Mineral Surface-Templated Self-Assembling Systems: Case Studies from Nanoscience and Surface Science towards Origins of Life Research
Abstract
1. Introduction
2. Case Studies
2.1. Simple Organic Molecules
2.2. Nucleic Acids
2.3. Peptides
2.4. Biomineralization
3. Prospective
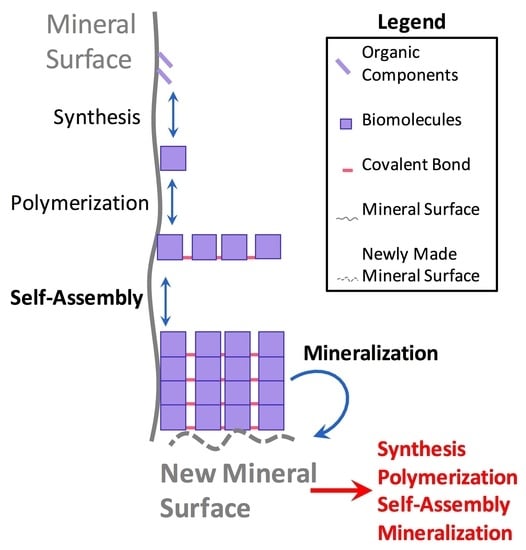
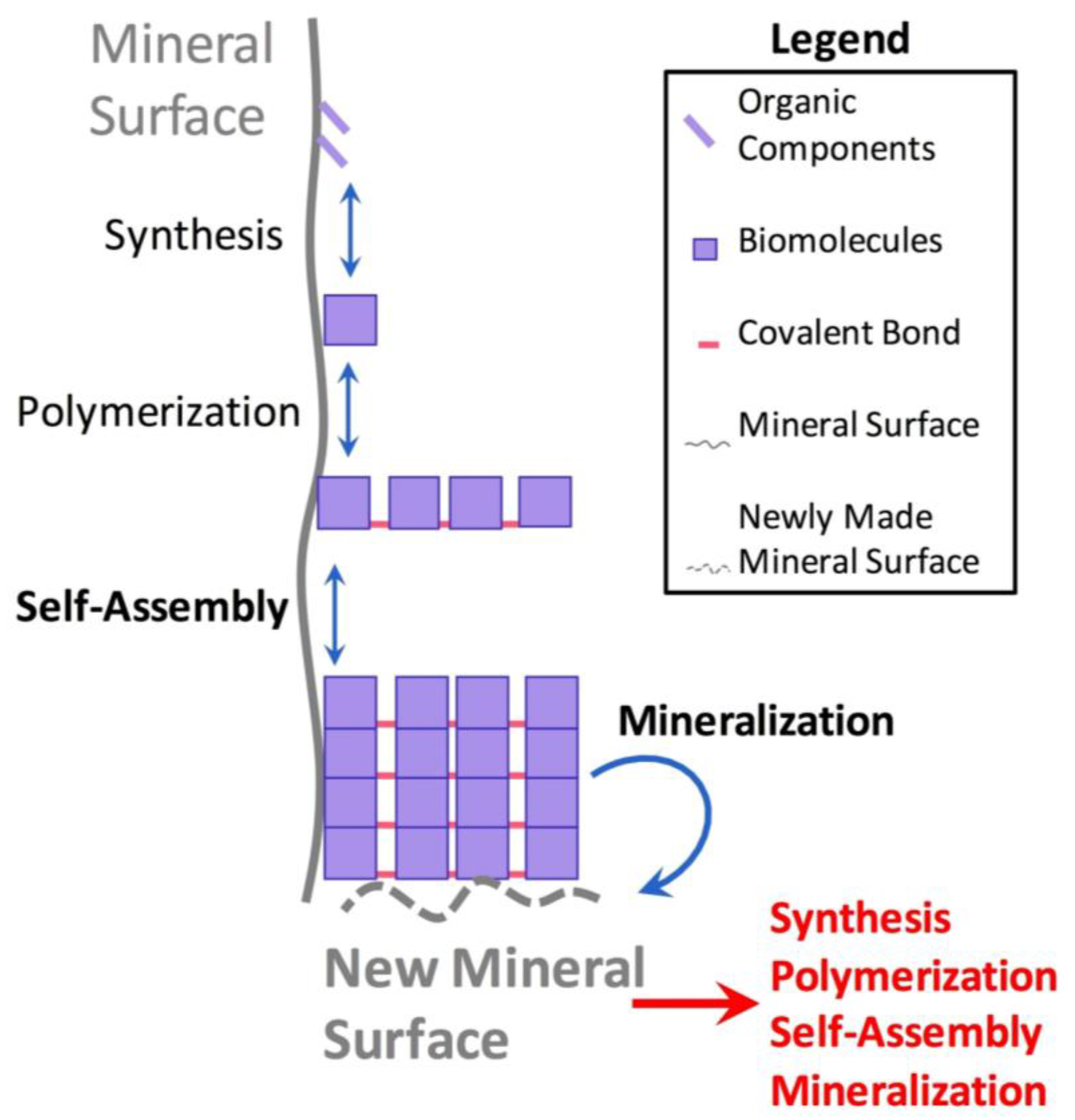
3.1. Synergistic Cyclical Model of Mineral-Templated Self-Assembling Systems Promoting Mineralization
3.2. Incorporating Recent Discoveries in Nanoscience and Surface Science into Origins of Life Research
Author Contributions
Acknowledgments
Conflicts of Interest
References
- Damineli, A.; Damineli, D.S.C. Origens da vida. Estudos Avançados 2007, 21, 263–284. [Google Scholar] [CrossRef]
- Kawai, S.; Haapasilta, V.; Lindner, B.D.; Tahara, K.; Spijker, P.; Buitendijk, J.A.; Pawlak, R.; Meier, T.; Tobe, Y.; Foster, A.S.; et al. Thermal control of sequential on-surface transformation of a hydrocarbon molecule on a copper surface. Nat. Commun. 2016, 7, 12711. [Google Scholar] [CrossRef] [PubMed]
- Johlin, J.M. Interfacial adsorption as a Function of the Concentration of Colloidal Solutions. J. Biol. Chem. 1929, 84, 543–551. [Google Scholar]
- Nič, M.; Jirát, J.; Košata, B.; Jenkins, A.; McNaught, A. Adsorption. In IUPAC Compendium of Chemical Terminology; Nič, M., Jirát, J., Košata, B., Jenkins, A., McNaught, A., Eds.; IUPAC: Zurich, Switzerland, 2009; ISBN 9780967855097. [Google Scholar]
- Pang, S.H.; Medlin, J.W. Controlling Catalytic Selectivity via Adsorbate Orientation on the Surface: From Furfural Deoxygenation to Reactions of Epoxides. J. Phys. Chem. Lett. 2015, 6, 1348–1356. [Google Scholar] [CrossRef] [PubMed]
- Chatterjee, D.; Akash, R.; Kamalnath, K.; Ahmad, R.; Singh, A.K.; Ravishankar, N. Orientation Selection during Heterogeneous Nucleation: Implications for Heterogeneous Catalysis. J. Phys. Chem. C 2017, 121, 10027–10037. [Google Scholar] [CrossRef]
- Nørskov, J.K. Electronic factors in catalysis. Prog. Surf. Sci. 1991, 38, 103–144. [Google Scholar] [CrossRef]
- Xu, Y.; Schoonen, M.A.A. The absolute energy positions of conduction and valence bands of selected semiconducting minerals. Am. Mineral. 2000, 85, 543–556. [Google Scholar] [CrossRef]
- Hazen, R.M.; Grew, E.S.; Downs, R.T.; Golden, J.; Hystad, G. Mineral Ecology: Chance and Necessity in the Mineral Diversity of Terrestrial Planets. Can. Mineral. 2015, 53, 295–324. [Google Scholar] [CrossRef]
- Schwartz, A.W.; Veen, M.; Bisseling, T.; Chittenden, G.J.F. Prebiotic nucleotide synthesis-demonstration of a geologically plausible pathway. Orig. Life 1975, 6, 163–168. [Google Scholar] [CrossRef] [PubMed]
- Pasek, M.A. Schreibersite on the early Earth: Scenarios for prebiotic phosphorylation. Geosci. Front. 2017, 8, 329–335. [Google Scholar] [CrossRef]
- Gull, M.; Mojica, M.A.; Fernández, F.M.; Gaul, D.A.; Orlando, T.M.; Liotta, C.L.; Pasek, M.A. Nucleoside phosphorylation by the mineral schreibersite. Sci. Rep. 2015, 5, 17198. [Google Scholar] [CrossRef] [PubMed]
- Saladino, R.; Barontini, M.; Cossetti, C.; Di Mauro, E.; Crestini, C. The effects of borate minerals on the synthesis of nucleic acid bases, amino acids and biogenic carboxylic acids from formamide. Orig. Life Evol. Biosph. 2011, 41, 317–330. [Google Scholar] [CrossRef] [PubMed]
- Ricardo, A.; Carrigan, M.A.; Olcott, A.N.; Benner, S.A. Borate minerals stabilize ribose. Science 2004, 303, 196. [Google Scholar] [CrossRef] [PubMed]
- Grew, E.S.; Bada, J.L.; Hazen, R.M. Borate minerals and origin of the RNA world. Orig. Life Evol. Biosph. 2011, 41, 307–316. [Google Scholar] [CrossRef] [PubMed]
- Lambert, J.-F. Adsorption and polymerization of amino acids on mineral surfaces: A review. Orig. Life Evol. Biosph. 2008, 38, 211–242. [Google Scholar] [CrossRef] [PubMed]
- Hashizume, H. Adsorption of Nucleic Acid Bases, Ribose, and Phosphate by Some Clay Minerals. Life 2015, 5, 637–650. [Google Scholar] [CrossRef] [PubMed]
- Hedges, J.I.; Hare, P.E. Amino acid adsorption by clay minerals in distilled water. Geochim. Cosmochim. Acta 1987, 51, 255–259. [Google Scholar] [CrossRef]
- Ferris, J.P. Montmorillonite-catalysed formation of RNA oligomers: The possible role of catalysis in the origins of life. Philos. Trans. Royal Soc. Lond. B Biol. Sci. 2006, 361, 1777–1786. [Google Scholar] [CrossRef] [PubMed]
- Ferris, J.P.; Hill, A.R., Jr.; Liu, R.; Orgel, L.E. Synthesis of long prebiotic oligomers on mineral surfaces. Nature 1996, 381, 59–61. [Google Scholar] [CrossRef] [PubMed]
- Ferris, J.; Ertem, G. Oligomerization of ribonucleotides on montmorillonite: Reaction of the 5′-phosphorimidazolide of adenosine. Science 1992, 257, 1387–1389. [Google Scholar] [CrossRef] [PubMed]
- Kawamura, K.; Ferris, J.P. Kinetic and Mechanistic Analysis of Dinucleotide and Oligonucleotide Formation from the 5′-Phosphorimidazolide of Adenosine on Na-Montmorillonite. J. Am. Chem. Soc. 1994, 116, 7564–7572. [Google Scholar] [CrossRef] [PubMed]
- Orgel, L.E. Polymerization on the rocks: Theoretical introduction. Orig. Life Evol. Biosph. 1998, 28, 227–234. [Google Scholar] [CrossRef] [PubMed]
- Hill, A.R., Jr.; Böhler, C.; Orgel, L.E. Polymerization on the rocks: Negatively-charged alpha-amino acids. Orig. Life Evol. Biosph. 1998, 28, 235–243. [Google Scholar] [CrossRef] [PubMed]
- Liu, R.; Orgel, L.E. Polymerization on the rocks: Beta-amino acids and arginine. Orig. Life Evol. Biosph. 1998, 28, 245–257. [Google Scholar] [CrossRef] [PubMed]
- Lahav, N.; White, D.; Chang, S. Peptide formation in the prebiotic era: Thermal condensation of glycine in fluctuating clay environments. Science 1978, 201, 67–69. [Google Scholar] [CrossRef] [PubMed]
- Kitadai, N.; Oonishi, H.; Umemoto, K.; Usui, T.; Fukushi, K.; Nakashima, S. Glycine Polymerization on Oxide Minerals. Orig. Life Evol. Biosph. 2017, 47, 123–143. [Google Scholar] [CrossRef] [PubMed]
- Hanczyc, M.M.; Fujikawa, S.M.; Szostak, J.W. Experimental models of primitive cellular compartments: Encapsulation, growth, and division. Science 2003, 302, 618–622. [Google Scholar] [CrossRef] [PubMed]
- Hanczyc, M.M.; Mansy, S.S.; Szostak, J.W. Mineral surface directed membrane assembly. Orig. Life Evol. Biosph. 2007, 37, 67–82. [Google Scholar] [CrossRef] [PubMed]
- Sahai, N.; Kaddour, H.; Dalai, P.; Wang, Z.; Bass, G.; Gao, M. Mineral Surface Chemistry and Nanoparticle-aggregation Control Membrane Self-Assembly. Sci. Rep. 2017, 7, 43418. [Google Scholar] [CrossRef] [PubMed]
- Ebisuzaki, T.; Maruyama, S. Nuclear geyser model of the origin of life: Driving force to promote the synthesis of building blocks of life. Geosci. Front. 2017, 8, 275–298. [Google Scholar] [CrossRef]
- Westall, F.; Brack, A. The Importance of Water for Life. Space Sci. Rev. 2018, 214, 50. [Google Scholar] [CrossRef]
- Alberts, B.; Johnson, A.; Lewis, J.; Morgan, D.; Raff, M.; Roberts, K.; Walter, P. Molecular Biology of the Cell; Garland Science: Oxford, UK, 2014; ISBN 9780815344322. [Google Scholar]
- Grover, M.; He, C.; Hsieh, M.-C.; Yu, S.-S. A Chemical Engineering Perspective on the Origins of Life. Processes 2015, 3, 309–338. [Google Scholar] [CrossRef]
- Mandal, D.; Nasrolahi Shirazi, A.; Parang, K. Self-assembly of peptides to nanostructures. Org. Biomol. Chem. 2014, 12, 3544–3561. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; Yang, W.; Chen, C.; Wang, J.; Zhang, L.; Xu, H. Rational design and self-assembly of short amphiphilic peptides and applications. Curr. Opin. Colloid Interface Sci. 2018, 35, 112–123. [Google Scholar] [CrossRef]
- Singh, N.; Kumar, M.; Miravet, J.F.; Ulijn, R.V.; Escuder, B. Peptide-Based Molecular Hydrogels as Supramolecular Protein Mimics. Chemistry 2017, 23, 981–993. [Google Scholar] [CrossRef] [PubMed]
- Aumiller, W.M., Jr.; Pir Cakmak, F.; Davis, B.W.; Keating, C.D. RNA-Based Coacervates as a Model for Membraneless Organelles: Formation, Properties, and Interfacial Liposome Assembly. Langmuir 2016, 32, 10042–10053. [Google Scholar] [CrossRef] [PubMed]
- Fung, S.Y.; Keyes, C.; Duhamel, J.; Chen, P. Concentration Effect on the Aggregation of a Self-Assembling Oligopeptide. Biophys. J. 2003, 85, 537–548. [Google Scholar] [CrossRef]
- Koga, S.; Williams, D.S.; Perriman, A.W.; Mann, S. Peptide-nucleotide microdroplets as a step towards a membrane-free protocell model. Nat. Chem. 2011, 3, 720–724. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.; He, L.; Yin, Y. Magnetic field guided colloidal assembly. Mater. Today 2013, 16, 110–116. [Google Scholar] [CrossRef]
- Yin, Y.; Niu, L.; Zhu, X.; Zhao, M.; Zhang, Z.; Mann, S.; Liang, D. Non-equilibrium behaviour in coacervate-based protocells under electric-field-induced excitation. Nat. Commun. 2016, 7, 10658. [Google Scholar] [CrossRef] [PubMed]
- Thomas, J.E.; Kelley, M.J. Interaction of mineral surfaces with simple organic molecules by diffuse reflectance IR spectroscopy (DRIFT). J. Colloid Interface Sci. 2008, 322, 516–526. [Google Scholar] [CrossRef] [PubMed]
- Hakim, S.S.; Olsson, M.H.M.; Sørensen, H.O.; Bovet, N.; Bohr, J.; Feidenhans’l, R.; Stipp, S.L.S. Interactions of the Calcite {10.4} Surface with Organic Compounds: Structure and Behaviour at Mineral—Organic Interfaces. Sci. Rep. 2017, 7, 7592. [Google Scholar] [CrossRef] [PubMed]
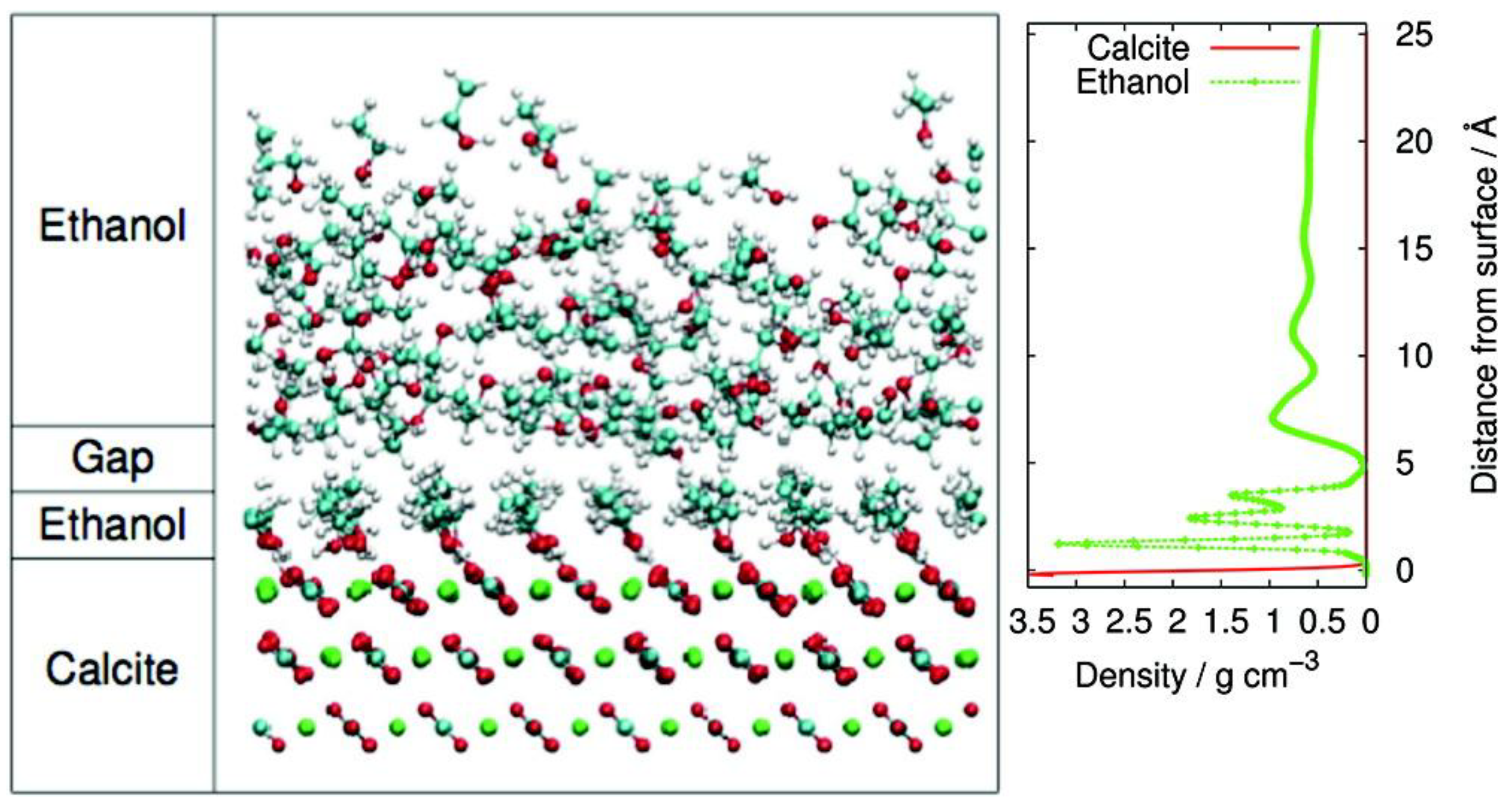
- Pasarín, I.S.; Yang, M.; Bovet, N.; Glyvradal, M.; Nielsen, M.M.; Bohr, J.; Feidenhans’l, R.; Stipp, S.L.S. Molecular ordering of ethanol at the calcite surface. Langmuir 2012, 28, 2545–2550. [Google Scholar] [CrossRef] [PubMed]
- Osman, M.A.; Ernst, M.; Meier, B.H.; Suter, U.W. Structure and Molecular Dynamics of Alkane Monolayers Self-Assembled on Mica Platelets. J. Phys. Chem. B 2002, 106, 653–662. [Google Scholar] [CrossRef]
- Duffy, D.M.; Harding, J.H. Simulation of organic monolayers as templates for the nucleation of calcite crystals. Langmuir 2004, 20, 7630–7636. [Google Scholar] [CrossRef] [PubMed]
- Sand, K.K.; Yang, M.; Makovicky, E.; Cooke, D.J.; Hassenkam, T.; Bechgaard, K.; Stipp, S.L.S. Binding of ethanol on calcite: The role of the OH bond and its relevance to biomineralization. Langmuir 2010, 26, 15239–15247. [Google Scholar] [CrossRef] [PubMed]
- Cooke, D.J.; Gray, R.J.; Sand, K.K.; Stipp, S.L.S.; Elliott, J.A. Interaction of ethanol and water with the {1014} surface of calcite. Langmuir 2010, 26, 14520–14529. [Google Scholar] [CrossRef] [PubMed]
- Yaalon, D.H. Mineral Composition of the Average Shale. Clay Miner. 1962, 5, 31–36. [Google Scholar] [CrossRef]
- Banerjee, S.; Abdulsattar, Z.R.; Agim, K.; Lane, R.H.; Hascakir, B. Mechanism of polymer adsorption on shale surfaces: Effect of polymer type and presence of monovalent and divalent salts. Petroleum 2017, 3, 384–390. [Google Scholar] [CrossRef]
- Carter, P.W. Adsorption of amino acid-containing organic matter by calcite and quartz. Geochim. Cosmochim. Acta 1978, 42, 1239–1242. [Google Scholar] [CrossRef]
- Lagaly, G.; Barrer, R.M.; Goulding, K. Clay-Organic Interactions. Philos. Trans. Royal Soc. A Math. Phys. Eng. Sci. 1984, 311, 315–332. [Google Scholar] [CrossRef]
- Mao, C. The emergence of complexity: Lessons from DNA. PLoS Biol. 2004, 2, e431. [Google Scholar] [CrossRef] [PubMed]
- Privalov, P.L.; Dragan, A.I.; Crane-Robinson, C.; Breslauer, K.J.; Remeta, D.P.; Minetti, C.A. What drives proteins into the major or minor grooves of DNA? J. Mol. Biol. 2007, 365, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Walter, N.G.; Engelke, D.R. Ribozymes: Catalytic RNAs that cut things, make things, and do odd and useful jobs. Biologist 2002, 49, 199–203. [Google Scholar] [PubMed]
- Robertson, M.P.; Joyce, G.F. The origins of the RNA world. Cold Spring Harb. Perspect. Biol. 2012, 4, a003608. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Prywes, N.; Tam, C.P.; O’Flaherty, D.K.; Lelyveld, V.S.; Izgu, E.C.; Pal, A.; Szostak, J.W. Enhanced Nonenzymatic RNA Copying with 2-Aminoimidazole Activated Nucleotides. J. Am. Chem. Soc. 2017, 139, 1810–1813. [Google Scholar] [CrossRef] [PubMed]
- Szostak, J.W. The eightfold path to non-enzymatic RNA replication. J. Syst. Chem. 2012, 3, 2. [Google Scholar] [CrossRef]
- Sowerby, S.J.; Heckl, W.M. The role of self-assembled monolayers of the purine and pyrimidine bases in the emergence of life. Orig. Life Evol. Biosph. 1998, 28, 283–310. [Google Scholar] [CrossRef] [PubMed]
- Kan, Y.; Tan, Q.; Wu, G.; Si, W.; Chen, Y. Study of DNA adsorption on mica surfaces using a surface force apparatus. Sci. Rep. 2015, 5, 8442. [Google Scholar] [CrossRef] [PubMed]
- Cai, P.; Huang, Q.-Y.; Zhang, X.-W. Interactions of DNA with Clay Minerals and Soil Colloidal Particles and Protection against Degradation by DNase. Environ. Sci. Technol. 2006, 40, 2971–2976. [Google Scholar] [CrossRef] [PubMed]
- Cleaves, H.J., II; Crapster-Pregont, E.; Jonsson, C.M.; Jonsson, C.L.; Sverjensky, D.A.; Hazen, R.A. The adsorption of short single-stranded DNA oligomers to mineral surfaces. Chemosphere 2011, 83, 1560–1567. [Google Scholar] [CrossRef] [PubMed]
- Perona, J.J. DNA Structure. In Encyclopedia of Genetics; Elsevier: New York, NY, USA, 2001; pp. 572–574. ISBN 9780122270802. [Google Scholar]
- Herbert, A.; Rich, A. The Biology of Left-handed Z.-DNA. J. Biol. Chem. 1996, 271, 11595–11598. [Google Scholar] [CrossRef] [PubMed]
- Freund, A.M.; Bichara, M.; Fuchs, R.P. Z-DNA-forming sequences are spontaneous deletion hot spots. Proc. Natl. Acad. Sci. USA 1989, 86, 7465–7469. [Google Scholar] [CrossRef] [PubMed]
- Wang, G.; Christensen, L.A.; Vasquez, K.M. Z-DNA-forming sequences generate large-scale deletions in mammalian cells. Proc. Natl. Acad. Sci. USA 2006, 103, 2677–2682. [Google Scholar] [CrossRef] [PubMed]
- Rich, A.; Zhang, S. Timeline: Z-DNA: The long road to biological function. Nat. Rev. Genet. 2003, 4, 566–572. [Google Scholar] [CrossRef] [PubMed]
- Berg, J.M.; Tymoczko, J.L.; Stryer, L. Biochemistry, Fifth Edition; W. H. Freeman: New York, NY, USA, 2002; ISBN 9780716730514. [Google Scholar]
- Swadling, J.B.; Coveney, P.V.; Christopher Greenwell, H. Stability of free and mineral-protected nucleic acids: Implications for the RNA world. Geochim. Cosmochim. Acta 2012, 83, 360–378. [Google Scholar] [CrossRef]
- Riazance, J.H.; Baase, W.A.; Johnson, W.C., Jr.; Hall, K.; Cruz, P.; Tinoco, I., Jr. Evidence for Z-form RNA by vacuum UV circular dichroism. Nucleic Acids Res. 1985, 13, 4983–4989. [Google Scholar] [CrossRef] [PubMed]
- Davis, P.W.; Adamiak, R.W.; Tinoco, I., Jr. Z-RNA: The solution NMR structure of r(CGCGCG). Biopolymers 1990, 29, 109–122. [Google Scholar] [CrossRef] [PubMed]
- D’Ascenzo, L.; Leonarski, F.; Vicens, Q.; Auffinger, P. “Z-DNA like” fragments in RNA: A recurring structural motif with implications for folding, RNA/protein recognition and immune response. Nucleic Acids Res. 2016, 44, 5944–5956. [Google Scholar] [CrossRef] [PubMed]
- Heuberger, B.D.; Pal, A.; Del Frate, F.; Topkar, V.V.; Szostak, J.W. Replacing uridine with 2-thiouridine enhances the rate and fidelity of nonenzymatic RNA primer extension. J. Am. Chem. Soc. 2015, 137, 2769–2775. [Google Scholar] [CrossRef] [PubMed]
- Franchi, M.; Ferris, J.P.; Gallori, E. Cations as mediators of the adsorption of nucleic acids on clay surfaces in prebiotic environments. Orig. Life Evol. Biosph. 2003, 33, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Hansma, H.G.; Laney, D.E. DNA binding to mica correlates with cationic radius: Assay by atomic force microscopy. Biophys. J. 1996, 70, 1933–1939. [Google Scholar] [CrossRef]
- Moore, E.K.; Hao, J.; Prabhu, A.; Zhong, H.; Jelen, B.I.; Meyer, M.; Hazen, R.M.; Falkowski, P.G. Geological and Chemical Factors that Impacted the Biological Utilization of Cobalt in the Archean Eon. J. Geophys. Res. Biogeosci. 2018, 123, 743–759. [Google Scholar] [CrossRef]
- Shannon, R.D. Revised effective ionic radii and systematic studies of interatomic distances in halides and chalcogenides. Acta Crystallogr. A 1976, 32, 751–767. [Google Scholar] [CrossRef]
- Crowe, S.A.; Jones, C.; Katsev, S.; Magen, C.; O’Neill, A.H.; Sturm, A.; Canfield, D.E.; Haffner, G.D.; Mucci, A.; Sundby, B.; et al. Photoferrotrophs thrive in an Archean Ocean analogue. Proc. Natl. Acad. Sci. USA 2008, 105, 15938–15943. [Google Scholar] [CrossRef] [PubMed]
- Sun, X.; Hyeon Ko, S.; Zhang, C.; Ribbe, A.E.; Mao, C. Surface-mediated DNA self-assembly. J. Am. Chem. Soc. 2009, 131, 13248–13249. [Google Scholar] [CrossRef] [PubMed]
- Woo, S.; Rothemund, P.W.K. Self-assembly of two-dimensional DNA origami lattices using cation-controlled surface diffusion. Nat. Commun. 2014, 5, 4889. [Google Scholar] [CrossRef] [PubMed]
- Perrault, S.D.; Shih, W.M. Virus-inspired membrane encapsulation of DNA nanostructures to achieve in vivo stability. ACS Nano 2014, 8, 5132–5140. [Google Scholar] [CrossRef] [PubMed]
- Wang, P.; Meyer, T.A.; Pan, V.; Dutta, P.K.; Ke, Y. The Beauty and Utility of DNA Origami. Chem 2017, 2, 359–382. [Google Scholar] [CrossRef]
- Akbari, E.; Mollica, M.Y.; Lucas, C.R.; Bushman, S.M.; Patton, R.A.; Shahhosseini, M.; Song, J.W.; Castro, C.E. Engineering Cell Surface Function with DNA Origami. Adv. Mater. 2017, 29. [Google Scholar] [CrossRef] [PubMed]
- Han, D.; Qi, X.; Myhrvold, C.; Wang, B.; Dai, M.; Jiang, S.; Bates, M.; Liu, Y.; An, B.; Zhang, F.; et al. Single-stranded DNA and RNA origami. Science 2017, 358, eaao2648. [Google Scholar] [CrossRef] [PubMed]
- Oi, H.; Fujita, D.; Suzuki, Y.; Sugiyama, H.; Endo, M.; Matsumura, S.; Ikawa, Y. Programmable formation of catalytic RNA triangles and squares by assembling modular RNA enzymes. J. Biochem. 2017, 161, 451–462. [Google Scholar] [CrossRef] [PubMed]
- Rahman, M.M.; Matsumura, S.; Ikawa, Y. Artificial RNA Motifs Expand the Programmable Assembly between RNA Modules of a Bimolecular Ribozyme Leading to Application to RNA Nanostructure Design. Biology 2017, 6, 37. [Google Scholar] [CrossRef] [PubMed]
- Van der Horst, G.; Christian, A.; Inoue, T. Reconstitution of a group I intron self-splicing reaction with an activator RNA. Proc. Natl. Acad. Sci. USA 1991, 88, 184–188. [Google Scholar] [CrossRef] [PubMed]
- Goedert, M.; Spillantini, M.G. A century of Alzheimer’s disease. Science 2006, 314, 777–781. [Google Scholar] [CrossRef] [PubMed]
- Hamley, I.W. The Amyloid Beta Peptide: A Chemist’s Perspective. Role in Alzheimer’s and Fibrillization. Chem. Rev. 2012, 112, 5147–5192. [Google Scholar] [CrossRef] [PubMed]
- Ferreira, S.T.; Vieira, M.N.N.; De Felice, F.G. Soluble protein oligomers as emerging toxins in Alzheimer’s and other amyloid diseases. IUBMB Life 2007, 59, 332–345. [Google Scholar] [CrossRef] [PubMed]
- Eisele, Y.S.; Monteiro, C.; Fearns, C.; Encalada, S.E.; Wiseman, R.L.; Powers, E.T.; Kelly, J.W. Targeting protein aggregation for the treatment of degenerative diseases. Nat. Rev. Drug Discov. 2015, 14, 759–780. [Google Scholar] [CrossRef] [PubMed]
- Ow, S.-Y.; Dunstan, D.E. A brief overview of amyloids and Alzheimer’s disease. Protein Sci. 2014, 23, 1315–1331. [Google Scholar] [CrossRef] [PubMed]
- Reches, M.; Gazit, E. Casting metal nanowires within discrete self-assembled peptide nanotubes. Science 2003, 300, 625–627. [Google Scholar] [CrossRef] [PubMed]
- Frederix, P.W.J.; Scott, G.G.; Abul-Haija, Y.M.; Kalafatovic, D.; Pappas, C.G.; Javid, N.; Hunt, N.T.; Ulijn, R.V.; Tuttle, T. Exploring the sequence space for (tri-)peptide self-assembly to design and discover new hydrogels. Nat. Chem. 2014, 7, 30–37. [Google Scholar] [CrossRef] [PubMed]
- Pappas, C.G.; Shafi, R.; Sasselli, I.R.; Siccardi, H.; Wang, T.; Narang, V.; Abzalimov, R.; Wijerathne, N.; Ulijn, R.V. Dynamic peptide libraries for the discovery of supramolecular nanomaterials. Nat. Nanotechnol. 2016, 11, 960–967. [Google Scholar] [CrossRef] [PubMed]
- Adler-Abramovich, L.; Gazit, E. The physical properties of supramolecular peptide assemblies: From building block association to technological applications. Chem. Soc. Rev. 2014, 43, 6881–6893. [Google Scholar] [CrossRef] [PubMed]
- Schnaider, L.; Brahmachari, S.; Schmidt, N.W.; Mensa, B.; Shaham-Niv, S.; Bychenko, D.; Adler-Abramovich, L.; Shimon, L.J.W.; Kolusheva, S.; DeGrado, W.F.; et al. Self-assembling dipeptide antibacterial nanostructures with membrane disrupting activity. Nat. Commun. 2017, 8, 1365. [Google Scholar] [CrossRef] [PubMed]
- Tao, K.; Makam, P.; Aizen, R.; Gazit, E. Self-assembling peptide semiconductors. Science 2017, 358, eaam9756. [Google Scholar] [CrossRef] [PubMed]
- Galit Fichman, E.G. Self-assembly of short peptides to form hydrogels: Design of building blocks, physical properties and technological applications. Acta Biomater. 2014, 10, 1671–1682. [Google Scholar] [CrossRef] [PubMed]
- Plankensteiner, K.; Reiner, H.; Schranz, B.; Rode, B.M. Prebiotic Formation of Amino Acids in a Neutral Atmosphere by Electric Discharge. Angew. Chem. Int. Ed. 2004, 43, 1886–1888. [Google Scholar] [CrossRef] [PubMed]
- Miller, S.L. The atmosphere of the primitive Earth and the prebiotic synthesis of amino acids. Orig. Life 1974, 5, 139–151. [Google Scholar] [CrossRef] [PubMed]
- Sephton, M.A. Organic compounds in carbonaceous meteorites. Nat. Prod. Rep. 2002, 19, 292–311. [Google Scholar] [CrossRef] [PubMed]
- Hennet, R.J.-C.; Holm, N.G.; Engel, M.H. Abiotic synthesis of amino acids under hydrothermal conditions and the origin of life: A perpetual phenomenon? Naturwissenschaften 1992, 79, 361–365. [Google Scholar] [CrossRef] [PubMed]
- Marshall, W.L. Hydrothermal synthesis of amino acids. Geochim. Cosmochim. Acta 1994, 58, 2099–2106. [Google Scholar] [CrossRef]
- Leman, L.J.; Huang, Z.-Z.; Reza Ghadiri, M. Peptide Bond Formation in Water Mediated by Carbon Disulfide. Astrobiology 2015, 15, 709–716. [Google Scholar] [CrossRef] [PubMed]
- Leman, L. Carbonyl Sulfide-Mediated Prebiotic Formation of Peptides. Science 2004, 306, 283–286. [Google Scholar] [CrossRef] [PubMed]
- Erastova, V.; Degiacomi, M.T.; Fraser, D.G.; Greenwell, H.C. Mineral surface chemistry control for origin of prebiotic peptides. Nat. Commun. 2017, 8, 2033. [Google Scholar] [CrossRef] [PubMed]
- Martra, G.; Deiana, C.; Sakhno, Y.; Barberis, I.; Fabbiani, M.; Pazzi, M.; Vincenti, M. The formation and self-assembly of long prebiotic oligomers produced by the condensation of unactivated amino acids on oxide surfaces. Angew. Chem. Int. Ed. Engl. 2014, 53, 4671–4674. [Google Scholar] [CrossRef] [PubMed]
- Kang, S.-G.; Li, H.; Huynh, T.; Zhang, F.; Xia, Z.; Zhang, Y.; Zhou, R. Molecular mechanism of surface-assisted epitaxial self-assembly of amyloid-like peptides. ACS Nano 2012, 6, 9276–9282. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Zhang, F.; Zhang, Y.; Ye, M.; Zhou, B.; Tang, Y.-Z.; Yang, H.-J.; Xie, M.-Y.; Chen, S.-F.; He, J.-H.; et al. Peptide diffusion and self-assembly in ambient water nanofilm on mica surface. J. Phys. Chem. B 2009, 113, 8795–8799. [Google Scholar] [CrossRef] [PubMed]
- Zhu, M.; Souillac, P.O.; Ionescu-Zanetti, C.; Carter, S.A.; Fink, A.L. Surface-catalyzed amyloid fibril formation. J. Biol. Chem. 2002, 277, 50914–50922. [Google Scholar] [CrossRef] [PubMed]
- Rout, S.K.; Friedmann, M.P.; Riek, R.; Greenwald, J. A prebiotic template-directed peptide synthesis based on amyloids. Nat. Commun. 2018, 9, 234. [Google Scholar] [CrossRef] [PubMed]
- Sun, L.; Narimatsu, T.; Tsuchiya, S.; Tanaka, T.; Li, P.; Hayamizu, Y. Water stability of self-assembled peptide nanostructures for sequential formation of two-dimensional interstitial patterns on layered materials. RSC Adv. 2016, 6, 96889–96897. [Google Scholar] [CrossRef]
- Sadler, K.; Tam, J.P. Peptide dendrimers: Applications and synthesis. Rev. Mol. Biotechnol. 2002, 90, 195–229. [Google Scholar] [CrossRef]
- Holliday, G.L.; Mitchell, J.B.O.; Thornton, J.M. Understanding the functional roles of amino acid residues in enzyme catalysis. J. Mol. Biol. 2009, 390, 560–577. [Google Scholar] [CrossRef] [PubMed]
- Dodson, G. Catalytic triads and their relatives. Trends Biochem. Sci. 1998, 23, 347–352. [Google Scholar] [CrossRef]
- Lagnoux, D.; Delort, E.; Douat-Casassus, C.; Esposito, A.; Reymond, J.-L. Synthesis and esterolytic activity of catalytic peptide dendrimers. Chemistry 2004, 10, 1215–1226. [Google Scholar] [CrossRef] [PubMed]
- Liao, H.-S.; Lin, J.; Liu, Y.; Huang, P.; Jin, A.; Chen, X. Self-assembly mechanisms of nanofibers from peptide amphiphiles in solution and on substrate surfaces. Nanoscale 2016, 8, 14814–14820. [Google Scholar] [CrossRef] [PubMed]
- Lin, J.; Luo, J.-B.; Yang, S.-T.; Zhou, Q.-H. Template-directed self-assembly of a designed amphiphilic hexapeptide on mica surface. Colloid Polym. Sci. 2013, 291, 2263–2270. [Google Scholar] [CrossRef]
- Cui, H.; Webber, M.J.; Stupp, S.I. Self-Assembly of Peptide Amphiphiles: From Molecules to Nanostructures to Biomaterials. Biopolymers 2010, 94, 1–18. [Google Scholar] [CrossRef] [PubMed]
- Mikhalevich, V.; Craciun, I.; Kyropoulou, M.; Palivan, C.G.; Meier, W. Amphiphilic Peptide Self-Assembly: Expansion to Hybrid Materials. Biomacromolecules 2017, 18, 3471–3480. [Google Scholar] [CrossRef] [PubMed]
- Hamley, I.W. Self-assembly of amphiphilic peptides. Soft Matter 2011, 7, 4122–4138. [Google Scholar] [CrossRef]
- Vauthey, S.; Santoso, S.; Gong, H.; Watson, N.; Zhang, S. Molecular self-assembly of surfactant-like peptides to form nanotubes and nanovesicles. Proc. Natl. Acad. Sci. USA 2002, 99, 5355–5360. [Google Scholar] [CrossRef] [PubMed]
- Hentrich, C.; Szostak, J.W. Controlled growth of filamentous fatty acid vesicles under flow. Langmuir 2014, 30, 14916–14925. [Google Scholar] [CrossRef] [PubMed]
- Adamala, K.; Szostak, J.W. Nonenzymatic template-directed RNA synthesis inside model protocells. Science 2013, 342, 1098–1100. [Google Scholar] [CrossRef] [PubMed]
- Hartgerink, J.D.; Beniash, E.; Stupp, S.I. Self-assembly and mineralization of peptide-amphiphile nanofibers. Science 2001, 294, 1684–1688. [Google Scholar] [CrossRef] [PubMed]
- Lai, P.; Moulton, K.; Krevor, S. Pore-scale heterogeneity in the mineral distribution and reactive surface area of porous rocks. Chem. Geol. 2015, 411, 260–273. [Google Scholar] [CrossRef]
- Jonker, A.M.; Dennis, W.P.; van Hest, J.C.M. Peptide- and Protein-Based Hydrogels. Chem. Mater. 2012, 24, 759–773. [Google Scholar] [CrossRef]
- Lei, Z.; Wang, Q.; Sun, S.; Zhu, W.; Wu, P. A Bioinspired Mineral Hydrogel as a Self-Healable, Mechanically Adaptable Ionic Skin for Highly Sensitive Pressure Sensing. Adv. Mater. 2017, 29, 1700321. [Google Scholar] [CrossRef] [PubMed]
- Feng, Y.; Lee, M.; Taraban, M.; Yu, Y.B. Diffusion of small molecules inside a peptide hydrogel. Chem. Commun. 2011, 47, 10455–10457. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Li, J.; Mooney, D.J. Designing hydrogels for controlled drug delivery. Nat. Rev. Mater. 2016, 1, 16071. [Google Scholar] [CrossRef] [PubMed]
- He, C.; Gállego, I.; Laughlin, B.; Grover, M.A.; Hud, N.V. A viscous solvent enables information transfer from gene-length nucleic acids in a model prebiotic replication cycle. Nat. Chem. 2016, 9, 318–324. [Google Scholar] [CrossRef] [PubMed]
- Caliari, S.R.; Burdick, J.A. A practical guide to hydrogels for cell culture. Nat. Methods 2016, 13, 405–414. [Google Scholar] [CrossRef] [PubMed]
- Gungormus, M.; Branco, M.; Fong, H.; Schneider, J.P.; Tamerler, C.; Sarikaya, M. Self assembled bi-functional peptide hydrogels with biomineralization-directing peptides. Biomaterials 2010, 31, 7266–7274. [Google Scholar] [CrossRef] [PubMed]
- Kasotakis, E.; Mossou, E.; Adler-Abramovich, L.; Mitchell, E.P.; Forsyth, V.T.; Gazit, E.; Mitraki, A. Design of metal-binding sites onto self-assembled peptide fibrils. Biopolymers 2009, 92, 164–172. [Google Scholar] [CrossRef] [PubMed]
- Jia, T.Z.; Hentrich, C.; Szostak, J.W. Rapid RNA exchange in aqueous two-phase system and coacervate droplets. Orig. Life Evol. Biosph. 2014, 44, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Aumiller, W.M., Jr.; Keating, C.D. Phosphorylation-mediated RNA/peptide complex coacervation as a model for intracellular liquid organelles. Nat. Chem. 2016, 8, 129–137. [Google Scholar] [CrossRef] [PubMed]
- Matsuura, K.; Nakamura, T.; Watanabe, K.; Noguchi, T.; Minamihata, K.; Kamiya, N.; Kimizuka, N. Self-assembly of Ni-NTA-modified β-annulus peptides into artificial viral capsids and encapsulation of His-tagged proteins. Org. Biomol. Chem. 2016, 14, 7869–7874. [Google Scholar] [CrossRef] [PubMed]
- Rufo, C.M.; Moroz, Y.S.; Moroz, O.V.; Stöhr, J.; Smith, T.A.; Hu, X.; DeGrado, W.F.; Korendovych, I.V. Short peptides self-assemble to produce catalytic amyloids. Nat. Chem. 2014, 6, 303–309. [Google Scholar] [CrossRef] [PubMed]
- Al-Garawi, Z.S.; McIntosh, B.A.; Neill-Hall, D.; Hatimy, A.A.; Sweet, S.M.; Bagley, M.C.; Serpell, L.C. The amyloid architecture provides a scaffold for enzyme-like catalysts. Nanoscale 2017, 9, 10773–10783. [Google Scholar] [CrossRef] [PubMed]
- Heuer, A.; Fink, D.; Laraia, V.; Arias, J.; Calvert, P.; Kendall, K.; Messing, G.; Blackwell, J.; Rieke, P.; Thompson, D.; et al. Innovative materials processing strategies: A biomimetic approach. Science 1992, 255, 1098–1105. [Google Scholar] [CrossRef] [PubMed]
- Addadi, L.; Weiner, S. Biomineralization: Mineral formation by organisms. Phys. Scr. 2014, 89, 098003. [Google Scholar] [CrossRef]
- Dhami, N.K.; Reddy, M.S.; Mukherjee, A. Biomineralization of calcium carbonates and their engineered applications: A review. Front. Microbiol. 2013, 4, 314. [Google Scholar] [CrossRef] [PubMed]
- Kirschvink, J.L. Magnetite biomineralization and geomagnetic sensitivity in higher animals: An update and recommendations for future study. Bioelectromagnetics 1989, 10, 239–259. [Google Scholar] [CrossRef] [PubMed]
- Tambutté, S.; Holcomb, M.; Ferrier-Pagès, C.; Reynaud, S.; Tambutté, É.; Zoccola, D.; Allemand, D. Coral biomineralization: From the gene to the environment. J. Exp. Mar. Biol. Ecol. 2011, 408, 58–78. [Google Scholar] [CrossRef]
- Fablet, R.; Pecquerie, L.; de Pontual, H.; Høie, H.; Millner, R.; Mosegaard, H.; Kooijman, S.A.L.M. Shedding light on fish otolith biomineralization using a bioenergetic approach. PLoS ONE 2011, 6, e27055. [Google Scholar] [CrossRef] [PubMed]
- Selenska-Pobell, S.; Merroun, M. Accumulation of Heavy Metals by Micro-organisms: Biomineralization and Nanocluster Formation. In Prokaryotic Cell Wall Compounds; Springer: Berlin, Germany, 2010; pp. 483–500. [Google Scholar]
- Pacton, M.; Wacey, D.; Corinaldesi, C.; Tangherlini, M.; Kilburn, M.R.; Gorin, G.E.; Danovaro, R.; Vasconcelos, C. Viruses as new agents of organomineralization in the geological record. Nat. Commun. 2014, 5, 4298. [Google Scholar] [CrossRef] [PubMed]
- Lin, W.; Paterson, G.A.; Zhu, Q.; Wang, Y.; Kopylova, E.; Li, Y.; Knight, R.; Bazylinski, D.A.; Zhu, R.; Kirschvink, J.L.; et al. Origin of microbial biomineralization and magnetotaxis during the Archean. Proc. Natl. Acad. Sci. USA 2017, 114, 2171–2176. [Google Scholar] [CrossRef] [PubMed]
- Butardo, V.M.; Sreenivasulu, N. Tailoring Grain Storage Reserves for a Healthier Rice Diet and its Comparative Status with Other Cereals. Int. Rev. Cell Mol. Biol. 2016, 323, 31–70. [Google Scholar] [PubMed]
- Adeva-Andany, M.M.; González-Lucán, M.; Donapetry-García, C.; Fernández-Fernández, C.; Ameneiros-Rodríguez, E. Glycogen metabolism in humans. BBA Clin. 2016, 5, 85–100. [Google Scholar] [CrossRef] [PubMed]
- Elieh-Ali-Komi, D.; Hamblin, M.R. Chitin and Chitosan: Production and Application of Versatile Biomedical Nanomaterials. Int. J. Adv. Res. (Indore) 2016, 4, 411–427. [Google Scholar]
- Hansma, H.G. Possible origin of life between mica sheets: Does life imitate mica? J. Biomol. Struct. Dyn. 2013, 31, 888–895. [Google Scholar] [CrossRef] [PubMed]
- Nidhin, M.; Sreeram, K.J.; Nair, B.U. Polysaccharide films as templates in the synthesis of hematite nanostructures with special properties. Appl. Surf. Sci. 2012, 258, 5179–5184. [Google Scholar] [CrossRef]
- Allen, C.C.; Westall, F.; Schelble, R.T. Importance of a martian hematite site for astrobiology. Astrobiology 2001, 1, 111–123. [Google Scholar] [CrossRef] [PubMed]
- Murai, K.; Kinoshita, T.; Nagata, K.; Higuchi, M. Mineralization of Calcium Carbonate on Multifunctional Peptide Assembly Acting as Mineral Source Supplier and Template. Langmuir 2016, 32, 9351–9359. [Google Scholar] [CrossRef] [PubMed]
- Nonoyama, T.; Tanaka, M.; Kinoshita, T.; Nagata, F.; Sato, K.; Kato, K. Morphology control of calcium phosphate by mineralization on the β-sheet peptide template. Chem. Commun. 2010, 46, 6983. [Google Scholar] [CrossRef] [PubMed]
- Ribeiro, S.; Radvar, E.; Shi, Y.; Borges, J.; Pirraco, R.P.; Leonor, I.B.; Mano, J.F.; Reis, R.L.; Mata, Á.; Azevedo, H.S. Nanostructured interfacial self-assembled peptide–polymer membranes for enhanced mineralization and cell adhesion. Nanoscale 2017, 9, 13670–13682. [Google Scholar] [CrossRef] [PubMed]
- Elham, R.; Remzi, B.C.; Helena, A. Self-assembled peptide-polymer hydrogels as a 3D nanofiber substrate for biomimetic hydroxyapatite mineralization. Front. Bioeng. Biotechnol. 2016, 4. [Google Scholar] [CrossRef]
- Li, Q.-L.; Ning, T.-Y.; Cao, Y.; Zhang, W.-B.; Mei, M.L.; Chu, C.H. A novel self-assembled oligopeptide amphiphile for biomimetic mineralization of enamel. BMC Biotechnol. 2014, 14, 32. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Ouyang, Z.; Ren, Z.; Li, J.; Zhang, P.; Wei, G.; Su, Z. Self-assembled peptide nanofibers on graphene oxide as a novel nanohybrid for biomimetic mineralization of hydroxyapatite. Carbon N. Y. 2015, 89, 20–30. [Google Scholar] [CrossRef]
- Chan, M.; Cleaves, H.J., II; Boston, P. What Would Earth Be Like Without Life? EOS 2018, 99. [Google Scholar] [CrossRef]

© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gillams, R.J.; Jia, T.Z. Mineral Surface-Templated Self-Assembling Systems: Case Studies from Nanoscience and Surface Science towards Origins of Life Research. Life 2018, 8, 10. https://doi.org/10.3390/life8020010
Gillams RJ, Jia TZ. Mineral Surface-Templated Self-Assembling Systems: Case Studies from Nanoscience and Surface Science towards Origins of Life Research. Life. 2018; 8(2):10. https://doi.org/10.3390/life8020010
Chicago/Turabian StyleGillams, Richard J., and Tony Z. Jia. 2018. "Mineral Surface-Templated Self-Assembling Systems: Case Studies from Nanoscience and Surface Science towards Origins of Life Research" Life 8, no. 2: 10. https://doi.org/10.3390/life8020010
APA StyleGillams, R. J., & Jia, T. Z. (2018). Mineral Surface-Templated Self-Assembling Systems: Case Studies from Nanoscience and Surface Science towards Origins of Life Research. Life, 8(2), 10. https://doi.org/10.3390/life8020010