. Acad. Sci. USA 2016, 113, E5588. [Google Scholar]
- Krishnakumar, R.; Prat, L.; Aerni, H.R.; Ling, J.; Merryman, C.; Glass, J.I.; Rinehart, J.; Soll, D. Transfer rna misidentification scrambles sense codon recoding. Chembiochem 2013, 14, 1967–1972. [Google Scholar] [CrossRef]
- Hong, S.H.; Kwon, Y.-C.; Jewett, M.C. Non-standard amino acid incorporation into proteins using escherichia coli cell-free protein synthesis. Front. Chem. 2014, 2, 34. [Google Scholar] [CrossRef]
- Cui, Z.; Mureev, S.; Polinkovsky, M.E.; Tnimov, Z.; Guo, Z.; Durek, T.; Jones, A.; Alexandrov, K. Combining sense and nonsense codon reassignment for site-selective protein modification with unnatural amino acids. ACS Synth. Biol. 2017, 6, 535–544. [Google Scholar] [CrossRef]
- Martin, R.W.; Des Soye, B.J.; Kwon, Y.-C.; Kay, J.; Davis, R.G.; Thomas, P.M.; Majewska, N.I.; Chen, C.X.; Marcum, R.D.; Weiss, M.G.; et al. Cell-free protein synthesis from genomically recoded bacteria enables multisite incorporation of noncanonical amino acids. Nat. Commun. 2018, 9, 1203. [Google Scholar] [CrossRef]
- Anderson, J.C.; Magliery, T.J.; Schultz, P.G. Exploring the limits of codon and anticodon size. Chem. Biol. 2002, 9, 237–244. [Google Scholar] [CrossRef]
- Anderson, J.C.; Wu, N.; Santoro, S.W.; Lakshman, V.; King, D.S.; Schultz, P.G. An expanded genetic code with a functional quadruplet codon. Proc. Natl. Acad. Sci. USA 2004, 101, 7566–7571. [Google Scholar] [CrossRef]
- Chatterjee, A.; Lajoie, M.J.; Xiao, H.; Church, G.M.; Schultz, P.G. A bacterial strain with a unique quadruplet codon specifying non-native amino acids. Chembiochem 2014, 15, 1782–1786. [Google Scholar] [CrossRef]
- Yamashige, R.; Kimoto, M.; Okumura, R.; Hirao, I. Visual detection of amplified DNA by polymerase chain reaction using a genetic alphabet expansion system. J. Am. Chem. Soc. 2018, 140, 14038–14041. [Google Scholar] [CrossRef]
- Okamoto, I.; Miyatake, Y.; Kimoto, M.; Hirao, I. High fidelity, efficiency and functionalization of ds–px unnatural base pairs in pcr amplification for a genetic alphabet expansion system. ACS Synth. Biol. 2016, 5, 1220–1230. [Google Scholar] [CrossRef]
- Malyshev, D.A.; Dhami, K.; Lavergne, T.; Chen, T.; Dai, N.; Foster, J.M.; Corrêa, I.R.; Romesberg, F.E. A semi-synthetic organism with an expanded genetic alphabet. Nature 2014, 509, 385–388. [Google Scholar] [CrossRef]
- Zhang, Y.; Ptacin, J.L.; Fischer, E.C.; Aerni, H.R.; Caffaro, C.E.; San Jose, K.; Feldman, A.W.; Turner, C.R.; Romesberg, F.E. A semi-synthetic organism that stores and retrieves increased genetic information. Nature 2017, 551, 644–647. [Google Scholar] [CrossRef]
- Zhang, Y.; Lamb, B.M.; Feldman, A.W.; Zhou, A.X.; Lavergne, T.; Li, L.; Romesberg, F.E. A semisynthetic organism engineered for the stable expansion of the genetic alphabet. Proc. Natl. Acad. Sci. USA 2017, 114, 1317–1322. [Google Scholar] [CrossRef]
- Betz, K.; Malyshev, D.A.; Lavergne, T.; Welte, W.; Diederichs, K.; Dwyer, T.J.; Ordoukhanian, P.; Romesberg, F.E.; Marx, A. Klentaq polymerase replicates unnatural base pairs by inducing a watson-crick geometry. Nat. Chem. Biol. 2012, 8, 612–614. [Google Scholar] [CrossRef]
- Malyshev, D.A.; Pfaff, D.A.; Ippoliti, S.I.; Hwang, G.T.; Dwyer, T.J.; Romesberg, F.E. Solution structure, mechanism of replication, and optimization of an unnatural base pair. Chem. Eur. J. 2010, 16, 12650–12659. [Google Scholar] [CrossRef]
- Hoshika, S.; Leal, N.A.; Kim, M.-J.; Kim, M.-S.; Karalkar, N.B.; Kim, H.-J.; Bates, A.M.; Watkins, N.E.; SantaLucia, H.A.; Meyer, A.J.; et al. Hachimoji DNA and rna: A genetic system with eight building blocks. Science 2019, 363, 884–887. [Google Scholar] [CrossRef]
- Kawakami, T.; Murakami, H.; Suga, H. Exploration of incorporation of nα-methylated amino acids into peptides by sense-suppression method. Nucleic Acids Symp. Ser. 2007, 51, 361–362. [Google Scholar] [CrossRef]
- Fujino, T.; Goto, Y.; Suga, H.; Murakami, H. Ribosomal synthesis of peptides with multiple β-amino acids. J. Am. Chem. Soc. 2016, 138, 1962–1969. [Google Scholar] [CrossRef]
- Dedkova, L.M.; Fahmi, N.E.; Golovine, S.Y.; Hecht, S.M. Construction of modified ribosomes for incorporation of d-amino acids into proteins. Biochemistry 2006, 45, 15541–15551. [Google Scholar] [CrossRef]
- Dedkova, L.M.; Fahmi, N.E.; Paul, R.; del Rosario, M.; Zhang, L.; Chen, S.; Feder, G.; Hecht, S.M. B-puromycin selection of modified ribosomes for in vitro incorporation of β-amino acids. Biochemistry 2012, 51, 401–415. [Google Scholar] [CrossRef] [PubMed]
- Hui, A.; de Boer, H.A. Specialized ribosome system: Preferential translation of a single mrna species by a subpopulation of mutated ribosomes in escherichia coli. Proc. Natl. Acad. Sci. USA 1987, 84, 4762–4766. [Google Scholar] [CrossRef] [PubMed]
- Rackham, O.; Chin, J.W. A network of orthogonal ribosome·mrna pairs. Nat. Chem. Biol. 2005, 1, 159–166. [Google Scholar] [CrossRef] [PubMed]
- Neumann, H.; Wang, K.; Davis, L.; Garcia-Alai, M.; Chin, J.W. Encoding multiple unnatural amino acids via evolution of a quadruplet-decoding ribosome. Nature 2010, 464, 441–444. [Google Scholar] [CrossRef] [PubMed]
- Orelle, C.; Carlson, E.D.; Szal, T.; Florin, T.; Jewett, M.C.; Mankin, A.S. Protein synthesis by ribosomes with tethered subunits. Nature 2015, 524, 119–124. [Google Scholar] [CrossRef] [PubMed]
- Schmied, W.H.; Tnimov, Z.; Uttamapinant, C.; Rae, C.D.; Fried, S.D.; Chin, J.W. Controlling orthogonal ribosome subunit interactions enables evolution of new function. Nature 2018, 564, 444–448. [Google Scholar] [CrossRef] [PubMed]
- Aleksashin, N.A.; Leppik, M.; Hockenberry, A.J.; Klepacki, D.; Vázquez-Laslop, N.; Jewett, M.C.; Remme, J.; Mankin, A.S. Assembly and functionality of the ribosome with tethered subunits. Nat. Commun. 2019, 10, 930. [Google Scholar] [CrossRef] [PubMed]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Arranz-Gibert, P.; Patel, J.R.; Isaacs, F.J. The Role of Orthogonality in Genetic Code Expansion. Life 2019, 9, 58. https://doi.org/10.3390/life9030058
Arranz-Gibert P, Patel JR, Isaacs FJ. The Role of Orthogonality in Genetic Code Expansion. Life. 2019; 9(3):58. https://doi.org/10.3390/life9030058
Chicago/Turabian StyleArranz-Gibert, Pol, Jaymin R. Patel, and Farren J. Isaacs. 2019. "The Role of Orthogonality in Genetic Code Expansion" Life 9, no. 3: 58. https://doi.org/10.3390/life9030058
APA StyleArranz-Gibert, P., Patel, J. R., & Isaacs, F. J. (2019). The Role of Orthogonality in Genetic Code Expansion. Life, 9(3), 58. https://doi.org/10.3390/life9030058




