The tRNA Elbow in Structure, Recognition and Evolution
Abstract
:1. Introduction
| Name | Type/Region of Polymer | Function | Mode of Interaction * | PDB Code | Ref. |
|---|---|---|---|---|---|
| Ribosome A site | 23S RNA; helix 38 and others | Translation | H,V | 4V6F | [13] |
| Ribosome P site | L5 protein and others | Translation | H,V | 4V51 | [10] |
| Ribosome E site | 23S RNA; L1 stalk and others | Translation | S | 4V4I | [11] |
| RNase P | RNA: J11/12–J12/11 | tRNA modification: 5′ end maturation | S | 3Q1Q | [14] |
| T-box riboswitch | RNA: Stem I distal region | Amino acid surveillance: tRNA binding, recognition of aminoacylation, and genetic switching | S | 4LCK | [15] |
| LeuRS | protein | tRNA aminoacylation | H, V | 2V0G | [16] |
| ValRS | protein | tRNA aminoacylation | H, V | 1GAX | [17] |
| GatDE | protein | Aminoacyl-tRNA transamidation | unknown | 2D6F | [18] |
| GatCAB | protein | Aminoacyl-tRNA transamidation | H, V | 3AL0 | [19] |
| RNase Z | protein | tRNA modification: 3′ end maturation | H, V | 4GCW | [20] |
| DusC | protein | tRNA modification: reduction of D-loop U16/U20 to dihydrouridine | H, V | 4YCP | [21] |
| TruB | protein | tRNA modification: pseudouridylation of T-loop U55 to Ψ | T-loop extraction | 1K8W | [22] |
| ArcTgt | protein | tRNA modification: transglycosylation of D-loop G15 to PreQ0. | D-loop extraction | 1J2B | [23] |
| CCA-adding enzyme | protein: tail domain | tRNA maturation; addition of 3′ CCA trinucleotide | H,V | 1SZ1 | [24] |
| CC-adding enzyme | protein: tail domain | tRNA maturation; addition of 3′ C nucleotides | H,V | 3WFR | [25] |
| A-adding enzyme | protein | tRNA maturation; addition of 3′ A nucleotide | H,V | 4X0B | [26] |
2. Anatomy of the tRNA Elbow
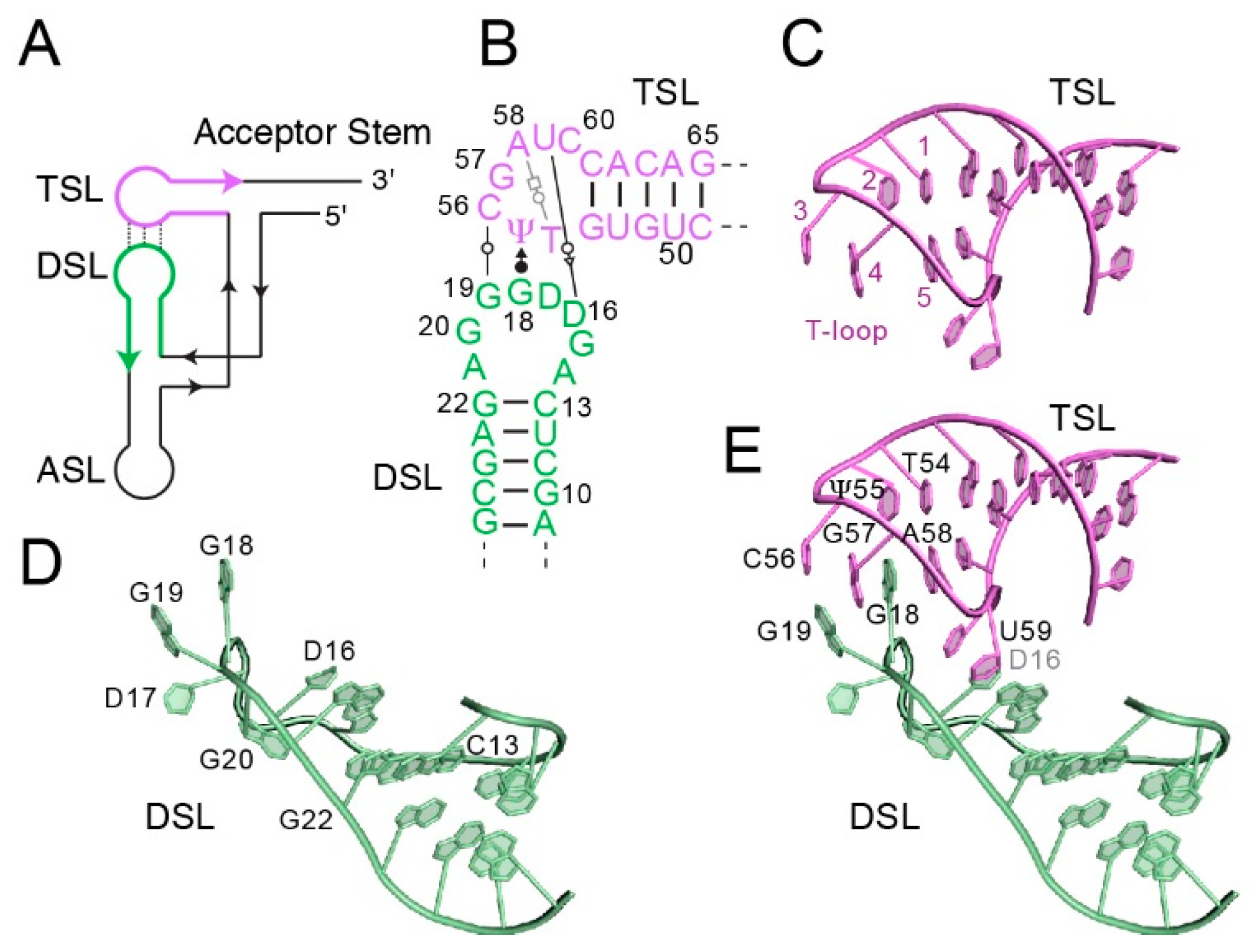
3. Engagement of the tRNA Elbow by the Ribosome
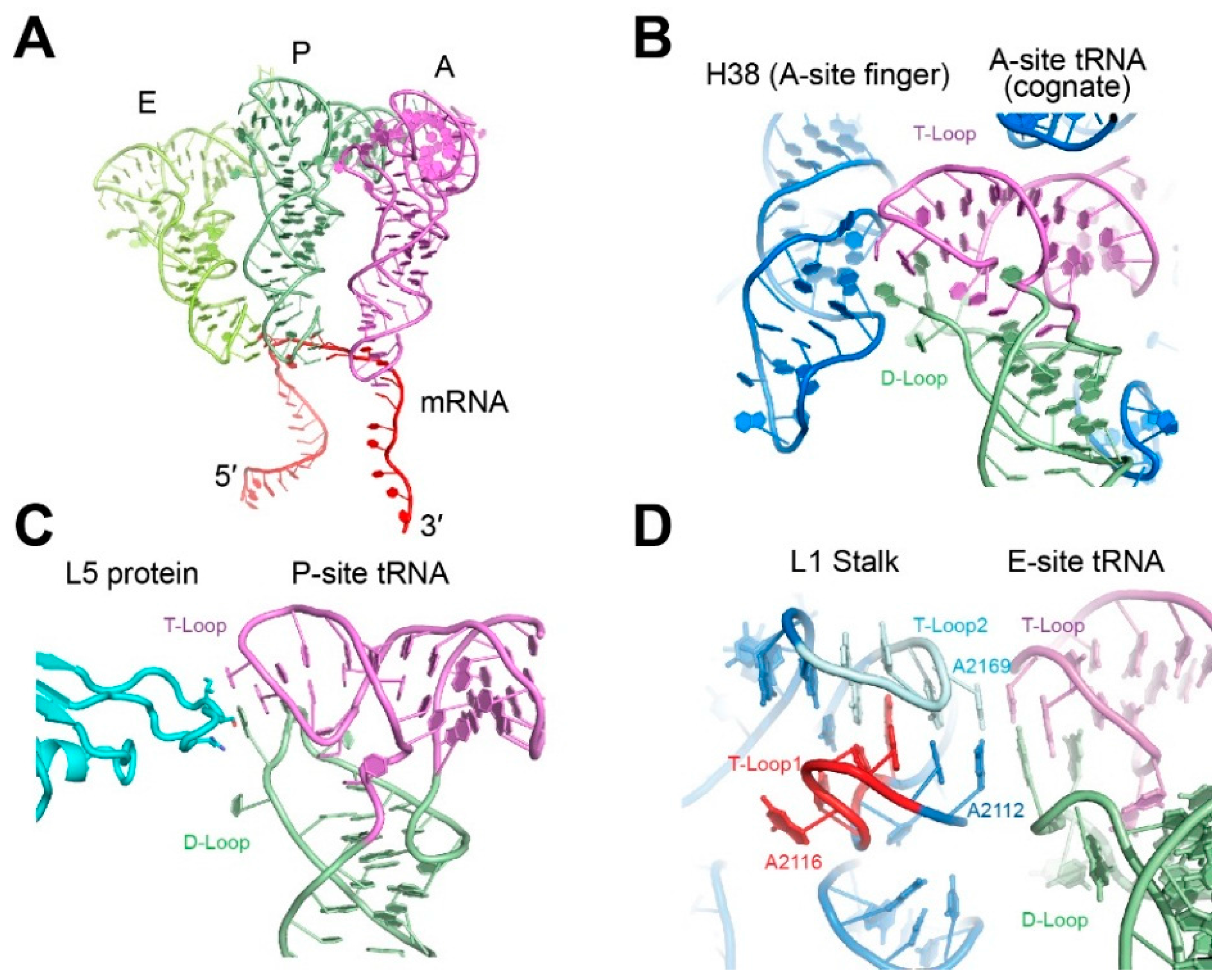
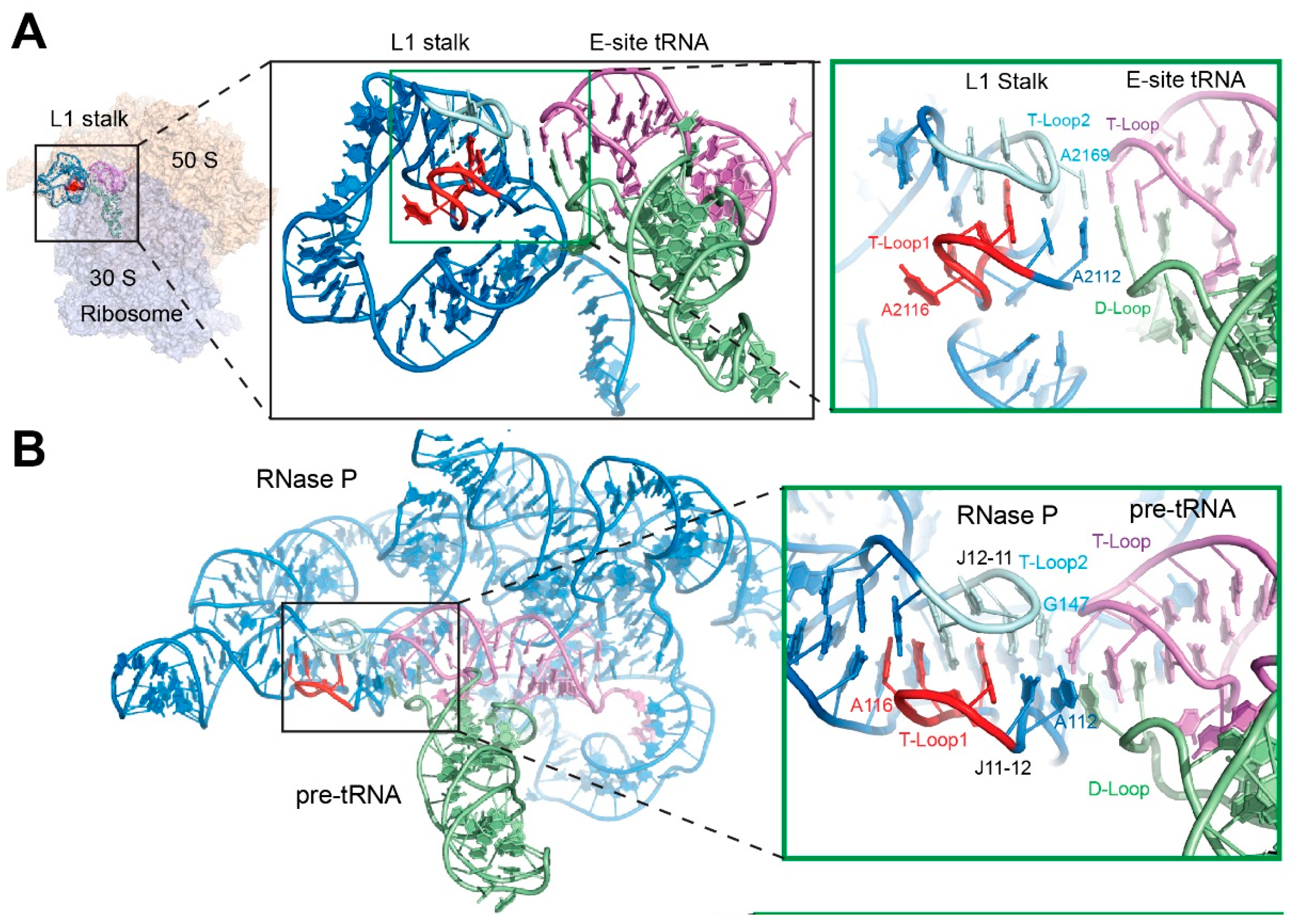
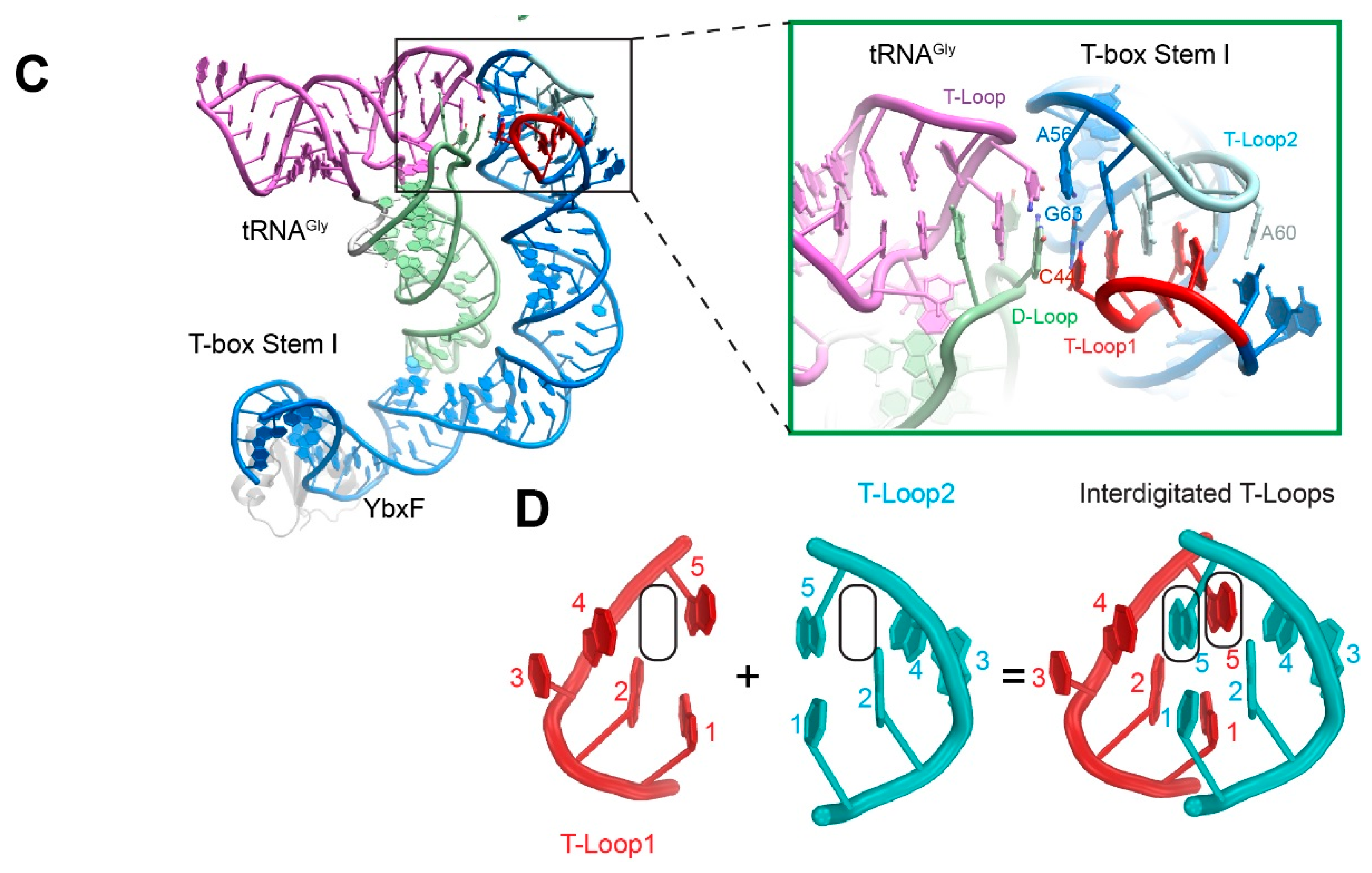
4. Recognition of the tRNA Elbow by Non-Coding RNAs
5. Diversity in Protein Recognition of the tRNA Elbow
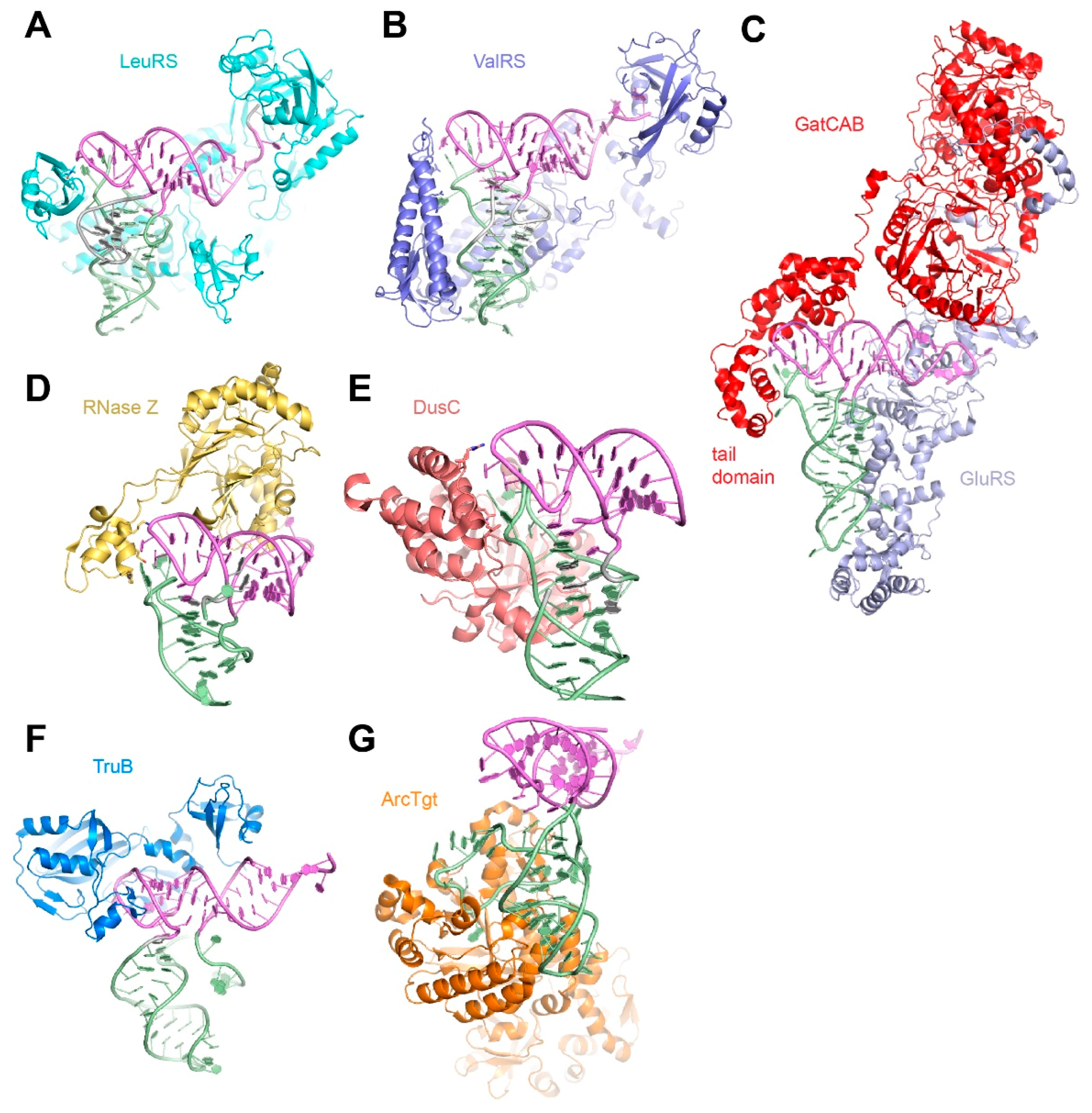
6. The tRNA Elbow in Evolution
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Kim, S.-H.; Suddath, F.L.; Quigley, G.J.; McPherson, A.; Sussman, J.L.; Wang, A.H.J.; Seeman, N.C.; Rich, A. Three-dimensional tertiary structure of yeast phenylalanine transfer RNA. Science 1974, 185, 435–440. [Google Scholar] [CrossRef] [PubMed]
- Robertus, J.D.; Ladner, J.E.; Finch, J.T.; Rhodes, D.; Brown, R.S.; Clark, B.F.; Klug, A. Structure of yeast phenylalanine tRNA at 3 Å resolution. Nature 1974, 250, 546–551. [Google Scholar] [CrossRef] [PubMed]
- Crick, F.H.C. The origin of the genetic code. J. Mol. Biol. 1968, 38, 367–379. [Google Scholar] [CrossRef]
- Colussi, T.M.; Costantino, D.A.; Hammond, J.A.; Ruehle, G.M.; Nix, J.C.; Kieft, J.S. The structural basis of transfer RNA mimicry and conformational plasticity by a viral RNA. Nature 2014, 511, 366–369. [Google Scholar] [CrossRef] [PubMed]
- Hammond, J.A.; Rambo, R.P.; Filbin, M.E.; Kieft, J.S. Comparison and functional implications of the 3D architectures of viral tRNA-like structures. RNA 2009, 15, 294–307. [Google Scholar] [CrossRef] [PubMed]
- Jones, C.P.; Cantara, W.A.; Olson, E.D.; Musier-Forsyth, K. Small-angle X-ray scattering-derived structure of the HIV-1 5′ UTR reveals 3D tRNA mimicry. Proc. Natl. Acad. Sci. USA 2014, 111, 3395–3400. [Google Scholar] [CrossRef] [PubMed]
- Zuo, X.; Wang, J.; Yu, P.; Eyler, D.; Xu, H.; Starich, M.R.; Tiede, D.M.; Simon, A.E.; Shapiro, B.A.; Wang, Y.X. Solution structure of the cap-independent translational enhancer and ribosome-binding element in the 3′ UTR of turnip crinkle virus. Proc. Natl. Acad. Sci. USA 2010, 107, 1385–1390. [Google Scholar] [CrossRef] [PubMed]
- Schlunzen, F.; Zarivach, R.; Harms, J.; Bashan, A.; Tocilj, A.; Albrecht, R.; Yonath, A.; Franceschi, F. Structural basis for the interaction of antibiotics with the peptidyl transferase centre in eubacteria. Nature 2001, 413, 814–821. [Google Scholar] [CrossRef] [PubMed]
- Lin, J.; Gagnon, M.G.; Bulkley, D.; Steitz, T.A. Conformational changes of elongation factor G on the ribosome during tRNA translocation. Cell 2015, 160, 219–227. [Google Scholar] [CrossRef] [PubMed]
- Selmer, M.; Dunham, C.M.; Murphy, F.V.T.; Weixlbaumer, A.; Petry, S.; Kelley, A.C.; Weir, J.R.; Ramakrishnan, V. Structure of the 70S ribosome complexed with mRNA and tRNA. Science 2006, 313, 1935–1942. [Google Scholar] [CrossRef] [PubMed]
- Korostelev, A.; Trakhanov, S.; Laurberg, M.; Noller, H. Crystal structure of a 70S ribosome-tRNA complex reveals functional interactions and rearrangements. Cell 2006, 126, 1065–1077. [Google Scholar] [CrossRef] [PubMed]
- Dunkle, J.A.; Wang, L.; Feldman, M.B.; Pulk, A.; Chen, V.B.; Kapral, G.J.; Noeske, J.; Richardson, J.S.; Blanchard, S.C.; Cate, J.H.D. Structures of the bacterial ribosome in classical and hybrid states of tRNA binding. Science 2011, 332, 981–984. [Google Scholar] [CrossRef] [PubMed]
- Jenner, L.; Demeshkina, N.; Yusupova, G.; Yusupov, M. Structural rearrangements of the ribosome at the tRNA proofreading step. Nat. Struct. Mol. Biol. 2010, 17, 1072–1078. [Google Scholar] [CrossRef] [PubMed]
- Reiter, N.J.; Osterman, A.; Torres-Larios, A.; Swinger, K.K.; Pan, T.; Mondragón, A. Structure of a bacterial ribonuclease P holoenzyme in complex with tRNA. Nature 2010, 468, 784–789. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Ferré-D’Amaré, A.R. Co-crystal structure of a T-box riboswitch stem I domain in complex with its cognate tRNA. Nature 2013, 500, 363–366. [Google Scholar] [CrossRef] [PubMed]
- Rock, F.L.; Mao, W.; Yaremchuk, A.; Tukalo, M.; Crepin, T.; Zhou, H.; Zhang, Y.K.; Hernandez, V.; Akama, T.; Alley, M.R.K.; et al. An antifungal agent inhibits an aminoacyl-tRNA synthetase by trapping tRNA in the editing eite. Science 2007, 316, 1759–1761. [Google Scholar] [CrossRef] [PubMed]
- Fukai, S.; Nureki, O.; Sekine, S.; Shimada, A.; Tao, J.; Vassylyev, D.G.; Yokoyama, S. Structural basis for double-sieve discrimination of L-valine from L-isoleucine and L-threonine by the complex of tRNA(Val) and valyl-tRNA synthetase. Cell 2000, 103, 793–803. [Google Scholar] [CrossRef]
- Oshikane, H.; Sheppard, K.; Fukai, S.; Nakamura, Y.; Ishitani, R.; Numata, T.; Sherrer, R.L.; Feng, L.; Schmitt, E.; Nureki, O.; et al. Structural basis of RNA-dependent recruitment of glutamine to the genetic code. Science 2006, 312, 1950–1954. [Google Scholar] [CrossRef] [PubMed]
- Ito, T.; Yokoyama, S. Two enzymes bound to one transfer RNA assume alternative conformations for consecutive reactions. Nature 2010, 467, 612–616. [Google Scholar] [CrossRef] [PubMed]
- Pellegrini, O.; Li de la Sierra-Gallay, I.; Piton, J.; Gilet, L.; Condon, C. Activation of tRNA maturation by downstream uracil residues in B. subtilis. Structure 2012, 20, 1769–1777. [Google Scholar] [CrossRef] [PubMed]
- Byrne, R.T.; Jenkins, H.T.; Peters, D.T.; Whelan, F.; Stowell, J.; Aziz, N.; Kasatsky, P.; Rodnina, M.V.; Koonin, E.V.; Antson, A.A.; et al. Major reorientation of tRNA substrates defines specificity of dihydrouridine synthases. Proc. Natl. Acad. Sci. USA 2015, 112, 6033–6037. [Google Scholar] [CrossRef] [PubMed]
- Hoang, C.; Ferré-D’Amaré, A.R. Cocrystal structure of a tRNA Y55 pseudouridine synthase: Nucleotide flipping by an RNA-modifying enzyme. Cell 2001, 107, 929–939. [Google Scholar] [CrossRef]
- Ishitani, R.; Nureki, O.; Nameki, N.; Okada, N.; Nishimura, S.; Yokoyama, S. Alternative tertiary structure of tRNA for recognition by a posttranscriptional modification enzyme. Cell 2003, 113, 383–394. [Google Scholar] [CrossRef]
- Xiong, Y.; Steitz, T.A. Mechanism of transfer RNA maturation by CCA-adding enzyme without using an oligonucleotide template. Nature 2004, 430, 640–645. [Google Scholar] [CrossRef] [PubMed]
- Yamashita, S.; Takeshita, D.; Tomita, K. Translocation and rotation of tRNA during template-independent RNA polymerization by tRNA nucleotidyltransferase. Structure 2014, 22, 315–325. [Google Scholar] [CrossRef] [PubMed]
- Yamashita, S.; Martinez, A.; Tomita, K. Measurement of Acceptor-TPsiC Helix Length of tRNA for Terminal A76-Addition by A-Adding Enzyme. Structure 2015, 23, 830–842. [Google Scholar] [CrossRef] [PubMed]
- Chan, C.W.; Chetnani, B.; Mondragón, A. Structure and function of the T-loop structural motif in noncoding RNAs. Wiley Interdiscip. Rev. RNA 2013, 4, 507–522. [Google Scholar] [CrossRef] [PubMed]
- Krasilnikov, A.S.; Mondragón, A. On the occurrence of the T-loop RNA folding motif in large RNA molecules. RNA 2003, 9, 640–643. [Google Scholar] [CrossRef] [PubMed]
- Leontis, N.B.; Westhof, E. Geometric nomenclature and classification of RNA base pairs. RNA 2001, 7, 499–512. [Google Scholar] [CrossRef] [PubMed]
- The PyMOL Molecular Graphics System, Version 1.8 Schrödinger, LLC. Available online: https://www.pymol.org/ (accessed on 6 January 2016).
- Pan, D.; Kirillov, S.; Zhang, C.M.; Hou, Y.M.; Cooperman, B.S. Rapid ribosomal translocation depends on the conserved 18–55 base pair in P-site transfer RNA. Nat. Struct. Mol. Biol. 2006, 13, 354–359. [Google Scholar] [CrossRef] [PubMed]
- Atkins, J.F.; Bjork, G.R. A gripping tale of ribosomal frameshifting: Extragenic suppressors of frameshift mutations spotlight P-site realignment. Microbiol. Mol. Biol. Rev. 2009, 73, 178–210. [Google Scholar] [CrossRef] [PubMed]
- Herr, A.J.; Atkins, J.F.; Gesteland, R.F. Mutations which alter the elbow region of tRNA2Gly reduce T4 gene 60 translational bypassing efficiency. EMBO J. 1999, 18, 2886–2896. [Google Scholar] [CrossRef] [PubMed]
- Cornish, P.V.; Ermolenko, D.N.; Staple, D.W.; Hoang, L.; Hickerson, R.P.; Noller, H.F.; Ha, T. Following movement of the L1 stalk between three functional states in single ribosomes. Proc. Natl. Acad. Sci. USA 2009, 106, 2571–2576. [Google Scholar] [CrossRef] [PubMed]
- Randau, L.; Schröder, I.; Söll, D. Life without RNase P. Nature 2008, 453, 120–123. [Google Scholar] [CrossRef] [PubMed]
- Grundy, F.J.; Henkin, T.M. tRNA as a positive regulator of transcription antitermination in B. subtilis. Cell 1993, 74, 475–482. [Google Scholar] [CrossRef]
- Zhang, J.; Ferré-D’Amaré, A.R. Structure and mechanism of the T-box riboswitches. Wiley Interdiscip. Rev. RNA 2015, 6, 419–433. [Google Scholar] [CrossRef] [PubMed]
- Grigg, J.C.; Ke, A. Structural Determinants for Geometry and Information Decoding of tRNA by T Box Leader RNA. Structure 2013, 21, 2025–2032. [Google Scholar] [CrossRef] [PubMed]
- Grigg, J.C.; Chen, Y.; Grundy, F.J.; Henkin, T.M.; Pollack, L.; Ke, A. T box RNA decodes both the information content and geometry of tRNA to affect gene expression. Proc. Natl. Acad. Sci. USA 2013, 110, 7240–7245. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Ferré-D’Amaré, A.R. Direct evaluation of tRNA aminoacylation status by the T-box riboswitch using tRNA-mRNA stacking and steric readout. Mol. Cell 2014, 55, 148–155. [Google Scholar] [CrossRef] [PubMed]
- Lehmann, J.; Jossinet, F.; Gautheret, D. A universal RNA structural motif docking the elbow of tRNA in the ribosome, RNase P and T-box leaders. Nucleic Acids Res. 2013, 41, 5494–5502. [Google Scholar] [CrossRef] [PubMed]
- Burley, S.K.; Petsko, G.A. Weakly polar interactions in proteins. Adv. Protein Chem. 1988, 39, 125–192. [Google Scholar] [PubMed]
- Burley, S.K.; Petsko, G.A. Aromatic-aromatic interaction: A mechanism of protein structure stabilization. Science 1985, 229, 23–28. [Google Scholar] [CrossRef] [PubMed]
- Ibba, M.; Soll, D. Aminoacyl-tRNA synthesis. Annu. Rev. Biochem. 2000, 69, 617–650. [Google Scholar] [CrossRef] [PubMed]
- Levinger, L.; Serjanov, D. Pathogenesis-related mutations in the T-loops of human mitochondrial tRNAs affect 3′ end processing and tRNA structure. RNA Biol. 2012, 9, 283–291. [Google Scholar] [CrossRef] [PubMed]
- Levinger, L.; Hopkinson, A.; Desetty, R.; Wilson, C. Effect of changes in the flexible arm on tRNase Z processing kinetics. J. Biol. Chem. 2009, 284, 15685–15691. [Google Scholar] [CrossRef] [PubMed]
- Hopkinson, A.; Levinger, L. Effects of conserved D/T loop substitutions in the pre-tRNA substrate on tRNase Z catalysis. RNA Biol. 2008, 5, 104–111. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Hopper, A.K.; Phizicky, E.M. tRNA transfers to the limelight. Genes Dev. 2003, 17, 162–180. [Google Scholar] [CrossRef] [PubMed]
- Sun, F.J.; Caetano-Anolles, G. The origin and evolution of tRNA inferred from phylogenetic analysis of structure. J. Mol. Evol. 2008, 66, 21–35. [Google Scholar] [CrossRef] [PubMed]
- De Farias, S.T.; do Rego, T.G.; Jose, M.V. Evolution of transfer RNA and the origin of the translation system. Front. Genet. 2014, 5. [Google Scholar] [CrossRef] [PubMed]
- Weiner, A.M.; Maizels, N. tRNA-like structures tag the 3′ ends of genomic RNA molecules for replication: Implications for the origin of protein synthesis. Proc. Natl. Acad. Sci. USA 1987, 84, 7383–7387. [Google Scholar] [CrossRef] [PubMed]
- Marquéz, V.; Nierhaus, K.H.; Ribas de Pouplana, L.; Schimmel, P. tRNA and Synthetases. In Protein Synthesis and Ribosome Structure; Wiley-VCH Verlag GmbH & Co. KGaA: Weinheim, Germany, 2006; pp. 145–184. [Google Scholar]
- Sharma, M.R.; Koc, E.C.; Datta, P.P.; Booth, T.M.; Spremulli, L.L.; Agrawal, R.K. Structure of the mammalian mitochondrial ribosome reveals an expanded functional role for its component proteins. Cell 2003, 115, 97–108. [Google Scholar] [CrossRef]
- Brown, A.; Amunts, A.; Bai, X.C.; Sugimoto, Y.; Edwards, P.C.; Murshudov, G.; Scheres, S.H.W.; Ramakrishnan, V. Structure of the large ribosomal subunit from human mitochondria. Science 2014, 346, 718–722. [Google Scholar] [CrossRef] [PubMed]
- Sharma, M.R.; Booth, T.M.; Simpson, L.; Maslov, D.A.; Agrawal, R.K. Structure of a mitochondrial ribosome with minimal RNA. Proc. Natl. Acad. Sci. USA 2009, 106, 9637–9642. [Google Scholar] [CrossRef] [PubMed]
- Rossmanith, W. Of P and Z: Mitochondrial tRNA processing enzymes. Biochim. Biophys. Acta 2012, 1819, 1017–1026. [Google Scholar] [CrossRef] [PubMed]
- Sherwood, A.V.; Grundy, F.J.; Henkin, T.M. T box riboswitches in Actinobacteria: Translational regulation via novel tRNA interactions. Proc. Natl. Acad. Sci. USA 2015, 112, 1113–1118. [Google Scholar] [CrossRef] [PubMed]
- Wilusz, J.E.; Freier, S.M.; Spector, D.L. 3′ end processing of a long nuclear-retained noncoding RNA yields a tRNA-like cytoplasmic RNA. Cell 2008, 135, 919–932. [Google Scholar] [CrossRef] [PubMed]
- Orgel, L.E. Evolution of the genetic apparatus. J. Mol. Biol. 1968, 38, 381–393. [Google Scholar] [CrossRef]
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons by Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, J.; Ferré-D’Amaré, A.R. The tRNA Elbow in Structure, Recognition and Evolution. Life 2016, 6, 3. https://doi.org/10.3390/life6010003
Zhang J, Ferré-D’Amaré AR. The tRNA Elbow in Structure, Recognition and Evolution. Life. 2016; 6(1):3. https://doi.org/10.3390/life6010003
Chicago/Turabian StyleZhang, Jinwei, and Adrian R. Ferré-D’Amaré. 2016. "The tRNA Elbow in Structure, Recognition and Evolution" Life 6, no. 1: 3. https://doi.org/10.3390/life6010003
APA StyleZhang, J., & Ferré-D’Amaré, A. R. (2016). The tRNA Elbow in Structure, Recognition and Evolution. Life, 6(1), 3. https://doi.org/10.3390/life6010003







