Emergence of a “Cyclosome” in a Primitive Network Capable of Building “Infinite” Proteins †
Abstract
1. Introduction
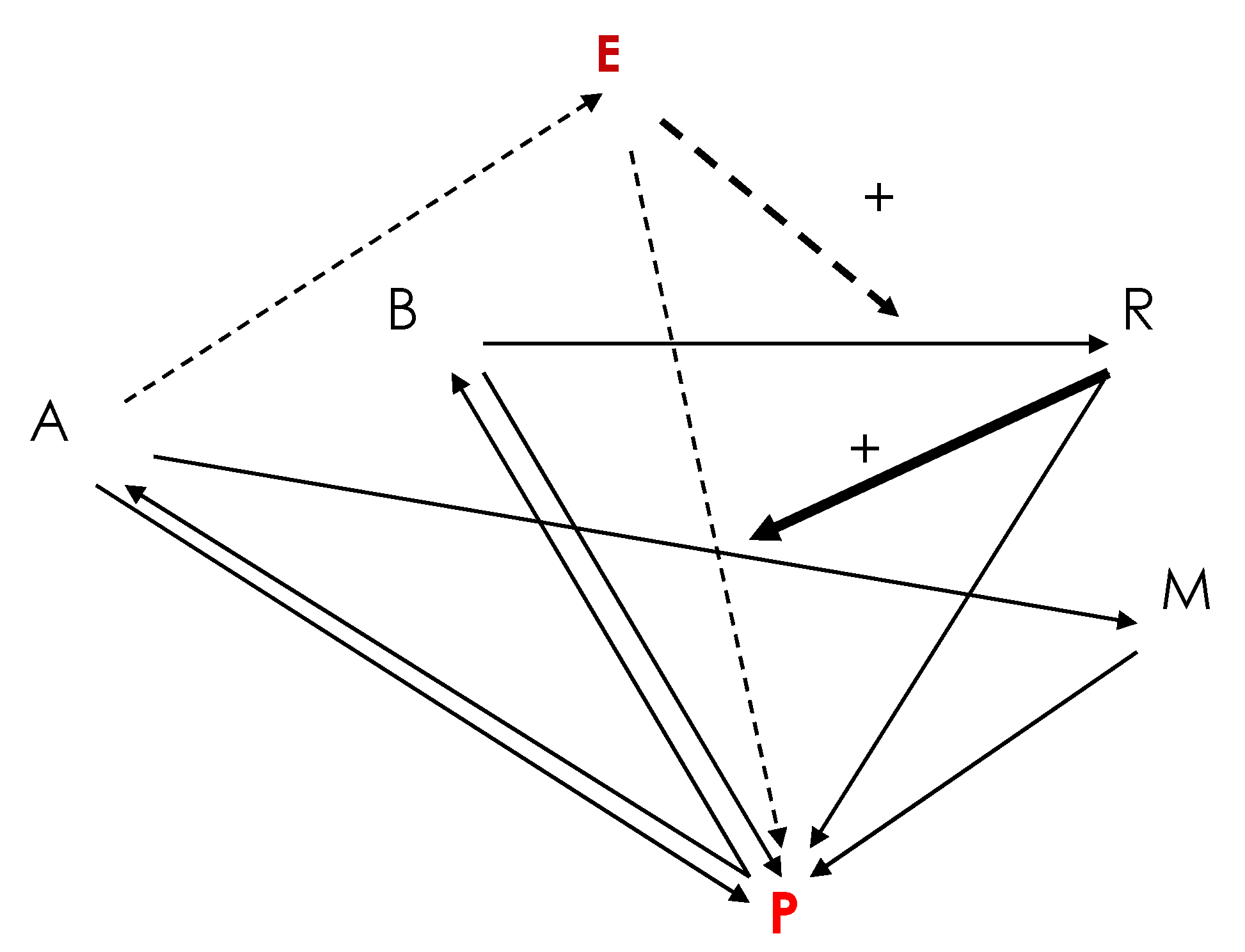
2. A Primitive Network at the Origin of Life
dA/dt = dA∆A + kPP − kARA − k’ERA
dB/dt = dB∆B + k’PP − kBB
dE/dt = dE∆E + k’ERA − kEE
dM/dt = dM∆M + kARA − kMM
dP/dt = dP∆P + kRR + kMM − k’PP − kPP
| −kR | 0 | kB | 0 | 0 |
| −K’kR/K | −K(kA+k’E)/kE | 0 | 0 | 0 |
| 0 | 0 | −kB | 0 | 0 |
| K’kRk’E/K(kA+k’E) | Kk’E/kR | 0 | −kE | |
| K’kRkA/K(kA+k’E) | KkA/kR | 0 | 0 | −kM |
3. Construction of the AL RNA Ring
- All dinucleotides should appear at least once (apart from CG because of CG suppression).
- Among rings satisfying the constraint “to be as short as possible and contain at least one codon of each amino acid synonymy class”, there is no solution for a length below 22 nucleotides. For length 22, 29,520 solutions contain the codon AUN twice, N being G for 52% of the solutions.
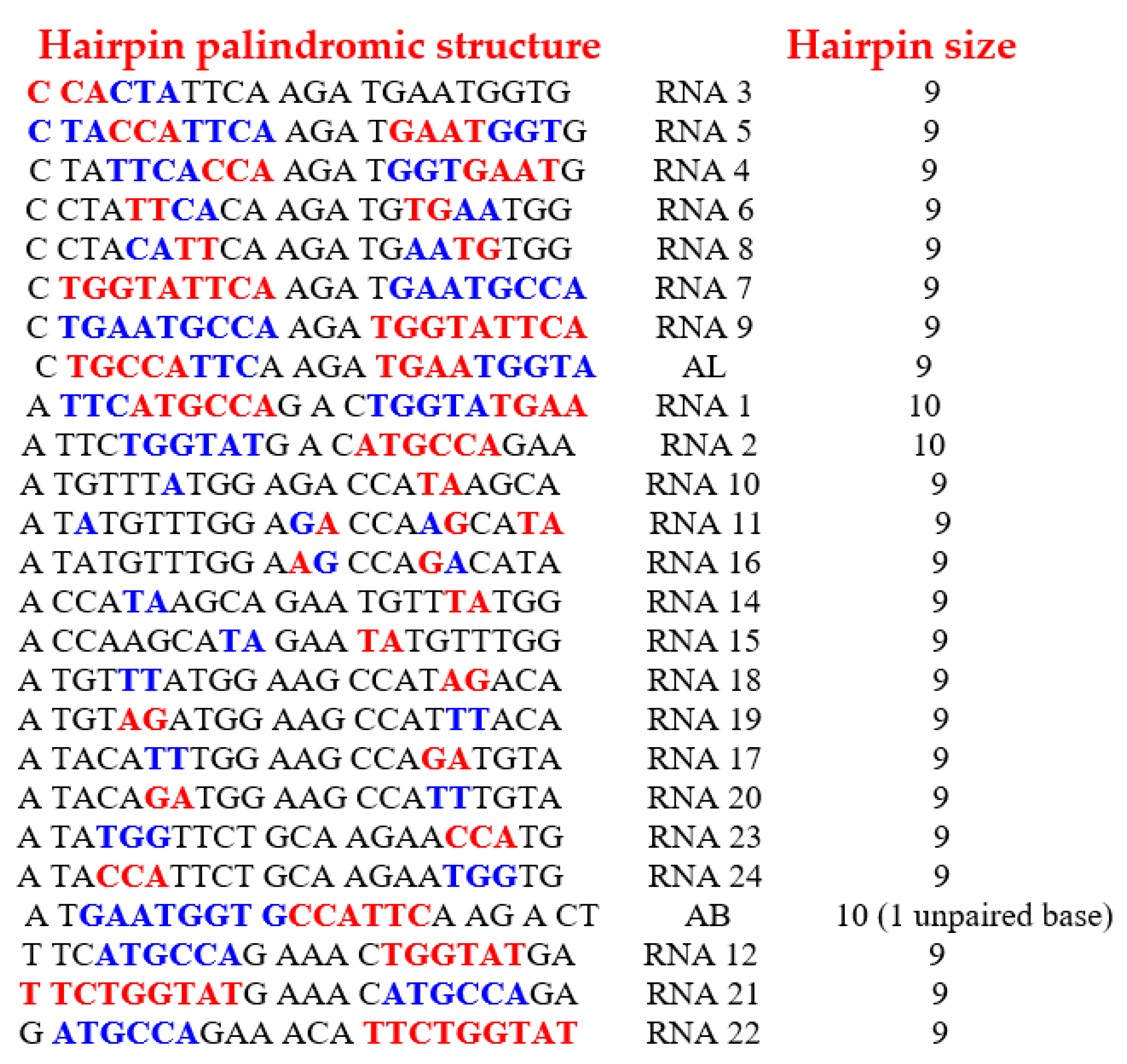
- From the 29,520 solutions, only 25 rings allow the formation of a hairpin at least 9-bases long.
- Of these 25 rings, 19 have both start and stop codons.
- Through calculation of the average genetic distances to the others (e.g., circular Hamming distance, permutation distance, and edit distance), one singular ring exhibits a minimum distance as compared to the others. Only one sequence, called AL (for ALpha) is thus acting as the barycenter of the set of the 18 others: 5′-AUGGUACUGCCAUUCAAGAUGA-3′.
4. Nucleo-Nucleic and Nucleo-Peptidic Mechanisms
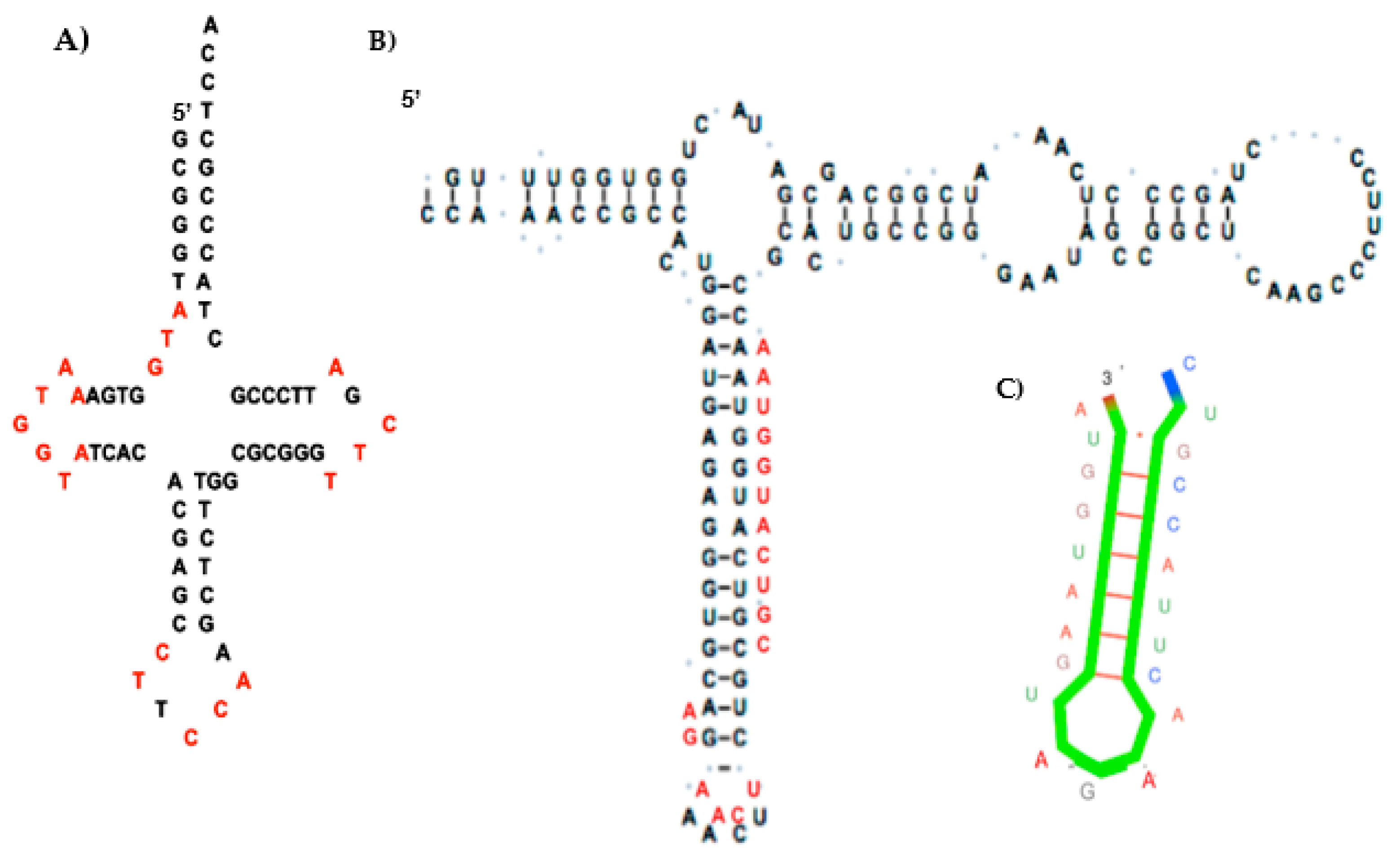
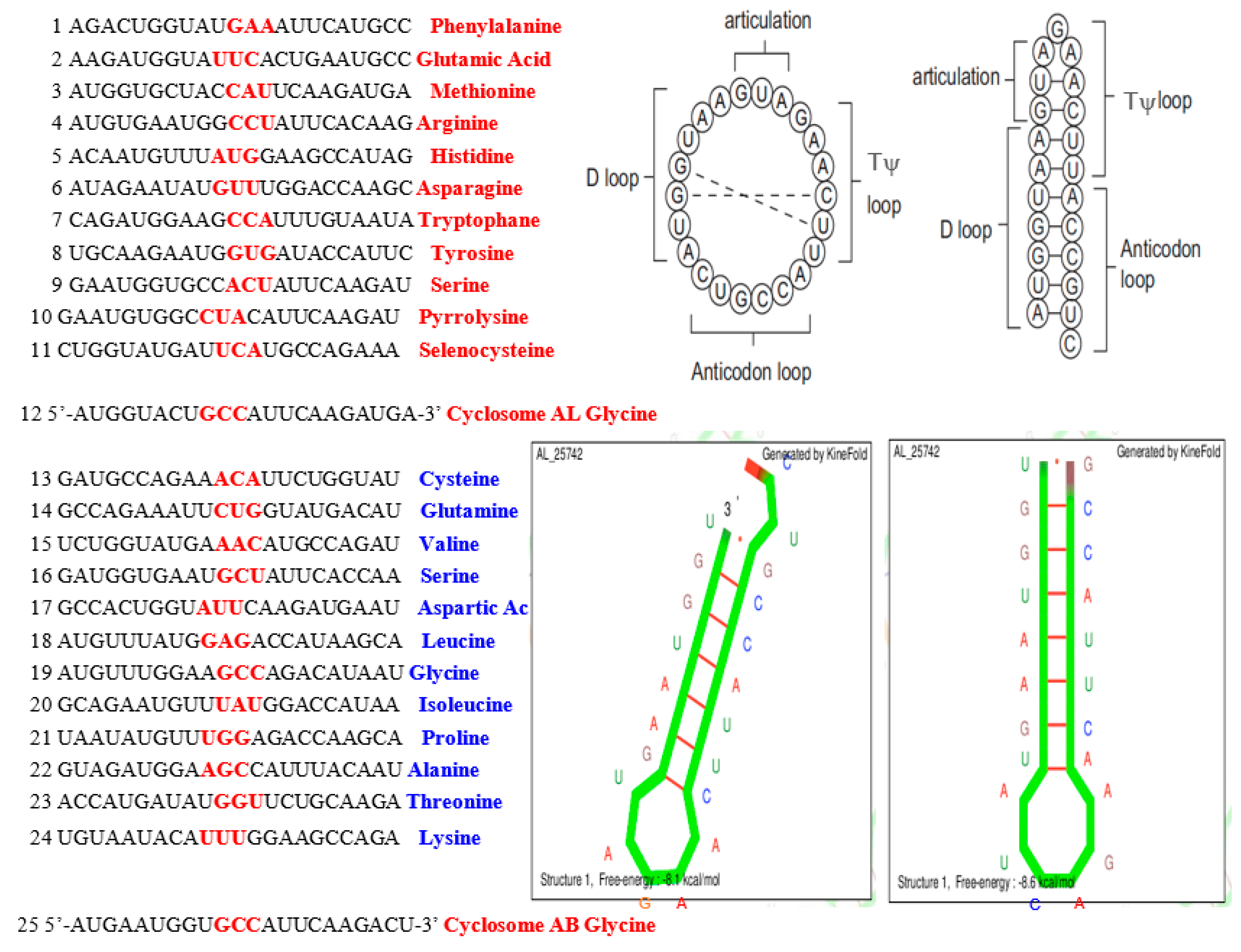
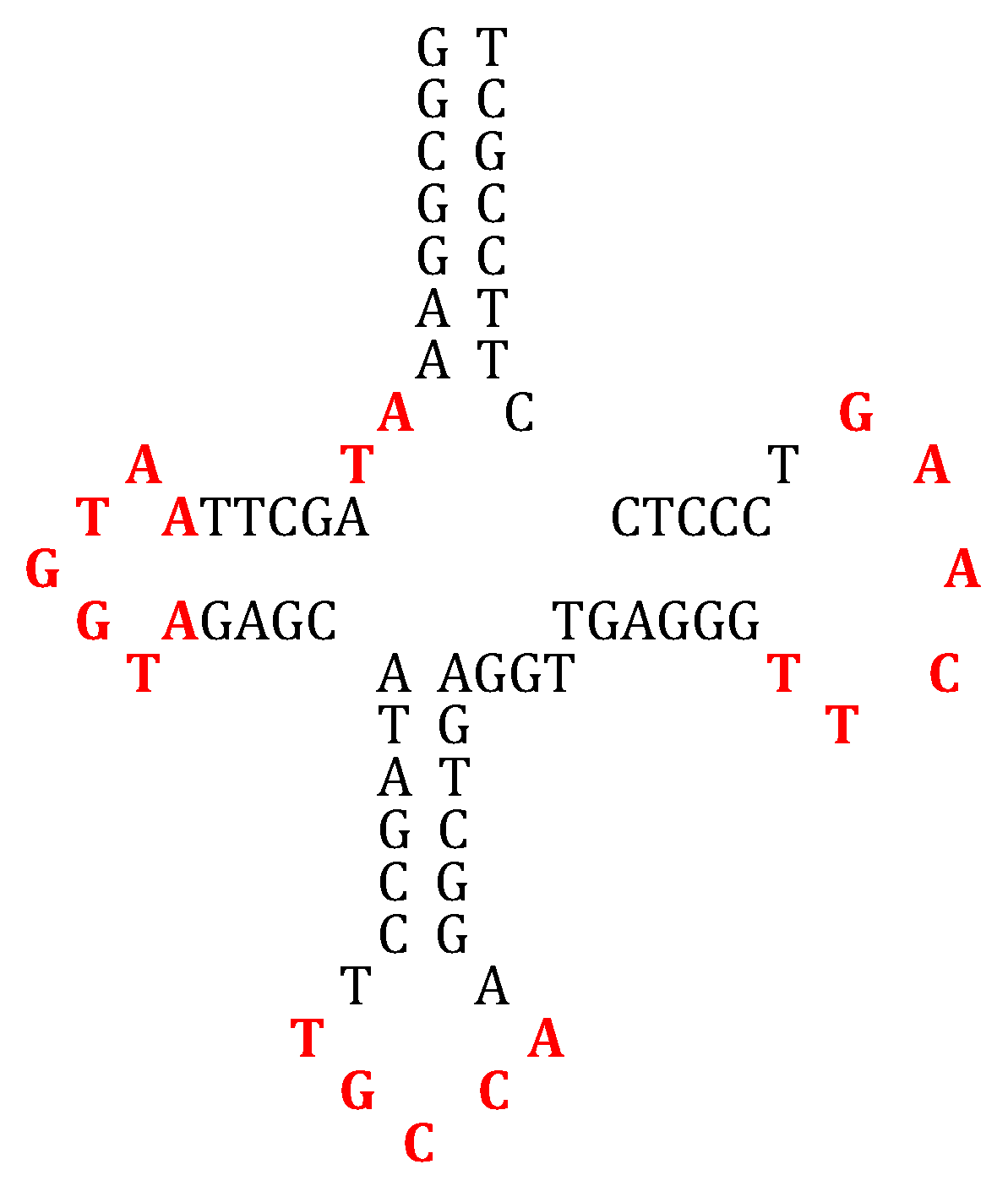
4.1. tRNA Loops
4.2. Giant Viruses
4.3. Circular RNAs
4.4. Ribozymes
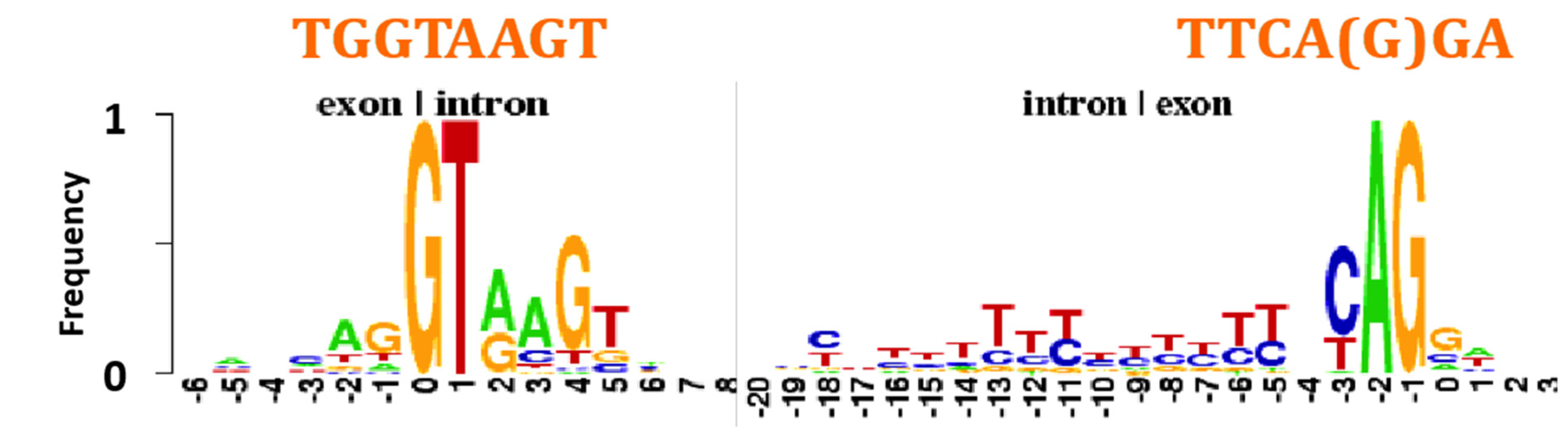
4.5. Intron-Exon Frontier
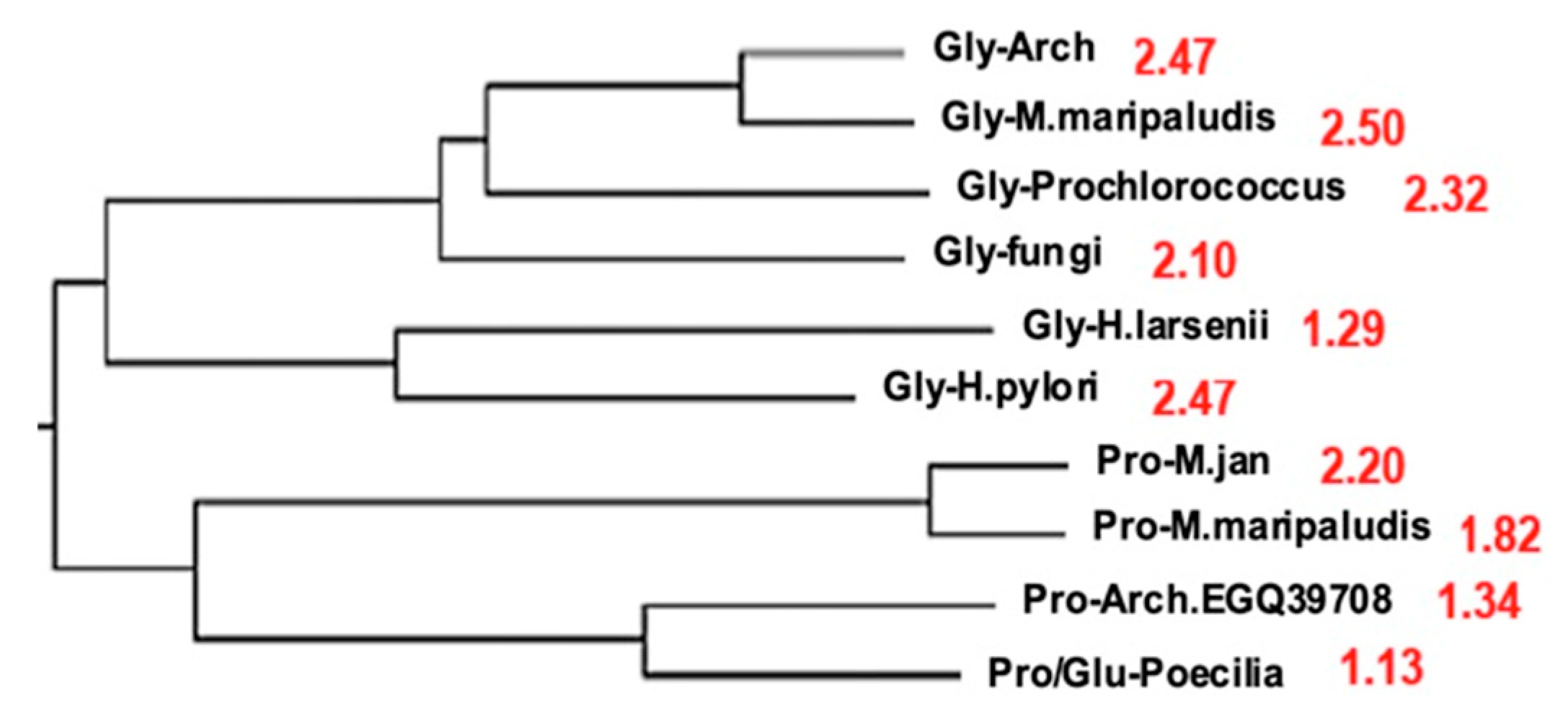
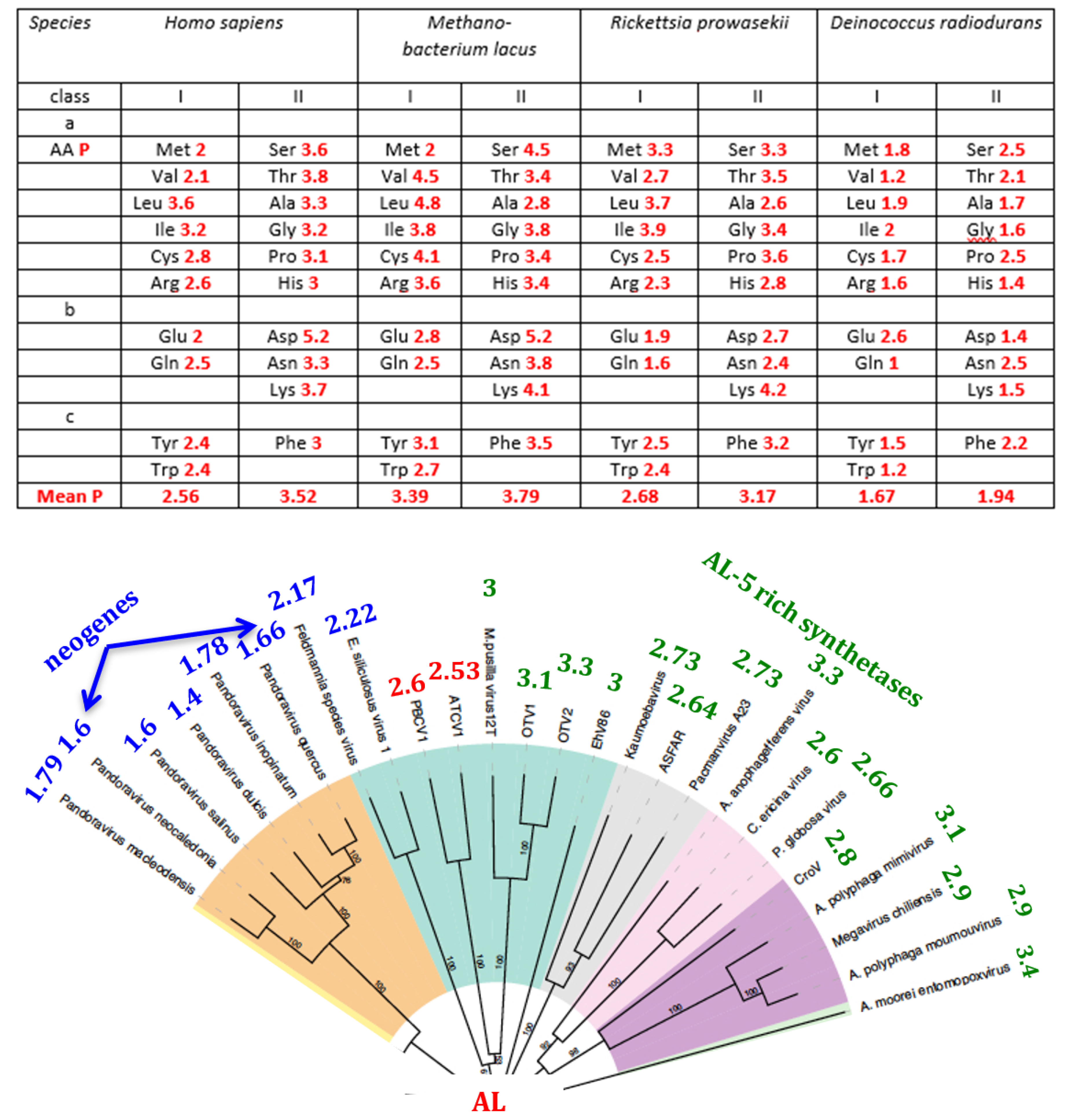
4.6. Synthetases
4.7. Small Acid-Soluble Spore Proteins (SASPs)
4.8. Defence Mechanisms
- p = 2.10−9 for 19 matches (with an insertion) between TIR and CRISPR using the binomial distribution B(1/4,22), p = 8.10−6 for 15 anti-matches between AL and CRISPR plus 1 quasi-anti-match G-T using the distribution B(1/4,21) × B(3/8,1),
- p = 7.10−4 for 13 matches between AL and consensus RSS using the binomial distribution B(1/4,22),
- p = 2.10−6 for 11 matches between AL and RSS spacer using the binomial distribution B(1/4,12).
- -
- Acanthamoeba castellanii mimivirus DNA, nearly complete genome, strain: Mimivirus kasaii GenBank: AP017644.1 457483-459936 R 350 Lambda-type exonuclease, with AL-proximity 3.02;
- -
- Acanthamoeba castellanii mimivirus DNA, nearly complete genome, strain: Mimivirus kasaii GenBank: AP017644.1 462878-464527 R 354 Lambda-type exonuclease, with AL-proximity 3.71.
4.9. Mitochondrial D-loop
4.10. 5S Ribosomal RNAs
4.11. Cytidine Deaminases
5. Discussion
5.1. Origins of the AL Ring
5.2. The AL-Pentamer Proximity as a Marker of Age of the Genome
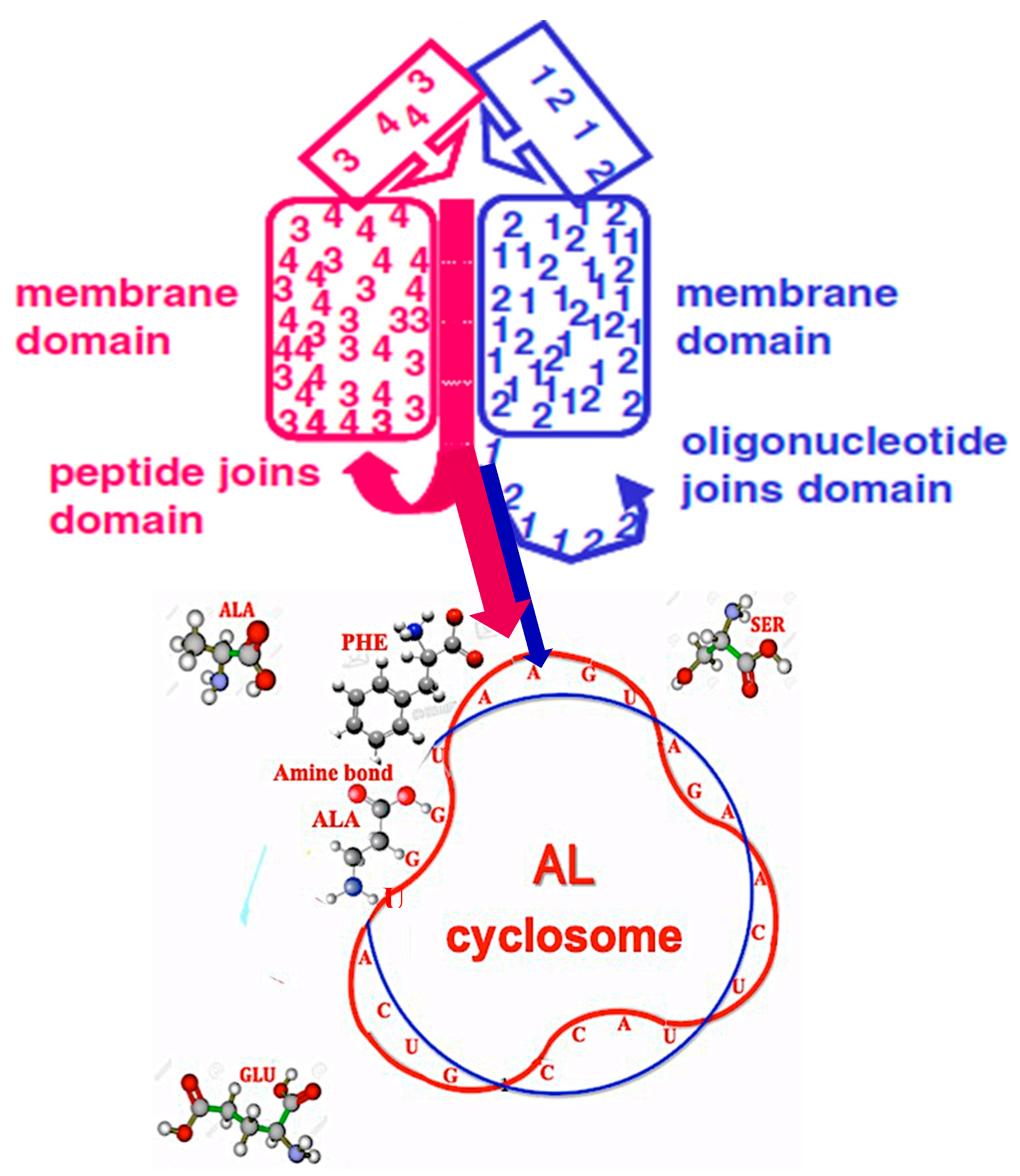
5.3. ‘Infinite’ Proteins
5.4. tRNA Building
6. Conclusions
- -
- Its subsequences (namely, pentamers) are observed as relics in many parts of modern genomes, especially in Archaea;
- -
- AL relics are often present in tRNA loops, and in mitochondrial D-loops;
- -
- An AL-heptamer constitutes the major part of the exon/intron boundary;
- -
- A scalar proximity to AL explains the relationships between polymerases and, more generally, between complete genomes in phylogenetic trees of Archaea. This proximity suggests a common origin for these genomes.
- -
- -
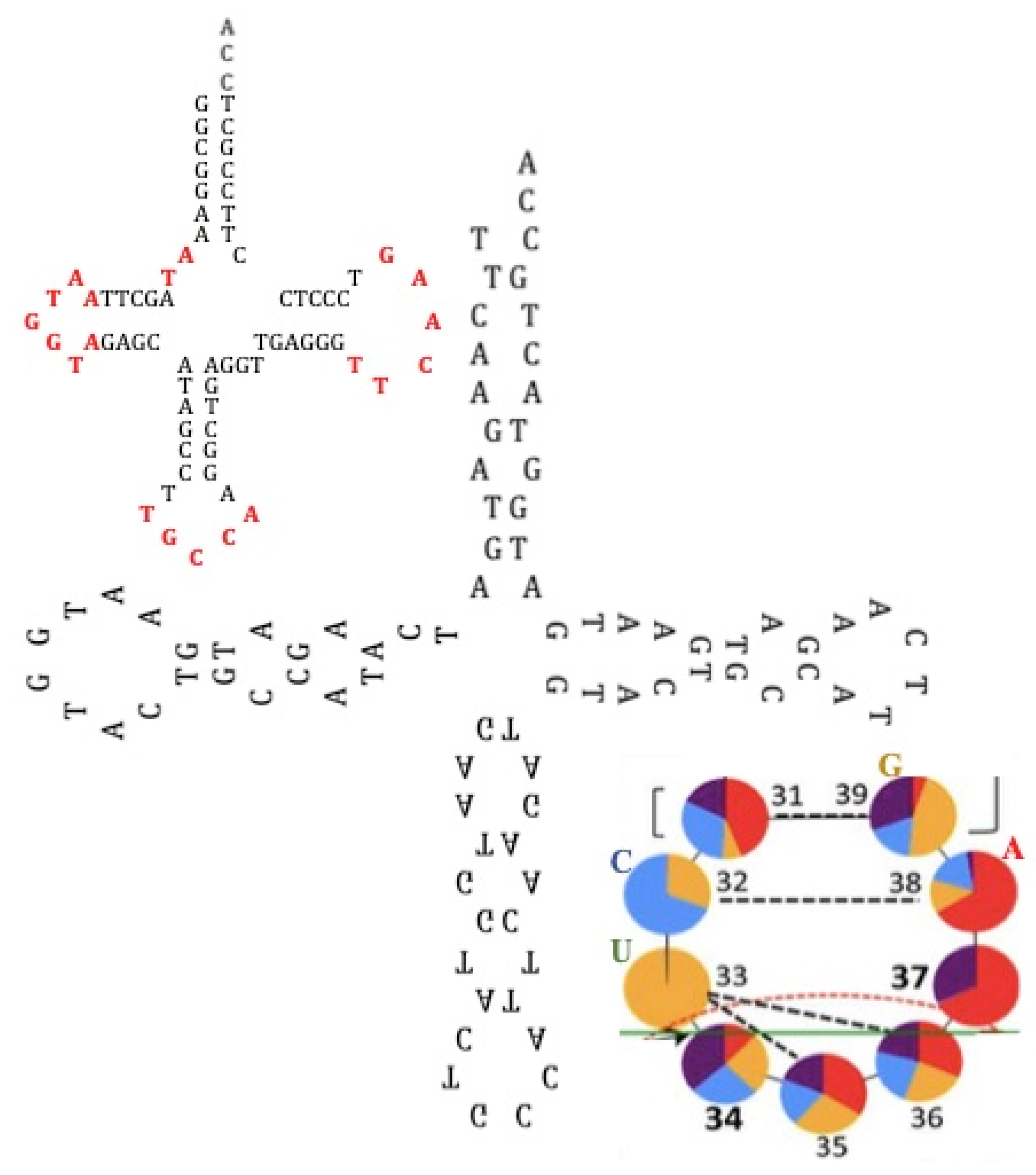
- A search for more AL relics in modern genomes at critical functional steps of the nuclear transcription/translation processes (notably when they are coupled as in Archaea [121], in which the Archaea tRNAGly presents the following sequence in its three successive loops: TGGTA CTGCCA TTCAA, that is a 16-mer from AL [122]), mitochondrial energetic or cellular immune receptor machineries);
- -
- -
- -
- The discovery of sequences linked to AL useful for synthetic biology and studies on “minimal cell” and its primitive genome, with original stable structures as those observed in the “cyclosome” (Figure 12);
- -
- The identification of genetic networks based on common sequences inherited from AL and appearing in regulatory RNAs like microRNAs or circular RNAs.
Supplementary Materials
Author Contributions
Acknowledgments
Conflicts of Interest
References
- Palade, G.E.; Siekevitz, P. Liver microsomes. An integrated morphological and biochemical study. J. Biophys. Biochem. Cytol. 1956, 2, 171–214. [Google Scholar] [CrossRef] [PubMed]
- Wolfram, S. A New Kind of Science; Wolfram Media, Inc.: Champaign, IL, USA, 2002. [Google Scholar]
- Gardner, M. Mathematical Games: The Fantastic Combinations of John Conway’s New Solitaire Game ‘Life’. Sci. Am. 1970, 22, 120–123. [Google Scholar] [CrossRef]
- Besson, J.; Gavaudan, P.; Schützenberger, M.P. Sur l’existence d’une certaine corrélation entre le poids moléculaire des acides aminés et le nombre de triplets intervenant dans leurs codages. C. R. Acad. Sci. 1969, 268, 1342–1344. [Google Scholar]
- Demongeot, J.; Hazgui, H. The Poitiers school of mathematical and theoretical biology: Besson-Gavaudan-Schützenberger’s conjectures on genetic code and RNA structures. Acta Biotheor. 2016, 64, 403–426. [Google Scholar] [CrossRef] [PubMed]
- Bloch, D.; McArthur, B.; Widdowson, R.; Spector, D.; Guimaraes, R.C.; Smith, J. tRNA-rRNA sequence homologies: A model for the origin of a common ancestral molecule, and prospects for its reconstruction. Orig. Life 1984, 14, 571–578. [Google Scholar] [CrossRef]
- Altman, S. Enzymatic cleavage of RNA by RNA. Biosci. Rep. 1990, 10, 317–337. [Google Scholar] [CrossRef] [PubMed]
- Cech, T.R. Self-splicing and enzymatic activity of an intervening sequence RNA from Tetrahymena. Biosci. Rep. 1990, 10, 239–261. [Google Scholar] [CrossRef]
- Agmon, I.; Auerbach, T.; Baram, D.; Bartels, H.; Bashan, A.; Berisio, R.; Fucini, P.; Hansen, H.; Harms, J.; Kessler, M.; et al. On peptide bond formation, translocation, nascent protein progression and the regulatory properties of ribosomes. Eur. J. Biochem. 2003, 270, 2543–2556. [Google Scholar] [CrossRef]
- Tamura, K.; Schimmel, P. Peptide synthesis with a template-like RNA guide and aminoacyl phosphate adaptors. Proc. Natl. Acad. Sci. USA 2003, 100, 8666–8669. [Google Scholar] [CrossRef]
- O’Donoghue, P.; Luthey-Schulten, Z. On the Evolution of Structure in Aminoacyl-tRNA Synthetases. Microbiol. Mol. Biol. Rev. 2003, 67, 550–573. [Google Scholar] [CrossRef]
- Agmon, I.; Bashan, A.; Zarivach, R.; Yonath, A. Symmetry at the active site of the ribosome: Structural and functional implications. Biol. Chem. 2005, 386, 833–844. [Google Scholar] [CrossRef] [PubMed]
- Tamura, K.; Schimmel, P.R. Chiral-selective aminoacylation of an RNA minihelix: Mechanistic features and chiral suppression. Proc. Natl. Acad. Sci. USA 2006, 103, 13750–13752. [Google Scholar] [CrossRef] [PubMed]
- Demongeot, J.; Glade, N.; Moreira, A.; Vial, L. RNA relics and origin of life. Int. J. Mol. Sci. 2009, 10, 3420–3441. [Google Scholar] [CrossRef] [PubMed]
- Demongeot, J.; Ben Amor, H.; Gillois, P.; Noual, M.; Sené, S. Robustness of regulatory networks. A Generic Approach with Applications at Different Levels: Physiologic, Metabolic and Genetic. Int. J. Mol. Sci. 2009, 10, 4437–4473. [Google Scholar] [CrossRef] [PubMed]
- Agmon, I. The Dimeric Proto-Ribosome: Structural Details and Possible Implications on the Origin of Life. Int. J. Mol. Sci. 2009, 10, 2921–2934. [Google Scholar] [CrossRef]
- Davidovich, C.; Belousoff, M.; Bashan, A.; Yonath, A. The evolving ribosome: From non-coded peptide bond formation to sophisticated translation machinery. Res. Microbiol. 2009, 160, 487–492. [Google Scholar] [CrossRef] [PubMed]
- Bokov, K.; Steinberg, S.V. A hierarchical model for evolution of 23S ribosomal RNA. Nature 2009, 457, 977–980. [Google Scholar] [CrossRef]
- Szostak, J.W. Origins of life: Systems chemistry on early Earth. Nature 2009, 459, 171. [Google Scholar] [CrossRef]
- Bashan, A.; Agmon, I.; Raz Zarivach, R.; Schluenzen, F.; Harms, J.; Berisio, R.; Bartels, H.; Franceschi, F.; Auerbach, T.; Hansen, H.A.S.; et al. Structural Basis of the Ribosomal Machinery for Peptide Bond Formation, Translocation, and Nascent Chain Progression. Mol. Cell 2013, 11, 91–102. [Google Scholar] [CrossRef]
- Huang, L.; Krupkin, M.; Bashan, A.; Yonath, A.; Massa, L. Protoribosome by quantum kernel energy method. Proc. Natl. Acad. Sci. USA 2013, 110, 14900–14905. [Google Scholar] [CrossRef]
- Bernhardt, H.S.; Tate, W.P. A Ribosome Without RNA. Front. Ecol. Evol. 2015, 3, 129. [Google Scholar] [CrossRef]
- Agmon, I. Could a Emerge Spontaneously in the Prebiotic World? Molecules 2016, 21, 1701. [Google Scholar] [CrossRef] [PubMed]
- Krupkin, M.; Wekselman, I.; Matzov, D.; Eyal, Z.; Diskin Posner, Y.; Rozenberg, H.; Zimmerman, E.; Bashan, A.; Yonath, A. Avilamycin and evernimicin induce structural changes in rProteins uL16 and CTC that enhance the inhibition of A-site tRNA binding. Proc. Natl. Acad. Sci. USA 2016, 113, 6796–6805. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.; Kowiatek, B.; Opron, K.; Burton, Z.F. Type-II tRNAs and Evolution of Translation Systems and the Genetic Code. Int. J. Mol. Sci. 2018, 19, 3275. [Google Scholar] [CrossRef] [PubMed]
- Norris, V. Why do bacteria divide? Front Microbiol. 2015, 6, 322. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Maturana, H.R.; Varela, F.J. Autopoiesis and Cognition: The Realization of the Living; Reidel: Boston, MA, USA, 1980. [Google Scholar]
- Bourgine, P.; Stewart, J. Autopoiesis and cognition. Artif. Life 2004, 10, 327–345. [Google Scholar] [CrossRef] [PubMed]
- Hunding, A.; Kepes, F.; Lancet, D.; Minsky, A.; Norris, V.; Raine, D.; Sriram, K.; Root-Bernstein, R. Compositional complementarity and prebiotic ecology in the origin of life. BioEssays 2006, 28, 399–412. [Google Scholar] [CrossRef] [PubMed]
- Robinson, R. Jump-starting a cellular world: Investigating the origin of life, from soup to networks. PLoS Biol. 2005, 3, e396. [Google Scholar] [CrossRef] [PubMed]
- Ono, N.; Ikegami, T. Self-maintenance and self-reproduction in an abstract cell model. J. Theor. Biol. 2000, 206, 243–253. [Google Scholar] [CrossRef]
- Ono, N.; Ikegami, T. Artificial chemistry: Computational studies on the emergence of selfreproducing units. In Proceedings of the 6th European Conference on Artificial Life (ECAL’01), Prague, Czech Republic, 10–14 September 2001; Kelemen, J., Sosik, S., Eds.; Springer: Berlin, Germany, 2001; pp. 186–195. [Google Scholar]
- Weil, G.; Heus, K.; Faraut, T.; Demongeot, J. An archetypal basic code for the primitive genome. Theor. Comput. Sci. 2004, 322, 313–334. [Google Scholar] [CrossRef]
- Demongeot, J.; Moreira, A. A circular RNA at the origin of life. J. Theor. Biol. 2007, 249, 314–324. [Google Scholar] [CrossRef] [PubMed]
- 5S RNAdb. Available online: http://www.combio.pl/rrna/alignment/ (accessed on 25 March 2019).
- Kinefold. Available online: http://kinefold.curie.fr (accessed on 25 March 2019).
- Fonville, N.C.; Velmurugan, K.R.; Tae, H.; Vaksman, Z.; McIver, L.J.; Garner, H.R. Genomic leftovers: Identifying novel microsatellites, over-represented motifs and functional elements in the human genome. Sci. Rep. 2016, 6, 27722. [Google Scholar] [CrossRef] [PubMed]
- Tóth, G.; Gáspári, Z.; Jurka, J. Microsatellites in different eukaryotic genomes: Survey and analysis. Genome Res. 2000, 10, 967–981. [Google Scholar] [CrossRef] [PubMed]
- Pemberton, T.J.; Sandefur, C.I.; Jakobsson, M.; Rosenberg, N.A. Sequence determinants of human microsatellite variability. BMC Genom. 2009, 10, 612. [Google Scholar] [CrossRef] [PubMed]
- Willard, H.F.; Waye, J.S. Chromosome-specific Subsets of Human Alpha Satellite DNA: Analysis of Sequence Divergence Within and Between Chromosomal Subsets and Evidence for an Ancestral Pentameric Repeat. J. Mol. Evol. 1987, 25, 207–214. [Google Scholar] [CrossRef]
- Zhuma, T.; Tyrrell, R.; Sekkali, B.; Skavdis, G.; Saveliev, A.; Tolaini, M.; Roderick, K.; Norton, T.; Smerdon, S.; Sedgwick, S.; et al. Human HMG box transcription factor HBP1: A role in hCD2 LCR function. EMBO J. 1999, 18, 6396–6406. [Google Scholar] [CrossRef]
- Presnyak, V.; Alhusaini, N.; Chen, Y.H.; Martin, S.; Morris, N.; Kline, N.; Olson, S.; Weinberg, D.; Baker, K.E.; Graveley, B.R.; et al. Codon optimality is a major determinant of mRNA stability. Cell 2015, 160, 1111–1124. [Google Scholar] [CrossRef]
- Hobish, M.K.; Wickramasinghe, N.S.; Ponnamperuma, C. Direct interaction between amino acids and nucleotides as a possible physicochemical basis for the origin of the genetic code. Adv. Space Res. 1995, 15, 365–382. [Google Scholar] [CrossRef]
- Yarus, M.; Widmann, J.J.; Knight, R. RNA-amino acid binding: A stereo chemical era for the genetic code. J. Mol. Evol. 2009, 69, 406–429. [Google Scholar] [CrossRef]
- Yarus, M. The Genetic Code and RNA-Amino Acid Affinities. Life 2017, 7, 13. [Google Scholar] [CrossRef]
- Polyansky, A.A.; Zagrovic, B. Evidence of direct complementary interactions between messenger RNAs and their cognate proteins. Nucleic Acids Res. 2013, 41, 8434–8443. [Google Scholar] [CrossRef] [PubMed]
- Zagrovic, B.; Bartonek, L.; Polyansky, A.A. RNA-protein interactions in an unstructured context. FEBS Lett. 2018, 592, 2901–2916. [Google Scholar] [CrossRef] [PubMed]
- Demongeot, J. Sur la possibilité de considérer le code génétique comme un code à enchaînement. Rev. De Biomaths 1978, 62, 61–66. [Google Scholar]
- Demongeot, J.; Besson, J. Code génétique et codes à enchaînement I. C. R. Acad. Sci. Série III 1983, 296, 807–810. [Google Scholar]
- Demongeot, J.; Besson, J. Genetic code and cyclic codes II. C. R. Acad. Sc. Série III 1996, 319, 520–528. [Google Scholar]
- Demongeot, J.; Drouet, E.; Moreira, A.; Rechoum, Y.; Sené, S. Micro-RNAs: Viral genome and robustness of the genes expression in host. Philos. Trans. R. Soc. A 2009, 367, 4941–4965. [Google Scholar] [CrossRef] [PubMed]
- Demongeot, J.; Glade, N.; Moreira, A. Evolution and RNA relics. A Systems Biology view. Acta Biotheoretica 2008, 56, 5–25. [Google Scholar] [CrossRef] [PubMed]
- Demongeot, J.; Hazgui, H.; Bandiera, S.; Cohen, O.; Henrion-Caude, A. MitomiRs, ChloromiRs and general modelling of the microRNA inhibition. Acta Biotheor. 2013, 61, 367–383. [Google Scholar] [CrossRef] [PubMed]
- Demongeot, J.; Cohen, O.; Henrion-Caude, A. MicroRNAs and Robustness in Biological Regulatory Networks. A Generic Approach with Applications at Different Levels: Physiologic, Metabolic, and Genetic. In Systems Biology of Metabolic and Signaling Networks; Aon, M.A., Saks, V., Schlattner, U., Eds.; Springer: Berlin, Germany, 2013; pp. 63–114. [Google Scholar]
- GtRNAdb. Available online: http://lowelab.ucsc.edu/GtRNAdb/Rhod_spha_ATCC_17029/rhodSpha_ ATCC17029-tRNAs. fa (accessed on 25 March 2019).
- Müller, M.; Legrand, C.; Tuorto, F.; Kelly, V.P.; Atlasi, Y.; Lyko, F.; Ehrenhofer-Murray, A.E. Queuine links translational control in eukaryotes to micronutrient from bacteria. Nucl. Acids Res. 2019, 47, 3711–3727. [Google Scholar] [CrossRef] [PubMed]
- Choi, H.; Otten, S.; McClain, W.H. Isolation of novel tRNAAla mutants by library selection in a tRNAAla knockout strain. Biochimie 2002, 84, 705–711. [Google Scholar] [CrossRef]
- Agris, P.F.; Vendeix, F.A.P.; Graham, W.D. tRNA’s Wobble Decoding of the Genome: 40 Years of Modification. J. Mol. Biol. 2007, 366, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Weixlbaumer, A.; Murphy, F.V.; Dziergowska, A.; Malkiewicz, A.; Vendeix, F.A.P.; Agris, P.F.; Ramakrishnan, V. Mechanism for expanding the decoding capacity of transfer RNAs by modification of uridines. Nat. Struct. Mol. Biol. 2007, 14, 498–502. [Google Scholar] [CrossRef] [PubMed]
- Targanski, I.; Cherkasova, V. Analysis of genomic tRNA sets from Bacteria, Archaea, and Eukarya points to anticodon–codon hydrogen bonds as a major determinant of tRNA compositional variations. RNA 2008, 14, 1095–1109. [Google Scholar] [CrossRef] [PubMed]
- Fergus, C.; Barnes, D.; Alqasem, M.A.; Kelly, V.P. The Queuine Micronutrient: Charting a Course from Microbe to Man. Nutrients 2015, 7, 2897–2929. [Google Scholar] [CrossRef] [PubMed]
- Choi, H.; Gabriel, K.; Schneider, J.; Otten, S.; McClain, W.H. Recognition of acceptor-stem structure of tRNAAsp by Escherichia coli aspartyl-tRNA synthetase. RNA 2003, 9, 386–393. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Root-Bernstein, R. Simultaneous origin of homochirality, the genetic code and its directionality. BioEssays 2007, 29, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Root-Bernstein, R.; Root-Bernstein, M. The ribosome as a missing link in prebiotic evolution II: Ribosomes encode ribosomal proteins that bind to common regions of their own mRNAs and rRNAs. J. Biol. 2016, 397, 115–127. [Google Scholar] [CrossRef]
- Root-Bernstein, R.; Root-Bernstein, M. The Ribosome as a Missing Link in Prebiotic Evolution III: Over-Representation of tRNA- and rRNA-Like Sequences and Plieofunctionality of Ribosome-Related Molecules Argues for the Evolution of Primitive Genomes from Ribosomal RNA Modules. Int. J. Mol. Sci. 2019, 20, 140. [Google Scholar] [CrossRef]
- Bartnik, E.; Borsuk, P. A glycine tRNA gene from lupine mitochondria. Nucleic Acids Res. 1986, 14, 2407. [Google Scholar] [CrossRef]
- Seligmann, H.; Raoult, D. Stem-Loop RNA Hairpins in Giant Viruses: Invading rRNA-Like Repeats and a Template Free RNA. Front Microbiol. 2018, 9, 101. [Google Scholar] [CrossRef]
- Seligmann, H.; Raoult, D. Unifying view of stem–loop hairpin RNA as origin of current and ancient parasitic and non-parasitic RNAs, including in giant viruses. Curr. Opin. Microbiol. 2016, 31, 1–8. [Google Scholar] [CrossRef] [PubMed]
- NCBI. Available online: https://blast.ncbi.nlm.nih.gov/ (accessed on 25 March 2019).
- Circbase. Available online: http://www.circbase.org/ (accessed on 25 March 2019).
- Ivanov, A.; Memczak, S.; Wyler, E.; Torti, F.; Porath, H.T.; Orejuela, M.R.; Piechotta, M.; Levanon, E.Y.; Landthaler, M.; Dieterich, C.; et al. Analysis of intron sequences reveals hallmarks of circular RNA biogenesis in animals. Cell Rep. 2014, 10, 170–177. [Google Scholar] [CrossRef] [PubMed]
- Liang, T.; Yang, C.; Li, P.; Liu, C.; Guo, L. Genetic analysis of loop sequences in the let-7 gene family reveal a relationship between loop evolution and multiple isomiRs. PLoS ONE 2014, 9, e113042. [Google Scholar] [CrossRef] [PubMed]
- Hampel, A.; Tritz, R. RNA Catalytic Properties of the Minimum (-)sTRSV Sequence. Biochemistry 1989, 28, 4929–4933. [Google Scholar] [CrossRef] [PubMed]
- Salter, J.; Krucinska, J.; Alam, S.; Grum-Tokars, V.; Wedekind, J.E. Water in the Active Site of an All-RNA Hairpin Ribozyme and Effects of Gua8 Base Variants on the Geometry of Phosphoryl Transfer. Biochemistry 2006, 45, 686–700. [Google Scholar] [CrossRef]
- Pérez-Ruiz, M.; Barroso-delJesus, A.; Berzal-Herranz, A. Specificity of the Hairpin Ribozyme. J. Biol. Chem. 1999, 274, 29376–29380. [Google Scholar] [CrossRef] [PubMed]
- Müller, U.F. Design and Experimental Evolution of trans-Splicing Group I Intron Ribozymes. Molecules 2017, 22, 75. [Google Scholar] [CrossRef]
- Paul, N.; Joyce, G.F. A self-replicating ligase ribozyme. Proc. Natl. Acad. Sci. USA 2002, 99, 12733–12740. [Google Scholar] [CrossRef]
- Perreault, J.; Weinberg, Z.; Roth, A.; Popescu, O.; Chartrand, P.; Ferbeyre, G.; Breaker, R.R. Identification of Hammerhead Ribozymes in All Domains of Life Reveals Novel Structural Variations. PLoS Comput. Biol. 2011, 7, e1002031. [Google Scholar] [CrossRef]
- Hammann, C.; Luptak, A.; Perreault, J.; De La Peña, M. The ubiquitous hammerhead ribozyme. RNA 2012, 18, 871–885. [Google Scholar] [CrossRef]
- Harris, K.A.; Lünse, C.E.; Li, S.; Brewer, K.I.; Breaker, R.R. Biochemical analysis of hatchet self-cleaving ribozymes. RNA 2015, 21, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Rupert, P.B.; Ferré-D’Amaré, A.R. Crystal structure of a hairpin ribozyme-inhibitor complex with implications for catalysis. Nature 2001, 410, 780–786. [Google Scholar] [CrossRef] [PubMed]
- Gebetsberger, J.; Micura, R. Unwinding the twister ribozyme: From structure to mechanism. WIREs RNA 2017, 8, e1402. [Google Scholar] [CrossRef] [PubMed]
- Chapple, K.E.; Bartel, D.P.; Unrau, P.J. Combinatorial minimization and secondary structure determination of a nucleotide synthase ribozyme. RNA 2003, 9, 1208–1220. [Google Scholar] [CrossRef] [PubMed]
- Luisi, P.L. The Emergence of Life: From Chemical Origins to Synthetic Biology; Cambridge University Press: Cambridge, UK, 2006. [Google Scholar]
- Alberts, B.; Johnson, A.; Lewis, J.; Raff, M.; Roberts, K.; Walter, P. Molecular Biology of the Cell; Garland Science: New York, NY, USA, 2002. [Google Scholar]
- Hartman, H. Speculations on the evolution of the genetic code I. Orig. Life 1975, 6, 423–427. [Google Scholar] [CrossRef]
- Hartman, H. Speculations on the evolution of the genetic code II. Orig. Life 1978, 9, 133–136. [Google Scholar] [CrossRef] [PubMed]
- Hartman, H. Speculations on the evolution of the genetic code III: The evolution of t-RNA. Orig. Life 1984, 14, 643–648. [Google Scholar] [CrossRef]
- Hartman, H. Speculations on the origin of the genetic code IV. J. Mol. Evol. 1995, 40, 541–544. [Google Scholar] [CrossRef]
- Hartman, H.; Favaretto, P.; Smith, T.F. The archaeal origins of the eukaryotic translational system. Archaea 2006, 2, 1–9. [Google Scholar] [CrossRef]
- Smith, T.F.; Lee, J.C.; Gutell, R.R.; Hartman, H. The origin and evolution of the ribosome. Biol. Direct 2008, 3, 16. [Google Scholar] [CrossRef]
- Hartman, H.; Smith, T.F. The evolution of the ribosome and the genetic code. Life 2014, 4, 227–249. [Google Scholar] [CrossRef] [PubMed]
- Smith, T.F.; Hartman, H. The evolution of Class II Aminoacyl-tRNA synthetases and the first code. FEBS Lett. 2015, 589, 3499–3507. [Google Scholar] [CrossRef] [PubMed]
- Lier, C.; Baticle, E.; Horvath, P.; Haguenoer, E.; Valentin, A.S.; Glaser, P.; Mereghetti, L.; Lanotte, P. Analysis of the type II-A CRISPR-Cas system of Streptococcus agalactiae reveals distinctive features according to genetic lineages. Front. Genet. 2015, 6, 214. [Google Scholar] [CrossRef] [PubMed]
- Horvath, P.; Romero, D.A.; Coûté-Monvoisin, A.C.; Richards, M.; Deveau, H.; Moineau, S.; Boyaval, P.; Fremaux, C.; Barrangou, R. Diversity, Activity, and Evolution of CRISPR Loci in Streptococcus thermophilus. J. Bacteriol. 2008, 190, 1401–1412. [Google Scholar] [CrossRef] [PubMed]
- Koonin, E.V.; Krupovic, M. Evolution of adaptive immunity from transposable elements combined with innate immune systems. Nat. Rev. Genet. 2015, 16, 184–192. [Google Scholar] [CrossRef] [PubMed]
- Koonin, E.V.; Makarova, K.S. Mobile Genetic Elements and Evolution of CRISPR-Cas Systems: All the Way There and Back. Genome Biol. Evol. 2017, 9, 2812–2825. [Google Scholar] [CrossRef] [PubMed]
- Lee, A.I.; Fugmann, S.D.; Cowell, L.G.; Ptaszek, L.M.; Kelsoe, G.; Schatz, D.G. A Functional Analysis of the Spacer of V(D)J Recombination Signal Sequences. PLos Biol. 2003, 1, 56–69. [Google Scholar] [CrossRef] [PubMed]
- Pasqual, N.; Gallagher, M.; Aude-Garcia, C.; Loiodice, M.; Thuderoz, F.; Demongeot, J.; Ceredig, R.; Marche, P.N.; Jouvin-Marche, E. Quantitative and Qualitative Changes in ADV-AJ Rearrangements During Mouse Thymocytes Differentiation: Implication for a Limited TCR ALPHA Chain Repertoire. J. Exp. Med. 2002, 196, 1163–1174. [Google Scholar] [CrossRef] [PubMed]
- Thuderoz, F.; Simonet, M.A.; Hansen, O.; Dariz, A.; Baum, T.P.; Hierle, V.; Demongeot, J.; Marche, P.N.; Jouvin-Marche, E. Numerical Modelling of the V-J Combinations of the T Cell Receptor TRA/TRD Locus. PLoS Comp. Biol. 2010, 6, e1000682. [Google Scholar] [CrossRef] [PubMed]
- Ramsden, D.A.; Baetz, K.; Wu, G.E. Conservation of sequence in recombination signal sequence spacers. Nucl. Acids Res. 1994, 22, 1785–1796. [Google Scholar] [CrossRef]
- Takeuchi, N.; Ishiguro, N.; Shinagawa, M. Molecular cloning and sequence analysis of bovine T-cell receptor gamma and delta chain genes. Immunogenetics 1992, 35, 89–96. [Google Scholar] [CrossRef] [PubMed]
- Liu, M.F.; Robbins, D.L.; Crowley, J.J.; Sinha, S.; Kozin, F.; Kipps, T.J.; Carson, D.A.; Chen, P.J. Characterization of four homologous L chain variable region genes that are related to 6B6.6 idiotype positive human rheumatoid factor L chains. J. Immunol. 1989, 142, 688–694. [Google Scholar]
- Levasseur, A.; Bekliz, M.; Chabrière, E.; Pontarotti, P.; La Scola, B.; Didier Raoult, D. MIMIVIRE is a defence system in mimivirus that confers resistance to virophage. Nature 2016, 531, 249–252. [Google Scholar] [CrossRef] [PubMed]
- Portugene. Available online: http://fpereira.portugene.com/research1.html (accessed on 25 March 2019).
- Woese, C. The biological significance of the genetic code. Prog. Mol. Subcell. Biol. 1969, 1, 5–46. [Google Scholar]
- Conticello, S.C.; Thomas, C.J.F.; Petersen-Mahrt, S.K.; Neuberger, M.S. Evolution of the AID/APOBEC Family of Polynucleotide (Deoxy)cytidine Deaminases. Mol. Biol. Evol. 2005, 22, 367–377. [Google Scholar] [CrossRef]
- Martinez, T.; Shapiro, M.; Bhaduri-McIntosh, S.; MacCarthy, T. Evolutionary effects of the AID/APOBEC family of mutagenic enzymes on human gamma-herpesviruses. Virus Evol. 2019, 5, vey040. [Google Scholar] [CrossRef]
- Kraft, M.L.; Weber, P.K.; Longo, M.L.; Hutcheon, I.D.; Boxwer, S.G. Phase separation of lipid membranes analyzed with high-resolution secondary ion mass spectrometry. Science 2006, 313, 1948–1951. [Google Scholar] [CrossRef]
- Raine, D.J.; Norris, V. Lipid domain boundaries as prebiotic catalysts of peptide bond formation. J. Theor. Biol. 2007, 246, 176–185. [Google Scholar] [CrossRef]
- Legendre, M.; Fabre, E.; Poirot, O.; Jeudy, S.; Lartigue, A.; Alempic, J.M.; Beucher, L.; Philippe, N.; Bertaux, L.; Christo-Foroux, E.; et al. Diversity and evolution of the emerging Pandoraviridae family. Nat. Commun. 2018, 9, 2285. [Google Scholar] [CrossRef]
- Di Giulio, M. On the Origin of Protein Synthesis: A Speculative Model Based on Hairpin tRNA Structures. J. Theor. Biol. 1994, 171, 303–308. [Google Scholar] [CrossRef]
- Tamura, K. Origins and Early Evolution of the tRNA Molecule. Life 2015, 5, 1687–1699. [Google Scholar] [CrossRef] [PubMed]
- Grosjean, H.; Westhof, E. An integrated, structure- and energy-based view of the genetic code. Nucleic Acids Res. 2016, 44, 8020–8040. [Google Scholar] [CrossRef] [PubMed]
- Fontecilla-Camps, J.C. The Stereochemical Basis of the Genetic Code and the (Mostly) Autotrophic Origin of Life. Life 2014, 4, 1013–1025. [Google Scholar] [CrossRef] [PubMed]
- Paecht-Horowitz, M.; Katchalsky, A. Synthesis of amino acyl-adenylates under prebiotic conditions. J. Mol. Evol. 1973, 2, 91–98. [Google Scholar] [CrossRef] [PubMed]
- Meyer, S.C.; Nelson, P.A. Can the origin of the genetic code be explained by direct RNA templating? Bio-Complex. 2011, 2011, 1–10. [Google Scholar] [CrossRef]
- Lancet, D.; Zidovetzki, R.; Markovitch, O. Systems protobiology: Origin of life in lipid catalytic networks. J. R. Soc. Interface 2018, 15, 20180159. [Google Scholar] [CrossRef]
- Hsiao, C.; Mohan, S.; Kalahar, B.K.; Williams, L.D. Peeling the onion: Ribosomes are ancient molecular fossils. Mol. Biol. Evol. 2009, 26, 2415–2425. [Google Scholar] [CrossRef]
- Opron, K.; Burton, Z.F. Ribosome Structure, Function, and Early Evolution. Int. J. Mol. Sci. 2018, 19, 40. [Google Scholar] [CrossRef]
- French, S.L.; Santangelo, T.J.; Beyer, A.L.; Reeve, J.N. Transcription and translation are coupled in Archaea. Mol. Biol. Evol. 2007, 24, 893–895. [Google Scholar] [CrossRef]
- Pak, D.; Du, N.; Kim, Y.; Sun, Y.; Burton, Z.F. Rooted tRNAomes and evolution of the genetic code. Transcription 2018, 9, 137–151. [Google Scholar] [CrossRef]
- Seligmann, H. Giant viruses as protein-coated amoeban mitochondria? Virus Res. 2018, 253, 77–86. [Google Scholar] [CrossRef] [PubMed]
- Seligmann, H. Protein sequences recapitulate genetic code evolution. Comput. Struct. Biotechnol. J. 2018, 16, 177–189. [Google Scholar] [CrossRef]
- Seligmann, H. Alignment-based and alignment-free methods converge with experimental data on amino acids coded by stop codons at split between nuclear and mitochondrial genetic codes. Biosystems 2018, 167, 33–46. [Google Scholar] [CrossRef] [PubMed]
- Demongeot, J.; Seligmann, H. Theoretical minimal RNA rings recapitulate the order of the genetic code’s codon-amino acid assignments. J. Theor. Biol. 2019, 471, 108–116. [Google Scholar] [CrossRef]
- Demongeot, J.; Seligmann, H. Spontaneous evolution of circular codes in theoretical minimal RNA rings. Gene 2019, 705, 95–102. [Google Scholar] [CrossRef] [PubMed]
- Demongeot, J.; Seligmann, H. Bias for 3’-dominant codon directional asymmetry in theoretical minimal RNA rings. J. Comput. Biol. 2019, 26. [Google Scholar] [CrossRef] [PubMed]
- Demongeot, J.; Seligmann, H. More pieces of ancient than recent theoretical minimal proto-tRNA-like RNA rings in genes coding for tRNA synthetases. J. Mol. Evol. 2019, 87, 1–23. [Google Scholar] [CrossRef] [PubMed]
- Torres de Farias, S.; Gaudêncio Rêgo, T.; José, M.V. Origin of the 16S Ribosomal Molecule from Ancestor tRNAs. Sci 2019, 1, 8. [Google Scholar] [CrossRef]



| Rhodobacter sphaeroides |
| AATGGTATTCCCATTCGATTTG tRNA-Gly (http://lowelab.ucsc.edu/GtRNAdb/Rhod_spha_ATCC_17029/rhodSpha_ATCC17029-tRNAs.fa) |
| AATGGTACTGCGTCTCAAGACG 5S rRNA (http://www.combio.pl/rrna/alignment/) |
| CCTGGAACTGCCATTGAAACTC 16S rRNA (https://www.ncbi.nlm.nih.gov/nuccore/636559472?report=fasta) |
| AATGGTACTGCCATTCAAGATG Consensus |
| Rhodospirillum rubrum |
| TGAATGGTACTTCCAATTCGAA tRNA-Gly (http://trna.ie.niigata-u.ac.jp/) |
| CCAATGGTACTGCGTCTTAAGG 5S rRNA (http://www.combio.pl/rrna/alignment/) |
| CTCCAGGTACTGCCCTTGATAC 16S rRNA (https://www.arb-silva.de/browser/) |
| CGAATGGTACTGCCATTTAAAA Consensus |
| Rubellimicrobium thermophilum |
| AGTGGTACTTCCATTCGACATG tRNA-Gly (http://trna.ie.niigata-u.ac.jp/) |
| AATGGTACTGCGCCTCAAGACG 5S rRNA (http://www.combio.pl/rrna/alignment/) |
| GATGGTCCAGGCGCTGCCGCTC 16S rRNA (https://www.arb-silva.de/browser/) |
| AATGGTACTGCCACTCAAGATG Consensus |
| Haematobacter missouriensis |
| AGGGGTATTGCCATTCGAATTA tRNA-Gly (http://trna.ie.niigata-u.ac.jp/cgi-bin/trnadb/whole_detail.cgi?SID=2138813) |
| TATGGTGCTTCCATTCCCGCTA tRNA-Gly (https://www.ncbi.nlm.nih.gov/nuccore/672903602?report=genbank) |
| AATGGTACTGCGTCTCAAGACG5S rRNA (http://www.combio.pl/rrna/alignment/) |
| AATGGTAGTGACAATGGGTTAA 16S rRNA (https://www.arb-silva.de/browser/) |
| AATGGTACTGCCATTCAAGATG Consensus |
| Paracoccus sp. S4493 |
| AATGGTACTTCCCTTCGATTTA tRNA-Gly (https://www.ncbi.nlm.nih.gov/nuccore/NZ_JXYF01000001.1?from=63131&to=63204&sat=19&sat_key=63645080&report=fasta&strand=2) |
| GATGGTACTGCGTCTTAAGACG5S rRNA (http://www.combio.pl/rrna/alignment/) |
| AATGGTGGTGACAGTGGGTTAA 16S rRNA (http://www.ebi.ac.uk/ena/data/view/FJ457300&display=fasta) |
| AATGGTACTGCCATTCAATTTA Consensus |
| Flavobacteria bacterium MS024-2A |
| CTGGTATTGCCATTCGAATCGC tRNA-Gly (http://gtrnadb.ucsc.edu/genomes/bacteria/Flav_bact_3519_10/flavBact_3519_10-tRNAs.fa) |
| ATGGTACTGCCATCCGGTGGGA 5S rRNA (http://www.combio.pl/rrna/alignment/) |
| ATGGTAACGGCATACCAAGGCA 16S RRNA (http://www.ebi.ac.uk/ena/data/view/AM931128&display=fasta) |
| ATGGTACTGCCATTCGAAGGGA Consensus |
| Methanococcus maripaludis |
| CTGGTACTTCCATTCAAATCGT tRNA-Gly (http://gtrnadb.ucsc.edu/genomes/archaea/Meth_mari_C5/methMari_C5_1-tRNAs.fa) |
| TAAGTACTGCCATCUGGUGGGA 5S rRNA (http://biobases.ibch.poznan.pl/htbins/getseq.cgi?name= Methanococcus%20maripaludis) |
| TCGGTACGGGCCTTGAGAGAGG 16S rRNA (http://www.ebi.ac.uk/ena/data/view/AB546258&display=fasta) |
| TTGGTACTGCCATTCAGAGAGA Consensus |
| Tremella mesenterica |
| GATCTGCGAAGTCAAGATGAAT 5S rRNA (http://www.combio.pl/rrna/alignment/) |
| GGTAATTCTAGAGCTAATACAT18S rRNA (https://www.ncbi.nlm.nih.gov/nuccore/256600119?report=fasta) |
| GTACCGTGAGGGAAAGATGAAA 28S rRNA (https://www.ncbi.nlm.nih.gov/nuccore/46402656?report=fasta) |
| GGTCCGTGAAGTCAAGATGAAT Consensus |
| Homo sapiens |
| GTGGTACTCCCATTCAATTTGG tRNA (http://trna.bioinf.uni-leipzig.de/DataOutput/Result) |
| ATGGTAGTCGCCGTGCCTACCA 18S rRNA (https://www.ncbi.nlm.nih.gov/nuccore/225637497?report=fasta) |
| ATGGTAATCCTGCTCAGTACGA 28S rRNA (https://www.ncbi.nlm.nih.gov/nuccore/1154886866?report=fasta) |
| ATGGTACTCCCATTCAATACGA Consensus |
| AATGGTACTGCCATTTAAAACG Consensus Bacteria |
| AATGGTACTGCCATTCAAGATG Consensus Bacteria |
| AATGGTACTGCCATTCAAGATG Consensus Bacteria |
| AATGGTACTGCCACTCAAGATG Consensus Bacteria |
| AATGGTACTGCCATTCAAGATG Consensus Bacteria |
| AATGGTACTGCCATTCAATTTA Consensus Bacteria |
| ATTGGTACTGCCATTCAGAGAG Consensus Archaea |
| AATGGTCCGTGAAGTCAAGATG Consensus Eukaryote AATGGTACTCCCATTCAATACG Consensus Eukaryote |
| AATGGTACTGCCATTCAAGATG Consensus consensorum |
| Species | Percentages |
|---|---|
| Archaea | 248/584 = 42.5% |
| Bacteria | 131983/155823 = 84.7% |
| Plant | 44/80 = 55% |
| Fungi | 106/115 = 92.2% |
| Virus | 6/18 = 33.3% |
| Phage | 67/276 = 24.3% |
| Chloroplast | 109/116 = 94% |
| AL-Pentamer | Observed Number |
|---|---|
| ATTCA | 43,219 * |
| TTCAA | 51,917 * |
| TCAAG | 44,233 * |
| CAAGA | 46,523 * |
| AAGAT | 43,189 * |
| AGATG | 48,717 * |
| GATGA | 34,600 |
| ATGAA | 51,794 * |
| TGAAT | 44,410 * |
| Homo sapiens mRNA for glycyl-tRNA synthetase, complete cds GenBank: D30658.1 36 × 100/2279 = 1.58 |
| Helicobacter pylori B38 complete genome, strain B38 NCBI Reference Sequence: NC_012973.1: c941829-940933 glycyl—tRNA ligase subunit alpha 15x100/893 = 1.68 |
| Methanococcus maripaludis, strain DSM 2067 chromosome, complete genome NCBI Reference Sequence: NZ_CP026606.1:782166-783890 glycyl-tRNA synthetase 32x100/1721 = 1.86 |
| Fusarium oxysporum f. sp. melonis 26406 unplaced genomic scaffold supercont1.3, whole genome shotgun sequence GenBank: JH659331.1: c2101550-2098309 glycyl-tRNA synthetase 68x100/3238 = 2.10 |
| Prochlorococcus marinus str. NATL1A, complete genome GenBank: CP000553.1: 728949-731111 glycyl-tRNA synthetase 50x100/2155 = 2.32 |
| Methanocaldococcus jannaschii DSM 2661, complete genome GenBank: L77117.1: 219104-220837 glycyl-tRNA synthetase 41x100/1730 = 2.37 |
| Archae Methanococcoides methylutens MM1, complete genome GenBank: CP009518.1: 1554853-1556598 glycyl-tRNA synthetase 43x100/1738 = 2.47 |
| Archae Candidatus Nanosalinarum sp. J07AB56 genomic scaffold scf_7180000039101, whole genome shotgun sequence GenBank: GL982569.1 571934-572506 prolyl-tRNA synthetase 32x100/1286 = 2.49 |
| Methanococcus maripaludis C5, complete genome GenBank: CP000609.1: c1395811-1394084 glycyl-tRNA synthetase 43x100/1720 = 2.50 |
| Methanobacterium formicicum strain BRM9, complete genome GenBank: CP006933.1 556622-558343 glycyl-tRNA synthetase 44x100/1718 = 2.56 |
| Rickettsia prowazekii strain Naples-1chromosome, complete genome GenBank: CP014865.1: c1072366-1071497 glycine-tRNA ligase subunit alpha 25x100/866 = 2.89 |
| TCAATCAAGATGAAGAGTACGT tRNA synthetase CCMP1335 XM_002286706.1 2.73 |
| AGAGTCAAGATGAATAGTAGTA glycyl-tRNA synthetase CCMP1335 XM_002286964.1 2.44 |
| CCATGCAAGATGAATGTGGGTG glycyl-tRNA synthetase CCMP1335 XM_002288084.1 1.92 |
| GCATTCAAGATGAATCTTCTTG arginyl-tRNA synthetase CCMP1335 XM_002288460.12.19 |
| TCCATCTCATGGAATGGTACTG methionyl-tRNA synthetase CCMP1335 XM_002292549.1 1.91 |
| CTACCTAGGATGAAGGGTCATG valyl-tRNA synthetase CCMP1335 XM_002295439.1 2.32 |
| GCATACAAGAGTAATGGATCTG cysteinyl-tRNA synthetase CCMP1335 XM_002286789.1 2.04 |
| CCATTCGAAATGTTTGGTATTG tRNA-Gly mitochondrion DQ186202.1 7.14 |
| CCATTGGTGTTGTATGGTAAAC 60S ribosomal protein CCMP1335 XM_002290416.1 1.83 |
| CCAAGGAGGATGCGCGAGACTG 60S ribosomal protein CCMP1335 XM_002290087.1 1.77 |
| CTAGTCAAGATGCCTCGTCTAG 40S ribosomal protein CCMP1335 XM_002290013.1 2.78 |
| AAATTGAAGATTAGTGGTGGAG 40S ribosomal protein CCMP1335 XM_002293773.1 2.97 |
| CCATGAATGTTTCATGCCTCTG 18S ribosomal protein Bc6EHU KP201658.1 1.55 |
| ACGTTCAACCACACTGGAACTG 16S ribosomal protein BFB575 KC545746.1 1.51 |
| CCATTCAAGATGAATGGTACTG CONSENSUS |
| 3′-ATACATCCC(C)TCTTAAGTTCCCTT-5′ (TIR) |
| 3′-TTCCATCCC -TCTTAAGTTCGATT-5′ (CRISPR) |
| 5′-ATGGTACTG -- CCATTCAAGATGA-3′ (AL) |
| 5′-GTGATACAG -- CCCTTAACAAAAA-3′ (murine consensus RSS) |
| 5′-ATTCAACATGAA-3′ (human RSS spacer) |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Demongeot, J.; Norris, V. Emergence of a “Cyclosome” in a Primitive Network Capable of Building “Infinite” Proteins. Life 2019, 9, 51. https://doi.org/10.3390/life9020051
Demongeot J, Norris V. Emergence of a “Cyclosome” in a Primitive Network Capable of Building “Infinite” Proteins. Life. 2019; 9(2):51. https://doi.org/10.3390/life9020051
Chicago/Turabian StyleDemongeot, Jacques, and Vic Norris. 2019. "Emergence of a “Cyclosome” in a Primitive Network Capable of Building “Infinite” Proteins" Life 9, no. 2: 51. https://doi.org/10.3390/life9020051
APA StyleDemongeot, J., & Norris, V. (2019). Emergence of a “Cyclosome” in a Primitive Network Capable of Building “Infinite” Proteins. Life, 9(2), 51. https://doi.org/10.3390/life9020051






