Bottom-Up Approaches to Synthetic Cooperation in Microbial Communities
Abstract
1. Introduction
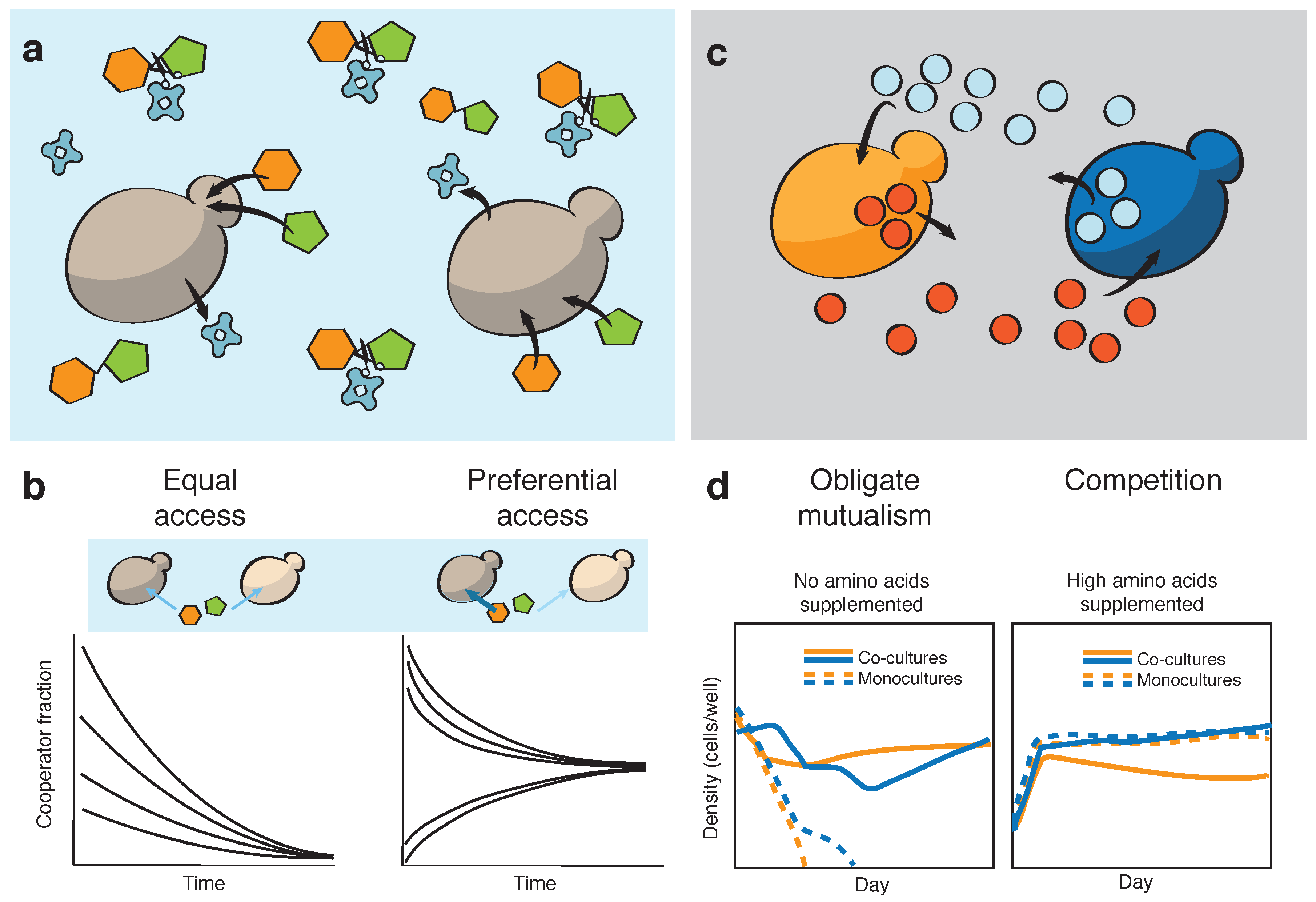
2. Engineering Microbial Cooperation, Mutualism and Cheating in Well-Mixed Environments
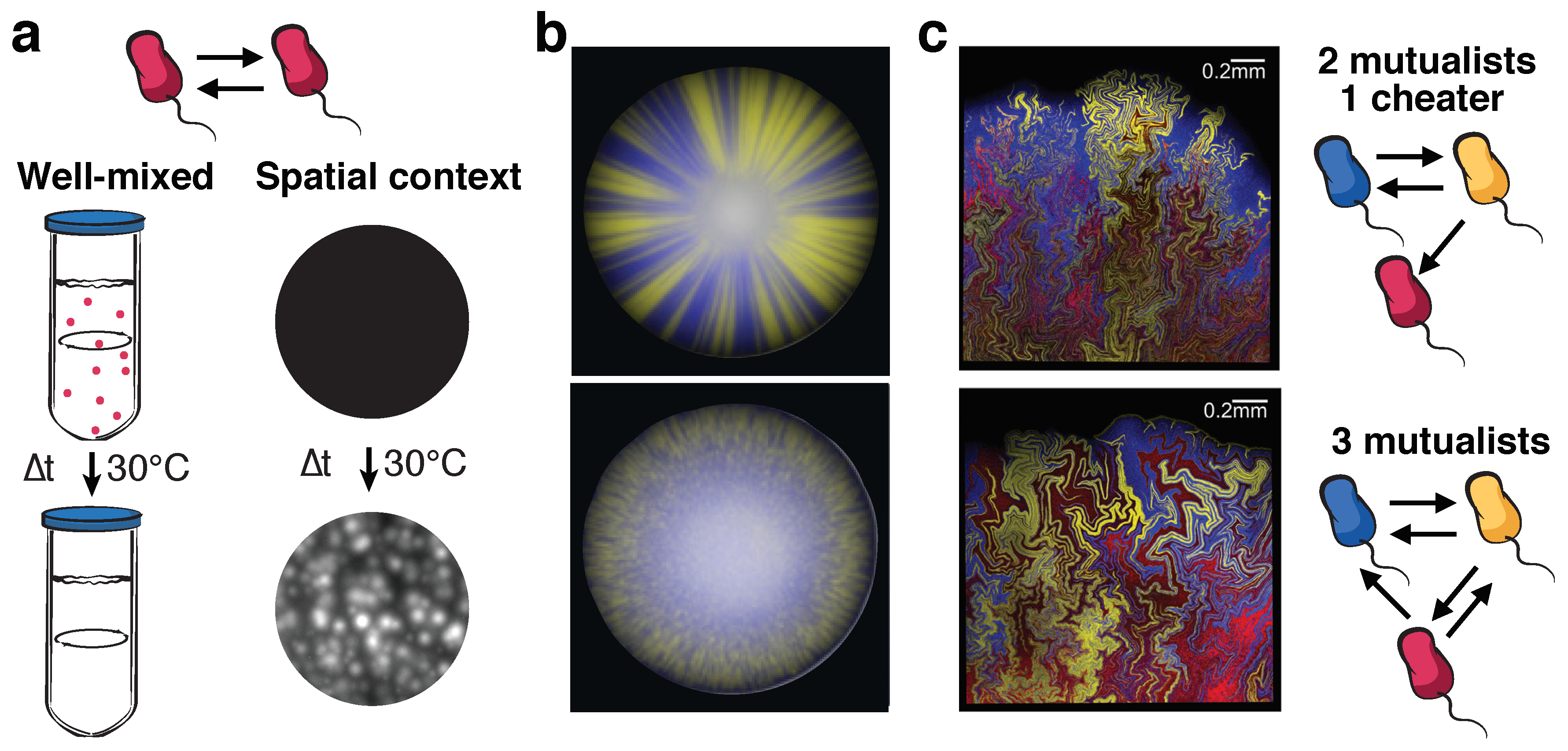
3. The Role of Spatial Structure
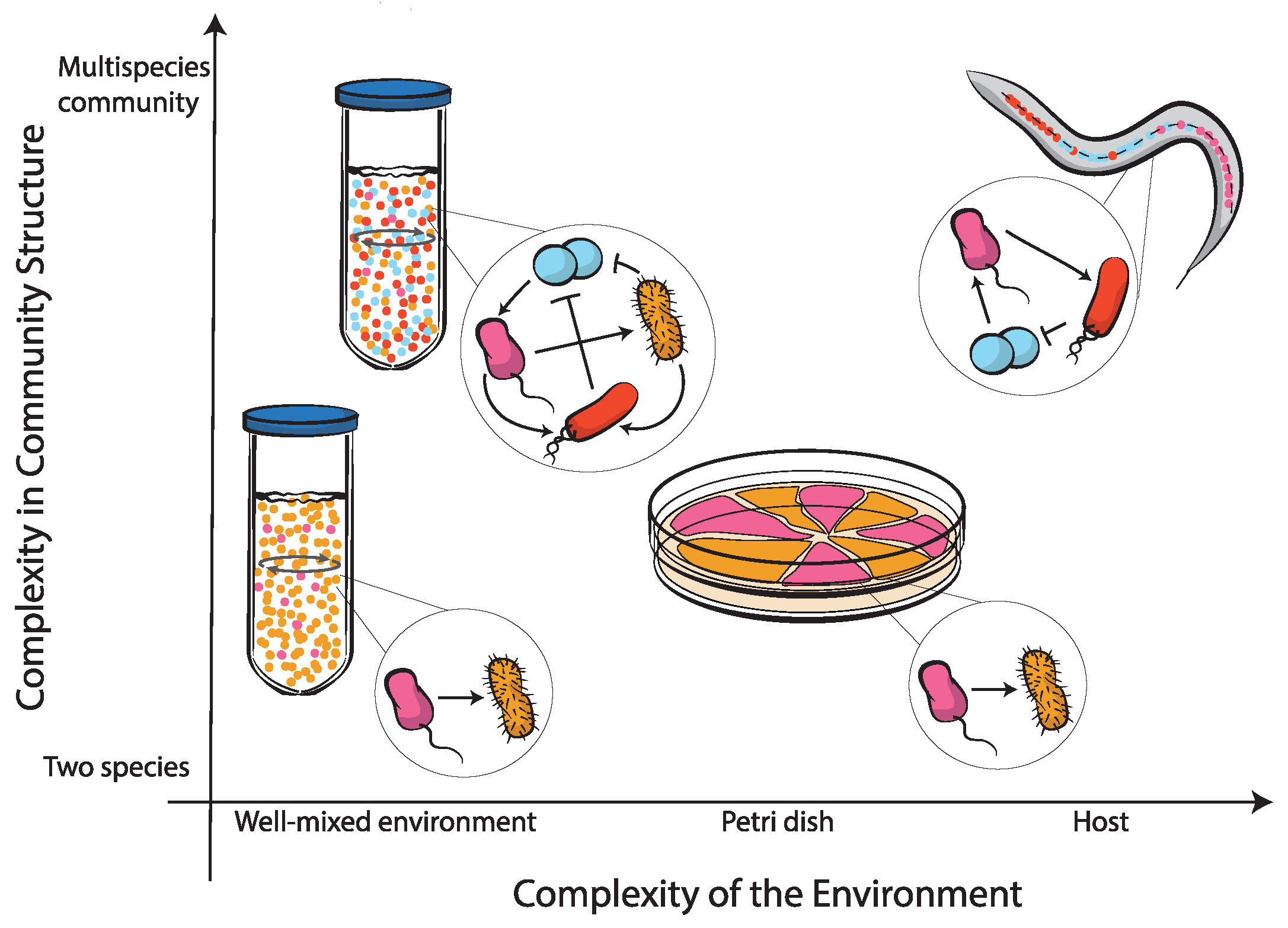
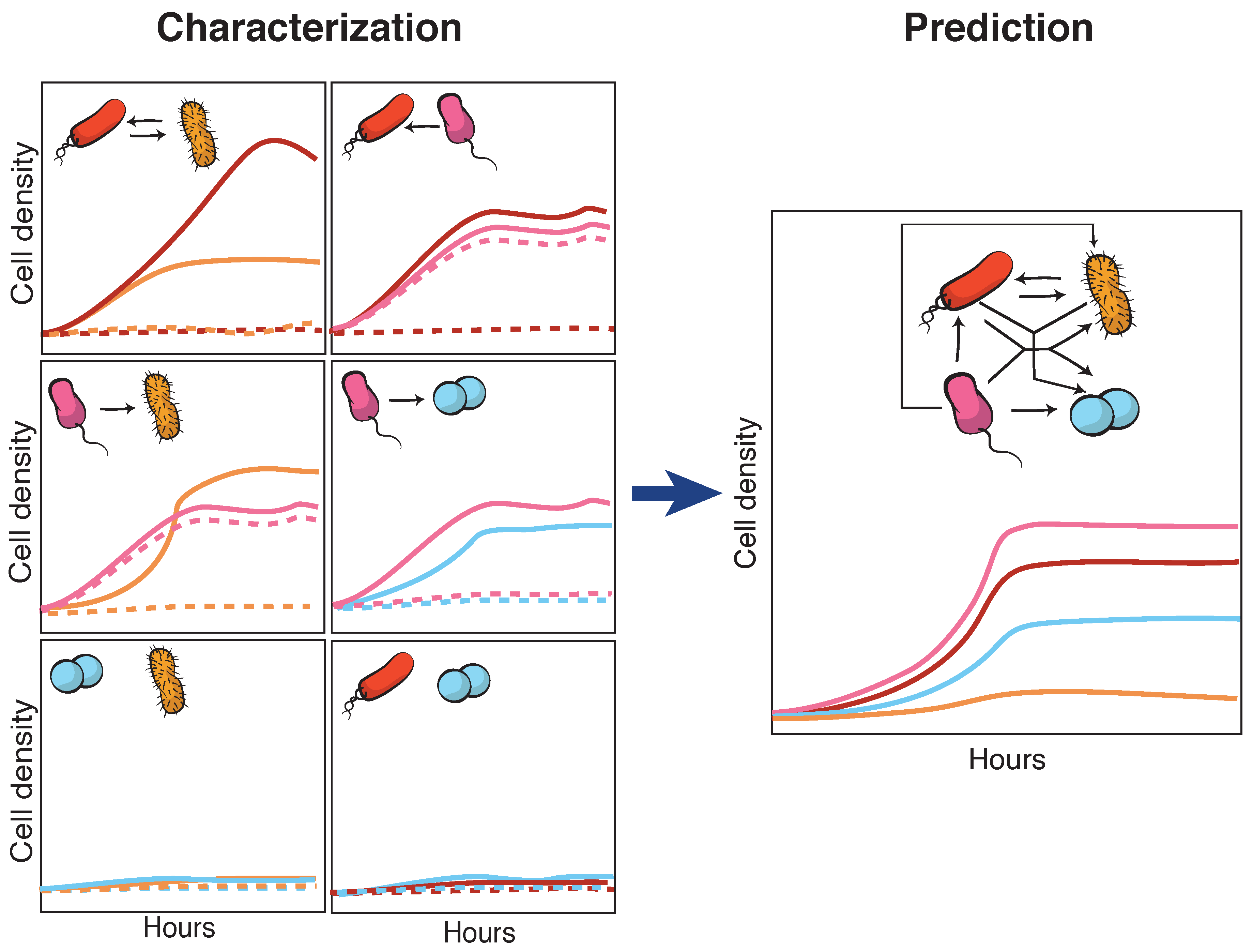
4. Towards Higher Complexity in Synthetic Microbial Communities
5. Synthetic Mutualism in Host-Associated Communities: The Nematode C. elegans as a Model System
6. Concluding Remarks
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Velicer, G.J. Social strife in the microbial world. Trends Microbiol. 2003, 11, 330–337. [Google Scholar] [CrossRef]
- West, S.A.; Griffin, A.S.; Gardner, A.; Diggle, S.P. Social evolution theory for microorganisms. Nat. Rev. Microbiol. 2006, 4, 597–607. [Google Scholar] [CrossRef] [PubMed]
- Rosenberg, E.; Keller, K.H.; Dworkin, M. Cell density-dependent growth of Myxococcus xanthus on casein. J. Bacteriol. 1977, 129, 770–777. [Google Scholar] [PubMed]
- Greig, D.; Travisano, M. The Prisoner’s Dilemma and polymorphism in yeast SUC genes. Proc. Biol. Sci. 2004, 271 (Suppl. 3), S25–S26. [Google Scholar] [CrossRef] [PubMed]
- Meredith, H.R.; Srimani, J.K.; Lee, A.J.; Lopatkin, A.J.; You, L. Collective antibiotic tolerance: Mechanisms, dynamics and intervention. Nat. Chem. Biol. 2015, 11, 182. [Google Scholar] [CrossRef] [PubMed]
- Strassmann, J.E.; Zhu, Y.; Queller, D.C. Altruism and social cheating in the social amoeba Dictyostelium discoideum. Nature 2000, 408, 965. [Google Scholar] [CrossRef] [PubMed]
- Mitri, S.; Richard Foster, K. The Genotypic View of Social Interactions in Microbial Communities. Annu. Rev. Genet. 2013, 47, 247–273. [Google Scholar] [CrossRef] [PubMed]
- Shou, W.; Ram, S.; Vilar, J.M.G. Synthetic cooperation in engineered yeast populations. Proc. Natl. Acad. Sci. USA 2007, 104, 1877–1882. [Google Scholar] [CrossRef] [PubMed]
- Yurtsev, E.A.; Conwill, A.; Gore, J. Oscillatory dynamics in a bacterial cross-protection mutualism. Proc. Natl. Acad. Sci. USA 2016. Available online: http://www.pnas.org/content/early/2016/05/17/1523317113.full.pdf (accessed on 14 February 2019). [CrossRef] [PubMed]
- West, S.A.; Cooper, G.A. Division of labour in microorganisms: An evolutionary perspective. Nat. Rev. Microbiol. 2016, 14, 716. [Google Scholar] [CrossRef] [PubMed]
- Nadell, C.D.; Drescher, K.; Foster, K.R. Spatial structure, cooperation and competition in biofilms. Nat. Rev. Microbiol. 2016, 14, 589. [Google Scholar] [CrossRef] [PubMed]
- Zomorrodi, A.R.; Segrè, D. Synthetic Ecology of Microbes: Mathematical Models and Applications. J. Mol. Biol. 2016, 428, 837–861. [Google Scholar] [CrossRef] [PubMed]
- Darwin, C. On the Origin of Species by Means of Natural Selection; Murray: London, UK, 1859. [Google Scholar]
- Kropotkin, P.A. Mutual Aid: A Factor of Evolution; W. Heinemann: London, UK, 1904. [Google Scholar]
- Hamilton, W. The genetical evolution of social behaviour. II. J. Theor. Biol. 1964, 7, 17–52. [Google Scholar] [CrossRef]
- Smith, J.M.; Price, G.R. The Logic of Animal Conflict. Nature 1973, 246, 15. [Google Scholar] [CrossRef]
- Axelrod, R.; Hamilton, W. The evolution of cooperation. Science 1981, 211, 1390–1396. Available online: http://science.sciencemag.org/content/211/4489/1390.full.pdf (accessed on 14 February 2019). [CrossRef] [PubMed]
- Nowak, M. Evolutionary Dynamics; Harvard University Press: Cambridge, MA, USA, 2006. [Google Scholar]
- Nowak, M.A. Five rules for the evolution of cooperation. Science 2006, 314, 1560–1563. [Google Scholar] [CrossRef] [PubMed]
- Gibson, D.G.; Hutchison, C.A.; Smith, H.O.; Venter, J.C. Synthetic Biology: Tools for Engineering Biological Systems; Cold Spring Harbor: New York, NY, USA, 2017. [Google Scholar]
- Gore, J.; Youk, H.; van Oudenaarden, A. Snowdrift game dynamics and facultative cheating in yeast. Nature 2009, 459, 253–256. [Google Scholar] [CrossRef] [PubMed]
- Estrela, S.; Libby, E.; Van Cleve, J.; Débarre, F.; Deforet, M.; Harcombe, W.R.; Peña, J.; Brown, S.P.; Hochberg, M.E. Environmentally Mediated Social Dilemmas. Trends Ecol. Evol. 2018. [Google Scholar] [CrossRef] [PubMed]
- Zomorrodi, A.R.; Segrè, D. Genome-driven evolutionary game theory helps understand the rise of metabolic interdependencies in microbial communities. Nat. Commun. 2017, 8, 1563. [Google Scholar] [CrossRef] [PubMed]
- Germerodt, S.; Bohl, K.; Lück, A.; Pande, S.; Schröter, A.; Kaleta, C.; Schuster, S.; Kost, C. Pervasive Selection for Cooperative Cross-Feeding in Bacterial Communities. PLOS Comput. Biol. 2016, 12, 1–21. [Google Scholar] [CrossRef] [PubMed]
- Dühring, S.; Ewald, J.; Germerodt, S.; Kaleta, C.; Dandekar, T.; Schuster, S. Modelling the host-pathogen interactions of macrophages and Candida albicans using Game Theory and dynamic optimization. J. R. Soc. Interface 2017, 14, 20170095. [Google Scholar] [CrossRef] [PubMed]
- Momeni, B.; Brileya, K.A.; Fields, M.W.; Shou, W. Strong inter-population cooperation leads to partner intermixing in microbial communities. eLife 2013, 2, e00230. [Google Scholar] [CrossRef] [PubMed]
- Kazamia, E.; Aldridge, D.C.; Smith, A.G. Synthetic ecology—A way forward for sustainable algal biofuel production? J. Biotechnol. 2012, 162, 163–169. [Google Scholar] [CrossRef]
- Wierckx, N.; Prieto, M.A.; Pomposiello, P.; de Lorenzo, V.; O’Connor, K.; Blank, L.M. Plastic waste as a novel substrate for industrial biotechnology. Microb. Biotechnol. 2015, 8, 900–903. [Google Scholar] [CrossRef] [PubMed]
- Solé, R. Bioengineering the biosphere? Ecol. Complex. 2015, 22, 40–49. [Google Scholar] [CrossRef]
- Sheth, R.U.; Cabral, V.; Chen, S.P.; Wang, H.H. Manipulating Bacterial Communities by in situ Microbiome Engineering. Trends Genet. 2016, 32, 189–200. [Google Scholar] [CrossRef] [PubMed]
- Arnold, J.W.; Roach, J.; Azcarate-Peril, M.A. Emerging Technologies for Gut Microbiome Research. Trends Microbiol. 2016, 24, 887–901. [Google Scholar] [CrossRef] [PubMed]
- McCarty, N.S.; Ledesma-Amaro, R. Synthetic Biology Tools to Engineer Microbial Communities for Biotechnology. Trends Biotechnol. 2018. [Google Scholar] [CrossRef] [PubMed]
- Brophy, J.A.N.; Voigt, C.A. Principles of genetic circuit design. Nat. Methods 2014, 11, 508. [Google Scholar] [CrossRef] [PubMed]
- Balagaddé, F.K.; Song, H.; Ozaki, J.; Collins, C.H.; Barnet, M.; Arnold, F.H.; Quake, S.R.; You, L. A synthetic Escherichia coli predator-prey ecosystem. Mol. Syst. Biol. 2008, 4, 187. [Google Scholar] [CrossRef] [PubMed]
- Sanchez, A.; Gore, J. Feedback between Population and Evolutionary Dynamics Determines the Fate of Social Microbial Populations. PLoS Biol. 2013, 11, e1001547. [Google Scholar] [CrossRef] [PubMed]
- Chen, A.; Sanchez, A.; Dai, L.; Gore, J. Dynamics of a producer-freeloader ecosystem on the brink of collapse. Nat. Commun. 2014, 5. [Google Scholar] [CrossRef] [PubMed]
- Hoek, T.A.; Axelrod, K.; Biancalani, T.; Yurtsev, E.A.; Liu, J.; Gore, J. Resource Availability Modulates the Cooperative and Competitive Nature of a Microbial Cross-Feeding Mutualism. PLOS Biol. 2016, 14, e1002540. [Google Scholar] [CrossRef] [PubMed]
- Wintermute, E.H.; Silver, P.A. Dynamics in the mixed microbial concourse. Genes Dev. 2010, 24, 2603–2614. [Google Scholar] [CrossRef] [PubMed]
- Wintermute, E.H.; Silver, P.A. Emergent cooperation in microbial metabolism. Mol. Syst. Biol. 2010, 6. [Google Scholar] [CrossRef] [PubMed]
- Cavaliere, M.; Feng, S.; Soyer, O.S.; Jiménez, J.I. Cooperation in microbial communities and their biotechnological applications. Environ. Microbiol. 2017, 19, 2949–2963. Available online: https://onlinelibrary.wiley.com/doi/pdf/10.1111/1462-2920.13767 (accessed on 14 February 2019). [CrossRef] [PubMed]
- Zengler, K.; Zaramela, L.S. The social network of microorganisms—How auxotrophies shape complex communities. Nat. Rev. Microbiol. 2018, 16, 383–390. [Google Scholar] [CrossRef] [PubMed]
- Mee, M.T.; Collins, J.J.; Church, G.M.; Wang, H.H. Syntrophic exchange in synthetic microbial communities. Proc. Natl. Acad. Sci. USA 2014, 111, E2149–E2156. [Google Scholar] [CrossRef] [PubMed]
- Pande, S.; Merker, H.; Bohl, K.; Reichelt, M.; Schuster, S.; de Figueiredo, L.F.; Kaleta, C.; Kost, C. Fitness and stability of obligate cross-feeding interactions that emerge upon gene loss in bacteria. ISME J. 2013, 8, 953. [Google Scholar] [CrossRef] [PubMed]
- Muller, M.J.I.; Neugeboren, B.I.; Nelson, D.R.; Murray, A.W. Genetic drift opposes mutualism during spatial population expansion. Proc. Natl. Acad. Sci. USA 2014, 111, 1037–1042. [Google Scholar] [CrossRef] [PubMed]
- Hosoda, K.; Suzuki, S.; Yamauchi, Y.; Shiroguchi, Y.; Kashiwagi, A.; Ono, N.; Mori, K.; Yomo, T. Cooperative Adaptation to Establishment of a Synthetic Bacterial Mutualism. PLoS ONE 2011, 6, e17105. [Google Scholar] [CrossRef] [PubMed]
- McNally, C.P.; Borenstein, E. Metabolic model-based analysis of the emergence of bacterial cross-feeding via extensive gene loss. BMC Syst. Biol. 2018, 12, 69. [Google Scholar] [CrossRef] [PubMed]
- Morris, J.J.; Lenski, R.E.; Zinser, E.R. The Black Queen Hypothesis: Evolution of Dependencies through Adaptive Gene Loss. mBio 2012, 3. Available online: https://mbio.asm.org/content/3/2/e00036-12.full.pdf (accessed on 14 February 2019). [CrossRef] [PubMed]
- Morris, J.J. Black Queen evolution: The role of leakiness in structuring microbial communities. Trends Genet. 2015, 31, 475–482. [Google Scholar] [CrossRef] [PubMed]
- Bajić, D.; Vila, J.C.C.; Blount, Z.D.; Sánchez, A. On the deformability of an empirical fitness landscape by microbial evolution. Proc. Natl. Acad. Sci. USA 2018, 115, 11286–11291. Available online: https://www.pnas.org/content/115/44/11286.full.pdf (accessed on 14 February 2019). [CrossRef]
- Pande, S.; Kaftan, F.; Lang, S.; Svatoš, A.; Germerodt, S.; Kost, C. Privatization of cooperative benefits stabilizes mutualistic cross-feeding interactions in spatially structured environments. ISME J. 2016, 10, 1413–1423. [Google Scholar] [CrossRef] [PubMed]
- Amor, D.R.; Montañez, R.; Duran-Nebreda, S.; Solé, R. Spatial dynamics of synthetic microbial mutualists and their parasites. PLOS Comput. Biol. 2017, 13, 1–22. [Google Scholar] [CrossRef] [PubMed]
- Ratzke, C.; Gore, J. Self-organized patchiness facilitates survival in a cooperatively growing Bacillus subtilis population. Nat. Microbiol. 2016, 1. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Prindle, A.; Humphries, J.; Gabalda-Sagarra, M.; Asally, M.; Lee, D.Y.D.; Ly, S.; Garcia-Ojalvo, J.; Süel, G.M. Metabolic co-dependence gives rise to collective oscillations within biofilms. Nature 2015, 523, 550–554. [Google Scholar] [CrossRef] [PubMed]
- Hallatschek, O.; Hersen, P.; Ramanathan, S.; Nelson, D.R. Genetic drift at expanding frontiers promotes gene segregation. Proc. Natl. Acad. Sci. USA 2007, 104, 19926–19930. [Google Scholar] [CrossRef] [PubMed]
- Hallatschek, O.; Nelson, D.R. Life at the front of an expanding population. Evolution 2010, 64, 193–206. [Google Scholar] [CrossRef] [PubMed]
- Nowak, M.A.; May, R.M. Evolutionary games and spatial chaos. Nature 1992, 359, 826. [Google Scholar] [CrossRef]
- Nadell, C.D.; Foster, K.R.; Xavier, J.B. Emergence of Spatial Structure in Cell Groups and the Evolution of Cooperation. PLoS Comput. Biol. 2010, 6, e1000716. [Google Scholar] [CrossRef] [PubMed]
- Boerlijst, M.; Hogeweg, P. Spiral wave structure in pre-biotic evolution: Hypercycles stable against parasites. Phys. D Nonlinear Phenomena 1991, 48, 17–28. [Google Scholar] [CrossRef]
- Sardanyés, J.; Solé, R.V. Spatio-temporal dynamics in simple asymmetric hypercycles under weak parasitic coupling. Phys. D Nonlinear Phenomena 2007, 231, 116–129. [Google Scholar] [CrossRef]
- Van Dyken, J.D.; Müller, M.J.I.; Mack, K.M.L.; Desai, M.M. Spatial Population Expansion Promotes the Evolution of Cooperation in an Experimental Prisoner’s Dilemma. Curr. Biol. 2013, 23, 919–923. [Google Scholar] [CrossRef] [PubMed]
- Datta, M.S.; Korolev, K.S.; Cvijovic, I.; Dudley, C.; Gore, J. Range expansion promotes cooperation in an experimental microbial metapopulation. Proc. Natl. Acad. Sci. USA 2013, 110, 7354–7359. [Google Scholar] [CrossRef] [PubMed]
- Momeni, B.; Waite, A.J.; Shou, W. Spatial self-organization favors heterotypic cooperation over cheating. eLife 2013, 2, e00960. [Google Scholar] [CrossRef] [PubMed]
- Widder, S.; Allen, R.J.; Pfeiffer, T.; Curtis, T.P.; Wiuf, C.; Sloan, W.T.; Cordero, O.X.; Brown, S.P.; Momeni, B.; Shou, W.; et al. Challenges in microbial ecology: building predictive understanding of community function and dynamics. ISME J. 2016, 10, 2557. [Google Scholar] [CrossRef] [PubMed]
- Kong, W.; Meldgin, D.R.; Collins, J.J.; Lu, T. Designing microbial consortia with defined social interactions. Nat. Chem. Biol. 2018, 14, 821–829. [Google Scholar] [CrossRef] [PubMed]
- Friedman, J.; Higgins, L.M.; Gore, J. Community structure follows simple assembly rules in microbial microcosms. Nat. Ecol. Evol. 2017, 1, 0109. [Google Scholar] [CrossRef] [PubMed]
- Higgins, L.M.; Friedman, J.; Shen, H.; Gore, J. Co-occurring soil bacteria exhibit a robust competitive hierarchy and lack of non-transitive interactions. bioRxiv 2017. Available online: https://www.biorxiv.org/content/early/2017/08/16/175737.full.pdf (accessed on 14 February 2019). [CrossRef]
- Grilli, J.; Barabás, G.; Michalska-Smith, M.J.; Allesina, S. Higher-order interactions stabilize dynamics in competitive network models. Nature 2017, 548, 210. [Google Scholar] [CrossRef] [PubMed]
- Cairns, J.; Jokela, R.; Hultman, J.; Tamminen, M.; Virta, M.; Hiltunen, T. Construction and Characterization of Synthetic Bacterial Community for Experimental Ecology and Evolution. Front. Genet. 2018, 9. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.J.; Boedicker, J.Q.; Choi, J.W.; Ismagilov, R.F. Defined spatial structure stabilizes a synthetic multispecies bacterial community. Proc. Natl. Acad. Sci. USA 2008, 105, 18188–18193. [Google Scholar] [CrossRef] [PubMed]
- Hays, S.G.; Yan, L.L.W.; Silver, P.A.; Ducat, D.C. Synthetic photosynthetic consortia define interactions leading to robustness and photoproduction. J. Biol. Eng. 2017, 11, 4. [Google Scholar] [CrossRef] [PubMed]
- Li, T.; Li, C.T.; Butler, K.; Hays, S.G.; Guarnieri, M.T.; Oyler, G.A.; Betenbaugh, M.J. Mimicking lichens: incorporation of yeast strains together with sucrose-secreting cyanobacteria improves survival, growth, ROS removal, and lipid production in a stable mutualistic co-culture production platform. Biotechnol. Biofuels 2017, 10, 55. [Google Scholar] [CrossRef] [PubMed]
- Zhou, K.; Qiao, K.; Edgar, S.; Stephanopoulos, G. Distributing a metabolic pathway among a microbial consortium enhances production of natural products. Nat. Biotechnol. 2015, 33, 77. [Google Scholar] [CrossRef] [PubMed]
- Thommes, M.; Wang, T.; Zhao, Q.; Paschalidis, I.C.; Segrè, D. Designing metabolic division of labor in microbial communities. bioRxiv 2018. Available online: https://www.biorxiv.org/content/early/2018/10/14/442376.full.pdf (accessed on 14 February 2019). [CrossRef]
- Ma, N.J.; Moonan, D.W.; Isaacs, F.J. Precise manipulation of bacterial chromosomes by conjugative assembly genome engineering. Nat. Protoc. 2014, 9, 2285. [Google Scholar] [CrossRef] [PubMed]
- Cong, L.; Ran, F.A.; Cox, D.; Lin, S.; Barretto, R.; Habib, N.; Hsu, P.D.; Wu, X.; Jiang, W.; Marraffini, L.A.; et al. Multiplex Genome Engineering Using CRISPR/Cas Systems. Science 2013, 339, 819–823. Available online: http://science.sciencemag.org/content/339/6121/819.full.pdf (accessed on 14 February 2019). [CrossRef] [PubMed]
- Solé, R.V.; Montañez, R.; Duran-Nebreda, S. Synthetic circuit designs for earth terraformation. Biol. Direct 2015, 10, 37. [Google Scholar] [CrossRef] [PubMed]
- Solé, R.V.; Montañez, R.; Duran-Nebreda, S.; Rodriguez-Amor, D.; Vidiella, B.; Sardanyés, J. Population dynamics of synthetic terraformation motifs. Open Sci. 2018, 5. Available online: http://rsos.royalsocietypublishing.org/content/5/7/180121.full.pdf (accessed on 14 February 2019). [CrossRef]
- Mallon, C.A.; Le Roux, X.; van Doorn, G.S.; Dini-Andreote, F.; Poly, F.; Salles, J.F. The impact of failure: Unsuccessful bacterial invasions steer the soil microbial community away from the invader’s niche. ISME J. 2018, 12, 728–741. [Google Scholar] [CrossRef] [PubMed]
- Costello, E.K.; Stagaman, K.; Dethlefsen, L.; Bohannan, B.J.M.; Relman, D.A. The Application of Ecological Theory Toward an Understanding of the Human Microbiome. Science 2012, 336, 1255–1262. Available online: http://science.sciencemag.org/content/336/6086/1255.full.pdf (accessed on 14 February 2019). [CrossRef] [PubMed]
- Soucy, S.M.; Huang, J.; Gogarten, J.P. Horizontal gene transfer: Building the web of life. Nat. Rev. Genet. 2015, 16, 472. [Google Scholar] [CrossRef] [PubMed]
- Levin, B.R.; Antia, R. Why we don’t get sick: The within-host population dynamics of bacterial infections. Science 2001, 292, 1112–1115. [Google Scholar] [CrossRef] [PubMed]
- Lozupone, C.A.; Stombaugh, J.I.; Gordon, J.I.; Jansson, J.K.; Knight, R. Diversity, stability and resilience of the human gut microbiota. Nature 2012, 489, 220–230. [Google Scholar] [CrossRef] [PubMed]
- Huttenhower, C.; Gevers, D.; Knight, R.; Abubucker, S.; Badger, J.H.; Chinwalla, A.T.; Creasy, H.H.; Earl, A.M.; Fitzgerald, M.G.; Fulton, R.S.; et al. Structure, function and diversity of the healthy human microbiome. Nature 2012, 486, 207–214. [Google Scholar] [CrossRef]
- Fukuda, S.; Toh, H.; Hase, K.; Oshima, K.; Nakanishi, Y.; Yoshimura, K.; Tobe, T.; Clarke, J.M.; Topping, D.L.; Suzuki, T.; et al. Bifidobacteria can protect from enteropathogenic infection through production of acetate. Nature 2011, 469, 543–547. [Google Scholar] [CrossRef] [PubMed]
- Kau, A.L.; Ahern, P.P.; Griffin, N.W.; Goodman, A.L.; Gordon, J.I. Human nutrition, the gut microbiome and the immune system. Nature 2011, 474, 327–336. [Google Scholar] [CrossRef] [PubMed]
- Lloyd-Price, J.; Abu-Ali, G.; Huttenhower, C. The healthy human microbiome. Genome Med. 2016, 8, 51. [Google Scholar] [CrossRef] [PubMed]
- Goodman, A.L.; Kallstrom, G.; Faith, J.J.; Reyes, A.; Moore, A.; Dantas, G.; Gordon, J.I. Extensive personal human gut microbiota culture collections characterized and manipulated in gnotobiotic mice. Proc. Natl. Acad. Sci. USA 2011, 108, 6252–6257. [Google Scholar] [CrossRef] [PubMed]
- Bashan, A.; Gibson, T.E.; Friedman, J.; Carey, V.J.; Weiss, S.T.; Hohmann, E.L.; Liu, Y.Y. Universality of human microbial dynamics. Nature 2016, 534, 259. [Google Scholar] [CrossRef] [PubMed]
- Mimee, M.; Tucker, A.; Voigt, C.; Lu, T. Programming a Human Commensal Bacterium, Bacteroides thetaiotaomicron, to Sense and Respond to Stimuli in the Murine Gut Microbiota. Cell Syst. 2015, 1, 62–71. [Google Scholar] [CrossRef] [PubMed]
- Leonard, S.P.; Perutka, J.; Powell, J.E.; Geng, P.; Richhart, D.D.; Byrom, M.; Kar, S.; Davies, B.W.; Ellington, A.D.; Moran, N.A.; et al. Genetic Engineering of Bee Gut Microbiome Bacteria with a Toolkit for Modular Assembly of Broad-Host-Range Plasmids. ACS Synth. Biol. 2018, 7, 1279–1290. [Google Scholar] [CrossRef] [PubMed]
- Claesen, J.; Fischbach, M.A. Synthetic microbes as drug delivery systems. ACS Synth. Biol. 2015, 4, 358–364. [Google Scholar] [CrossRef] [PubMed]
- Wu, H.C.; Tsao, C.Y.; Quan, D.N.; Cheng, Y.; Servinsky, M.D.; Carter, K.K.; Jee, K.J.; Terrell, J.L.; Zargar, A.; Rubloff, G.W.; et al. Autonomous bacterial localization and gene expression based on nearby cell receptor density. Mol. Syst. Biol. 2013, 9, 636. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Yang, B.; Cheng, X.; Qiao, Y.; Tang, B.; Chen, G.; Wei, J.; Liu, X.; Cheng, W.; Du, P.; et al. Salmonella-mediated tumor-targeting TRAIL gene therapy significantly suppresses melanoma growth in mouse model. Cancer Sci. 2012, 103, 325–333. [Google Scholar] [CrossRef] [PubMed]
- Anderson, J.C.; Clarke, E.J.; Arkin, A.P.; Voigt, C.A. Environmentally controlled invasion of cancer cells by engineered bacteria. J. Mol. Biol. 2006, 355, 619–627. [Google Scholar] [CrossRef] [PubMed]
- Archer, E.J.; Robinson, A.B.; Süel, G.M. Engineered E. coli that detect and respond to gut inflammation through nitric oxide sensing. ACS Synth. Biol. 2012, 1, 451–457. [Google Scholar] [CrossRef] [PubMed]
- Saeidi, N.; Wong, C.K.; Lo, T.M.; Nguyen, H.X.; Ling, H.; Leong, S.S.J.; Poh, C.L.; Chang, M.W. Engineering microbes to sense and eradicate Pseudomonas aeruginosa, a human pathogen. Mol. Syst. Biol. 2011, 7, 521. [Google Scholar] [CrossRef] [PubMed]
- Hwang, I.Y.; Koh, E.; Wong, A.; March, J.C.; Bentley, W.E.; Lee, Y.S.; Chang, M.W. Engineered probiotic Escherichia coli can eliminate and prevent Pseudomonas aeruginosa gut infection in animal models. Nat. Commun. 2017, 8, 15028. [Google Scholar] [CrossRef] [PubMed]
- Kostic, A.D.; Howitt, M.R.; Garrett, W.S. Exploring host-microbiota interactions in animal models and humans. Genes Dev. 2013, 27, 701–718. [Google Scholar] [CrossRef] [PubMed]
- Yilmaz, L.S.; Walhout, A.J. Worms, bacteria, and micronutrients: an elegant model of our diet. Trends Genet. 2014, 30, 496–503. [Google Scholar] [CrossRef] [PubMed]
- Vega, N.M.; Gore, J. Stochastic assembly produces heterogeneous communities in the Caenorhabditis elegans intestine. PLoS Biol. 2017, 15, e2000633. [Google Scholar] [CrossRef] [PubMed]
- Scott, T.A.; Quintaneiro, L.M.; Norvaisas, P.; Lui, P.P.; Wilson, M.P.; Leung, K.Y.; Herrera-Dominguez, L.; Sudiwala, S.; Pessia, A.; Clayton, P.T.; et al. Host-Microbe Co-metabolism Dictates Cancer Drug Efficacy in C. elegans. Cell 2017, 169, 442–456. [Google Scholar] [CrossRef] [PubMed]
- The C. elegans Sequencing Consortium. Genome sequence of the nematode C. elegans: A platform for investigating biology. Science 1998, 282, 2012–2018. [Google Scholar] [CrossRef]
- Yan, G.; Vértes, P.E.; Towlson, E.K.; Chew, Y.L.; Walker, D.S.; Schafer, W.R.; Barabási, A.L. Network control principles predict neuron function in the Caenorhabditis elegans connectome. Nature 2017, 550, 519–523. [Google Scholar] [CrossRef] [PubMed]
- Brenner, S. The genetics of Caenorhabditis elegans. Genetics 1974, 77, 71–94. [Google Scholar] [PubMed]
- Corsi, A.K.; Wightman, B.; Chalfie, M. A transparent window into biology: A primer on Caenorhabditis elegans. Genetics 2015, 200, 387–407. [Google Scholar] [CrossRef] [PubMed]
- Avery, L.; You, Y.J. C. elegans feeding. WormBook 2012, 1–23. [Google Scholar] [CrossRef]
- Medlock, G.L.; Carey, M.A.; McDuffie, D.G.; Mundy, M.B.; Giallourou, N.; Swann, J.R.; Kolling, G.L.; Papin, J.A. Inferring Metabolic Mechanisms of Interaction within a Defined Gut Microbiota. Cell Syst. 2018, 7, 245–257. [Google Scholar] [CrossRef] [PubMed]
- Berg, M.; Stenuit, B.; Ho, J.; Wang, A.; Parke, C.; Knight, M.; Alvarez-Cohen, L.; Shapira, M. Assembly of the Caenorhabditis elegans gut microbiota from diverse soil microbial environments. ISME J. 2016, 10, 1998–2009. [Google Scholar] [CrossRef] [PubMed]
- Ortiz Lopez, A.; Vega, N.M.; Gore, J. Interspecies bacterial competition determines community assembly in the C. elegans intestine. bioRxiv 2019. Available online: https://www.biorxiv.org/content/early/2019/01/30/535633.full.pdf (accessed on 14 February 2019). [CrossRef]
- Portal-Celhay, C.; Blaser, M.J. Competition and resilience between founder and introduced bacteria in the Caenorhabditis elegans gut. Infect. Immun. 2012, 80, 1288–1299. [Google Scholar] [CrossRef] [PubMed]
- Burns, A.R.; Stephens, W.Z.; Stagaman, K.; Wong, S.; Rawls, J.F.; Guillemin, K.; Bohannan, B.J. Contribution of neutral processes to the assembly of gut microbial communities in the zebrafish over host development. ISME J. 2016, 10, 655–664. [Google Scholar] [CrossRef] [PubMed]
- Hubbell, S. The Unified Neutral Theory of Biodiversity and Biogeography (MPB-32); Monographs in Population Biology, Princeton University Press: Princeton, NJ, USA, 2001. [Google Scholar]
- Schulenburg, H.; Félix, M.A. The natural biotic environment of Caenorhabditis elegans. Genetics 2017, 206, 55–86. [Google Scholar] [CrossRef] [PubMed]
- Watson, E.; MacNeil, L.; Ritter, A.; Yilmaz, L.; Rosebrock, A.; Caudy, A.; Walhout, A. Interspecies Systems Biology Uncovers Metabolites Affecting C. elegans Gene Expression and Life History Traits. Cell 2014, 156, 759–770. [Google Scholar] [CrossRef] [PubMed]
- Montalvo-Katz, S.; Huang, H.; Appel, M.D.; Berg, M.; Shapira, M. Association with soil bacteria enhances p38-dependent infection resistance in Caenorhabditis elegans. Infect. Immun. 2013, 81, 514–520. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.; Mylonakis, E. Caenorhabditis elegans immune conditioning with the probiotic bacterium Lactobacillus acidophilus strain ncfm enhances gram-positive immune responses. Infect. Immun. 2012, 80, 2500–2508. [Google Scholar] [CrossRef] [PubMed]
- Gusarov, I.; Gautier, L.; Smolentseva, O.; Shamovsky, I.; Eremina, S.; Mironov, A.; Nudler, E. Bacterial Nitric Oxide Extends the Lifespan of C. elegans. Cell 2013, 152, 818–830. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; Yu, X.; Jia, R.; Yang, R.; Rui, Q.; Wang, D. Lactic acid bacteria protects caenorhabditis elegans from toxicity of graphene oxide by maintaining normal intestinal permeability under different genetic backgrounds. Sci. Rep. 2015, 5, 17233. [Google Scholar] [CrossRef] [PubMed]
- Donia, M. A Toolbox for Microbiome Engineering. Cell Syst. 2015, 1, 21–23. [Google Scholar] [CrossRef] [PubMed]
- Bulleri, F.; Eriksson, B.K.; Queirós, A.; Airoldi, L.; Arenas, F.; Arvanitidis, C.; Bouma, T.J.; Crowe, T.P.; Davoult, D.; Guizien, K.; et al. Harnessing positive species interactions as a tool against climate-driven loss of coastal biodiversity. PLOS Biol. 2018, 16, e2006852. [Google Scholar] [CrossRef] [PubMed]
| Motif | Interaction |
|---|---|
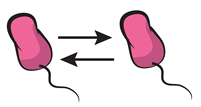 | Cooperation: interaction that increases the fitness of neighboring cells. Homotypic cooperation, more specifically, refers to cooperative interactions happening between cells sharing a given genotype. |
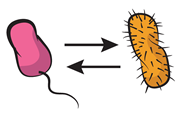 | Mutualism: cooperative interaction occurring between different genotypes, i.e., heterotypic cooperation. |
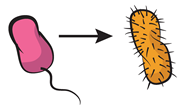 | Commensalism: interaction that increases the fitness of a given genotype, with no cost or benefit for the donor. |
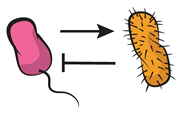 | Cheating (or parasitism): one of the members benefits from the interaction at the expenses of the donor, or cooperator. This interaction motif is also known as parasitism. This same interaction motif is known as predation [34] when additional inhibitory effects, other than competition for available resources, are played against the donor organism. |
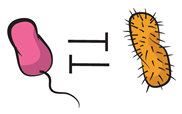 | Competition: both members experience a reduced fitness as a result of the interaction. |
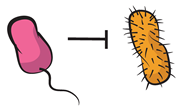 | Amensalism: one of the partners is negatively affected by the presence of another, the latter experiencing neither cost nor benefit. |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rodríguez Amor, D.; Dal Bello, M. Bottom-Up Approaches to Synthetic Cooperation in Microbial Communities. Life 2019, 9, 22. https://doi.org/10.3390/life9010022
Rodríguez Amor D, Dal Bello M. Bottom-Up Approaches to Synthetic Cooperation in Microbial Communities. Life. 2019; 9(1):22. https://doi.org/10.3390/life9010022
Chicago/Turabian StyleRodríguez Amor, Daniel, and Martina Dal Bello. 2019. "Bottom-Up Approaches to Synthetic Cooperation in Microbial Communities" Life 9, no. 1: 22. https://doi.org/10.3390/life9010022
APA StyleRodríguez Amor, D., & Dal Bello, M. (2019). Bottom-Up Approaches to Synthetic Cooperation in Microbial Communities. Life, 9(1), 22. https://doi.org/10.3390/life9010022





