Next-Generation-Sequencing-Based Simple Sequence Repeat (SSR) Marker Development and Linkage Mapping in Lentil (Lens culinaris L.)
Abstract
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Material
2.2. DNA Isolation and Sequencing
2.3. SSR Detection, Annotation, and Primer Design
2.4. Validation and Selection of SSR Primer Pairs
2.5. Construction of Genetic Linkage Maps
3. Results
3.1. Sequence Assembly and Simple Sequence Repeat Identification
3.2. Primer Design and SSR Validation
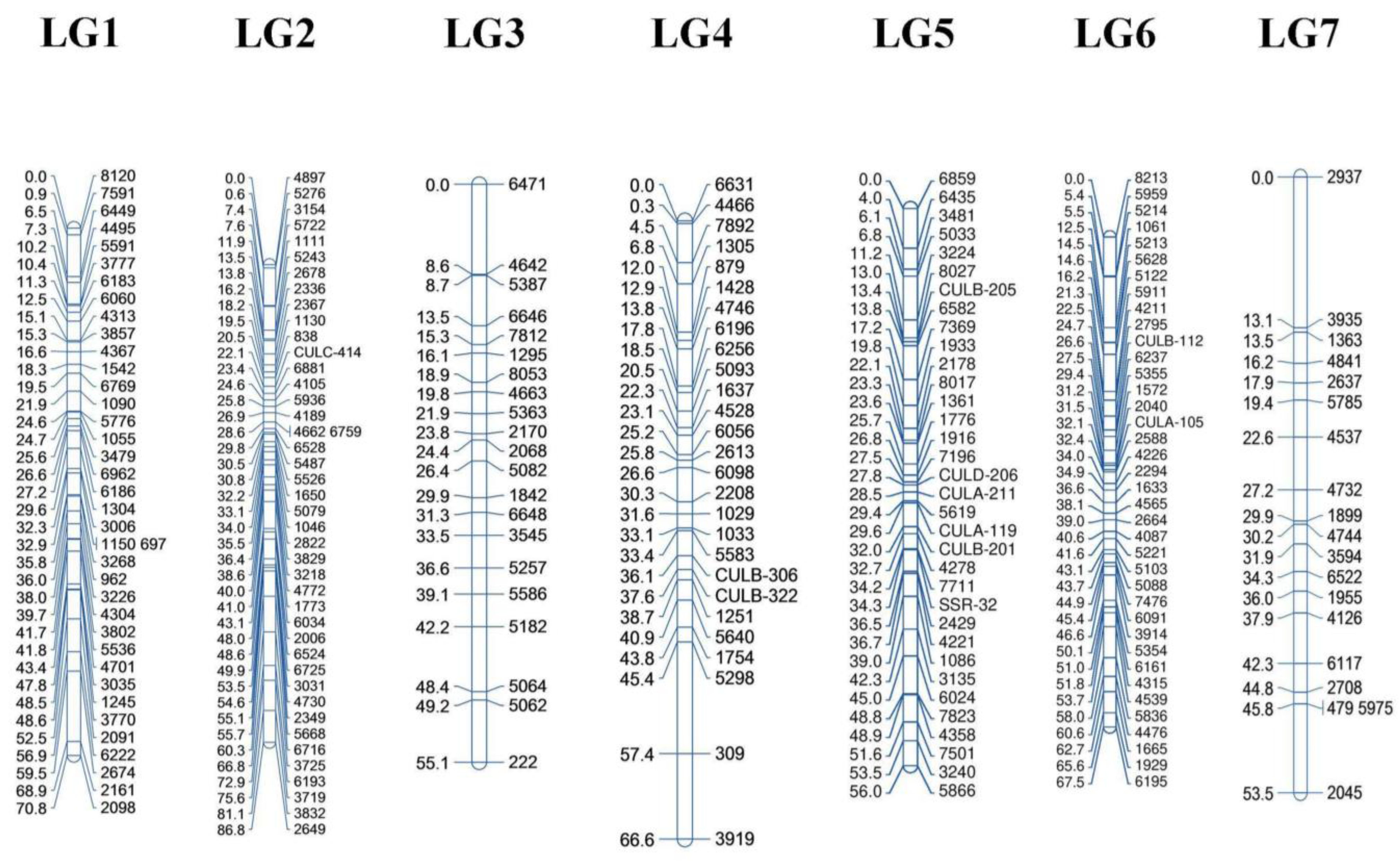
3.3. Construction of Genetic Linkage Map
4. Discussion
4.1. Identification of Novel SSR Markers
4.2. Construction of Genetic Linkage Map Using New SSR Markers
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Ford, R.; Taylor, P. Construction of an intraspecific linkage map of lentil (Lens culinaris ssp. culinaris). Theor. Appl. Genet. 2003, 107, 910–916. [Google Scholar]
- Gupta, M.; Verma, B.; Kumar, N.; Chahota, R.K.; Rathour, R.; Sharma, S.K.; Bhatia, S.; Sharma, T.R. Construction of intersubspecific molecular genetic map of lentil based on ISSR, RAPD and SSR markers. J. Genet. 2012, 91, 279–287. [Google Scholar] [CrossRef] [PubMed]
- Kumar, J.; Basu, P.; Srivastava, E.; Chaturvedi, S.; Nadarajan, N.; Kumar, S. Phenotyping of traits imparting drought tolerance in lentil. Crop Pasture Sci. 2012, 63, 547–554. [Google Scholar] [CrossRef]
- Idrissi, O.; Udupa, S.M.; Houasli, C.; De Keyser, E.; Van Damme, P.; De Riek, J. Genetic diversity analysis of Moroccan lentil (Lens culinaris Medik.) landraces using Simple Sequence Repeat and Amplified Fragment Length Polymorphisms reveals functional adaptation towards agro-environmental origins. Plant Breed. 2015, 134, 322–332. [Google Scholar] [CrossRef]
- Kaale, L.D.; Siddiq, M.; Hooper, S. Lentil (Lens culinaris Medik) as nutrient-rich and versatile food legume: A review. Legume Sci. 2022, 5, e169. [Google Scholar] [CrossRef]
- Coyne, C.J.; Kumar, S.; Wettberg, E.J.B.; Marques, E.; Berger, J.D.; Redden, R.J.; Ellis, T.H.N.; Brus, J.; Zablatzká, L.; Smýkal, P. Potential and limits of exploitation of crop wild relatives for pea, lentil, and chickpea improvement. Legume Sci. 2020, 2, e36. [Google Scholar] [CrossRef]
- Jiang, G.-L. Molecular Markers and Marker-Assisted Breeding in Plants; IntechOpen: London, UK, 2013. [Google Scholar] [CrossRef]
- Udupa, S.M.; Robertson, L.; Weigand, F.; Baum, M.; Kahl, G. Allelic variation at (TAA) n microsatellite loci in a world collection of chickpea (Cicer arietinum L.) germplasm. Mol. Gen. Genet. MGG 1999, 261, 354–363. [Google Scholar] [CrossRef]
- Hamwieh, A.; Udupa, S.M.; Sarker, A.; Jung, C.; Baum, M. Development of new microsatellite markers and their application in the analysis of genetic diversity in lentils. Breed. Sci. 2009, 59, 77–86. [Google Scholar] [CrossRef]
- Khazaei, H.; Podder, R.; Caron, C.T.; Kundu, S.S.; Diapari, M.; Vandenberg, A.; Bett, K.E. Marker–trait association analysis of iron and zinc concentration in lentil (Lens culinaris Medik.) seeds. Plant Genome 2017, 10, plantgenome2017.02.0007. [Google Scholar] [CrossRef]
- Reddy, M.R.K.; Rathour, R.; Kumar, N.; Katoch, P.; Sharma, T.R. Cross-genera legume SSR markers for analysis of genetic diversity in Lens species. Plant Breed. 2010, 129, 514–518. [Google Scholar] [CrossRef]
- Choudhury, A.; Deb, S.; Kharbyngar, B.; Rajpal, V.R.; Rao, S.R. Dissecting the plant genome: Through new generation molecular markers. Genet. Resour. Crop Evol. 2022, 69, 2661–2698. [Google Scholar] [CrossRef]
- Toklu, F.; Karaköy, T.; Haklı, E.; Bicer, T.; Brandolini, A.; Kilian, B.; Özkan, H. Genetic variation among lentil (Lens culinaris Medik) landraces from Southeast Turkey. Plant Breed. 2009, 128, 178–186. [Google Scholar] [CrossRef]
- Zaccardelli, M.; Lupo, F.; Piergiovanni, A.R.; Laghetti, G.; Sonnante, G.; Daminati, M.G.; Sparvoli, F.; Lioi, L. Characterization of Italian lentil (Lens culinaris Medik.) germplasm by agronomic traits, biochemical and molecular markers. Genet. Resour. Crop Evol. 2012, 59, 727–738. [Google Scholar] [CrossRef]
- Alghamdi, S.S.; Khan, A.M.; Ammar, M.H.; El-Harty, E.H.; Migdadi, H.M.; El-Khalik, S.M.A.; Al-Shameri, A.M.; Javed, M.M.; Al-Faifi, S.A. Phenological, Nutritional and Molecular Diversity Assessment among 35 Introduced Lentil (Lens culinaris Medik.) Genotypes Grown in Saudi Arabia. Int. J. Mol. Sci. 2014, 15, 277–295. [Google Scholar] [CrossRef] [PubMed]
- Tewari, K.; Dikshit, H.; Jain, N.; Kumari, J.; Singh, D. Genetic differentiation of wild and cultivated Lens based on molecular markers. J. Plant Biochem. Biotechnol. 2012, 21, 198–204. [Google Scholar] [CrossRef]
- Joshi, M.; Verma, S.; Singh, J.; Barh, A. Genetic diversity assessment in lentil (Lens culinaris Medikus) genotypes through ISSR marker. Bioscan 2013, 8, 1529–1532. [Google Scholar]
- Khazaei, H.; Caron, C.T.; Fedoruk, M.; Diapari, M.; Vandenberg, A.; Coyne, C.J.; McGee, R.; Bett, K.E. Genetic diversity of cultivated lentil (Lens culinaris Medik.) and its relation to the world’s agro-ecological zones. Front. Plant Sci. 2016, 7, 1093. [Google Scholar] [CrossRef]
- Sharma, R.; Chaudhary, L.; Kumar, M.; Panwar, N. Analysis of genetic parameters and trait relationship for seed yield and its attributing components in lentil (Lens culinaris Medik.). Legume Res. Int. J. 2022, 45, 1344–1350. [Google Scholar] [CrossRef]
- Abo-Elwafa, A.; Murai, K.; Shimada, T. Intra-and inter-specific variations in Lens revealed by RAPD markers. Theor. Appl. Genet. 1995, 90, 335–340. [Google Scholar] [CrossRef]
- Datta, S.; Tiwari, S.; Kaashyap, M.; Gupta, P.P.; Choudhury, P.R.; Kumari, J.; Kumar, S. Genetic similarity analysis in lentil using cross-genera legume sequence tagged microsatellite site markers. Crop Sci. 2011, 51, 2412–2422. [Google Scholar] [CrossRef]
- Varshney, R.; Bertioli, D.; Moretzsohn, M.d.C.; Vadez, V.; Krishnamurthy, L.; Aruna, R.; Nigam, S.; Moss, B.; Seetha, K.; Ravi, K. The first SSR-based genetic linkage map for cultivated groundnut (Arachis hypogaea L.). Theor. Appl. Genet. 2009, 118, 729–739. [Google Scholar] [CrossRef] [PubMed]
- Hamwieh, A.; Udupa, S.; Choumane, W.; Sarker, A.; Dreyer, F.; Jung, C.; Baum, M. A genetic linkage map of Lens sp. based on microsatellite and AFLP markers and the localization of fusarium vascular wilt resistance. Theor. Appl. Genet. 2005, 110, 669–677. [Google Scholar] [CrossRef] [PubMed]
- Verma, P.; Sharma, T.R.; Srivastava, P.S.; Abdin, M.Z.; Bhatia, S. Exploring genetic variability within lentil (Lens culinaris Medik.) and across related legumes using a newly developed set of microsatellite markers. Mol. Biol. Rep. 2014, 41, 5607–5625. [Google Scholar] [CrossRef] [PubMed]
- Andeden, E.E.; Baloch, F.S.; Çakır, E.; Toklu, F.; Özkan, H. Development, characterization and mapping of microsatellite markers for lentil (Lens culinaris Medik.). Plant Breed. 2015, 134, 589–598. [Google Scholar] [CrossRef]
- Bakir, M.; Kahraman, A. Development of New SSR (Simple Sequence Repeat) Markers for Lentils (Lens culinaris Medik.) from Genomic Library Enriched with AG and AC Microsatellites. Biochem. Genet. 2019, 57, 338–353. [Google Scholar] [CrossRef]
- Havey, M.; Muehlbauer, F. Linkages between restriction fragment length, isozyme, and morphological markers in lentil. Theor. Appl. Genet. 1989, 77, 395–401. [Google Scholar] [CrossRef]
- Eujayl, I.; Baum, M.; Powell, W.; Erskine, W.; Pehu, E. A genetic linkage map of lentil (Lens sp.) based on RAPD and AFLP markers using recombinant inbred lines. Theor. Appl. Genet. 1998, 97, 83–89. [Google Scholar] [CrossRef]
- Rubeena, T.P.; Ford, R.; Taylor, P. Molecular mapping the lentil (Lens culinaris ssp. culinaris) genome. TAG Theor. Appl. Genet. Theor. Angew. Genet. 2003, 107, 910–916. [Google Scholar] [CrossRef]
- Duran, Y.; Fratini, R.; Garcia, P.; Perez De La Vega, M. An intersubspecific genetic map of Lens. Theor. Appl. Genet. 2004, 108, 1265–1273. [Google Scholar] [CrossRef]
- Tullu, A.; Tar’an, B.; Warkentin, T.; Vandenberg, A. Construction of an intraspecific linkage map and QTL analysis for earliness and plant height in lentil. Crop Sci. 2008, 48, 2254–2264. [Google Scholar] [CrossRef]
- Tanyolac, B.; Ozatay, S.; Kahraman, A.; Muehlbauer, F. Linkage mapping of lentil (Lens culinaris L.) genome using recombinant inbred lines revealed by AFLP, ISSR, RAPD and some morphologic markers. J. Agric. Biotech. Sustain. Dev. 2010, 2, 1–6. [Google Scholar]
- Kaur, S.; Cogan, N.O.I.; Stephens, A.; Noy, D.; Butsch, M.; Forster, J.W.; Materne, M. EST-SNP discovery and dense genetic mapping in lentil (Lens culinaris Medik.) enable candidate gene selection for boron tolerance. Theor. Appl. Genet. 2014, 127, 703–713. [Google Scholar] [CrossRef] [PubMed]
- Aldemir, S.; Ateş, D.; TEMEL, H.Y.; Yağmur, B.; Alsaleh, A.; Kahriman, A.; Özkan, H.; Vandenberg, A.; Tanyolac, M.B. QTLs for iron concentration in seeds of the cultivated lentil (Lens culinaris Medic.) via genotyping by sequencing. Turk. J. Agric. For. 2017, 41, 243–255. [Google Scholar] [CrossRef]
- Mane, R.; Katoch, M.; Singh, M.; Sharma, R.; Sharma, T.; Chahota, R. Identification of genomic regions associated with early plant vigour in lentil (Lens culinaris). J. Genet. 2020, 99, 21. [Google Scholar] [CrossRef] [PubMed]
- Haile, T.A.; Stonehouse, R.; Weller, J.L.; Bett, K.E. Genetic basis for lentil adaptation to summer cropping in northern temperate environments. Plant Genome 2021, 14, e20144. [Google Scholar] [CrossRef]
- Gela, T.S.; Koh, C.S.; Caron, C.T.; Chen, L.-A.; Vandenberg, A.; Bett, K.E. QTL mapping of lentil anthracnose (Colletotrichum lentis) resistance from Lens ervoides accession IG 72815 in an interspecific RIL population. Euphytica 2021, 217, 70. [Google Scholar] [CrossRef]
- Kumar, J.; Sen Gupta, D.; Baum, M.; Varshney, R.K.; Kumar, S. Genomics-assisted lentil breeding: Current status and future strategies. Legume Sci. 2021, 3, e71. [Google Scholar] [CrossRef]
- Bett, K.; Ramsay, L.; Sharpe, A.; Cook, D.; Penmetsa, R.; Verma, N. Lentil genome sequencing: Establishing a comprehensive platform for molecular breeding. In Proceedings of International Food Legumes Research Conference (IFLRC-VI) and ICCLG-VII; Crop Development Center: Saskatoon, SK, Canada, 2014. [Google Scholar]
- Winter, P.; Benko-Iseppon, A.-M.; Hüttel, B.; Ratnaparkhe, M.; Tullu, A.; Sonnante, G.; Pfaff, T.; Tekeoglu, M.; Santra, D.; Sant, V. A linkage map of the chickpea (Cicer arietinum L.) genome based on recombinant inbred lines from a C. arietinum × C. reticulatum cross: Localization of resistance genes for fusarium wilt races 4 and 5. Theor. Appl. Genet. 2000, 101, 1155–1163. [Google Scholar] [CrossRef]
- Kujur, A.; Bajaj, D.; Saxena, M.S.; Tripathi, S.; Upadhyaya, H.D.; Gowda, C.L.; Singh, S.; Jain, M.; Tyagi, A.K.; Parida, S.K. Functionally relevant microsatellite markers from chickpea transcription factor genes for efficient genotyping applications and trait association mapping. DNA Res. 2013, 20, 355–374. [Google Scholar] [CrossRef]
- Zalapa, J.E.; Cuevas, H.; Zhu, H.; Steffan, S.; Senalik, D.; Zeldin, E.; McCown, B.; Harbut, R.; Simon, P. Using next-generation sequencing approaches to isolate simple sequence repeat (SSR) loci in the plant sciences. Am. J. Bot. 2012, 99, 193–208. [Google Scholar] [CrossRef]
- Kaur, S.; Cogan, N.O.; Pembleton, L.W.; Shinozuka, M.; Savin, K.W.; Materne, M.; Forster, J.W. Transcriptome sequencing of lentil based on second-generation technology permits large-scale unigene assembly and SSR marker discovery. BMC Genom. 2011, 12, 265. [Google Scholar] [CrossRef] [PubMed]
- Doyle, J.J.; Doyle, J.L. A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem. Bull. 1987, 19, 11–15. [Google Scholar]
- You, F.M.; Huo, N.; Gu, Y.Q.; Luo, M.-c.; Ma, Y.; Hane, D.; Lazo, G.R.; Dvorak, J.; Anderson, O.D. BatchPrimer3: A high throughput web application for PCR and sequencing primer design. BMC Bioinform. 2008, 9, 253. [Google Scholar] [CrossRef]
- Schuelke, M. An economic method for the fluorescent labeling of PCR fragments. Nat. Biotechnol. 2000, 18, 233–234. [Google Scholar] [CrossRef] [PubMed]
- Van Ooijen, J. Multipoint maximum likelihood mapping in a full-sib family of an outbreeding species. Genet. Res. 2011, 93, 343–349. [Google Scholar] [CrossRef] [PubMed]
- Kosambi, D. Efficient mapping of a female sterile gene in wheat (Triticum aestivum L.). Ann Eugen. 1944, 12, 172–175. [Google Scholar] [CrossRef]
- Voorrips, R.E. MapChart: Software for the graphical presentation of linkage maps and QTLs. J. Hered. 2002, 93, 77–78. [Google Scholar] [CrossRef]
- Sharma, S. Prebreeding Using Wild Species for Genetic Enhancement of Grain Legumes at ICRISAT. Crop Sci. 2017, 57, 1132–1144. [Google Scholar] [CrossRef]
- Wu, X.; Li, N.; Hao, J.; Hu, J.; Zhang, X.; Blair, M.W. Genetic Diversity of Chinese and Global Pea (Pisum sativum L.) Collections. Crop Sci. 2017, 57, 1574–1584. [Google Scholar] [CrossRef]
- Abdallah, F.; Kumar, S.; Amri, A.; Mentag, R.; Kehel, Z.; Mejri, R.K.; Triqui, Z.E.A.; Hejjaoui, K.; Baum, M.; Amri, M. Wild Lathyrus species as a great source of resistance for introgression into cultivated grass pea (Lathyrus sativus L.) against broomrape weeds (Orobanche crenata Forsk. and Orobanche foetida Poir.). Crop Sci. 2021, 61, 263–276. [Google Scholar] [CrossRef]
- Semagn, K.; Bjornstad, A.; Xu, Y. The genetic dissection of quantitative traits in crops. Electron. J. Biotechnol. 2010, 13, 16–17. [Google Scholar] [CrossRef]
- Desta, Z.A.; Ortiz, R. Genomic selection: Genome-wide prediction in plant improvement. Trends Plant Sci. 2014, 19, 592–601. [Google Scholar] [CrossRef] [PubMed]
- Semagn, K.; Bjornstad, A.; Ndjiondjop, M.N. An overview of molecular marker methods for plants. Afr. J. Biotechnol. 2006, 5, 2540–2568. [Google Scholar]
- Singh, D.; Singh, C.K.; Tribuvan, K.U.; Tyagi, P.; Taunk, J.; Tomar, R.S.S.; Kumari, S.; Tripathi, K.; Kumar, A.; Gaikwad, K.; et al. Development, Characterization, and Cross Species/Genera Transferability of Novel EST-SSR Markers in Lentil, with Their Molecular Applications. Plant Mol. Biol. Rep. 2020, 38, 114–129. [Google Scholar] [CrossRef]
- Bakir, M. Transferability of newly developed genomic lentil SSR markers to Cicer species. Legume Res. 2019, 42, 479–484. [Google Scholar] [CrossRef]
- Verma, P.; Shah, N.; Bhatia, S. Development of an expressed gene catalogue and molecular markers from the de novo assembly of short sequence reads of the lentil (Lens culinaris Medik.) transcriptome. Plant Biotechnol. J. 2013, 11, 894–905. [Google Scholar] [CrossRef]
- Verma, P.; Goyal, R.; Chahota, R.K.; Sharma, T.R.; Abdin, M.Z.; Bhatia, S. Construction of a Genetic Linkage Map and Identification of QTLs for Seed Weight and Seed Size Traits in Lentil (Lens culinaris Medik.). PLoS ONE 2015, 10, e0139666. [Google Scholar] [CrossRef]



| Type of SSRs | Number of SSRs | % |
|---|---|---|
| Dinucleotides | 4643 | 53.38 |
| Trinucleotides | 2643 | 30.38 |
| Tetranucleotides | 527 | 6.59 |
| Pentanucleotides | 278 | 3.19 |
| Hexanucleotides | 606 | 6.96 |
| Total | 8697 | ~100.00 |
| a SSR Motifs | Number of SSRs (F ≥ 10) | Motif Frequency (%) |
|---|---|---|
| AG | 727 | 15.65 |
| AT | 507 | 10.91 |
| CT | 587 | 12.64 |
| GA | 815 | 17.55 |
| TC | 1013 | 21.81 |
| AAT | 279 | 10.55 |
| TTA | 285 | 10.78 |
| AAAT | 80 | 15.18 |
| Linkage Groups | No. of SSRs | Pct. of SSRs (%) | Length (cM) | Marker Density (cM) a | Max. Gap Length (cM) b |
|---|---|---|---|---|---|
| LG1 | 38 | 17.27 | 70.79 | 1.86 | 9.36 |
| LG2 | 43 | 19.55 | 86.84 | 2.01 | 6.73 |
| LG3 | 21 | 9.55 | 55.13 | 2.62 | 8.64 |
| LG4 | 27 | 12.27 | 66.61 | 2.46 | 11.92 |
| LG5 | 34 | 15.45 | 56.00 | 1.64 | 4.41 |
| LG6 | 38 | 17.27 | 67.50 | 1.77 | 7.07 |
| LG7 | 19 | 8.64 | 53.46 | 2.81 | 13.11 |
| Total-Mean | 220 | 100.00 | 456.33 | 2.16 * | 8.74 * |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Topu, M.; Sesiz, U.; Bektaş, H.; Toklu, F.; Özkan, H. Next-Generation-Sequencing-Based Simple Sequence Repeat (SSR) Marker Development and Linkage Mapping in Lentil (Lens culinaris L.). Life 2023, 13, 1579. https://doi.org/10.3390/life13071579
Topu M, Sesiz U, Bektaş H, Toklu F, Özkan H. Next-Generation-Sequencing-Based Simple Sequence Repeat (SSR) Marker Development and Linkage Mapping in Lentil (Lens culinaris L.). Life. 2023; 13(7):1579. https://doi.org/10.3390/life13071579
Chicago/Turabian StyleTopu, Mustafa, Uğur Sesiz, Harun Bektaş, Faruk Toklu, and Hakan Özkan. 2023. "Next-Generation-Sequencing-Based Simple Sequence Repeat (SSR) Marker Development and Linkage Mapping in Lentil (Lens culinaris L.)" Life 13, no. 7: 1579. https://doi.org/10.3390/life13071579
APA StyleTopu, M., Sesiz, U., Bektaş, H., Toklu, F., & Özkan, H. (2023). Next-Generation-Sequencing-Based Simple Sequence Repeat (SSR) Marker Development and Linkage Mapping in Lentil (Lens culinaris L.). Life, 13(7), 1579. https://doi.org/10.3390/life13071579






