The Fungal and Bacterial Interface in the Respiratory Mycobiome with a Focus on Aspergillus spp.
Abstract
1. Introduction
2. Lung and Lower Respiratory Tract Microbiome
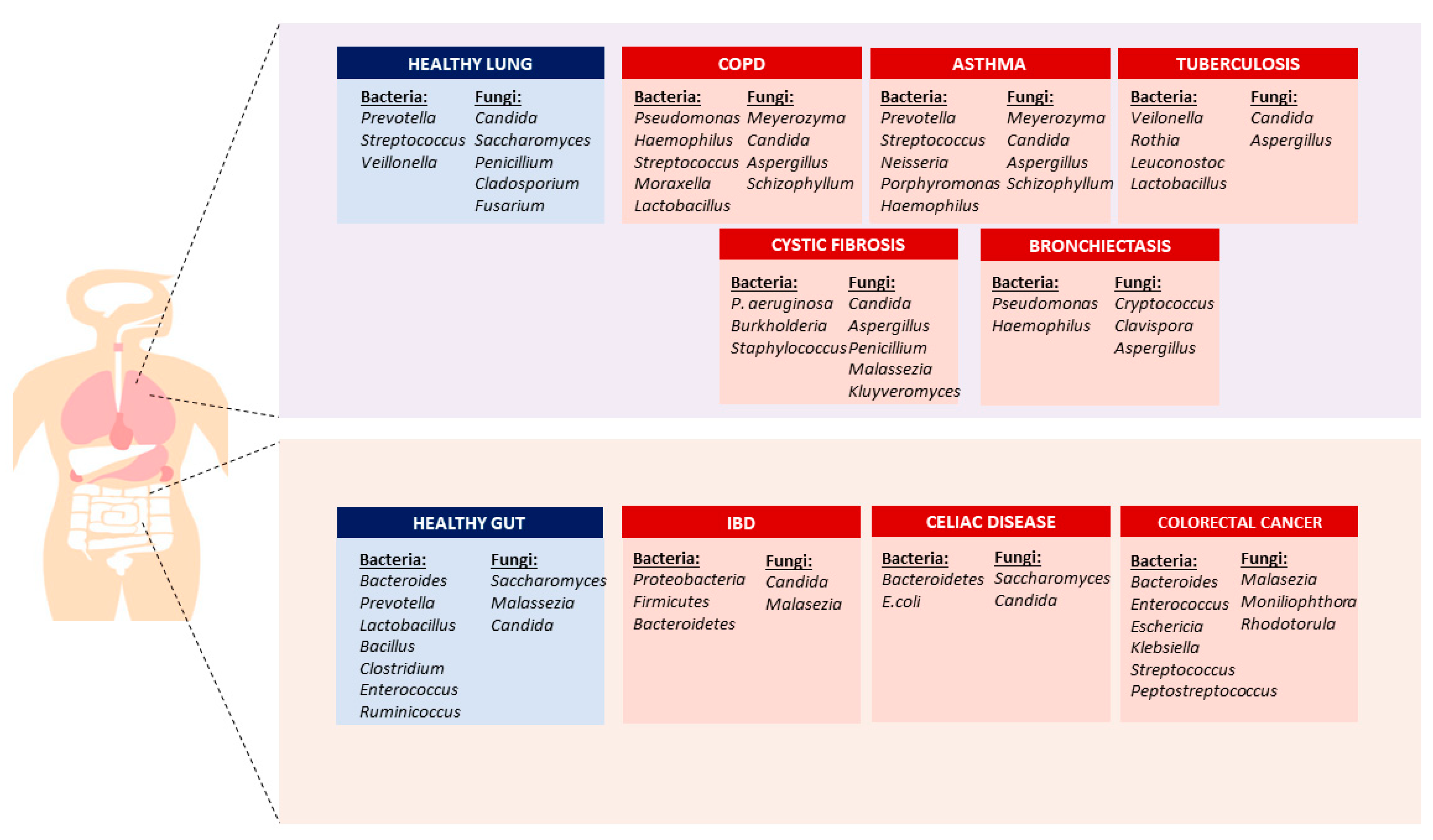
2.1. Bacterial Microbiome
2.2. Fungal Microbiome (Mycobiome)
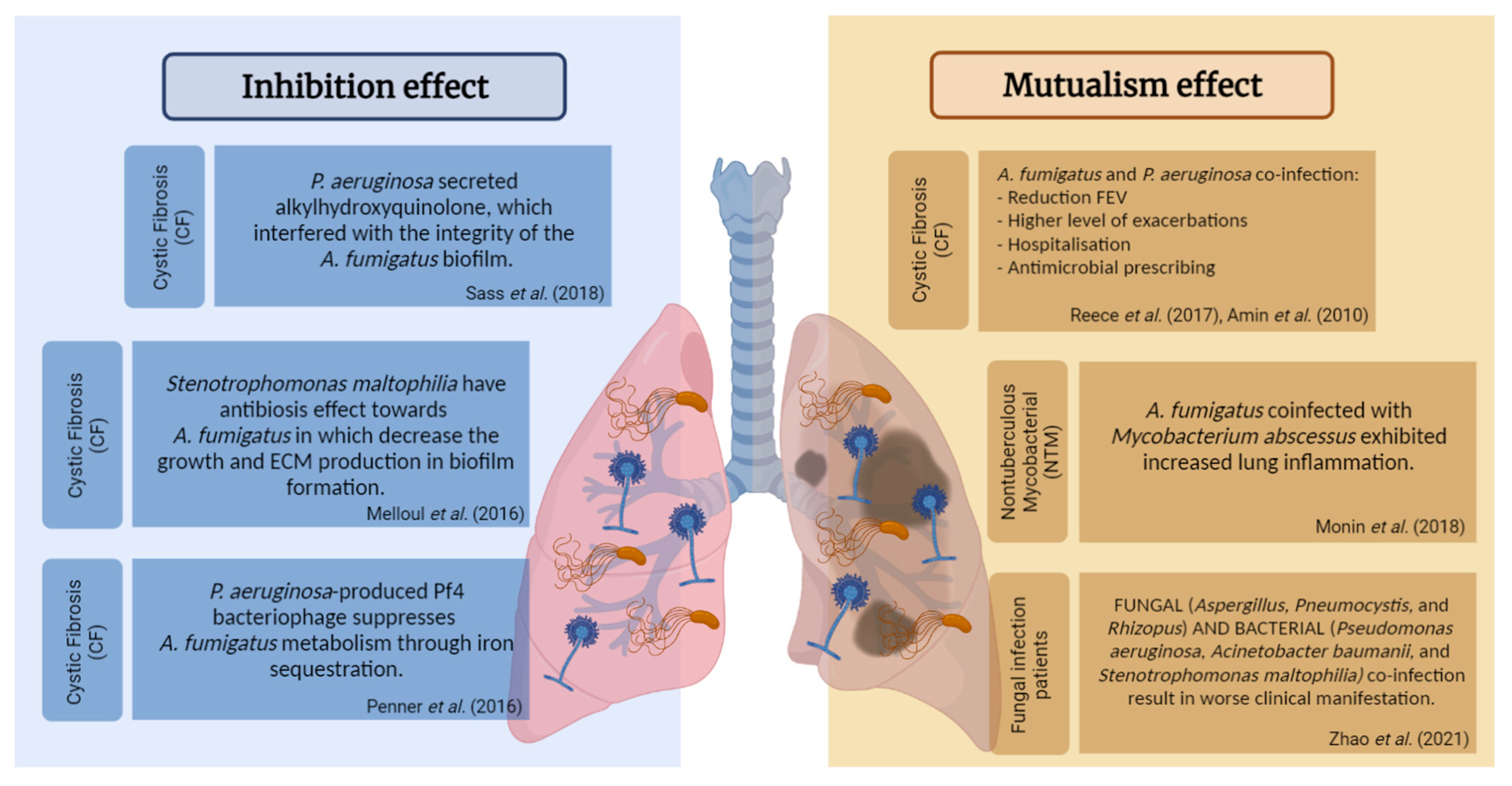
2.3. Bacterial–Fungal Interaction
| No. | Mode of Interaction | Source of Isolates | Reference |
|---|---|---|---|
| 1. | Pseudomonas aeruginosa inhibited Aspergillus fumigatus biofilms from conidia in CF isolates compared to non-CF. | CF and non-CF patients | [99] |
| 2. | Four typical phenazines released by Pseudomonas aeruginosa suppressed the growth of Aspergillus fumigatus by inducing ROS and NOS. | Murine aspergillosis models | [100] |
| 3. | Pyoverdine, a chemical made by P. aeruginosa, may be able to collect iron from the environment, which would prevent A. fumigatus from growing as a result of nutritional shortage. | A. fumigatus isolates were from ATCC (ATCC 90240), ATCC 46645 sidA ftrA mutant, while the P. aeruginosa isolates were obtained from CF patients. | [101] |
| 4. | P. aeruginosa produced dirhamnolipids that promoted the secretion of dihydroxynaphthalene (DHN) and pyo-melanin from A. fumigatus, which surrounded their hyphae to facilitate the P. aeruginosa binding, leading to the inhibition of the fungi growth through the blocking of β1,3 glucan synthase (GS) activity. | Murine aspergillosis models | [102] |
| 5. | P. aeruginosa secreted alkylhydroxyquinolones, which interfered with the integrity of the A. fumigatus biofilm. | CF patients (pediatric) | [103] |
| 6. | P. aeruginosa-produced Pf4 bacteriophage suppressed the A. fumigatus metabolism through iron sequestration. | CF and non-CF patients | [104] |
| 7. | A. fumigatus inhibited the biofilm formation of P. aeruginosa through gliotoxin production. | CF patients | [105] |
| 8. | A. fumigatus overcame iron starvation by releasing its hydroxamate siderophores, thus promoting iron and depriving P. aeruginosa. | Isolates 10 AF, AF13073, AfΔsidA, AF46645, AfΔsidC, AfΔsidF, AfS77, PA14, pvdD-, pvdD-pchE- | [106] |
| 9. | P. aeruginosa may encourage fungal growth by secreting volatile organic chemicals. | Aspergillus fumigatus CBS144-89, Pseudomonas aeruginosa PAO1 | [107] |
| 10. | The coexistence of A. fumigatus may enhance P. aeruginosa’s phenotypic and genetic changes, increasing bacterial virulences. | A. fumigatus 53470 (AF53470), A. fumigatus ATCC 36607 (AF36607), P. aeruginosa 56402 (PA56402) and P. aeruginosa ATCC27853 (PA27853) were used in this study. | [108] |
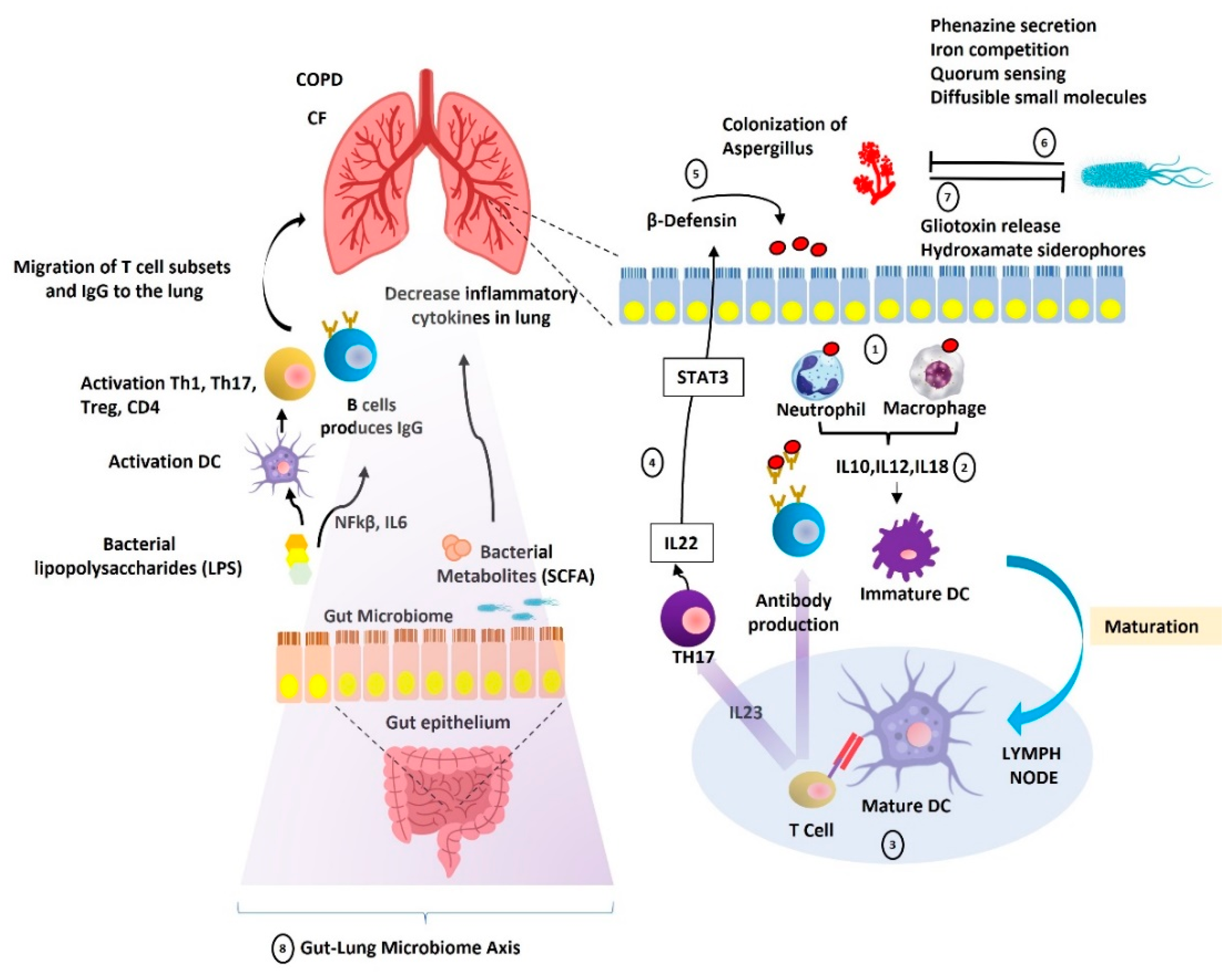
3. Gut-Lung Microbiome Axis
4. Discussion
4.1. The Fungal and Bacterial Interface in Specific Respiratory Diseases Entities
4.1.1. Chronic Obstructive Pulmonary Disease (COPD)
4.1.2. Cystic Fibrosis (CF)
4.1.3. Chronic Pulmonary Aspergillosis (CPA)
4.2. Challenges in the Detection of the Lung Microbiome
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Kolwijck, E.; van de Veerdonk, F.L. The potential impact of the pulmonary microbiome on immunopathogenesis of Aspergillus-related lung disease. Eur. J. Immunol. 2014, 44, 3156–3165. [Google Scholar] [CrossRef] [PubMed]
- Hérivaux, A.; Willis, J.R.; Mercier, T.; Lagrou, K.; Gonçalves, S.M.; Gonçales, R.A.; Maertens, J.; Carvalho, A.; Gabaldón, T.; Cunha, C. Lung microbiota predict invasive pulmonary aspergillosis and its outcome in immunocompromised patients. Thorax 2022, 77, 283–291. [Google Scholar] [CrossRef] [PubMed]
- Dickson, R.P.; Erb-Downward, J.R.; Martinez, F.J.; Huffnagle, G.B. The microbiome and the respiratory Tract. Annu. Rev. Physiol. 2016, 78, 481–504. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Wang, K.; Wu, W.; Giannoulatou, E.; Ho, J.W.K.; Li, L. Host and microbiome multi-omics integration: Applications and methodologies. Biophys. Rev. 2019, 11, 55–65. [Google Scholar] [CrossRef]
- Beck, J.M.; Young, V.B.; Huffnagle, G.B. The microbiome of the lung. Transl. Res. 2012, 160, 258–266. [Google Scholar] [CrossRef]
- Morgan, X.C.; Huttenhower, C. Chapter 12: Human microbiome analysis. PLoS Comput. Biol. 2012, 8, e1002808. [Google Scholar] [CrossRef]
- Dickson, R.P.; Erb-Downward, J.R.; Freeman, C.M.; McCloskey, L.; Falkowski, N.R.; Huffnagle, G.B.; Curtis, J.L. Bacterial Topography of the Healthy Human Lower Respiratory Tract. mBio 2017, 8, e02287-16. [Google Scholar] [CrossRef]
- Gusareva, E.S.; Acerbi, E.; Lau, K.J.X.; Luhung, I.; Premkrishnan, B.N.V.; Kolundžija, S.; Purbojati, R.W.; Wong, A.; Houghton, J.N.I.; Miller, D.; et al. Microbial communities in the tropical air ecosystem follow a precise diel cycle. Proc. Natl. Acad. Sci. USA 2019, 116, 23299–23308. [Google Scholar] [CrossRef]
- Filho, F.S.L.; Alotaibi, N.M.; Ngan, D.; Tam, S.; Yang, J.; Hollander, Z.; Chen, V.; Fitzgerald, J.M.; Nislow, C.; Leung, J.M.; et al. Sputum Microbiome Is Associated with 1-Year Mortality after Chronic Obstructive Pulmonary Disease Hospitalizations. Am. J. Respir. Crit. Care Med. 2019, 199, 1205–1213. [Google Scholar] [CrossRef]
- Acosta, N.; Heirali, A.; Somayaji, R.; Surette, M.G.; Workentine, M.L.; Sibley, C.D.; Rabin, H.R.; Parkins, M.D. Sputum microbiota is predictive of long-term clinical outcomes in young adults with cystic fibrosis. Thorax 2018, 73, 1016–1025. [Google Scholar] [CrossRef]
- Rogers, G.B.; Zain, N.M.M.; Bruce, K.D.; Burr, L.D.; Chen, A.C.; Rivett, D.W.; McGuckin, M.A.; Serisier, D.J. A Novel Microbiota Stratification System Predicts Future Exacerbations in Bronchiectasis. Ann. Am. Thorac. Soc. 2014, 11, 496–503. [Google Scholar] [CrossRef]
- Richardson, M.; Bowyer, P.; Sabino, R. The human lung and Aspergillus: You are what you breathe in? Med. Mycol. 2019, 57, S145–S154. [Google Scholar] [CrossRef] [PubMed]
- Kwon-Chung, K.J.; Sugui, J.A. Aspergillus fumigatus—What Makes the Species a Ubiquitous Human Fungal Pathogen? PLoS Pathog. 2013, 9, e1003743. [Google Scholar] [CrossRef] [PubMed]
- Arastehfar, A.; Carvalho, A.; Houbraken, J.; Lombardi, L.; Garcia-Rubio, R.; Jenks, J.D.; Rivero-Menendez, O.; Aljohani, R.; Jacobsen, I.D.; Berman, J.; et al. Aspergillus fumigatus and aspergillosis: From basics to clinics. Stud. Mycol. 2021, 100, 100115. [Google Scholar] [CrossRef] [PubMed]
- Denning, D.W. Chronic forms of pulmonary aspergillosis. Clin. Microbiol. Infect. 2001, 7 (Suppl. S2), 25–31. [Google Scholar] [CrossRef] [PubMed]
- Latgé, J.-P. The pathobiology of Aspergillus fumigatus. Trends Microbiol. 2001, 9, 382–389. [Google Scholar] [CrossRef] [PubMed]
- Bertuzzi, M.; Hayes, G.E.; Icheoku, U.J.; van Rhijn, N.; Denning, D.W.; Osherov, N.; Bignell, E.M. Anti-Aspergillus Activities of the Respiratory Epithelium in Health and Disease. J. Fungi 2018, 4, 8. [Google Scholar] [CrossRef]
- Latgé, J.-P.; Chamilos, G. Aspergillus fumigatus and Aspergillosis in 2019. Clin. Microbiol. Rev. 2019, 33, e00140-18. [Google Scholar] [CrossRef]
- Denning, D.W.; Pashley, C.; Hartl, D.; Wardlaw, A.; Godet, C.; Del Giacco, S.; Delhaes, L.; Sergejeva, S. Fungal allergy in asthma–state of the art and research needs. Clin. Transl. Allergy 2014, 4, 14. [Google Scholar] [CrossRef]
- Menzies, D.; Holmes, L.; McCumesky, G.; Prys-Picard, C.; Niven, R. Aspergillus sensitization is associated with airflow limitation and bronchiectasis in severe asthma. Allergy Eur. J. Allergy Clin. Immunol. 2011, 66, 679–685. [Google Scholar] [CrossRef]
- Tiew, P.Y.; Dicker, A.J.; Keir, H.R.; Poh, M.E.; Pang, S.L.; Mac Aogáin, M.; Chua, B.Q.Y.; Tan, J.L.; Xu, H.; Koh, M.S.; et al. A high-risk airway mycobiome is associated with frequent exacerbation and mortality in COPD. Eur. Respir. J. 2021, 57, 2002050. [Google Scholar] [CrossRef] [PubMed]
- Tiew, P.Y.; Thng, K.X.; Chotirmall, S.H. Clinical Aspergillus Signatures in COPD and Bronchiectasis. J. Fungi 2022, 8, 480. [Google Scholar] [CrossRef] [PubMed]
- Vandenborght, L.-E.; Enaud, R.; Coron, N.; Denning, D.; Delhaes, L. From culturomics to metagenomics: The mycobiome in chronic respiratory diseases. ERS Monogr. 2019, 2019, 88–118. [Google Scholar] [CrossRef]
- Denning, D.W.; Cadranel, J.; Beigelman-Aubry, C.; Ader, F.; Chakrabarti, A.; Blot, S.; Ullmann, A.J.; Dimopoulos, G.; Lange, C. Chronic pulmonary aspergillosis: Rationale and clinical guidelines for diagnosis and management. Eur. Respir. J. 2016, 47, 45–68. [Google Scholar] [CrossRef] [PubMed]
- Smith, N.L.; Denning, D.W. Underlying conditions in chronic pulmonary aspergillosis including simple aspergilloma. Eur. Respir. J. 2011, 37, 865–872. [Google Scholar] [CrossRef]
- Rapaka, R.R.; Kolls, J.K. Pathogenesis of allergic bronchopulmonary aspergillosis in cystic fibrosis: Current understanding and future directions. Med. Mycol. 2009, 47 (Suppl. S1), S331–S337. [Google Scholar] [CrossRef]
- Isa, K.N.M.; Jalaludin, J.; Elias, S.M.; Than, L.T.L.; Jabbar, M.A.; Saudi, A.S.M.; Norbäck, D.; Hashim, J.H.; Hashim, Z. Metagenomic characterization of indoor dust fungal associated with allergy and lung inflammation among school children. Ecotoxicol. Environ. Saf. 2021, 221, 112430. [Google Scholar] [CrossRef]
- Andreu, Z.; Yáñez-Mó, M. Tetraspanins in Extracellular Vesicle Formation and Function. Front. Immunol. 2014, 5, 442. [Google Scholar] [CrossRef]
- Almohammai, A.; Rahbarghazi, R.; Keyhanmanesh, R.; Rezaie, J.; Ahmadi, M. Asthmatic condition induced the activity of exosome secretory pathway in rat pulmonary tissues. J. Inflamm. 2021, 18, 14. [Google Scholar] [CrossRef]
- Huffnagle, G.B.; Noverr, M.C. The emerging world of the fungal microbiome. Trends Microbiol. 2013, 21, 334–341. [Google Scholar] [CrossRef]
- Cohen, R.; Roth, F.J.; Delgado, E.; Ahearn, D.G.; Kalser, M.H. Fungal Flora of the Normal Human Small and Large Intestine. N. Engl. J. Med. 1969, 280, 638–641. [Google Scholar] [CrossRef] [PubMed]
- Li, X.V.; Leonardi, I.; Putzel, G.G.; Semon, A.; Fiers, W.D.; Kusakabe, T.; Lin, W.-Y.; Gao, I.H.; Doron, I.; Gutierrez-Guerrero, A.; et al. Immune regulation by fungal strain diversity in inflammatory bowel disease. Nature 2022, 603, 672–678. [Google Scholar] [CrossRef]
- Abreu, N.A.; Nagalingam, N.A.; Song, Y.; Roediger, F.C.; Pletcher, S.D.; Goldberg, A.N.; Lynch, S.V. Sinus Microbiome Diversity Depletion and Corynebacterium tuberculostearicum Enrichment Mediates Rhinosinusitis. Sci. Transl. Med. 2012, 4, 151ra124. [Google Scholar] [CrossRef]
- Turek, E.M.; Cox, M.J.; Hunter, M.; Hui, J.; James, P.; Willis-Owen, S.A.G.; Cuthbertson, L.; James, A.; Musk, A.W.; Moffatt, M.F.; et al. Airway microbial communities, smoking and asthma in a general population sample. Ebiomedicine 2021, 71, 103538. [Google Scholar] [CrossRef]
- Ramsheh, M.Y.; Haldar, K.; Esteve-Codina, A.; Purser, L.F.; Richardson, M.; Müller-Quernheim, J.; Greulich, T.; Nowinski, A.; Barta, I.; Stendardo, M.; et al. Lung microbiome composition and bronchial epithelial gene expression in patients with COPD versus healthy individuals: A bacterial 16S rRNA gene sequencing and host transcriptomic analysis. Lancet Microbe 2021, 2, e300–e310. [Google Scholar] [CrossRef] [PubMed]
- Hilty, M.; Burke, C.; Pedro, H.; Cardenas, P.; Bush, A.; Bossley, C.; Davies, J.; Ervine, A.; Poulter, L.; Pachter, L.; et al. Disordered microbial communities in asthmatic airways. PLoS ONE 2010, 5, e8578. [Google Scholar] [CrossRef]
- Charlson, E.S.; Bittinger, K.; Haas, A.R.; Fitzgerald, A.S.; Frank, I.; Yadav, A.; Bushman, F.D.; Collman, R.G. Topographical Continuity of Bacterial Populations in the Healthy Human Respiratory Tract. Am. J. Respir. Crit. Care Med. 2011, 184, 957–963. [Google Scholar] [CrossRef] [PubMed]
- Morris, A.; Beck, J.M.; Schloss, P.D.; Campbell, T.B.; Crothers, K.; Curtis, J.L.; Flores, S.C.; Fontenot, A.P.; Ghedin, E.; Huang, L.; et al. Comparison of the respiratory microbiome in healthy nonsmokers and smokers. Am. J. Respir. Crit. Care Med. 2013, 187, 1067–1075. [Google Scholar] [CrossRef] [PubMed]
- Gleeson, K.; Eggli, D.F.; Maxwell, S.L. Quantitative aspiration during sleep in normal subjects. Chest 1997, 111, 1266–1272. [Google Scholar] [CrossRef] [PubMed]
- Venkataraman, A.; Bassis, C.M.; Beck, J.M.; Young, V.B.; Curtis, J.L.; Huffnagle, G.B.; Schmidt, T.M. Application of a neutral community model to assess structuring of the human lung microbiome. mBio 2015, 6, e02284-14. [Google Scholar] [CrossRef]
- de Dios Caballero, J.; Vida, R.; Cobo, M.; Máiz, L.; Suárez, L.; Galeano, J.; Baquero, F.; Cantón, R.; del Campo, R.; Soléand, A.; et al. Individual Patterns of Complexity in Cystic Fibrosis Lung Microbiota, Including Predator Bacteria, over a 1-Year Period. mBio 2017, 8, e00959-17. [Google Scholar] [CrossRef] [PubMed]
- Zemanick, E.T.; Wagner, B.D.; Robertson, C.E.; Ahrens, R.C.; Chmiel, J.F.; Clancy, J.P.; Gibson, R.L.; Harris, W.T.; Kurland, G.; Laguna, T.A.; et al. Airway microbiota across age and disease spectrum in cystic fibrosis. Eur. Respir. J. 2017, 50, 1700832. [Google Scholar] [CrossRef] [PubMed]
- Segal, L.N.; Blaser, M.J. A Brave New World: The Lung Microbiota in an Era of Change. Ann. Am. Thorac. Soc. 2014, 11 (Suppl. S1), S21–S27. [Google Scholar] [CrossRef] [PubMed]
- Dickson, R.P.; Huffnagle, G.B. The Lung Microbiome: New Principles for Respiratory Bacteriology in Health and Disease. PLoS Pathog. 2015, 11, e1004923. [Google Scholar] [CrossRef] [PubMed]
- Ericsson, A.C.; Personett, A.R.; Grobman, M.E.; Rindt, H.; Reinero, C.R. Composition and Predicted Metabolic Capacity of Upper and Lower Airway Microbiota of Healthy Dogs in Relation to the Fecal Microbiota. PLoS ONE 2016, 11, e0154646. [Google Scholar] [CrossRef] [PubMed]
- Yu, G.; Gail, M.H.; Consonni, D.; Carugno, M.; Humphrys, M.; Pesatori, A.C.; Caporaso, N.E.; Goedert, J.J.; Ravel, J.; Landi, M.T. Characterizing human lung tissue microbiota and its relationship to epidemiological and clinical features. Genome Biol. 2016, 17, 163. [Google Scholar] [CrossRef] [PubMed]
- Nicola, I.; Cerutti, F.; Grego, E.; Bertone, I.; Gianella, P.; D’Angelo, A.; Peletto, S.; Bellino, C. Characterization of the upper and lower respiratory tract microbiota in Piedmontese calves. Microbiome 2017, 5, 152. [Google Scholar] [CrossRef]
- Siqueira, F.M.; Pérez-Wohlfeil, E.; Carvalho, F.M.; Trelles, O.; Schrank, I.S.; Vasconcelos, A.T.R.; Zaha, A. Microbiome overview in swine lungs. PLoS ONE 2017, 12, e0181503. [Google Scholar] [CrossRef]
- Vientos-Plotts, A.I.; Ericsson, A.C.; Rindt, H.; Reinero, C.R. Oral Probiotics Alter Healthy Feline Respiratory Microbiota. Front. Microbiol. 2017, 8, 1287. [Google Scholar] [CrossRef]
- Biesbroek, G.; Sanders, E.A.M.; Roeselers, G.; Wang, X.; Caspers, M.P.M.; Trzcinski, K.; Bogaert, D.; Keijser, B.J.F. Deep Sequencing Analyses of Low Density Microbial Communities: Working at the Boundary of Accurate Microbiota Detection. PLoS ONE 2012, 7, e32942. [Google Scholar] [CrossRef]
- Miquel, S.; Leclerc, M.; Martin, R.; Chain, F.; Lenoir, M.; Raguideau, S.; Hudault, S.; Bridonneau, C.; Northene, T.; Bowene, B.; et al. Identification of Metabolic Signatures Linked to Anti-Inflammatory Effects of Faecalibacterium prausnitzii. mBio 2015, 6, e00300-15. [Google Scholar] [CrossRef] [PubMed]
- Belvoncikova, P.; Splichalova, P.; Videnska, P.; Gardlik, R. The Human Mycobiome: Colonization, Composition and the Role in Health and Disease. J. Fungi 2022, 8, 1046. [Google Scholar] [CrossRef] [PubMed]
- Zhang, I.; Pletcher, S.D.; Goldberg, A.N.; Barker, B.M.; Cope, E.K. Fungal Microbiota in Chronic Airway Inflammatory Disease and Emerging Relationships with the Host Immune Response. Front. Microbiol. 2017, 8, 2477. [Google Scholar] [CrossRef] [PubMed]
- Bittinger, K.; Charlson, E.S.; Loy, E.; Shirley, D.J.; Haas, A.R.; Laughlin, A.; Yi, Y.; Wu, G.D.; Lewis, J.D.; Frank, I.; et al. Improved Characterization of Medically Relevant Fungi in the Human Respiratory Tract Using Next-Generation Sequencing. Genome Biol. 2014, 15, 487. [Google Scholar] [CrossRef] [PubMed]
- Charlson, E.S.; Diamond, J.M.; Bittinger, K.; Fitzgerald, A.S.; Yadav, A.; Haas, A.R.; Bushman, F.D.; Collman, R.G. Lung-enriched Organisms and Aberrant Bacterial and Fungal Respiratory Microbiota after Lung Transplant. Am. J. Respir. Crit. Care Med. 2012, 186, 536–545. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.H.; Clark, S.T.; Surendra, A.; Copeland, J.K.; Wang, P.W.; Ammar, R.; Collins, C.; Tullis, D.E.; Nislow, C.; Hwang, D.M.; et al. Global Analysis of the Fungal Microbiome in Cystic Fibrosis Patients Reveals Loss of Function of the Transcriptional Repressor Nrg1 as a Mechanism of Pathogen Adaptation. PLoS Pathog. 2015, 11, e1005308. [Google Scholar] [CrossRef]
- Liu, H.; Liang, Z.; Cao, N.; Yi, X.; Tan, X.; Liu, Z.; Wang, F.; Yang, Y.; Li, C.; Xiang, Z.; et al. Airway bacterial and fungal microbiome in chronic obstructive pulmonary disease. Med. Microecol. 2021, 7, 100035. [Google Scholar] [CrossRef]
- Rick, E.M.; Woolnough, K.F.; Seear, P.J.; Fairs, A.; Satchwell, J.; Richardson, M.; Monteiro, W.R.; Craner, M.; Bourne, M.; Wardlaw, A.J.; et al. The airway fungal microbiome in asthma. Clin. Exp. Allergy 2020, 50, 1325–1341. [Google Scholar] [CrossRef]
- Mac Aogáin, M.; Chandrasekaran, R.; Hou Lim, A.Y.; Low, T.B.; Tan, G.L.; Hassan, T.; Ong, T.H.; Qi Ng, A.H.; Bertrand, D.; Koh, J.Y.; et al. Immunological corollary of the pulmonary mycobiome in bronchiectasis: The CAMEB study. Eur. Respir. J. 2018, 52, 1800766. [Google Scholar] [CrossRef]
- mac Aogáin, M.; Tiew, P.Y.; Lim, A.Y.H.; Low, T.B.; Tan, G.L.; Hassan, T.; Ong, T.H.; Pang, S.L.; Lee, Z.Y.; Gwee, X.W.; et al. Distinct “Immunoallertypes” of Disease and High Frequencies of Sensitization in Non-Cystic Fibrosis Bronchiectasis. Am. J. Respir. Crit. Care Med. 2019, 199, 842–853. [Google Scholar] [CrossRef]
- Dickson, R.P.; Martinez, F.J.; Huffnagle, G.B. The role of the microbiome in exacerbations of chronic lung diseases. Lancet 2014, 384, 691–702. [Google Scholar] [CrossRef] [PubMed]
- Romani, L. Immunity to fungal infections. Nat. Rev. Immunol. 2011, 11, 275–288. [Google Scholar] [CrossRef] [PubMed]
- Iwasaki, A.; Medzhitov, R. Control of adaptive immunity by the innate immune system. Nat. Immunol. 2015, 16, 343–353. [Google Scholar] [CrossRef]
- Tipton, L.; Ghedin, E.; Morris, A. The lung mycobiome in the next-generation sequencing era. Virulence 2017, 8, 334–341. [Google Scholar] [CrossRef]
- Arumugam, M.; Raes, J.; Pelletier, E.; Le Paslier, D.; Yamada, T.; Mende, D.R.; Fernandes, G.R.; Tap, J.; Bruls, T.; Batto, J.M.; et al. Enterotypes of the human gut microbiome. Nature 2011, 473, 174–180. [Google Scholar] [CrossRef] [PubMed]
- Huang, C.; Yu, Y.; Du, W.; Liu, Y.; Dai, R.; Tang, W.; Wang, P.; Zhang, C.; Shi, G. Fungal and bacterial microbiome dysbiosis and imbalance of trans-kingdom network in asthma. Clin. Transl. Allergy 2020, 10, 42. [Google Scholar] [CrossRef] [PubMed]
- Sharma, A.; Laxman, B.; Naureckas, E.T.; Hogarth, D.K.; Sperling, A.I.; Solway, J.; Ober, C.; Gilbert, J.A.; White, S.R. Associations between fungal and bacterial microbiota of airways and asthma endotypes. J. Allergy Clin. Immunol. 2019, 144, 1214–1227.e7. [Google Scholar] [CrossRef]
- Soret, P.; Vandenborght, L.-E.; Francis, F.; Coron, N.; Enaud, R.; Avalos, M.; Schaeverbeke, T.; Berger, P.; Fayon, M.; Thiebaut, R.; et al. Respiratory mycobiome and suggestion of inter-kingdom network during acute pulmonary exacerbation in cystic fibrosis. Sci. Rep. 2020, 10, 3589. [Google Scholar] [CrossRef]
- Nash, A.K.; Auchtung, T.A.; Wong, M.C.; Smith, D.P.; Gesell, J.R.; Ross, M.C.; Stewart, C.J.; Metcalf, G.A.; Muzny, D.M.; Gibbs, R.A.; et al. The gut mycobiome of the Human Microbiome Project healthy cohort. Microbiome 2017, 5, 153. [Google Scholar] [CrossRef]
- Wang, Z.; Yang, Y.; Yan, Z.; Liu, H.; Chen, B.; Liang, Z.; Wang, F.; Miller, B.E.; Tal-Singer, R.; Yi, X.; et al. Multi-omic meta-analysis identifies functional signatures of airway microbiome in chronic obstructive pulmonary disease. ISME J. 2020, 14, 2748–2765. [Google Scholar] [CrossRef]
- Durack, J.; Lynch, S.V.; Nariya, S.; Bhakta, N.R.; Beigelman, A.; Castro, M.; Dyer, A.-M.; Israel, E.; Kraft, M.; Martin, R.J.; et al. Features of the bronchial bacterial microbiome associated with atopy, asthma, and responsiveness to inhaled corticosteroid treatment. J. Allergy Clin. Immunol. 2017, 140, 63–75. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.-Y.; Li, C.X.; Liang, Z.-Y.; Zhang, S.-Y.; Yang, W.-Y.; Ye, Y.-M.; Lin, Y.-X.; Chen, R.-C.; Zhou, H.-W.; Su, J. The Interactions of Airway Bacterial and Fungal Communities in Clinically Stable Asthma. Front. Microbiol. 2020, 11, 1647. [Google Scholar] [CrossRef] [PubMed]
- Eshetie, S.; van Soolingen, D. The Respiratory Microbiota: New Insights into Pulmonary Tuberculosis. BMC Infect. Dis. 2019, 19, 92. [Google Scholar] [CrossRef] [PubMed]
- Botero, L.E.; Delgado-Serrano, L.; Cepeda, M.L.; Bustos, J.R.; Anzola, J.M.; Del Portillo, P.; Robledo, J.; Zambrano, M.M. Respiratory tract clinical sample selection for microbiota analysis in patients with pulmonary tuberculosis. Microbiome 2014, 2, 29. [Google Scholar] [CrossRef] [PubMed]
- Van Der Gast, C.J.; Walker, A.W.; Stressmann, F.A.; Rogers, G.B.; Scott, P.; Daniels, T.W.; Carroll, M.P.; Parkhill, J.; Bruce, K.D. Partitioning core and satellite taxa from within cystic fibrosis lung bacterial communities. ISME J. 2010, 5, 780–791. [Google Scholar] [CrossRef] [PubMed]
- Willger, S.D.; Grim, S.L.; Dolben, E.L.; Shipunova, A.; Hampton, T.H.; Morrison, H.G.; Filkins, L.M.; O’Toole, G.A.; Moulton, L.A.; Ashare, A.; et al. Characterization and quantification of the fungal microbiome in serial samples from individuals with cystic fibrosis. Microbiome 2014, 2, 40. [Google Scholar] [CrossRef]
- Woo, T.E.; Lim, R.; Heirali, A.A.; Acosta, N.; Rabin, H.R.; Mody, C.H.; Somayaji, R.; Surette, M.G.; Sibley, C.D.; Storey, D.G.; et al. A longitudinal characterization of the Non-Cystic Fibrosis Bronchiectasis airway microbiome. Sci. Rep. 2019, 9, 6871. [Google Scholar] [CrossRef]
- Alam, M.T.; Amos, G.C.A.; Murphy, A.R.J.; Murch, S.; Wellington, E.M.H.; Arasaradnam, R.P. Microbial imbalance in inflammatory bowel disease patients at different taxonomic levels. Gut Pathog. 2020, 12, 1. [Google Scholar] [CrossRef]
- Underhill, D.M.; Braun, J. Fungal microbiome in inflammatory bowel disease: A critical assessment. J. Clin. Investig. 2022, 132, e155786. [Google Scholar] [CrossRef]
- Nadal, I.; Donant, E.; Ribes-Koninckx, C.; Calabuig, M.; Sanz, Y. Imbalance in the composition of the duodenal microbiota of children with coeliac disease. J. Med. Microbiol. 2007, 56, 1669–1674. [Google Scholar] [CrossRef]
- El Mouzan, M.; Al-Hussaini, A.; Fanelli, B.; Assiri, A.; AlSaleem, B.; Al Mofarreh, M.; Al Sarkhy, A.; Alasmi, M. Fungal Dysbiosis in Children with Celiac Disease. Dig. Dis. Sci. 2022, 67, 216–223. [Google Scholar] [CrossRef] [PubMed]
- Wang, T.; Cai, G.; Qiu, Y.; Fei, N.; Zhang, M.; Pang, X.; Jia, W.; Cai, S.; Zhao, L. Structural segregation of gut microbiota between colorectal cancer patients and healthy volunteers. ISME J. 2011, 6, 320–329. [Google Scholar] [CrossRef] [PubMed]
- Coker, O.O.; Nakatsu, G.; Dai, R.Z.; Wu, W.K.K.; Wong, S.H.; Ng, S.C.; Chan, F.K.L.; Sung, J.J.Y.; Yu, J. Enteric fungal microbiota dysbiosis and ecological alterations in colorectal cancer. Gut 2018, 68, 654–662. [Google Scholar] [CrossRef]
- De Sordi, L.; Mühlschlegel, F.A. Quorum sensing and fungal-bacterial interactions in Candida albicans: A communicative network regulating microbial coexistence and virulence. FEMS Yeast Res. 2009, 9, 990–999. [Google Scholar] [CrossRef]
- Dixon, E.F.; Hall, R.A. Noisy neighbourhoods: Quorum sensing in fungal-polymicrobial infections. Cell Microbiol. 2015, 17, 1431–1441. [Google Scholar] [CrossRef]
- Braga, R.M.; Dourado, M.N.; Araújo, W.L. Microbial interactions: Ecology in a molecular perspective. Braz. J. Microbiol. 2016, 47 (Suppl. S1), 86–98. [Google Scholar] [CrossRef] [PubMed]
- Padder, S.A.; Prasad, R.; Shah, A.H. Quorum sensing: A less known mode of communication among fungi. Microbiol. Res. 2018, 210, 51–58. [Google Scholar] [CrossRef]
- Rodrigues, C.F.; Černáková, L. Farnesol and Tyrosol: Secondary Metabolites with a Crucial quorum-sensing Role in Candida Biofilm Development. Genes 2020, 11, 444. [Google Scholar] [CrossRef]
- Cugini, C.; Calfee, M.W.; Farrow, J.M.; Morales, D.K.; Pesci, E.C.; Hogan, D.A. Farnesol, a common sesquiterpene, inhibits PQS production in Pseudomonas aeruginosa. Mol. Microbiol. 2007, 65, 896–906. [Google Scholar] [CrossRef]
- Peleg, A.Y.; Tampakakis, E.; Fuchs, B.B.; Eliopoulos, G.M.; Moellering, R.C.; Mylonakis, E. Prokaryote–eukaryote interactions identified by using Caenorhabditis elegans. Proc. Natl. Acad. Sci. USA 2008, 105, 14585–14590. [Google Scholar] [CrossRef]
- de Dios Caballero, J.; Cantón, R.; Ponce-Alonso, M.; García-Clemente, M.M.; de la Pedrosa, E.G.G.; López-Campos, J.L.; Máiz, L.; del Campo, R.; Martínez-García, M.Á. The Human Mycobiome in Chronic Respiratory Diseases: Current Situation and Future Perspectives. Microorganisms 2022, 10, 810. [Google Scholar] [CrossRef] [PubMed]
- Narunsky-Haziza, L.; Sepich-Poore, G.D.; Livyatan, I.; Asraf, O.; Martino, C.; Nejman, D.; Gavert, N.; Stajich, J.E.; Amit, G.; González, A.; et al. Pan-cancer analyses reveal cancer-type-specific fungal ecologies and bacteriome interactions. Cell 2022, 185, 3789–3806.e17. [Google Scholar] [CrossRef] [PubMed]
- Moll, H.; Sonesson, A.; Jantzen, E.; Marre, R.; Zähringer, U. Identification of 27-oxo-octacosanoic acid and heptacosane-1,27-dioic acid in Legionella pneumophila. FEMS Microbiol. Lett. 1992, 76, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Hogan, D.A.; Vik, Å.; Kolter, R. A Pseudomonas aeruginosa quorum-sensing molecule influences Candida albicans morphology. Mol. Microbiol. 2004, 54, 1212–1223. [Google Scholar] [CrossRef]
- Inoue, Y.; Shiraishi, A.; Hada, T.; Hirose, K.; Hamashima, H.; Shimada, J. The antibacterial effects of terpene alcohols on Staphylococcus aureus and their mode of action. FEMS Microbiol. Lett. 2004, 237, 325–331. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.-H.; He, Y.; Gao, Y.; Wu, J.E.; Dong, Y.-H.; He, C.; Wang, S.X.; Weng, L.-X.; Xu, J.-L.; Tay, L.; et al. A bacterial cell-cell communication signal with cross-kingdom structural analogues. Mol. Microbiol. 2004, 51, 903–912. [Google Scholar] [CrossRef] [PubMed]
- Boon, C.; Deng, Y.; Wang, L.-H.; He, Y.; Xu, J.-L.; Fan, Y.; Pan, S.Q.; Zhang, L.-H. A novel DSF-like signal from Burkholderia cenocepacia interferes with Candida albicans morphological transition. ISME J. 2008, 2, 27–36. [Google Scholar] [CrossRef]
- Davis-Hanna, A.; Piispanen, A.E.; Stateva, L.I.; Hogan, D.A. Farnesol and dodecanol effects on the Candida albicans Ras1-cAMP signalling pathway and the regulation of morphogenesis. Mol. Microbiol. 2008, 67, 47–62. [Google Scholar] [CrossRef]
- Ferreira, J.A.G.; Penner, J.C.; Moss, R.B.; Haagensen, J.A.J.; Clemons, K.V.; Spormann, A.M.; Nazik, H.; Cohen, K.; Banaei, N.; Carolino, E.; et al. Inhibition of Aspergillus fumigatus and Its Biofilm by Pseudomonas aeruginosa Is Dependent on the Source, Phenotype and Growth Conditions of the Bacterium. PLoS ONE 2015, 10, e0134692. [Google Scholar] [CrossRef]
- Briard, B.; Bomme, P.; Lechner, B.E.; Mislin, G.L.A.; Lair, V.; Prévost, M.-C.; Latgé, J.-P.; Haas, H.; Beauvais, A. Pseudomonas aeruginosa manipulates redox and iron homeostasis of its microbiota partner Aspergillus fumigatus via phenazines. Sci. Rep. 2015, 5, 8220. [Google Scholar] [CrossRef]
- Sass, G.; Nazik, H.; Penner, J.; Shah, H.; Ansari, S.R.; Clemons, K.V.; Groleau, M.-C.; Dietl, A.-M.; Visca, P.; Haas, H.; et al. Studies of Pseudomonas aeruginosa Mutants Indicate Pyoverdine as the Central Factor in Inhibition of Aspergillus fumigatus Biofilm. J. Bacteriol. 2018, 200, e00345-17. [Google Scholar] [CrossRef]
- Briard, B.; Rasoldier, V.; Bomme, P.; ElAouad, N.; Guerreiro, C.; Chassagne, P.; Muszkieta, L.; Latgé, J.-P.; Mulard, L.; Beauvais, A. Dirhamnolipids secreted from Pseudomonas aeruginosa modify anjpegungal susceptibility of Aspergillus fumigatus by inhibiting β1,3 glucan synthase activity. ISME J. 2017, 11, 1578–1591. [Google Scholar] [CrossRef] [PubMed]
- Jerry Reen, F.; Phelan, J.P.; Woods, D.F.; Shanahan, R.; Cano, R.; Clarke, S.; McGlacken, G.P.; O’Gara, F. Harnessing Bacterial Signals for Suppression of Biofilm Formation in the Nosocomial Fungal Pathogen Aspergillus fumigatus. Front. Microbiol. 2016, 7, 2074. [Google Scholar] [CrossRef]
- Penner, J.C.; Ferreira, J.A.G.; Secor, P.R.; Sweere, J.M.; Birukova, M.K.; Joubert, L.-M.; Haagensen, J.A.J.; Garcia, O.; Malkovskiy, A.V.; Kaber, G.; et al. Pf4 bacteriophage produced by Pseudomonas aeruginosa inhibits Aspergillus fumigatus metabolism via iron sequestration. Microbiology 2016, 162, 1583–1594. [Google Scholar] [CrossRef] [PubMed]
- Reece, E.; Doyle, S.; Greally, P.; Renwick, J.; McClean, S. Aspergillus fumigatus Inhibits Pseudomonas aeruginosa in Co-culture: Implications of a Mutually Antagonistic Relationship on Virulence and Inflammation in the CF Airway. Front. Microbiol. 2018, 9, 1205. [Google Scholar] [CrossRef]
- Sass, G.; Ansari, S.R.; Dietl, A.-M.; Déziel, E.; Haas, H.; Stevens, D.A. Intermicrobial interaction: Aspergillus fumigatus siderophores protect against competition by Pseudomonas aeruginosa. PLoS ONE 2019, 14, e0216085. [Google Scholar] [CrossRef]
- Briard, B.; Heddergott, C.; Latgé, J.-P. Volatile Compounds Emitted by Pseudomonas aeruginosa Stimulate Growth of the Fungal Pathogen Aspergillus fumigatus. mBio 2016, 7, e00219-16. [Google Scholar] [CrossRef]
- Manavathu, E.K.; Vager, D.L.; Vazquez, J.A. Development and antimicrobial susceptibility studies of in vitro monomicrobial and polymicrobial biofilm models with Aspergillus fumigatus and Pseudomonas aeruginosa. BMC Microbiol. 2014, 14, 53. [Google Scholar] [CrossRef]
- Melloul, E.; Luiggi, S.; Anaïs, L.; Arné, P.; Costa, J.M.; Fihman, V.; Briard, B.; Dannaoui, E.; Guillot, J.; Decousser, J.W.; et al. Characteristics of Aspergillus fumigatus in Association with Stenotrophomonas maltophilia in an In Vitro Model of Mixed Biofilm. PLoS ONE 2016, 11, e0166325. [Google Scholar] [CrossRef]
- Reece, E.; Segurado, R.; Jackson, A.; McClean, S.; Renwick, J.; Greally, P. Co-colonisation with Aspergillus fumigatus and Pseudomonas aeruginosa is associated with poorer health in cystic fibrosis patients: An Irish registry analysis. BMC Pulm. Med. 2017, 17, 70. [Google Scholar] [CrossRef]
- Amin, R.; Dupuis, A.; Aaron, S.D.; Ratjen, F. The Effect of Chronic Infection with Aspergillus fumigatus on Lung Function and Hospitalization in Patients with Cystic Fibrosis. Chest 2010, 137, 171–176. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Z.; Song, J.; Yang, C.; Yang, L.; Chen, J.; Li, X.; Wang, Y.; Feng, J. Prevalence of Fungal and Bacterial Co-Infection in Pulmonary Fungal Infections: A Metagenomic Next Generation Sequencing-Based Study. Front. Cell Infect. Microbiol. 2021, 11, 749905. [Google Scholar] [CrossRef]
- Monin, L.; Mehta, S.; Elsegeiny, W.; Gopal, R.; McAleer, J.P.; Oury, T.D.; Kolls, J.; Khader, S.A. Aspergillus fumigatus Preexposure Worsens Pathology and Improves Control of Mycobacterium abscessus Pulmonary Infection in Mice. Infect. Immun. 2018, 86, e00859-17. [Google Scholar] [CrossRef]
- Hogan, D.A.; Kolter, R. Pseudomonas—Candida Interactions: An Ecological Role for Virulence Factors. Science 2002, 296, 2229–2232. [Google Scholar] [CrossRef] [PubMed]
- Tampakakis, E.; Peleg, A.Y.; Mylonakis, E. Interaction of Candida albicans with an Intestinal Pathogen, Salmonella enterica Serovar Typhimurium. Eukaryot. Cell 2009, 8, 732–737. [Google Scholar] [CrossRef] [PubMed]
- Peleg, A.Y.; Hogan, D.A.; Mylonakis, E. Medically important bacterial–fungal interactions. Nat. Rev. Gews Microbiol. 2010, 8, 340–349. [Google Scholar] [CrossRef]
- Fitzsimmons, N.; Berry, D.R. Inhibition of Candida albicans by Lactobacillus acidophilus: Evidence for the involvement of a peroxidase system. Microbios 1994, 80, 125–133. [Google Scholar]
- Buffo, J.; Herman, M.A.; Soll, D.R. A characterization of pH-regulated dimorphism in Candida albicans. Mycopathologia 1984, 85, 21–30. [Google Scholar] [CrossRef]
- Borman, A.M.; Palmer, M.; Delhaes, L.; Carrère, J.; Favennec, L.; Ranque, S.; Gangneux, J.-P.; Horré, R.; Bouchara, J.-P. Lack of standardization in the procedures for mycological examination of sputum samples from CF patients: A possible cause for variations in the prevalence of filamentous fungi. Med. Mycol. 2010, 48 (Suppl. S1), S88–S97. [Google Scholar] [CrossRef]
- Delhaes, L.; Monchy, S.; Fréalle, E.; Hubans, C.; Salleron, J.; Leroy, S.; Prevotat, A.; Wallet, F.; Wallaert, B.; Dei-Cas, E.; et al. The Airway Microbiota in Cystic Fibrosis: A Complex Fungal and Bacterial Community—Implications for Therapeutic Management. PLoS ONE 2012, 7, e36313. [Google Scholar] [CrossRef]
- Pragman, A.A.; Kim, H.B.; Reilly, C.S.; Wendt, C.; Isaacson, R.E. The Lung Microbiome in Moderate and Severe Chronic Obstructive Pulmonary Disease. PLoS ONE 2012, 7, e47305. [Google Scholar] [CrossRef] [PubMed]
- Botterel, F.; Angebault, C.; Cabaret, O.; Stressmann, F.A.; Costa, J.-M.; Wallet, F.; Wallaert, B.; Bruce, K.; Delhaes, L. Fungal and Bacterial Diversity of Airway Microbiota in Adults with Cystic Fibrosis: Concordance Between Conventional Methods and Ultra-Deep Sequencing, and Their Practical use in the Clinical Laboratory. Mycopathologia 2018, 183, 171–183. [Google Scholar] [CrossRef] [PubMed]
- Delhaes, L.; Touati, K.; Faure-Cognet, O.; Cornet, M.; Botterel, F.; Dannaoui, E.; Morio, F.; Le Pape, P.; Grenouillet, F.; Favennec, L.; et al. Prevalence, geographic risk factor, and development of a standardized protocol for fungal isolation in cystic fibrosis: Results from the international prospective study “MFIP”. J. Cyst. Fibros. 2019, 18, 212–220. [Google Scholar] [CrossRef] [PubMed]
- Shirazi, F.; Ferreira, J.A.G.; Stevens, D.A.; Clemons, K.V.; Kontoyiannis, D.P. Biofilm Filtrates of Pseudomonas aeruginosa Strains Isolated from Cystic Fibrosis Patients Inhibit Preformed Aspergillus fumigatus Biofilms via Apoptosis. PLoS ONE 2016, 11, e0150155. [Google Scholar] [CrossRef]
- Loussert, C.; Schmitt, C.; Prevost, M.-C.; Balloy, V.; Fadel, E.; Philippe, B.; Kauffmann-Lacroix, C.; Latgé, J.P.; Beauvais, A. In vivo biofilm composition of Aspergillus fumigatus. Cell Microbiol. 2010, 12, 405–410. [Google Scholar] [CrossRef]
- Jakobi, M.; Winkelmann, G.; Kaiser, D.; Kempter, C.; Jung, G.; Berg, G.; Bahl, H. Maltophilin: A New Antifungal Compound Produced by Stenotrophomonas maltophilia R3089. J. Antibiot. 1996, 49, 1101–1104. [Google Scholar] [CrossRef]
- VAN Gennip, M.; Christensen, L.D.; Alhede, M.; Phipps, R.; Jensen, P.; Christophersen, L.; Pamp, S.J.; Moser, C.; Mikkelsen, P.J.; Koh, A.Y.; et al. Inactivation of the rhlA gene in Pseudomonas aeruginosa prevents rhamnolipid production, disabling the protection against polymorphonuclear leukocytes. Apmis 2009, 117, 537–546. [Google Scholar] [CrossRef]
- Li, S.; Calvo, A.M.; Yuen, G.Y.; DU, L.; Harris, S.D. Induction of Cell Wall Thickening by the Antifungal Compound Dihydromaltophilin Disrupts Fungal Growth and is Mediated by Sphingolipid Biosynthesis. J. Eukaryot. Microbiol. 2009, 56, 182–187. [Google Scholar] [CrossRef]
- Smith, K.; Rajendran, R.; Kerr, S.; Lappin, D.F.; Mackay, W.G.; Williams, C.; Ramage, G. Aspergillus fumigatus enhances elastase production in Pseudomonas aeruginosa co-cultures. Med. Mycol. 2015, 53, 645–655. [Google Scholar] [CrossRef]
- Briard, B.; Mislin, G.L.A.; Latgé, J.P.J.-P.; Beauvais, A. Interactions between Aspergillus fumigatus and Pulmonary Bacteria: Current State of the Field, New Data, and Future Perspective. J. Fungi 2019, 5, 48. [Google Scholar] [CrossRef]
- Sass, G.; Nazik, H.; Penner, J.; Shah, H.; Ansari, S.R.; Clemons, K.V.; Groleau, M.-C.; Dietl, A.-M.; Visca, P.; Haas, H.; et al. Aspergillus-Pseudomonas interaction, relevant to competition in airways. Med. Mycol. 2019, 57, S228–S232. [Google Scholar] [CrossRef]
- Mowat, E.; Rajendran, R.; Williams, C.; McCulloch, E.; Jones, B.; Lang, S.; Ramage, G. Pseudomonas aeruginosa and their small diffusible extracellular molecules inhibit Aspergillus fumigatus biofilm formation. FEMS Microbiol. Lett. 2010, 313, 96–102. [Google Scholar] [CrossRef] [PubMed]
- Buffie, C.G.; Pamer, E.G. Microbiota-mediated colonization resistance against intestinal pathogens. Nat. Rev. Immunol. 2013, 13, 790–801. [Google Scholar] [CrossRef] [PubMed]
- Bowerman, K.L.; Rehman, S.F.; Vaughan, A.; Lachner, N.; Budden, K.F.; Kim, R.Y.; Wood, D.L.A.; Gellatly, S.L.; Shukla, S.D.; Wood, L.G.; et al. Disease-associated gut microbiome and metabolome changes in patients with chronic obstructive pulmonary disease. Nat. Commun. 2020, 11, 5886. [Google Scholar] [CrossRef] [PubMed]
- Sun, Z.; Zhu, Q.; Shen, Y.; Yan, T.; Zhou, X. Dynamic changes of gut and lung microorganisms during chronic obstructive pulmonary disease exacerbations. Kaohsiung J. Med. Sci. 2020, 36, 107–113. [Google Scholar] [CrossRef] [PubMed]
- Zhang, D.; Wang, Y.; He, K.; Yang, Q.; Gong, M.; Ji, M.; Chen, L. Wolbachia limits pathogen infections through induction of host innate immune responses. PLoS ONE 2020, 15, e0226736. [Google Scholar] [CrossRef]
- Budden, K.F.; Shukla, S.D.; Rehman, S.F.; Bowerman, K.L.; Keely, S.; Hugenholtz, P.; Armstrong-James, D.P.H.; Adcock, I.M.; Chotirmall, S.H.; Chung, K.F.; et al. Functional effects of the microbiota in chronic respiratory disease. Lancet Respir. Med. 2019, 7, 907–920. [Google Scholar] [CrossRef]
- Qu, L.; Cheng, Q.; Wang, Y.; Mu, H.; Zhang, Y. COPD and Gut–Lung Axis: How Microbiota and Host Inflammasome Influence COPD and Related Therapeutics. Front. Microbiol. 2022, 13, 868086. [Google Scholar] [CrossRef]
- Southam, D.S.; Dolovich, M.; O’Byrne, P.M.; Inman, M.D. Distribution of intranasal instillations in mice: Effects of volume, time, body position, and anesthesia. Am. J. Physiol. Cell Mol. Physiol. 2002, 282, L833–L839. [Google Scholar] [CrossRef]
- Enaud, R.; Prevel, R.; Ciarlo, E.; Beaufils, F.; Wieërs, G.; Guery, B.; Delhaes, L. The Gut-Lung Axis in Health and Respiratory Diseases: A Place for Inter-Organ and Inter-Kingdom Crosstalks. Front. Cell Infect. Microbiol. 2020, 10, 9. [Google Scholar] [CrossRef]
- Skelly, A.N.; Sato, Y.; Kearney, S.; Honda, K. Mining the microbiota for microbial and metabolite-based immunotherapies. Nat. Rev. Immunol. 2019, 19, 305–323. [Google Scholar] [CrossRef]
- Mazmanian, S.K.; Liu, C.H.; Tzianabos, A.O.; Kasper, D.L. An Immunomodulatory Molecule of Symbiotic Bacteria Directs Maturation of the Host Immune System. Cell 2005, 122, 107–118. [Google Scholar] [CrossRef] [PubMed]
- Neish, A.S.; Gewirtz, A.T.; Zeng, H.; Young, A.N.; Hobert, M.E.; Karmali, V.; Rao, A.S.; Madara, J.L. Prokaryotic Regulation of Epithelial Responses by Inhibition of Ikappa B-alpha Ubiquitination. Science 2000, 289, 1560–1563. [Google Scholar] [CrossRef] [PubMed]
- Gollwitzer, E.S.; Saglani, S.; Trompette, A.; Yadava, K.; Sherburn, R.; McCoy, K.D.; Nicod, L.P.; Lloyd, C.M.; Marsland, B.J. Lung microbiota promotes tolerance to allergens in neonates via PD-L1. Nat. Med. 2014, 20, 642–647. [Google Scholar] [CrossRef] [PubMed]
- Herbst, T.; Sichelstiel, A.; Schär, C.; Yadava, K.; Bürki, K.; Cahenzli, J.; McCoy, K.; Marsland, B.J.; Harris, N.L. Dysregulation of allergic airway inflammation in the absence of microbial colonization. Am. J. Respir. Crit. Care Med. 2011, 184, 198–205. [Google Scholar] [CrossRef]
- Russell, S.L.; Gold, M.J.; Hartmann, M.; Willing, B.P.; Thorson, L.; Wlodarska, M.; Gill, N.; Blanchet, M.; Mohn, W.W.; McNagny, K.M.; et al. Early life antibiotic-driven changes in microbiota enhance susceptibility to allergic asthma. EMBO Rep. 2012, 13, 440–447. [Google Scholar] [CrossRef] [PubMed]
- Allie, S.R.; Bradley, J.E.; Mudunuru, U.; Schultz, M.D.; Graf, B.A.; Lund, F.E.; Randall, T.D. The establishment of resident memory B cells in the lung requires local antigen encounter. Nat. Immunol. 2018, 20, 97–108. [Google Scholar] [CrossRef] [PubMed]
- Samuelson, D.R.; Welsh, D.A.; Shellito, J.E. Regulation of lung immunity and host defense by the intestinal microbiota. Front. Microbiol. 2015, 6, 1085. [Google Scholar] [CrossRef]
- Abreu, M.T. Toll-like receptor signalling in the intestinal epithelium: How bacterial recognition shapes intestinal function. Nat. Rev. Immunol. 2010, 10, 131–143. [Google Scholar] [CrossRef]
- Round, J.L.; Lee, S.M.; Li, J.; Tran, G.; Jabri, B.; Chatila, T.A.; Mazmanian, S.K. The Toll-Like Receptor 2 Pathway Establishes Colonization by a Commensal of the Human Microbiota. Science 2011, 332, 974–977. [Google Scholar] [CrossRef]
- Fagundes, C.T.; Amaral, F.A.; Vieira, A.T.; Soares, A.C.; Pinho, V.; Nicoli, J.R.; Vieira, L.Q.; Teixeira, M.M.; Souza, D.G. Transient TLR Activation Restores Inflammatory Response and Ability to Control Pulmonary Bacterial Infection in Germfree Mice. J. Immunol. 2012, 188, 1411–1420. [Google Scholar] [CrossRef] [PubMed]
- Sigmundsdottir, H.; Butcher, E.C. Environmental cues, dendritic cells and the programming of tissue-selective lymphocyte trafficking. Nat. Immunol. 2008, 9, 981–987. [Google Scholar] [CrossRef] [PubMed]
- Mikhak, Z.; Strassner, J.P.; Luster, A.D. Lung dendritic cells imprint T cell lung homing and promote lung immunity through the chemokine receptor CCR4. J. Exp. Med. 2013, 210, 1855–1869. [Google Scholar] [CrossRef] [PubMed]
- Ruane, D.; Brane, L.; Reis, B.S.; Cheong, C.; Poles, J.; Do, Y.; Zhu, H.; Velinzon, K.; Choi, J.-H.; Studt, N.; et al. Lung dendritic cells induce migration of protective T cells to the gastrointestinal tract. J. Exp. Med. 2013, 210, 1871–1888. [Google Scholar] [CrossRef] [PubMed]
- Bacher, P.; Hohnstein, T.; Beerbaum, E.; Röcker, M.; Blango, M.G.; Kaufmann, S.; Röhmel, J.; Eschenhagen, P.; Grehn, C.; Seidel, K.; et al. Human Anti-fungal Th17 Immunity and Pathology Rely on Cross-Reactivity against Candida albicans. Cell 2019, 176, 1340–1355.e15. [Google Scholar] [CrossRef]
- Cai, L.; Gao, P.; Wang, Z.; Dai, C.; Ning, Y.; Ilkit, M.; Xue, X.; Xiao, J.; Chen, C. Lung and gut microbiomes in pulmonary aspergillosis: Exploring adjunctive therapies to combat the disease. Front. Immunol. 2022, 13, 988708. [Google Scholar] [CrossRef]
- Noverr, M.C.; Huffnagle, G.B. Does the Microbiota Regulate Immune Responses Outside the Gut? Trends Microbiol. 2004, 12, 562–568. [Google Scholar] [CrossRef]
- Noverr, M.C.; Falkowski, N.R.; McDonald, R.A.; McKenzie, A.N.; Huffnagle, G.B. Development of Allergic Airway Disease in Mice following Antibiotic Therapy and Fungal Microbiota Increase: Role of Host Genetics, Antigen, and Interleukin-13. Infect. Immun. 2005, 73, 30–38. [Google Scholar] [CrossRef]
- Smeekens, S.P.; Van De Veerdonk, F.L.; Van Der Meer, J.W.M.; Kullberg, B.J.; Joosten, L.A.B.; Netea, M.G. The Candida Th17 response is dependent on mannan- and -glucan-induced prostaglandin E2. Int. Immunol. 2010, 22, 889–895. [Google Scholar] [CrossRef]
- Kim, Y.-G.; Udayanga, K.G.S.; Totsuka, N.; Weinberg, J.B.; Núñez, G.; Shibuya, A. Gut Dysbiosis Promotes M2 Macrophage Polarization and Allergic Airway Inflammation via Fungi-Induced PGE2. Cell Host Microbe 2014, 15, 95–102. [Google Scholar] [CrossRef]
- Noverr, M.C.; Noggle, R.M.; Toews, G.B.; Huffnagle, G.B. Role of Antibiotics and Fungal Microbiota in Driving Pulmonary Allergic Responses. Infect. Immun. 2004, 72, 4996–5003. [Google Scholar] [CrossRef]
- Hill, D.A.; Siracusa, M.C.; Abt, M.C.; Kim, B.S.; Kobuley, D.; Kubo, M.; Kambayashi, T.; LaRosa, D.F.; Renner, E.D.; Orange, J.S.; et al. Commensal bacteria–derived signals regulate basophil hematopoiesis and allergic inflammation. Nat. Med. 2012, 18, 538–546. [Google Scholar] [CrossRef] [PubMed]
- Han, M.L.K.; Huang, Y.J.; Lipuma, J.J.; Boushey, H.A.; Boucher, R.C.; Cookson, W.O.; Curtis, J.L.; Erb-Downward, J.; Lynch, S.V.; Sethi, S.; et al. Significance of the microbiome in obstructive lung disease. Thorax 2012, 67, 456–463. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Bafadhel, M.; Haldar, K.; Spivak, A.; Mayhew, D.; Miller, B.E.; Tal-Singer, R.; Johnston, S.L.; Ramsheh, M.Y.; Barer, M.R.; et al. Lung microbiome dynamics in COPD exacerbations. Eur. Respir. J. 2016, 47, 1082–1092. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Singh, R.; Miller, B.E.; Tal-Singer, R.; Van Horn, S.; Tomsho, L.; Mackay, A.; Allinson, J.P.; Webb, A.J.; Brookes, A.J.; et al. Sputum microbiome temporal variability and dysbiosis in chronic obstructive pulmonary disease exacerbations: An analysis of the COPDMAP study. Thorax 2018, 73, 331–338. [Google Scholar] [CrossRef] [PubMed]
- Mayhew, D.; Devos, N.; Lambert, C.; Brown, J.R.; Clarke, S.C.; Kim, V.L.; Magid-Slav, M.; Miller, B.E.; Ostridge, K.K.; Patel, R.; et al. Longitudinal profiling of the lung microbiome in the AERIS study demonstrates repeatability of bacterial and eosinophilic COPD exacerbations. Thorax 2018, 73, 422–430. [Google Scholar] [CrossRef]
- Sinha, R.; Weissenburger-Moser, L.A.; Clarke, J.L.; Smith, L.M.; Heires, A.J.; Romberger, D.J.; LeVan, T.D. Short term dynamics of the sputum microbiome among COPD patients. PLoS ONE 2018, 13, e0191499. [Google Scholar] [CrossRef] [PubMed]
- Dima, E.; Kyriakoudi, A.; Kaponi, M.; Vasileiadis, I.; Stamou, P.; Koutsoukou, A.; Koulouris, N.G.; Rovina, N. The lung microbiome dynamics between stability and exacerbation in chronic obstructive pulmonary disease (COPD): Current perspectives. Respir. Med. 2019, 157, 1–6. [Google Scholar] [CrossRef]
- Singh, R.; Mackay, A.J.; Patel, A.R.C.; Garcha, D.S.; Kowlessar, B.S.; Brill, S.E.; Donnelly, L.E.; Barnes, P.J.; Donaldson, G.C.; Wedzicha, J.A. Inflammatory thresholds and the species-specific effects of colonising bacteria in stable chronic obstructive pulmonary disease. Respir. Res. 2014, 15, 114. [Google Scholar] [CrossRef]
- Barker, B.L.; Haldar, K.; Patel, H.; Pavord, I.D.; Barer, M.R.; Brightling, C.E.; Bafadhel, M. Association Between Pathogens Detected Using Quantitative Polymerase Chain Reaction with Airway Inflammation in COPD at Stable State and Exacerbations. Chest 2015, 147, 46–55. [Google Scholar] [CrossRef]
- Sze, M.A.; Dimitriu, P.A.; Suzuki, M.; McDonough, J.E.; Campbell, J.D.; Brothers, J.F.; Erb-Downward, J.R.; Huffnagle, G.B.; Hayashi, S.; Elliott, W.M.; et al. Host Response to the Lung Microbiome in Chronic Obstructive Pulmonary Disease. Am. J. Respir. Crit. Care Med. 2015, 192, 438–445. [Google Scholar] [CrossRef] [PubMed]
- Segal, L.N.; Alekseyenko, A.V.; Clemente, J.C.; Kulkarni, R.; Wu, B.; Chen, H.; Berger, K.I.; Goldring, R.M.; Rom, W.N.; Blaser, M.J.; et al. Enrichment of lung microbiome with supraglottic taxa is associated with increased pulmonary inflammation. Microbiome 2013, 1, 19. [Google Scholar] [CrossRef] [PubMed]
- Segal, L.N.; Clemente, J.C.; Tsay, J.-C.J.; Koralov, S.B.; Keller, B.C.; Wu, B.G.; Alison, M.; Shen, N.; Ghedin, E.; Morris, A.; et al. Enrichment of the lung microbiome with oral taxa is associated with lung inflammation of a Th17 phenotype. Nat. Microbiol. 2016, 1, 16031. [Google Scholar] [CrossRef] [PubMed]
- Jones, B.; Donovan, C.; Liu, G.; Gomez, H.M.; Chimankar, V.; Harrison, C.L.; Wiegman, C.H.; Adcock, I.M.; Knight, D.A.; Hirota, J.A.; et al. Animal models of COPD: What do they tell us? Respirology 2017, 22, 21–32. [Google Scholar] [CrossRef]
- Afessa, B.; Morales, I.J.; Scanlon, P.D.; Peters, S.G. Prognostic factors, clinical course, and hospital outcome of patients with chronic obstructive pulmonary disease admitted to an intensive care unit for acute respiratory failure. Crit. Care Med. 2002, 30, 1610–1615. [Google Scholar] [CrossRef]
- Guinea, J.; Torres-Narbona, M.; Gijón, P.; Muñoz, P.; Pozo, F.; Peláez, T.; de Miguel, J.; Bouza, E. Pulmonary aspergillosis in patients with chronic obstructive pulmonary disease: Incidence, risk factors, and outcome. Clin. Microbiol. Infect. 2010, 16, 870–877. [Google Scholar] [CrossRef]
- Huerta, A.; Soler, N.; Esperatti, M.; Guerrero, M.; Menendez, R.; Gimeno, A.; Zalacaín, R.; Mir, N.; Aguado, J.M.; Torres, A. Importance of Aspergillus spp. isolation in Acute exacerbations of severe COPD: Prevalence, factors and follow-up: The FUNGI-COPD study. Respir. Res. 2014, 15, 17. [Google Scholar] [CrossRef]
- Cui, L.; Lucht, L.; Tipton, L.; Rogers, M.B.; Fitch, A.; Kessinger, C.; Camp, D.; Kingsley, L.; Leo, N.; Greenblatt, R.M.; et al. Topographic Diversity of the Respiratory Tract Mycobiome and Alteration in HIV and Lung Disease. Am. J. Respir. Crit. Care Med. 2015, 191, 932–942. [Google Scholar] [CrossRef]
- Tawfik, O.W.; Papasian, C.J.; Dixon, A.Y.; Potter, L.M.P. Saccharomyces cerevisiae pneumonia in a patient with acquired immune deficiency syndrome. J. Clin. Microbiol. 1989, 27, 1689–1691. [Google Scholar] [CrossRef]
- Chowdhary, A.; Agarwal, K.; Kathuria, S.; Singh, P.K.; Roy, P.; Gaur, S.N.; de Hoog, G.S.; Meis, J.F. Clinical Significance of Filamentous Basidiomycetes Illustrated by Isolates of the Novel Opportunist Ceriporia lacerata from the Human Respiratory Tract. J. Clin. Microbiol. 2013, 51, 585–590. [Google Scholar] [CrossRef]
- de la Cámara, R.; Pinilla, I.; Muñoz, E.; Buendía, B.; Steegmann, J.L.; Fernández-Rañada, J.M. Penicillium Brevicompactum as the Cause of a Necrotic Lung Ball in an Allogeneic Bone Marrow Transplant Recipient. Bone Marrow Transpl. 1996, 18, 1189–1193. [Google Scholar]
- Christenson, S.A.; van den Berge, M.; Faiz, A.; Inkamp, K.; Bhakta, N.; Bonser, L.R.; Zlock, L.T.; Barjaktarevic, I.Z.; Barr, R.G.; Bleecker, E.R.; et al. An airway epithelial IL-17A response signature identifies a steroid-unresponsive COPD patient subgroup. J. Clin. Investig. 2019, 129, 169–181. [Google Scholar] [CrossRef]
- Rouzéa, A.; Cottereau, A.; Nseir, S. Chronic obstructive pulmonary disease and the risk for ventilator-associated pneumonia. Curr. Opin. Crit. Care 2014, 20, 525–531. [Google Scholar] [CrossRef]
- Tiew, P.Y.; San Ko, F.W.; Pang, S.L.; Matta, S.A.; Sio, Y.Y.; Poh, M.E.; Lau, K.J.X.; Mac Aogáin, M.; Jaggi, T.K.; Ivan, F.X.; et al. Environmental fungal sensitisation associates with poorer clinical outcomes in COPD. Eur. Respir. J. 2020, 56, 2000418. [Google Scholar] [CrossRef] [PubMed]
- Valenza, G.; Tappe, D.; Turnwald, D.; Frosch, M.; König, C.; Hebestreit, H.; Abele-Horn, M. Prevalence and antimicrobial susceptibility of microorganisms isolated from sputa of patients with cystic fibrosis. J. Cyst. Fibros. 2008, 7, 123–127. [Google Scholar] [CrossRef] [PubMed]
- Filkins, L.M.; Hampton, T.H.; Gifford, A.H.; Gross, M.J.; Hogan, D.A.; Sogin, M.L.; Morrison, H.G.; Paster, B.J.; O’Toole, G.A. Prevalence of Streptococci and Increased Polymicrobial Diversity Associated with Cystic Fibrosis Patient Stability. J. Bacteriol. 2012, 194, 4709–4717. [Google Scholar] [CrossRef] [PubMed]
- Eickmeier, O.; Rieber, N.; Eckrich, J.; Hector, A.; Graeppler-Mainka, U.; Hartl, D. Immune Response, Diagnosis and Treatment of Allergic Bronchopulmonary Aspergillosis in Cystic Fibrosis Lung Disease. Curr. Pharm. Des. 2013, 19, 3669–3678. [Google Scholar] [CrossRef]
- Iliev, I.D.; Funari, V.A.; Taylor, K.D.; Nguyen, Q.; Reyes, C.N.; Strom, S.P.; Brown, J.; Becker, C.A.; Fleshner, P.R.; Dubinsky, M.; et al. Interactions Between Commensal Fungi and the C-Type Lectin Receptor Dectin-1 Influence Colitis. Science 2012, 336, 1314–1317. [Google Scholar] [CrossRef]
- Chai, L.Y.A.; De Boer, M.G.J.; Van Der Velden, W.J.F.M.; Plantinga, T.S.; van Spriel, A.B.; Jacobs, C.; Halkes, C.J.M.; Vonk, A.G.; Blijlevens, N.M.; Van Dissel, J.T.; et al. The Y238X Stop Codon Polymorphism in the Human β-Glucan Receptor Dectin-1 and Susceptibility to Invasive Aspergillosis. J. Infect. Dis. 2011, 203, 736–743. [Google Scholar] [CrossRef]
- Sainz, J.; Lupiáñez, C.B.; Segura-Catena, J.; Vazquez, L.; Ríos, R.; Oyonarte, S.; Hemminki, K.; Försti, A.; Jurado, M. Dectin-1 and DC-SIGN Polymorphisms Associated with Invasive Pulmonary Aspergillosis Infection. PLoS ONE 2012, 7, e32273. [Google Scholar] [CrossRef]
- Hunter, R.C.; Klepac-Ceraj, V.; Lorenzi, M.M.; Grotzinger, H.; Martin, T.R.; Newman, D.K. Phenazine Content in the Cystic Fibrosis Respiratory Tract Negatively Correlates with Lung Function and Microbial Complexity. Am. J. Respir. Cell Mol. Biol. 2012, 47, 738–745. [Google Scholar] [CrossRef] [PubMed]
- Sass, G.; Nazik, H.; Chatterjee, P.; Shrestha, P.; Groleau, M.-C.; Déziel, E.; Stevens, D.A. Altered Pseudomonas Strategies to Inhibit Surface Aspergillus Colonies. Front. Cell Infect. Microbiol. 2021, 11, 734296. [Google Scholar] [CrossRef] [PubMed]
- Kerr, J. Inhibition of fungal growth by Pseudomonas aeruginosa and Pseudomonas cepacia isolated from patients with cystic fibrosis. J. Infect. 1994, 28, 305–310. [Google Scholar] [CrossRef] [PubMed]
- Moree, W.J.; Phelan, V.V.; Wu, C.-H.; Bandeira, N.; Cornett, D.S.; Duggan, B.M.; Dorrestein, P.C. Interkingdom metabolic transformations captured by microbial imaging mass spectrometry. Proc. Natl. Acad. Sci. USA 2012, 109, 13811–13816. [Google Scholar] [CrossRef]
- Paugam, A.; Baixench, M.-T.; Demazes-Dufeu, N.; Burgel, P.-R.; Sauter, E.; Kanaan, R.; Dusser, D.; Dupouy-Camet, J.; Hubert, D. Characteristics and consequences of airway colonization by filamentous fungi in 201 adult patients with cystic fibrosis in France. Med. Mycol. 2010, 48 (Suppl. S1), S32–S36. [Google Scholar] [CrossRef]
- Phelan, V.V.; Moree, W.J.; Aguilar, J.; Cornett, D.S.; Koumoutsi, A.; Noble, S.M.; Pogliano, K.; Guerrero, C.A.; Dorrestein, P.C. Impact of a Transposon Insertion in phzF2 on the Specialized Metabolite Production and Interkingdom Interactions of Pseudomonas aeruginosa. J. Bacteriol. 2014, 196, 1683–1693. [Google Scholar] [CrossRef]
- Keown, K.; Reid, A.; Moore, J.E.; Taggart, C.C.; Downey, D.G. Coinfection with Pseudomonas aeruginosa and Aspergillus fumigatus in cystic fibrosis. Eur. Respir. Rev. 2020, 29, 200011. [Google Scholar] [CrossRef]
- Godet, C.; Philippe, B.; Laurent, F.; Cadranel, J. Chronic Pulmonary Aspergillosis: An Update on Diagnosis and Treatment. Respiration 2014, 88, 162–174. [Google Scholar] [CrossRef]
- Kosmidis, C.; Denning, D.W. The clinical spectrum of pulmonary aspergillosis. Thorax 2015, 70, 270–277. [Google Scholar] [CrossRef]
- Oliveira, M.; Pinto, M.; Simões, H.; Gomes, J.; Veríssimo, C.; Sabino, R. Molecular detection of Aspergillus in respiratory samples collected from patients at higher risk of chronic pulmonary aspergillosis. Infect. Dis. Now 2022, 53, 104633. [Google Scholar] [CrossRef]
- Wardlaw, A.J.; Rick, E.M.; Ozyigit, L.P.; Scadding, A.; Gaillard, E.A.; Pashley, C.H. New Perspectives in the Diagnosis and Management of Allergic Fungal Airway Disease. J. Asthma Allergy 2021, 14, 557–573. [Google Scholar] [CrossRef] [PubMed]
- Cui, L.; Morris, A.; Ghedin, E. The human mycobiome in health and disease. Genome Med. 2013, 5, 63. [Google Scholar] [CrossRef]
- Kwizera, R.; Katende, A.; Bongomin, F.; Nakiyingi, L.; Kirenga, B.J. Misdiagnosis of chronic pulmonary aspergillosis as pulmonary tuberculosis at a tertiary care center in Uganda: A case series. J. Med. Case Rep. 2021, 15, 140. [Google Scholar] [CrossRef]
- Marsland, B.J.; Gollwitzer, E.S. Host–microorganism interactions in lung diseases. Nat. Rev. Immunol. 2014, 14, 827–835. [Google Scholar] [CrossRef] [PubMed]
- Rogers, G.B.; Shaw, D.; Marsh, R.L.; Carroll, M.P.; Serisier, D.J.; Bruce, K.D. Respiratory microbiota: Addressing clinical questions, informing clinical practice. Thorax 2015, 70, 74–81. [Google Scholar] [CrossRef] [PubMed]
- Gensollen, T.; Iyer, S.S.; Kasper, D.L.; Blumberg, R.S. How colonization by microbiota in early life shapes the immune system. Science 2016, 352, 539–544. [Google Scholar] [CrossRef] [PubMed]
- Faner, R.; Sibila, O.; Agustí, A.; Bernasconi, E.; Chalmers, J.D.; Huffnagle, G.B.; Manichanh, C.; Molyneaux, P.L.; Paredes, R.; Brocal, V.P.; et al. The microbiome in respiratory medicine: Current challenges and future perspectives. Eur. Respir. J. 2017, 49, 1602086. [Google Scholar] [CrossRef]
- Waldor, M.K.; Tyson, G.; Borenstein, E.; Ochman, H.; Moeller, A.; Finlay, B.B.; Kong, H.H.; Gordon, J.I.; Nelson, K.E.; Dabbagh, K.; et al. Where Next for Microbiome Research? PLoS Biol. 2015, 13, e1002050. [Google Scholar] [CrossRef]
- Clooney, A.G.; Fouhy, F.; Sleator, R.D.; O’Driscoll, A.; Stanton, C.; Cotter, P.D.; Claesson, M.J. Comparing Apples and Oranges? Next Generation Sequencing and Its Impact on Microbiome Analysis. PLoS ONE 2016, 11, e0148028. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rozaliyani, A.; Antariksa, B.; Nurwidya, F.; Zaini, J.; Setianingrum, F.; Hasan, F.; Nugrahapraja, H.; Yusva, H.; Wibowo, H.; Bowolaksono, A.; et al. The Fungal and Bacterial Interface in the Respiratory Mycobiome with a Focus on Aspergillus spp. Life 2023, 13, 1017. https://doi.org/10.3390/life13041017
Rozaliyani A, Antariksa B, Nurwidya F, Zaini J, Setianingrum F, Hasan F, Nugrahapraja H, Yusva H, Wibowo H, Bowolaksono A, et al. The Fungal and Bacterial Interface in the Respiratory Mycobiome with a Focus on Aspergillus spp. Life. 2023; 13(4):1017. https://doi.org/10.3390/life13041017
Chicago/Turabian StyleRozaliyani, Anna, Budhi Antariksa, Fariz Nurwidya, Jamal Zaini, Findra Setianingrum, Firman Hasan, Husna Nugrahapraja, Humaira Yusva, Heri Wibowo, Anom Bowolaksono, and et al. 2023. "The Fungal and Bacterial Interface in the Respiratory Mycobiome with a Focus on Aspergillus spp." Life 13, no. 4: 1017. https://doi.org/10.3390/life13041017
APA StyleRozaliyani, A., Antariksa, B., Nurwidya, F., Zaini, J., Setianingrum, F., Hasan, F., Nugrahapraja, H., Yusva, H., Wibowo, H., Bowolaksono, A., & Kosmidis, C. (2023). The Fungal and Bacterial Interface in the Respiratory Mycobiome with a Focus on Aspergillus spp. Life, 13(4), 1017. https://doi.org/10.3390/life13041017








