The Reticulon-4 3-bp Deletion/Insertion Polymorphism Is Associated with Structural mRNA Changes and the Risk of Breast Cancer: A Population-Based Case–Control Study with Bioinformatics Analysis
Abstract
1. Introduction
2. Materials and Methods
2.1. Subjects
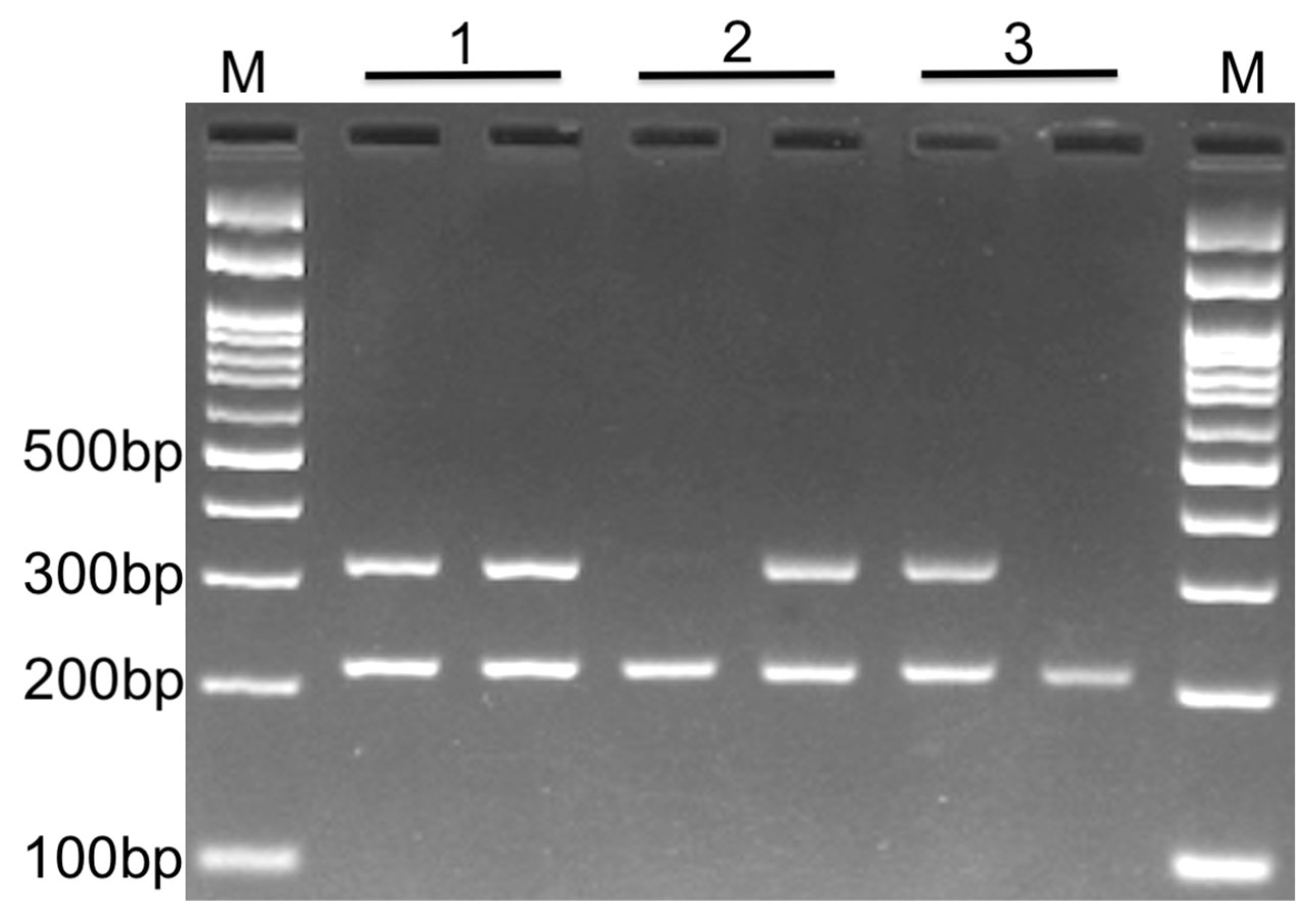
2.2. DNA Extraction and Genotyping
- -
- Deletion allele (F1): CAA-F 5′-GTCTGTGCAATGAAATTGATGTTGGA-3′;
- -
- Insertion allele (F2): CAA-F 5′-GTCTGTGCAATGAAATTGATGTTGTT-3′;
- -
- Generic reverse primer: 5′ACCGGTAAAGCAGGAATGACAA-3′.
2.3. Computational Analyses
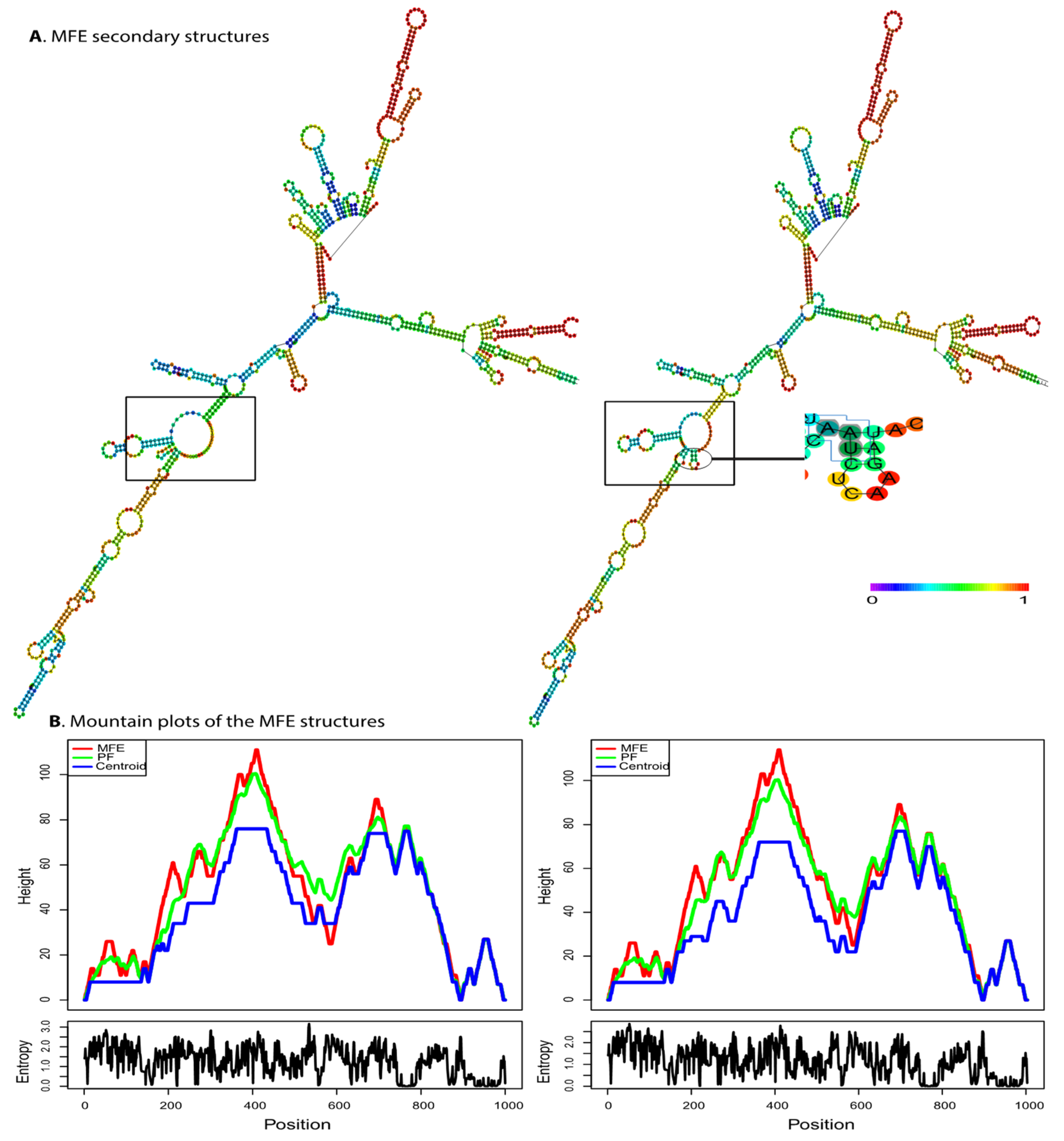
2.3.1. mRNA Secondary Structure Prediction
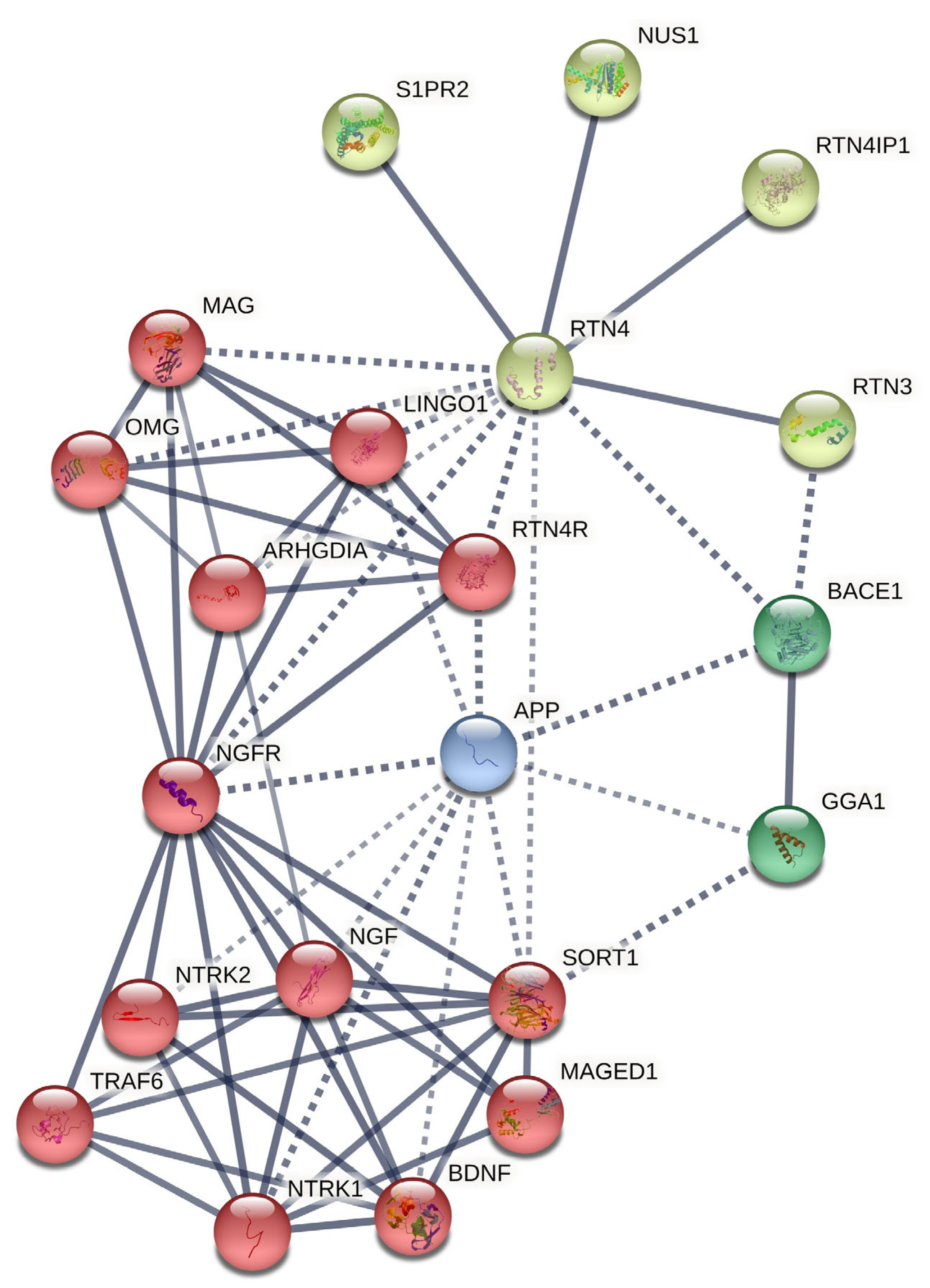
2.3.2. Protein–Protein Interaction Network
2.4. Statistical Analysis
3. Results
3.1. Genotype and Allele Frequency
3.2. Computational Findings
3.2.1. mRNA Second Structure Prediction
3.2.2. Protein–Protein Interaction Network (PPIN)
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- DeSantis, C.E.; Ma, J.; Gaudet, M.M.; Newman, L.A.; Miller, K.D.; Sauer, A.G.; Jemal, A.; Siegel, R.L. Breast cancer statistics, 2019. CA Cancer J. Clin. 2019, 69, 438–451. (In English) [Google Scholar] [CrossRef] [PubMed]
- DeSantis, C.E.; Bray, F.; Ferlay, J.; Lortet-Tieulent, J.; Anderson, B.O.; Jemal, A. International Variation in Female Breast Cancer Incidence and Mortality Rates. Cancer Epidemiol. Biomark. Prev. 2015, 24, 1495–1506. (In English) [Google Scholar] [CrossRef] [PubMed]
- Sarabandi, S.; Sattarifard, H.; Kiumarsi, M.; Karami, S.; Taheri, M.; Hashemi, M.; Bahari, G.; Ghavami, S. Association between Genetic Polymorphisms of miR-1307, miR-1269, miR-3117 and Breast Cancer Risk in a Sample of South East Iranian Women. Asian Pac. J. Cancer Prev. 2021, 22, 201–208. [Google Scholar] [CrossRef] [PubMed]
- Karami, S.; Sattarifard, H.; Kiumarsi, M.; Sarabandi, S.; Taheri, M.; Hashemi, M.; Bahari, G.; Ghavami, S. Evaluating the Possible Association between PD-1 (Rs11568821, Rs2227981, Rs2227982) and PD-L1 (Rs4143815, Rs2890658) Polymorphisms and Susceptibility to Breast Cancer in a Sample of Southeast Iranian Women. Asian Pac. J. Cancer Prev. 2020, 21, 3115–3123. [Google Scholar] [CrossRef] [PubMed]
- Angahar, L. An overview of breast cancer epidemiology, risk factors, pathophysiology, and cancer risks reduction. MOJ Biol. Med. 2017, 1, 92–96. [Google Scholar] [CrossRef]
- Alizadeh, J.; Zeki, A.A.; Mirzaei, N.; Tewary, S.; Moghadam, A.R.; Glogowska, A.; Nagakannan, P.; Eftekharpour, E.; Wiechec, E.; Gordon, J.W.; et al. Mevalonate Cascade Inhibition by Simvastatin Induces the Intrinsic Apoptosis Pathway via Depletion of Isoprenoids in Tumor Cells. Sci. Rep. 2017, 7, 44841. [Google Scholar] [CrossRef]
- Mavaddat, N.; Antoniou, A.C.; Easton, D.F.; Garcia-Closas, M. Genetic susceptibility to breast cancer. Mol. Oncol. 2010, 4, 174–191. (In English) [Google Scholar] [CrossRef]
- Easton, D.F.; Pharoah, P.D.; Antoniou, A.C.; Tischkowitz, M.; Tavtigian, S.V.; Nathanson, K.L.; Devilee, P.; Meindl, A.; Couch, F.J.; Southey, M.; et al. Gene-panel sequencing and the prediction of breast-cancer risk. N. Engl. J. Med. 2015, 372, 2243–2257. [Google Scholar] [CrossRef]
- Wendt, C.; Margolin, S. Identifying breast cancer susceptibility genes—A review of the genetic background in familial breast cancer. Acta Oncol. 2019, 58, 135–146. [Google Scholar] [CrossRef]
- Yang, Y.S.; Strittmatter, S.M. The reticulons: A family of proteins with diverse functions. Genome Biol. 2007, 8, 234. [Google Scholar] [CrossRef]
- Oertle, T.; Huber, C.; van der Putten, H.; Schwab, M.E. Genomic structure and functional characterisation of the promoters of human and mouse nogo/rtn4. J. Mol. Biol. 2003, 325, 299–323. (In English) [Google Scholar] [CrossRef]
- Voeltz, G.K.; Prinz, W.A.; Shibata, Y.; Rist, J.M.; Rapoport, T.A. A class of membrane proteins shaping the tubular endoplasmic reticulum. Cell 2006, 124, 573–586. (In English) [Google Scholar] [CrossRef]
- Teng, F.Y.; Tang, B.L. Cell autonomous function of Nogo and reticulons: The emerging story at the endoplasmic reticulum. J. Cell Physiol. 2008, 216, 303–308. (In English) [Google Scholar] [CrossRef]
- Dodd, D.A.; Niederoest, B.; Bloechlinger, S.; Dupuis, L.; Loeffler, J.-P.; Schwab, M.E. Nogo-A, -B, and -C Are Found on the Cell Surface and Interact Together in Many Different Cell Types. J. Biol. Chem. 2005, 280, 12494–12502. [Google Scholar] [CrossRef]
- Oertle, T.; Merkler, D.; Schwab, M.E. Do cancer cells die because of Nogo-B? Oncogene 2003, 22, 1390–1399. (In English) [Google Scholar] [CrossRef]
- Watari, A.; Yutsudo, M. Multi-functional gene ASY/Nogo/RTN-X/RTN4: Apoptosis, tumor suppression, and inhibition of neuronal regeneration. Apoptosis 2003, 8, 5–9. (In English) [Google Scholar] [CrossRef]
- Chen, Y.; Tang, X.; Cao, X.; Chen, H.; Zhang, X. Human Nogo-C overexpression induces HEK293 cell apoptosis via a mechanism that involves JNK-c-Jun pathway. Biochem. Biophys. Res. Commun. 2006, 348, 923–928. (In English) [Google Scholar] [CrossRef]
- Li, Q.; Qi, B.; Oka, K.; Shimakage, M.; Yoshioka, N.; Inoue, H.; Hakura, A.; Kodama, K.; Stanbridge, E.J.; Yutsudo, M. Link of a new type of apoptosis-inducing gene ASY/Nogo-B to human cancer. Oncogene 2001, 20, 3929–3936. (In English) [Google Scholar] [CrossRef]
- Shimakage, M.; Inoue, N.; Ohshima, K.; Kawahara, K.; Oka, T.; Yasui, K.; Matsumoto, K.; Inoue, H.; Watari, A.; Higashiyama, S. Down-regulation of ASY/Nogo transcription associated with progression of adult T-cell leukemia/lymphoma. Int. J. Cancer 2006, 119, 1648–1653. [Google Scholar] [CrossRef]
- Hu, W.; Liu, Z.; Salato, V.; North, P.E.; Bischoff, J.; Kumar, S.N.; Fang, Z.; Rajan, S.; Hussain, M.M.; Miao, Q.R. NOGOB receptor-mediated RAS signaling pathway is a target for suppressing proliferating hemangioma. JCI Insight 2021, 6, e142299. (In English) [Google Scholar] [CrossRef]
- Zhao, B.; Hu, W.; Kumar, S.; Gonyo, P.; Rana, U.; Liu, Z.; Wang, B.; Duong, W.Q.; Yang, Z.; Williams, C.L.; et al. The Nogo-B receptor promotes Ras plasma membrane localization and activation. Oncogene 2017, 36, 3406–3416. (In English) [Google Scholar] [CrossRef] [PubMed]
- Pathak, G.P.; Shah, R.; Kennedy, B.E.; Murphy, J.P.; Clements, D.; Konda, P.; Giacomantonio, M.; Xu, Z.; Schlaepfer, I.R.; Gujar, S. RTN4 Knockdown Dysregulates the AKT Pathway, Destabilizes the Cytoskeleton, and Enhances Paclitaxel-Induced Cytotoxicity in Cancers. Mol. Ther. 2018, 26, 2019–2033. (In English) [Google Scholar] [CrossRef] [PubMed]
- Yang, F.; Yang, S.; Liu, J.; Pang, X.; Shi, F.; Qin, H.; Wang, J.; Tang, R. Impact of RTN4 gene polymorphism and its plasma level on susceptibility to nasopharyngeal carcinoma: A case-control study. Medicine 2019, 98, e17831. (In English) [Google Scholar] [CrossRef] [PubMed]
- Novak, G.; Tallerico, T. Nogo A, B and C expression in schizophrenia, depression and bipolar frontal cortex, and correlation of Nogo expression with CAA/TATC polymorphism in 3′-UTR. Brain Res. 2006, 1120, 161–171. (In English) [Google Scholar] [CrossRef] [PubMed]
- Barrett, L.W.; Fletcher, S.; Wilton, S.D. Regulation of eukaryotic gene expression by the untranslated gene regions and other non-coding elements. Cell Mol. Life Sci. 2012, 69, 3613–3634. (In English) [Google Scholar] [CrossRef]
- Chan, J.J.; Tabatabaeian, H.; Tay, Y. 3′UTR heterogeneity and cancer progression. Trends Cell Biol. 2022, 3, 568–582. [Google Scholar] [CrossRef]
- Zhou, B.; Rao, L.; Li, Y.; Gao, L.; Li, C.; Chen, Y.; Xue, H.; Liang, W.; Lv, M.; Song, Y.; et al. The association between dilated cardiomyopathy and RTN4 3′UTR insertion/deletion polymorphisms. Clin. Chim. Acta 2009, 400, 21–24. (In English) [Google Scholar] [CrossRef]
- Chen, Y.; Zhou, B.; Li, H.; Peng, Y.; Wang, Y.; Rao, L. Analysis of RTN4 3′UTR insertion/deletion polymorphisms in ventricular septal defect in a Chinese Han population. DNA Cell Biol. 2011, 30, 323–327. (In English) [Google Scholar] [CrossRef]
- Zhang, K.; Bai, P.; Shi, S.; Zhou, B.; Wang, Y.; Song, Y.; Rao, L.; Zhang, L. Association of genetic variations in RTN4 3′-UTR with risk of uterine leiomyomas. Pathol. Oncol. Res. 2013, 19, 475–479. (In English) [Google Scholar] [CrossRef]
- Shi, S.; Zhou, B.; Wang, Y.; Chen, Y.; Zhang, K.; Wang, K.; Quan, Y.; Song, Y.; Rao, L.; Zhang, L. Genetic variation in RTN4 3′-UTR and susceptibility to cervical squamous cell carcinoma. DNA Cell Biol. 2012, 31, 1088–1094. (In English) [Google Scholar] [CrossRef]
- Lu, D.Y.; Mao, X.H.; Zhou, Y.H.; Yan, X.L.; Wang, W.P.; Zheng, Y.B.; Xiao, J.J.; Zhang, P.; Wang, J.G.; Ashwani, N.; et al. RTN4 3′-UTR insertion/deletion polymorphism and susceptibility to non-small cell lung cancer in Chinese Han population. Asian Pac. J. Cancer Prev. 2014, 15, 5249–5252. (In English) [Google Scholar] [CrossRef]
- Pu, Y.; Chen, P.; Zhou, B.; Zhang, P.; Wang, Y.; Song, Y.; Zhang, L. Association of genetic variations in RTN4 3′-UTR with risk for clear cell renal cell carcinoma. Fam. Cancer 2018, 17, 129–134. (In English) [Google Scholar] [CrossRef]
- Wang, N.; Chen, K.; Xu, J.; Yuan, F.; Li, H.; Deng, F.; Zhang, L. Association of CAA and TATC Insertion/Deletion Genetic Polymorphisms in RTN4 3′-UTR with Hepatocellular Carcinoma Risk. Pathol. Oncol. Res. 2018, 24, 31–34. (In English) [Google Scholar] [CrossRef]
- Eskandari-Nasab, E.; Hashemi, M.; Ebrahimi, M.; Amininia, S.; Bahari, G.; Mashhadi, M.A.; Taheri, M. Evaluation of CCL5 -403 G>A and CCR5 Δ32 gene polymorphisms in patients with breast cancer. Cancer Biomark. 2014, 14, 343–351. (In English) [Google Scholar] [CrossRef]
- Eskandari-Nasab, E.; Hashemi, M.; Ebrahimi, M.; Amininia, S. The functional 4-bp insertion/deletion ATTG polymorphism in the promoter region of NF-KB1 reduces the risk of BC. Cancer Biomark. 2016, 16, 109–115. (In English) [Google Scholar] [CrossRef]
- Gaudet, M.; Fara, A.-G.; Beritognolo, I.; Sabatti, M. Allele-Specific PCR in SNP Genotyping. In Single Nucleotide Polymorphisms: Methods and Protocols; Komar, A.A., Ed.; Humana Press: Totowa, NJ, USA, 2009; pp. 415–424. [Google Scholar]
- Gruber, A.R.; Lorenz, R.; Bernhart, S.H.; Neuböck, R.; Hofacker, I.L. The Vienna RNA websuite. Nucleic Acids Res. 2008, 36, W70–W74. (In English) [Google Scholar] [CrossRef]
- Ding, Y.; Chan, C.Y.; Lawrence, C.E. RNA secondary structure prediction by centroids in a Boltzmann weighted ensemble. Rna 2005, 11, 1157–1166. (In English) [Google Scholar] [CrossRef]
- Zuker, M.; Stiegler, P. Optimal computer folding of large RNA sequences using thermodynamics and auxiliary information. Nucleic Acids Res. 1981, 9, 133–148. (In English) [Google Scholar] [CrossRef]
- Szklarczyk, D.; Gable, A.L.; Lyon, D.; Junge, A.; Wyder, S.; Huerta-Cepas, J.; Simonovic, M.; Doncheva, N.T.; Morris, J.H.; Bork, P.; et al. STRING v11: Protein–protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 2018, 47, D607–D613. [Google Scholar] [CrossRef]
- Tabasi, F.; Hasanpour, V.; Sarhadi, S.; Kaykhaei, M.A.; Pourzand, P.; Heravi, M.; Langari, A.A.; Bahari, G.; Taheri, M.; Hashemi, M.; et al. Association of miR-499 Polymorphism and Its Regulatory Networks with Hashimoto Thyroiditis Susceptibility: A Population-Based Case-Control Study. Int. J. Mol. Sci. 2021, 22, 10094. [Google Scholar] [CrossRef]
- Shatoff, E.; Bundschuh, R. Single nucleotide polymorphisms affect RNA-protein interactions at a distance through modulation of RNA secondary structures. PLOS Comput. Biol. 2020, 16, e1007852. [Google Scholar] [CrossRef] [PubMed]
- Chi, C.; Liu, N.; Yue, L.; Qi, W.W.; Xu, L.L.; Qiu, W.S. RTN4/Nogo is an independent prognostic marker for gastric cancer: Preliminary results. Eur. Rev. Med. Pharmacol. Sci. 2015, 19, 241–246. (In English) [Google Scholar] [PubMed]
- Xiong, N.X.; Zhao, H.Y.; Zhang, F.C.; He, Z.Q. Negative correlation of Nogo-A with the malignancy of oligodendroglial tumor. Neurosci. Bull. 2007, 23, 41–45. (In English) [Google Scholar] [CrossRef] [PubMed]
- Wu, D.; Zhao, B.; Qi, X.; Peng, F.; Fu, H.; Chi, X.; Miao, Q.R.; Shao, S. Nogo-B receptor promotes epithelial-mesenchymal transition in non-small cell lung cancer cells through the Ras/ERK/Snail1 pathway. Cancer Lett. 2018, 418, 135–146. (In English) [Google Scholar] [CrossRef] [PubMed]
- Zhu, B.; Chen, S.; Hu, X.; Jin, X.; Le, Y.; Cao, L.; Yuan, Z.; Lin, Z.; Jiang, S.; Sun, L.; et al. Knockout of the Nogo-B Gene Attenuates Tumor Growth and Metastasis in Hepatocellular Carcinoma. Neoplasia 2017, 19, 583–593. (In English) [Google Scholar] [CrossRef]
- Xiao, W.; Zhou, S.; Xu, H.; Li, H.; He, G.; Liu, Y.; Qi, Y. Nogo-B promotes the epithelial-mesenchymal transition in HeLa cervical cancer cells via Fibulin-5. Oncol. Rep. 2013, 29, 109–116. (In English) [Google Scholar] [CrossRef]
- Kawaguchi, N.; Tashiro, K.; Taniguchi, K.; Kawai, M.; Tanaka, K.; Okuda, J.; Hayashi, M.; Uchiyama, K. Nogo-B (Reticulon-4B) functions as a negative regulator of the apoptotic pathway through the interaction with c-FLIP in colorectal cancer cells. Biochim. Biophys. Acta Mol. Basis Dis. 2018, 1864, 2600–2609. (In English) [Google Scholar] [CrossRef]
- Wang, J.; Zhong, Q.; Zhang, H.; Liu, S.; Li, S.; Xia, T.; Xiao, Z.; Chen, R.; Ye, Y.; Liang, F.; et al. Nogo-B promotes invasion and metastasis of nasopharyngeal carcinoma via RhoA-SRF-MRTFA pathway. Cell Death Dis. 2022, 13, 76. [Google Scholar] [CrossRef]
- Pula, B.; Olbromski, M.; Owczarek, T.; Ambicka, A.; Witkiewicz, W.; Ugorski, M.; Rys, J.; Zabel, M.; Dziegiel, P.; Podhorska-Okolow, M. Nogo-B receptor expression correlates negatively with malignancy grade and ki-67 antigen expression in invasive ductal breast carcinoma. Anticancer Res. 2014, 34, 4819–4828. (In English) [Google Scholar]
- Wang, B.; Zhao, B.; North, P.; Kong, A.; Huang, J.; Miao, Q.R. Expression of NgBR is highly associated with estrogen receptor alpha and survivin in breast cancer. PLoS ONE 2013, 8, e78083. (In English) [Google Scholar] [CrossRef]
- Jin, Y.; Hu, W.; Liu, T.; Rana, U.; Aguilera-Barrantes, I.; Kong, A.; Kumar, S.N.; Wang, B.; Gao, P.; Wang, X.; et al. Nogo-B receptor increases the resistance of estrogen receptor positive breast cancer to paclitaxel. Cancer Lett. 2018, 419, 233–244. (In English) [Google Scholar] [CrossRef]
- Koboldt, D.C.; Fulton, R.S.; McLellan, M.D.; Schmidt, H.; Kalicki-Veizer, J.; McMichael, J.F.; Fulton, L.L.; Dooling, D.J.; Ding, L.; Mardis, E.R.; et al. Comprehensive molecular portraits of human breast tumours. Nature 2012, 490, 61–70. [Google Scholar]
- Dai, X.; Xiang, L.; Li, T.; Bai, Z. Cancer Hallmarks, Biomarkers and Breast Cancer Molecular Subtypes. J. Cancer 2016, 7, 1281–1294. (In English) [Google Scholar] [CrossRef]
- Li, H.; Prever, L.; Hirsch, E.; Gulluni, F. Targeting PI3K/AKT/mTOR Signaling Pathway in Breast Cancer. Cancers 2021, 13, 3517. (In English) [Google Scholar] [CrossRef]
- Dittrich, A.; Gautrey, H.; Browell, D.; Tyson-Capper, A. The HER2 Signaling Network in Breast Cancer--Like a Spider in its Web. J. Mammary Gland Biol. Neoplasia 2014, 19, 253–270. (In English) [Google Scholar] [CrossRef]
- Galiè, M. RAS as Supporting Actor in Breast Cancer. Front. Oncol. 2019, 9, 1199. (In English) [Google Scholar] [CrossRef]
- Zhao, B.; Xu, B.; Hu, W.; Song, C.; Wang, F.; Liu, Z.; Ye, M.; Zou, H.; Miao, Q.R. Comprehensive proteome quantification reveals NgBR as a new regulator for epithelial-mesenchymal transition of breast tumor cells. J. Proteom. 2015, 112, 38–52. (In English) [Google Scholar] [CrossRef]
- Miao, R.Q.; Gao, Y.; Harrison, K.D.; Prendergast, J.; Acevedo, L.M.; Yu, J.; Hu, F.; Strittmatter, S.M.; Sessa, W.C. Identification of a receptor necessary for Nogo-B stimulated chemotaxis and morphogenesis of endothelial cells. Proc. Natl. Acad. Sci. USA 2006, 103, 10997–11002. [Google Scholar] [CrossRef]
- Li, Y.-K.; Xie, Y.J.; Wu, D.-C.; Long, S.-L.; Tang, S.; Mo, Z.-C. Nogo-B receptor in relevant carcinoma: Current achievements, challenges and aims (Review). Int. J. Oncol. 2018, 53, 1827–1835. [Google Scholar] [CrossRef]
- Pyne, S.; Edwards, J.; Ohotski, J.; Pyne, N.J. Sphingosine 1-phosphate receptors and sphingosine kinase 1: Novel biomarkers for clinical prognosis in breast, prostate, and hematological cancers. Front. Oncol. 2012, 2, 168. [Google Scholar] [CrossRef]
- Hii, L.-W.; Chung, F.F.-L.; Mai, C.-W.; Ng, P.Y.; Leong, C.-O. Sphingosine Kinase 1 Signaling in Breast Cancer: A Potential Target to Tackle Breast Cancer Stem Cells. Front. Mol. Biosci. 2021, 8, 748470. [Google Scholar] [CrossRef] [PubMed]
- Ohotski, J.; Rosen, H.; Bittman, R.; Pyne, S.; Pyne, N.J. Sphingosine kinase 2 prevents the nuclear translocation of sphingosine 1-phosphate receptor-2 and tyrosine 416 phosphorylated c-Src and increases estrogen receptor negative MDA-MB-231 breast cancer cell growth: The role of sphingosine 1-phosphate receptor-4. Cell. Signal. 2014, 26, 1040–1047. [Google Scholar] [PubMed]
- Cabeza, C.; Figueroa, A.; Lazo, O.M.; Galleguillos, C.; Pissani, C.; Klein, A.; Gonzalez-Billault, C.; Inestrosa, N.C.; Alvarez, A.R.; Zanlungo, S. Cholinergic abnormalities, endosomal alterations and up-regulation of nerve growth factor signaling in Niemann-Pick type C disease. Mol. Neurodegener. 2012, 7, 1–18. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Wang, R.; Thrimawithana, T.; Little, P.J.; Xu, J.; Feng, Z.-P.; Zheng, W. The Nerve Growth Factor Signaling and Its Potential as Therapeutic Target for Glaucoma. BioMed Res. Int. 2014, 2014, 759473. [Google Scholar] [CrossRef] [PubMed]
- Tajbakhsh, A.; Mokhtari-Zaer, A.; Rezaee, M.; Afzaljavan, F.; Rivandi, M.; Hassanian, S.M.; Ferns, G.A.; Pasdar, A.; Avan, A. Therapeutic Potentials of BDNF/TrkB in Breast Cancer; Current Status and Perspectives. J. Cell Biochem. 2017, 118, 2502–2515. (In English) [Google Scholar] [CrossRef]
- Uren, R.T.; Turnley, A.M. Regulation of neurotrophin receptor (Trk) signaling: Suppressor of cytokine signaling 2 (SOCS2) is a new player. Front. Mol. Neurosci. Rev. 2014, 7, 39. (In English) [Google Scholar] [CrossRef]
- Vanhecke, E.; Adriaenssens, E.; Verbeke, S.; Meignan, S.; Germain, E.; Berteaux, N.; Nurcombe, V.; Le Bourhis, X.; Hondermarck, H. Brain-Derived Neurotrophic Factor and Neurotrophin-4/5 Are Expressed in Breast Cancer and Can Be Targeted to Inhibit Tumor Cell SurvivalBDNF and NT-4/5 Promote Breast Tumor Growth. Clin. Cancer Res. 2011, 17, 1741–1752. [Google Scholar] [CrossRef]
- LaFerla, F.M.; Green, K.N.; Oddo, S. Intracellular amyloid-β in Alzheimer’s disease. Nat. Rev. Neurosci. 2007, 8, 499–509. [Google Scholar] [CrossRef]
- Zhai, K.; Huang, Z.; Huang, Q.; Tao, W.; Fang, X.; Zhang, A.; Li, X.; Stark, G.R.; Hamilton, T.A.; Bao, S. Pharmacological inhibition of BACE1 suppresses glioblastoma growth by stimulating macrophage phagocytosis of tumor cells. Nat. Cancer 2021, 2, 1136–1151. (In English) [Google Scholar] [CrossRef]
- Yaghoobi, H.; Azizi, H.; Banitalebi-Dehkordi, M.; Mohammad Rezaei, F.; Arsang-Jnag, S.; Taheri, M.; Ghafouri-Fard, S. Beta-Secretase 1 (BACE1) Is Down-Regulated in Invasive Ductal Carcinoma of Breast. Rep. Biochem. Mol. Biol. 2019, 8, 200–207. (In English) [Google Scholar]
- Baumkötter, F.; Schmidt, N.; Vargas, C.; Schilling, S.; Weber, R.; Wagner, K.; Fiedler, S.; Klug, W.; Radzimanowski, J.; Nickolaus, S.; et al. Amyloid precursor protein dimerization and synaptogenic function depend on copper binding to the growth factor-like domain. J. Neurosci. 2014, 34, 11159–11172. (In English) [Google Scholar] [CrossRef]
- Takagi, K.; Ito, S.; Miyazaki, T.; Miki, Y.; Shibahara, Y.; Ishida, T.; Watanabe, M.; Inoue, S.; Sasano, H.; Suzuki, T. Amyloid precursor protein in human breast cancer: An androgen-induced gene associated with cell proliferation. Cancer Sci. 2013, 104, 1532–1538. [Google Scholar] [CrossRef]
- Wu, X.; Chen, S.; Lu, C. Amyloid precursor protein promotes the migration and invasion of breast cancer cells by regulating the MAPK signaling pathway. Int. J. Mol. Med. 2020, 45, 162–174. (In English) [Google Scholar] [CrossRef]
- Reyhaneh, F.; Rakesh, N. The Role of Apoptosis as a Double-Edge Sword in Cancer. In Regulation and Dysfunction of Apoptosis; Yusuf, T., Ed.; IntechOpen: Rijeka, Croatia, 2021; Chapter 1. [Google Scholar]
- Karande, P.; Sande, A.; Karande, N. Apoptosis—A Beginning of Programmed Ending; B P International: Kowloon, Hong Kong, 2023; pp. 1–35. [Google Scholar] [CrossRef]
- Wan, Q.; Kuang, E.; Dong, W.; Zhou, S.; Xu, H.; Qi, Y.; Liu, Y. Reticulon 3 mediates Bcl-2 accumulation in mitochondria in response to endoplasmic reticulum stress. Apoptosis 2007, 12, 319–328. (In English) [Google Scholar] [CrossRef]
- Wallach, D. The Tumor Necrosis Factor Family: Family Conventions and Private Idiosyncrasies. Cold Spring Harb. Perspect. Biol. 2018, 10, a028431. (In English) [Google Scholar] [CrossRef]
- Truzzi, F.; Marconi, A.; Lotti, R.; Dallaglio, K.; French, L.E.; Hempstead, B.L.; Pincelli, C. Neurotrophins and their receptors stimulate melanoma cell proliferation and migration. J. Investig. Dermatol. 2008, 128, 2031–2040. (In English) [Google Scholar] [CrossRef]
- Khwaja, F.; Tabassum, A.; Allen, J.; Djakiew, D. The p75(NTR) tumor suppressor induces cell cycle arrest facilitating caspase mediated apoptosis in prostate tumor cells. Biochem. Biophys. Res. Commun. 2006, 341, 1184–1192. (In English) [Google Scholar] [CrossRef]
- Nykjaer, A.; Lee, R.; Teng, K.K.; Jansen, P.; Madsen, P.; Nielsen, M.S.; Jacobsen, C.; Kliemannel, M.; Schwarz, E.; Willnow, T.E. Sortilin is essential for proNGF-induced neuronal cell death. Nature 2004, 427, 843–848. [Google Scholar] [CrossRef]
- Descamps, S.; Toillon, R.-A.; Adriaenssens, E.; Pawlowski, V.; Cool, S.M.; Nurcombe, V.; Le Bourhis, X.; Boilly, B.; Peyrat, J.-P.; Hondermarck, H. Nerve growth factor stimulates proliferation and survival of human breast cancer cells through two distinct signaling pathways. J. Biol. Chem. 2001, 276, 17864–17870. [Google Scholar] [CrossRef]
- Demont, Y.; Corbet, C.; Page, A.; Ataman-Önal, Y.; Choquet-Kastylevsky, G.; Fliniaux, I.; Le Bourhis, X.; Toillon, R.A.; Bradshaw, R.A.; Hondermarck, H. Pro-nerve growth factor induces autocrine stimulation of breast cancer cell invasion through tropomyosin-related kinase A (TrkA) and sortilin protein. J. Biol. Chem. 2012, 287, 1923–1931. (In English) [Google Scholar] [CrossRef]
- Boulter, E.; Garcia-Mata, R.; Guilluy, C.; Dubash, A.; Rossi, G.; Brennwald, P.J.; Burridge, K. Regulation of Rho GTPase crosstalk, degradation and activity by RhoGDI1. Nat. Cell Biol. 2010, 12, 477–483. [Google Scholar] [CrossRef] [PubMed]
- Crosas-Molist, E.; Samain, R.; Kohlhammer, L.; Orgaz, J.L.; George, S.L.; Maiques, O.; Barcelo, J.; Sanz-Moreno, V. Rho GTPase signaling in cancer progression and dissemination. Physiol. Rev. 2022, 102, 455–510. [Google Scholar] [CrossRef] [PubMed]
- De, P.; Rozeboom, B.J.; Aske, J.C.; Dey, N. Active RAC1 Promotes Tumorigenic Phenotypes and Therapy Resistance in Solid Tumors. Cancers 2020, 12, 1541. (In English) [Google Scholar] [CrossRef]
- Hein, A.L.; Post, C.M.; Sheinin, Y.M.; Lakshmanan, I.; Natarajan, A.; Enke, C.A.; Batra, S.K.; Ouellette, M.M.; Yan, Y. RAC1 GTPase promotes the survival of breast cancer cells in response to hyper-fractionated radiation treatment. Oncogene 2016, 35, 6319–6329. (In English) [Google Scholar] [CrossRef] [PubMed]
- Jiang, W.G.; Watkins, G.; Lane, J.; Cunnick, G.H.; Douglas-Jones, A.; Mokbel, K.; Mansel, R.E. Prognostic value of rho GTPases and rho guanine nucleotide dissociation inhibitors in human breast cancers. Clin. Cancer Res. 2003, 9, 6432–6440. [Google Scholar]
- Cho, H.J.; Kim, J.T.; Baek, K.E.; Kim, B.Y.; Lee, H.G. Regulation of Rho GTPases by RhoGDIs in Human Cancers. Cells 2019, 8, 1037. (In English) [Google Scholar] [CrossRef]
- Novak, G.; Kim, D.; Seeman, P.; Tallerico, T. Schizophrenia and Nogo: Elevated mRNA in cortex, and high prevalence of a homozygous CAA insert. Brain Res. Mol. Brain Res. 2002, 107, 183–189. (In English) [Google Scholar] [CrossRef]
- Chatterjee, S.; Pal, J.K. Role of 5′- and 3′-untranslated regions of mRNAs in human diseases. Biol. Cell 2009, 101, 251–262. (In English) [Google Scholar] [CrossRef]
| Description | Nodes | Edges | Expected Number of Edges | Avg. Node Degree | Avg. Local Clustering Coefficient | Inflation Parameter (MCL) | Enrichment p-Value |
|---|---|---|---|---|---|---|---|
| RTN4 PPIN | 21 | 65 | 22 | 6.19 | 0.781 | 6 | 5.06 × 10−14 |
| Characteristics | Patients, n (SD) | Control, n (SD) | p * | |
|---|---|---|---|---|
| Age | Mean (SD) | 48.34 (10.82) | 49.30 (11.96) | 0.92 |
| Median (IQR) | 48.00 (16.0) | 46.0 (17.0) | ||
| Polymorphism | Group | Genotypes, n (%) | Allele, n (%) | HWE | ||||
|---|---|---|---|---|---|---|---|---|
| Del/Del | Ins/Del | Ins/Ins | Del | Ins | χ2 | p | ||
| rs34917480 | Patients | 134 (63.2) | 60 (28.3) | 18 (8.5) | 328 (77.4) | 96 (22.6) | 16.15 | <0.0001 |
| Controls | 116 (51.6) | 88 (39.1) | 21 (9.3) | 320 (71.1) | 130 (28.9) | 0.05 | 0.81 | |
| Polymorphisms | Genetic Models | Patients, n (%) | Control, n (%) | OR (95% CI) * | p ** |
|---|---|---|---|---|---|
| RTN4 rs34917480 | Codominant | ||||
| Del/Del | 134 (63.2) | 116 (51.6) | 1.00 | - | |
| Ins/Del | 60 (28.3) | 88 (39.1) | 0.59 (0.39–0.89) | 0.012 | |
| Ins/Ins | 18 (8.5) | 21 (9.3) | 0.74 (0.378–1.46) | 0.388 | |
| Dominant | |||||
| Del/Del | 134 (63.2) | 116 (51.6) | 1.00 | - | |
| Ins/Del + Ins/Ins | 78 (36.8) | 109 (48.4) | 0.62 (0.43–0.91) | 0.015 | |
| Recessive | |||||
| Del/Del + Ins/Del | 194 (91.5) | 204 (90.7) | 1.00 | - | |
| Ins/Ins | 18 (8.5) | 21 (9.3) | 0.92 (0.47–1.78) | 0.8 | |
| Over-dominant | |||||
| Del/Del + Ins/Ins | 152 (71.7) | 137 (60.9) | 1.00 | - | |
| Ins/Del | 60 (28.3) | 88 (39.1) | 0.62 (0.41–0.92) | 0.017 | |
| Alleles | |||||
| Del | 328 (77.4) | 320 (71.1) | 1.00 | - | |
| Ins | 96 (22.6) | 130 (28.9) | 0.72 (0.53–0.99) | 0.035 |
| Characteristic of Patients | Genotypes | p | ||
|---|---|---|---|---|
| Del/Del n (%) | Ins/Del n (%) | Ins/Ins n (%) | ||
| Age, years | 0.32 | |||
| ≤50 | 86 (67.2) | 32 (25.0) | 10 (7.8) | |
| >50 | 48 (57.1) | 28 (33.30 | 8 (9.5) | |
| Tumor size, cm | 0.58 | |||
| ≤2 | 41 (59.9) | 23 (33.3) | 5 (7.2) | |
| >2 | 90 (66.2) | 36 (26.5) | 10 (7.4) | |
| Histology | 0.085 | |||
| Adenocarcinoma | 22 (54.2) | 19 (45.2) | 1 (2.4) | |
| Ductal carcinoma | 94 (68.1) | 33 (23.9) | 11 (8.0) | |
| Lobular carcinoma | 5 (83.3) | 0 (0) | 1 (16.7) | |
| Mucinous carcinoma | 11 (61.1) | 5 (27.8) | 2 (11.1) | |
| Grade | 0.094 | |||
| I | 23 (63.9) | 6 (16.7) | 7 (19.4) | |
| II | 68 (63.6) | 33 (30.8) | 6 (5.6) | |
| III+IV | 24 (66.7) | 8 (22.2) | 4 (11.1) | |
| Stage | 0.88 | |||
| I | 22 (61.1) | 12 (33.3) | 2 (5.6) | |
| II | 48 (62.3) | 24 (31.2) | 5 (6.5) | |
| III | 40 (63.5) | 16 (25.4) | 7 911.1) | |
| IV | 23 (67.6) | 8 (23.5) | 3 (8.8) | |
| Estrogen receptor status | 0.16 | |||
| Positive | 84 (67.7) | 27 (21.8) | 13 (19.5) | |
| Negative | 46 (62.2) | 24 (32.4) | 4 (5.4) | |
| Progesterone receptor status | 0.031 | |||
| Positive | 84 (70.6) | 23 (19.3) | 12 (10.1) | |
| Negative | 45 (57.7) | 28 (35.9) | 5 (6.4) | |
| HER2 status | 0.66 | |||
| Positive | 68 (62.4) | 33 (30.3) | 8 (7.3) | |
| Negative | 66 (66.0) | 25 (25.0) | 9 (9.0) | |
| Description | Observed | Background | Strength | FDR * |
|---|---|---|---|---|
| Selected biological process (GO) † | ||||
| Regulation of plasma membrane-bounded cell projection organization | 13 | 687 | 1.25 | 9.31 × 10−11 |
| Regulation of cell development | 14 | 956 | 1.13 | 9.31 × 10−11 |
| Neurotrophin trk receptor signaling pathway | 6 | 20 | 2.45 | 2.89 × 10−10 |
| Regulation of cell differentiation | 16 | 1874 | 0.9 | 8.70 × 10−10 |
| Regulation of multicellular organismal development | 16 | 2096 | 0.85 | 3.49 × 10−9 |
| Regulation of the developmental process | 17 | 2648 | 0.78 | 5.33 × 10−9 |
| Regulation of multicellular organismal process | 17 | 3227 | 0.69 | 8.35 × 10−8 |
| Regulation of anatomical structure morphogenesis | 12 | 1095 | 1.01 | 9.42 × 10−8 |
| Regulation of apoptotic process | 13 | 1550 | 0.89 | 2.67 × 10−7 |
| Negative regulation of cell differentiation | 10 | 728 | 1.11 | 5.29 × 10−7 |
| Positive regulation of cell development | 9 | 556 | 1.18 | 9.80 × 10−7 |
| Selected molecular function (GO) †† | ||||
| Nerve growth factor receptor activity | 2 | 2 | 2.97 | 0.0035 |
| Ganglioside gt1b binding | 2 | 3 | 2.79 | 0.0051 |
| Nerve growth factor receptor binding | 2 | 5 | 2.57 | 0.0069 |
| Tumor necrosis factor receptor superfamily binding | 3 | 48 | 1.77 | 0.0069 |
| Amide binding | 5 | 369 | 1.1 | 0.0105 |
| Signaling receptor binding | 8 | 1581 | 0.67 | 0.0332 |
| Cytokine receptor binding | 4 | 264 | 1.15 | 0.0345 |
| Protein binding | 16 | 7026 | 0.33 | 0.0384 |
| Selected KEGG pathway ††† | ||||
| MAPK signaling pathway | 6 | 288 | 1.29 | 8.20 × 10−5 |
| Ras signaling pathway | 5 | 226 | 1.31 | 0.00043 |
| PI3K-Akt signaling pathway | 5 | 350 | 1.12 | 0.0026 |
| Selected Reactome pathway †††† | ||||
| p75 NTR receptor-mediated signaling | 10 | 96 | 1.99 | 5.08 × 10−15 |
| Signal transduction | 15 | 2741 | 0.71 | 1.65 × 10−6 |
| Activation of TRKA receptors | 3 | 6 | 2.67 | 2.76 × 10−5 |
| p75NTR recruits signaling complexes | 3 | 13 | 2.33 | 0.00016 |
| NF-kB is activated and signals survival | 3 | 13 | 2.33 | 0.00016 |
| NRIF signals cell death from the nucleus | 3 | 16 | 2.24 | 0.00022 |
| Cell death signaling via NRAGE, NRIF, and NADE | 4 | 75 | 1.7 | 0.00025 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pourzand, P.; Tabasi, F.; Fayazbakhsh, F.; Sarhadi, S.; Bahari, G.; Mohammadi, M.; Jomepour, S.; Nafeli, M.; Mosayebi, F.; Heravi, M.; et al. The Reticulon-4 3-bp Deletion/Insertion Polymorphism Is Associated with Structural mRNA Changes and the Risk of Breast Cancer: A Population-Based Case–Control Study with Bioinformatics Analysis. Life 2023, 13, 1549. https://doi.org/10.3390/life13071549
Pourzand P, Tabasi F, Fayazbakhsh F, Sarhadi S, Bahari G, Mohammadi M, Jomepour S, Nafeli M, Mosayebi F, Heravi M, et al. The Reticulon-4 3-bp Deletion/Insertion Polymorphism Is Associated with Structural mRNA Changes and the Risk of Breast Cancer: A Population-Based Case–Control Study with Bioinformatics Analysis. Life. 2023; 13(7):1549. https://doi.org/10.3390/life13071549
Chicago/Turabian StylePourzand, Pouria, Farhad Tabasi, Fariba Fayazbakhsh, Shamim Sarhadi, Gholamreza Bahari, Mohsen Mohammadi, Sahar Jomepour, Mohammad Nafeli, Fatemeh Mosayebi, Mehrdad Heravi, and et al. 2023. "The Reticulon-4 3-bp Deletion/Insertion Polymorphism Is Associated with Structural mRNA Changes and the Risk of Breast Cancer: A Population-Based Case–Control Study with Bioinformatics Analysis" Life 13, no. 7: 1549. https://doi.org/10.3390/life13071549
APA StylePourzand, P., Tabasi, F., Fayazbakhsh, F., Sarhadi, S., Bahari, G., Mohammadi, M., Jomepour, S., Nafeli, M., Mosayebi, F., Heravi, M., Taheri, M., Hashemi, M., & Ghavami, S. (2023). The Reticulon-4 3-bp Deletion/Insertion Polymorphism Is Associated with Structural mRNA Changes and the Risk of Breast Cancer: A Population-Based Case–Control Study with Bioinformatics Analysis. Life, 13(7), 1549. https://doi.org/10.3390/life13071549







