Abstract
The recent pandemic of COVID-19 caused by the SARS-CoV-2 virus has brought upon the world an unprecedented challenge. During its acute dissemination, a rush for vaccines started, making the scientific community come together and contribute to the development of efficient therapeutic agents and vaccines. Natural products have been used as sources of individual molecules and extracts capable of inhibiting/neutralizing several microorganisms, including viruses. Natural extracts have shown effective results against the coronavirus family, when first tested in the outbreak of SARS-CoV-1, back in 2002. In this review, the relationship between natural extracts and SARS-CoV is discussed, while also providing insight into misinformation regarding the use of plants as possible therapeutic agents. Studies with plant extracts on coronaviruses are presented, as well as the main inhibition assays and trends for the future regarding the yet unknown long-lasting effects post-infection with SARS-CoV-2.
4. Plant Metabolites: Debunking Misconceptions
The COVID-19 pandemic showed that even in the face of worldwide unrest, millions of infections and deaths, certain denier movements and organizations were still questioning the pandemic, its origins, and, in some cases, proposing homemade remedies and questionable medications to fight the virus or infection symptoms. These groups were fueled by misinformation mainly spread through the Internet, specifically social media [27]. Among the proposed remedies, many of them based on empirical knowledge, using simple logic, it was assumed that if the symptoms of COVID-19 were those of the common flu then the same homemade remedies could help and cure the infection. These remedies and Chinese medicine-based papers have been published [28,29,30,31,32,33,34] alongside others that claim that specific vitamins and other remedies may improve the recovery or boost the immune system against SARS-CoV-2, but not without controversy [35,36,37,38]. One example of these misconceptions was the use of chloroquine and hydroxychloroquine (antimalarial drugs) against covid, which after much research was deemed to not have any positive effect on overall mortality, ventilation needs and hospitalization, despite promising results in vitro. Thus, it remains to be confirmed if it has any effect on the early stages of the disease [39,40], and one report event accounts for a worsening of the clinical status [41]. Another proposed drug that was proposed as a treatment for COVID-19 was ivermectin, an antiparasitic treatment. Several publications have also debunked its use, claiming that the drug did not result in a lower incidence of admission to hospitals and prolonged emergency observation [42]. Furthermore, in another clinical trial, neither ivermectin or metformin and fluvoxamine showed effects on hypoxemia, hospitalization or death of patients with COVID-19 [43].
While there is generalized scientific support that plants and their metabolites may play a role in drugs against COVID, its prophylaxis, or be used as active agents in fomites or textiles, the approach to using natural molecules should be the same as any other molecule. A big misconception present in most societies is that natural molecules are always safer, more effective, and sustainable, although this is not always the case [44].
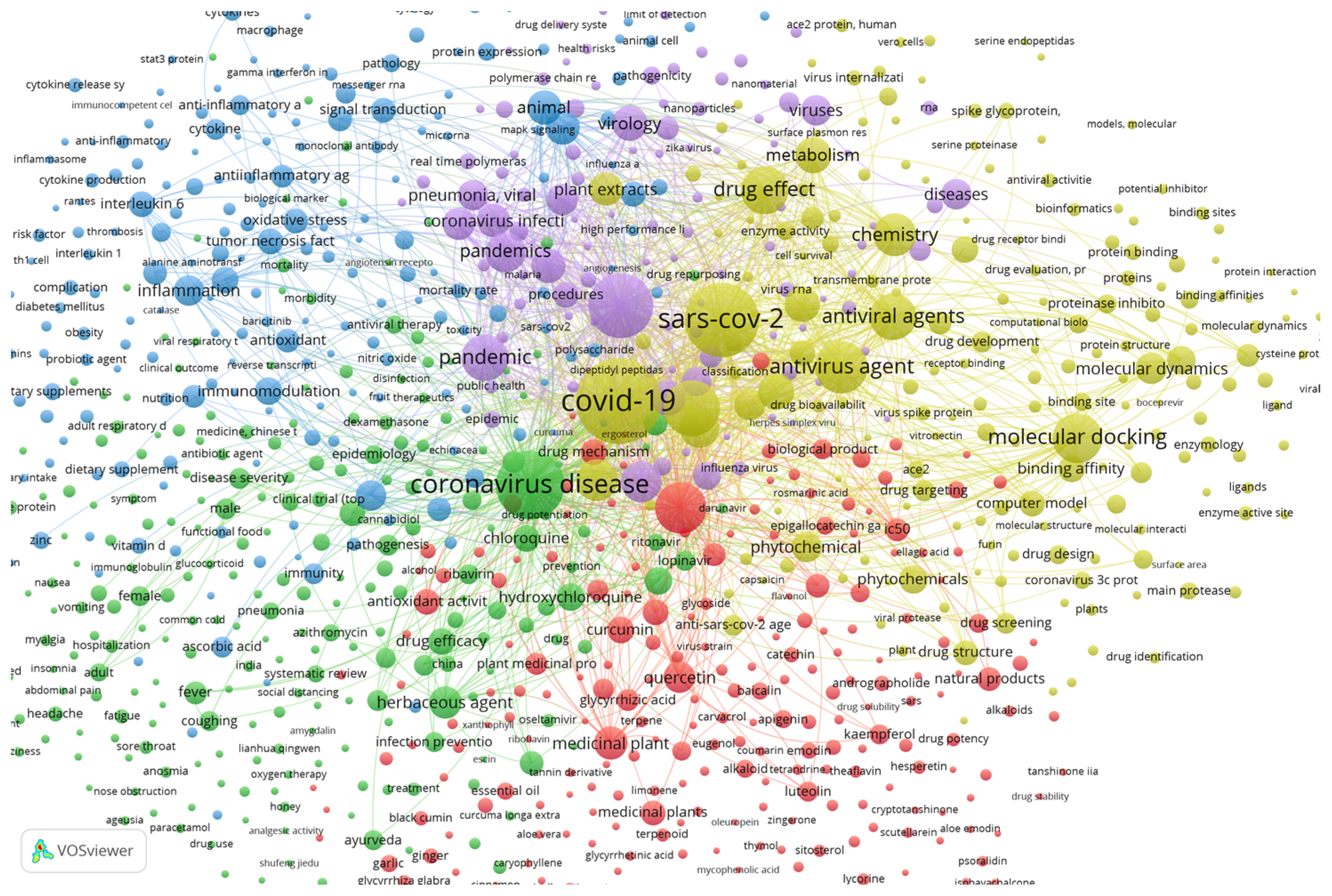
Using the SCOPUS database and VOSviewer software, a brief relation between plants and SARS-CoV-2 can be established. About 1504 documents show up using the keywords “coronavirus”, “covid”, “sars”, “plant” and “extract” from 2020 to 2022 within the SCOPUS search engine. Documents that contain at least one of the searched keywords were clustered by VOSviewer according to their relevance and relationship. The plot of the relationships between the documents with these keywords is shown in Figure 3.
Figure 3.
Plot from VOSviewer showing the bibliographic data from specific keywords between 2020 and 2022, totaling 1504 documents grouped by co-occurrence.
Figure 3 was obtained by analyzing the co-occurrence of bibliographic data obtained from the previously mentioned keywords. The relatedness of items is determined based on the number of documents in which they occur together, with a minimum of 5 occurrences of a keyword to be considered. The visualization weight was set at the total link strength and showed a maximum of 1000. Regarding the relationship between the keywords and plants and metabolites, some specific compounds have connections with other keywords, showing that considerable amounts of studies used those compounds as potential antiviral candidates.
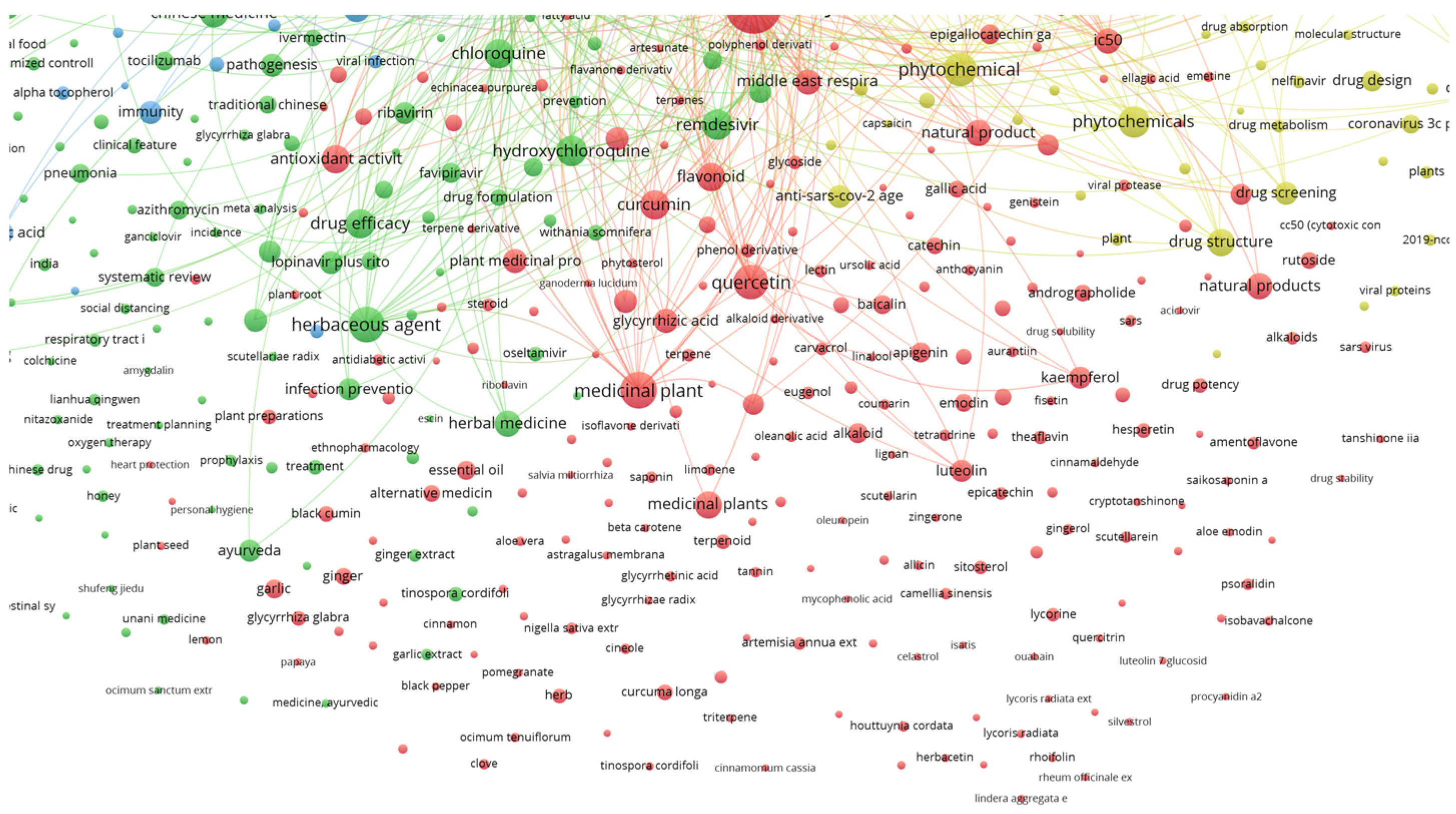
The lines connecting the keywords show a strong connection between them, while isolated circles mean that although the keywords occur together, there are fewer documents with both, implying a lower correlation between them, and thus these compounds have less importance. This can also be observed by the physical distance between the circles. Examples of compounds highly related to the defined keywords and with connecting lines are quercetin, luteolin, kaempferol, carvacrol and gallic acid, shown in Figure 4.
Figure 4.
Detail of relationships between plant secondary metabolites and the searched keywords.
While specific metabolites do occur in Figure 4, there could be many others that are still under research and have not been published yet. The time frame set to build Figure 3 and Figure 4 was set between 2020 and 2022, which is a very short period considering the time the research takes to achieve the results, followed by the writing and publication of the study. Still, the fact that some individual compounds already show bibliographic correlation is impressive and reveals that there is potential. Furthermore, keywords such as medicinal plant, flavonoid, phytochemicals, natural product, flavanone derivative and phytotherapy are all correlated with the keywords, but do not reveal the specific used metabolite or extract, thus demanding further rummage and increasing the potential of natural products in fighting SARS-CoV-2.
7. Future and Trends
The relationship between viruses and humans has always been about balancing infectiousness and lethality, and while outbreaks of highly contagious and lethal viruses have been few, recent studies have shown that they are expected to increase in coming years. Improvement of hygiene and knowledge of viruses has helped mankind to protect itself from viruses, but some pockets of densely packed population and wild animals has created the perfect concoction for spillover of viruses and other pathogens from animals to humans, and while this could be relatively easy to overcome, other changes are more cumbersome.
Climate change, which represents a huge issue on its own [100], is probably the most important factor that will increase the pressure for viruses such as the Coronaviridae to jump from animals to humans [101]. Deforestation increases pressure on wild ecosystems, making wild animals share habitats with humans. Beyond this, floods increase the prevalence of viral vectors, and droughts change ecosystems, making them abruptly shift, allowing for new connections between animals, resulting in a mixture of viruses that can easily infect humans.
Beyond the grim scenario of climate change helping to increase the prevalence of viruses, the unknown long-standing symptoms, and opportunistic diseases in patients previously infected with COVID-19 is another matter of concern. Autoimmune diseases are manifesting themselves in some patients after recovering from mild to severe COVID-19, which shows the long road ahead for satisfactory knowledge of this illness and its sequelae over time [102,103,104].
While mitigating climate change may help reduce the probability of higher occurrences of a global pandemic, it will not avoid them. In fact, it is hardly possible to completely avoid contained outbreaks of viral pandemics, and thus, mankind can only work to reduce their widespread and be prepared to quickly implement vaccines, treatments and measures to mitigate their effects, which could control or even preview further outbreaks. These efforts must be carried out by all branches of society, and all possibilities should be pondered.
Plants, being a widely disseminated and endless resource, and are in some cases the cure or treatment for many previous diseases and pathogenic vectors, are once again paramount to fend of this coronavirus, and will be effective in the coming pandemics.
The use of plant extracts as adjuvants in antiviral treatments, as accessory treatment for symptomatic relief, as application in protective equipment for health professionals, and being mixed into disinfectant solutions and other applications is still being studied, and results of publications and intellectual property are expected to be rolled out in the coming months and years.
Author Contributions
Writing—review and editing, S.A.H.; writing—original draft, M.C.; investigation, S.A.H., M.C., F.S.R. and T.C.S.P.P.; funding acquisition, I.C.F.R.F.; project administration, L.B. and M.P. All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by the Portuguese Fundação para Ciência e Tecnologia (FCT), which financed the Centro de Investigação de Montanha through (UIDB/00690/2020), as well as the researchers S. Heleno and M. Carocho (CEECIND/00831/2018, CEECIND/03040/2017) as well as L. Barros. The research was also funded by the Programa Operacional Regional Norte 2020, within the “PlantCovid” project, NORTE-01-02B7-FEDER-054870.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
No new data was produced.
Acknowledgments
The authors are grateful to the Foundation for Science and Technology (FCT, Portugal) for financial support through national funds FCT/MCTES to the CIMO (UIDB/00690/2020). Acknowledgments to the European Fund for Regional Development through the Programa Operacional Regional Norte 2020, within the “PlantCovid” project, NORTE-01-02B7-FEDER-054870. S. Heleno and M. Carocho thank FCT for their individual employment program-contract (CEECIND/00831/2018, CEECIND/03040/2017). L. Barros also thanks the national funding by FCT, through the institutional scientific employment program-contract for her contract.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Adedeji, A.O.; Severson, W.; Jonsson, C.; Singh, K.; Weiss, S.R.; Sarafianos, S.G. Novel inhibitors of severe acute respiratory syndrome coronavirus entry that act by three distinct mechanisms. J. Virol. 2013, 87, 8017–8028. [Google Scholar] [CrossRef]
- Al-Sanea, M.M.; Abelyan, N.; Abdelgawad, M.A.; Musa, A.; Ghoneim, M.M.; Al-Warhi, T.; Aljaeed, N.; Alotaibi, O.J.; Alnusaire, T.S.; Abdelwahab, S.F.; et al. Strawberry and ginger silver nanoparticles as potential inhibitors for SARS-CoV-2 sssisted by in silico modeling and metabolic profiling. Antibiotics 2021, 10, 824. [Google Scholar] [CrossRef]
- Artika, I.M.; Dewantari, A.K.; Wiyatno, A. Molecular biology of coronaviruses: Current knowledge. Heliyon 2020, 6, e04743. [Google Scholar] [CrossRef] [PubMed]
- Berger, A.; Drosten, C.; Doerr, H.; Stürmer, M.; Preiser, W. Severe acute respiratory syndrome (SARS)-paradigm of an emerging viral infection. J. Clin. Virol. 2004, 29, 13–22. [Google Scholar] [CrossRef]
- Raj, V.S.; Osterhaus, A.D.; Fouchier, R.A.; Haagmans, B.L. MERS: Emergence of a novel human coronavirus. Curr. Opin. Virol. 2014, 5, 58–62. [Google Scholar] [CrossRef] [PubMed]
- Harrison, A.G.; Lin, T.; Wang, P. Mechanisms of SARS-CoV-2 transmission and pathogenesis. Trends Immunol. 2020, 41, 1100–1115. [Google Scholar] [CrossRef]
- World Health Organization—WHO. Middle East Respiratory Syndrome Coronavirus (MERS-CoV). 2020. Available online: https://www.who.int/news/item/27-07-2017-countries-agree-next-steps-to-combat-global-health-threat-by-mers-cov/ (accessed on 26 July 2021).
- Chen, Z.; Boon, S.S.; Wang, M.H.; Chan, R.W.Y.; Chan, P.K.S. Genomic and evolutionary comparison between SARS-CoV-2 and other human coronaviruses. J. Virol. Methods 2021, 289, 114032. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Nyodu, R.; Maurya, V.K.; Saxena, S.K. Chapter 3—Morphology, genome organization, replication, and pathogenesis of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2). In Coronavirus Disease 2019 (COVID-19); Saxena, S.K., Ed.; Springer: Singapore, 2020. [Google Scholar]
- Schoeman, D.; Fielding, B.C. Coronavirus envelope protein: Current knowledge. Virol. J. 2019, 16, 69. [Google Scholar] [CrossRef]
- De Wit, E.; Doremalen, N.; Falzarano Munster, V.J. SARS and MERS: Recent insights into emerging coronaviruses. Nat. Rev. Microbiol. 2016, 14, 523–534. [Google Scholar] [CrossRef]
- Low, J.S.; Jerak, J.; Tortorici, M.A.; McCallum, M.; Pinto, D.; Cassotta, A.; Foglierini, M.; Mele, F.; Abdelnabi, R.; Weynand, B.; et al. ACE-2-biinding exposes the SARS-CoV-2 fusion peptide to broadly neutralizing coronavisus antibodies. Science 2022, 377, 735–742. [Google Scholar] [CrossRef]
- Munir, M.; Tandiabang, P.; Setyawati, T.; Basry, A.; Cyio, A.; Rahman, N. Bioethical perspective of convalescent plasma therapy for COVID-19: A systematic review. Transfus. Clin. Et Biol. 2021, 28, 271–275. [Google Scholar] [CrossRef] [PubMed]
- Pérez-Alba, E.; Nuzzolo-Shihadeh, L.; Aguirre-García, G.M.; Espinosa-Mora, J.; Lecona-Garcia, J.D.; Flores-Pérez, R.O.; Mendoza-Garza, M.; Camacho-Ortiz, A. Baricitinib plus dexamethasone compared to dexamethasone for the treatment of severe COVID-19 pneumonia: A retrospective analysis. J. Microbiol. Immunol. Infect. 2021, 54, 787–793. [Google Scholar] [CrossRef] [PubMed]
- Snow, T.A.C.; Saleem, N.; Ambler, G.; Nastouli, E.; McCoy, L.E.; Singer, M.; Arulkumaran, N. Convalescent plasma for COVID-19: A meta-analysis, trial sequential analysis, and meta-regression. Br. J. Anaesth. 2021, 127, 834–844. [Google Scholar] [CrossRef] [PubMed]
- Bravo, J.P.K.; Dangerfield, T.L.; Taylor, D.W.; Johnson, K.A. Remdesivir is a delayed translocation inhibitor of SARS-CoV-2 replication. Mol. Cell 2021, 81, 1548–1552. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.R.; Zhang, Y.N.; Li, X.D.; Zhang, H.Q.; Xiao, S.Q.; Deng, F.; Yuan, Z.M.; Ye, H.Q.; Zhang, B. A cell-based large-scale screening of natural compounds for inhibitors of SARS-CoV-2. Signal Transduct. Target. Ther. 2020, 5, 218. [Google Scholar] [CrossRef]
- Kyriazopoulou, E.; Huet, T.; Cavalli, G.; Gori, A.; Kyprianou, M.; Pickkers, P.; Eugen-Olsen, J.; Clerici, M.; Veas, F.; Chatellier, G.; et al. Effect of anakinra on mortality in patients with COVID-19: A systematic review and patient-level meta-analysis. Lancet Rheumatol. 2021, 10, E690–E697. [Google Scholar] [CrossRef]
- Taramasso, L.; Magnasco, L.; Portunato, F.; Briano, F.; Vena, A.; Giacobbe, D.R.; Dentone, C.; Robba, C.; Ball, L.; Loconte, M.; et al. Clinical presentation of secondary infectious complications in COVID-19 patients with intensive care unit treated with tocilizumab or standard of care. Eur. J. Intern. Med. 2021, 94, 39–44. [Google Scholar] [CrossRef]
- Dougan, M.; Nirula, A.; Azizad, M.; Mocherla, B.; Gottlieb, R.L.; Chen, P.; Hebert, C.; Perry, R.; Boscia, J.; Heller, B.; et al. Bamlanivimab plus etesevimab in mild or moderate Covid-19. N. Engl. J. Med. 2021, 385, 1382–1392. [Google Scholar] [CrossRef]
- Kim, J.Y.; Jang, Y.R.; Hong, J.H.; Jung, J.G.; Park, J.; Streinu-Cercel, A.; Streinu-Cercel, A.; Săndulescu, O.; Lee, S.J.; Kim, S.H.; et al. Safety, virologic efficacy, and pharmacokinetics of CT-P59, a neutralizing monoclonal antibody against SARS-CoV-2 spike receptor-binding protein: Two randomized, placebo-controlled phase 1 studies in healthy subjects and patients with mild SARS-CoV-2 infection. Clin. Ther. 2021, 43, 1706–1727. [Google Scholar]
- Razonable, R.R.; Pawlowski, C.; O'Horo, J.C.; Arndt, L.L.; Arndt, R.; Bierle, D.M.; Borgen, M.D.; Hanson, S.N.; Hedin, M.C.; Lenehan, P.; et al. Casirivimab-Imdevimab treatment is associated with reduced rates of hospitalization among high-risk patients with mild to moderate coronavirus disease-19. EClinicalMecdicine 2021, 40, 101102. [Google Scholar] [CrossRef]
- Baral, P.K.; Yin, J.; James, M.N.G. Treatment and prevention strategies for the COVID 19 pandemic: A review of immunotherapeutics approaches for neutralizing SARS-CoV-2. Int. J. Biol. Macromol. 2021, 186, 490–500. [Google Scholar] [CrossRef] [PubMed]
- Hoffmann, M.; Hofmann-Winkler, H.; Krüger, N.; Kempf, A.; Nehlmeier, I.; Graichen, L.; Arora, P.; Sidarovich, A.; Moldenhauer, A.-S.; Winkler, M.S.; et al. SARS-CoV-2 variant B.1.617 is resistant to bamlanivimab and evades antibodies induced by infection and vaccination. Cell Rep. 2021, 36, 109415. [Google Scholar] [CrossRef] [PubMed]
- Rosa, S.S.; Prazeres, D.M.F.; Azevedo, A.M.; Marques, M.P.C. mRNA vaccines manufacturing: Challenges and bottlenecks. Vaccine 2021, 39, 2190–2200. [Google Scholar] [CrossRef] [PubMed]
- Nagy, A.; Alhatlani, B. An overview of current COVID-19 vaccine platforms. Comput. Struct. Biotechnol. J. 2021, 19, 2508–2517. [Google Scholar] [CrossRef]
- Ceron, W.; De-Lima-Santos, M.-F.; Quiles, M.G. Fake news agendas in the era of COVID-19: Identifying trends through fact-checking content. Online Soc. Netw. Media 2021, 21, 100116. [Google Scholar] [CrossRef]
- Gautam, S.; Gautam, A.; Chhetri, S.; Bhattarai, U. Immunity against COVID-19: Potential role of Ayush Kwath. J. Ayurveda Integr. Med. 2020, 13, 100350. [Google Scholar] [CrossRef]
- Hu, H.; Ji, Z.; Feng, C.; Pang, W.; Chen, Z.; Zhang, J.; Wang, H. PROSPERO’s systematic review protocols of traditional Chinese medicine for COVID-19: An overview. Integr. Med. Res. 2021, 10, 100774. [Google Scholar] [CrossRef]
- Liang, S.-B.; Fang, M.; Liang, C.-H.; Lan, H.-D.; Shen, C.; Yan, L.-J.; Hu, X.-Y.; Han, M.; Robinson, N.; Liu, J.-P. Therapeutic effects and safety of oral Chinese patent medicine for COVID-19: A rapid systematic review and meta-analysis of randomized controlled trials. Complement. Ther. Med. 2021, 60, 102744. [Google Scholar] [CrossRef]
- Orisakwe, O.E.; Orish, C.N.; Nwanaforo, E.O. Coronavirus disease (COVID-19) and Africa: Acclaimed home remedies. Sci. Afr. 2020, 10, e00620. [Google Scholar] [CrossRef]
- Ren, W.; Liang, P.; Ma, Y.; Sun, Q.; Pu, Q.; Dong, L.; Luo, G.; Mazhar, M.; Liu, J.; Wang, R.; et al. Research progress of traditional Chinese medicine against COVID-19. Biomed. Pharmacother. 2021, 137, 111310. [Google Scholar] [CrossRef]
- Yin, X.; Cai, S.-B.; Tao, L.-T.; Chen, L.-M.; Zhang, Z.-D.; Xiao, S.-H.; Fan, A.Y.; Zou, X. Recovery of a patient with severe COVID-19 by acupuncture and Chinese herbal medicine adjuvant to standard care. J. Integr. Med. 2021, 19, 460–466. [Google Scholar] [PubMed]
- Zhu, W.; Xu, M.; Chen, C.Z.; Guo, H.; Shen, M.; Hu, X.; Shinn, P.; Klumpp-Thomas, C.; Michael, S.G.; Zheng, W. Identification of SARS-CoV-2 3CL protease inhibitors by a quantitative high-throughput screening. ACS Pharmacol. Transl. Sci. 2020, 3, 1008–1016. [Google Scholar] [CrossRef] [PubMed]
- Feyaerts, A.F.; Luyten, W. Vitamin C as prophylaxis and adjunctive medical treatment for COVID-19? Nutrition 2020, 79, 110948. [Google Scholar] [CrossRef] [PubMed]
- Islam, A.T.M.R.; Ferdousi, J.; Shahinozzaman, M. Previously published ethnopharmacological reports reveal the potentiality of plants and plant-derived products used as traditional home remedies by Bangladeshi COVID-19 patients to combat SARS-CoV-2. Saudi J. Biol. Sci. 2021, 28, 6653–6673. [Google Scholar] [CrossRef] [PubMed]
- Lyu, M.; Fan, G.; Xiao, G.; Wang, T.; Xu, D.; Gao, J.; Ge, S.; Li, Q.; Ma, Y.; Zhang, H.; et al. Traditional chinese medicine in COVID-19. Acta Pharm. Sin. B 2021, 11, 3337–3363. [Google Scholar] [CrossRef] [PubMed]
- Shakoor, H.; Feehan, J.; Dhaheri, A.S.A.; Ali, H.I.; Platat, C.; Ismail, L.C.; Apostopoulos, V.; Stojanovska, L. Immune-boosting role of vitamins D, C, E, zinc, selenium and moega-3 fatty acids: Could they help against COVID-19? Maturitas 2021, 143, 1–9. [Google Scholar] [CrossRef]
- Gasmi, A.; Peana, M.; Noor, S.; Lysiuk, R.; Menzel, A.; Benahmed, A.G.; Bjørklund, G. Chloroquine and hydroxychloroquine in the treatment of COVID-19: The never-ending story. Appl. Microbiol. Biotechnol. 2021, 105, 1333–1343. [Google Scholar] [CrossRef]
- Di Stefano, L.; Ogburn, E.L.; Ram, M.; Scharfstein, D.O.; Li, T.; Khanal, P.; Baksh, S.N.; McBee, N.; Gruber, J.; Gildea, M.R.; et al. Hydroxychlorooquine/chloroquine for the treatment of hospitalized patients with COVID-19: An individual participant data meta-analysis. PLoS ONE 2022, 17, e0273526. [Google Scholar] [CrossRef]
- Réa-Neto, Á.; Bernardelli, R.S.; Câmara, B.M.D.; Reese, F.B.; Queiroga, M.V.O.; Oliveira, M.C. An open-label randomized controlled trial evaluating the efficacy of cholorquine/hydroxychloroquine in severe COVID-19 patients. Sci. Rep. 2021, 11, 9023. [Google Scholar] [CrossRef]
- Reis, G.; Silva, E.A.S.M.; Silva, D.C.M.; Thabane, L.; Milagres, A.C.; Ferreira, T.S.; Santos, C.V.Q.; Campos, V.H.S.; Nogueira, A.M.R.; Almeida, A.P.F.G.; et al. Effect of early treatment with ivermectin among patients with Codid-19. N. Engl. J. Med. 2022, 386, 1721–1731. [Google Scholar] [CrossRef]
- Bramante, C.T.; Huling, J.D.; Tignanelli, C.J.; Buse, J.B.; Liebovitz, D.M.; Nicklas, J.M.; Kenneth Cohen, K.; Puskarich, M.A.; Belani, H.K.; Proper, J.L.; et al. Randomized trial of metformin, ivermectin, and fluvoxamine for Covid-19. N. Engl. J. Med. 2022, 387, 599–610. [Google Scholar] [CrossRef] [PubMed]
- Firn, R. Are NP’S different from synthetic chemicals? In Nature’s Chemicals: The Natural Products That Shaped Our World; Oxford University Press: Oxford, UK, 2010. [Google Scholar]
- Islam, M.T.; Sarkar, C.; El-Kersh, D.M.; Jamaddar, S.; Uddin, S.J.; Shilpi, J.A.; Mubarak, M.S. Natural products and their derivatives against coronavirus: A review of the non-clinical and pre-clinical data. Phytother. Res. 2020, 34, 2471–2492. [Google Scholar] [CrossRef] [PubMed]
- Van Hoof, L.; Totté, J.; Corthout, J.; Pieters, L.; Mertens, F.; Berghe, D.A.V.; Vlietinck, A.J.; Dommisse, R.; Esmans, E. Plant Antiviral Agents, VI. Plant Antiviral Agents, VI. Isolation of Antiviral Phenolic Glucosides from Populus Cultivar Beaupre by Droplet Counter-Current Chromatography. J. Nat. Prod. 1989, 52, 875–878. [Google Scholar]
- Chávez, J.H.; Leal, P.C.; Yunes, R.A.; Nunes, R.J.; Barardi, C.R.; Pinto, A.R.; Simões, C.M.; Zanetti, C.R. Evaluation of antiviral activity of phenolic compounds and derivatives against rabies virus. Vet. Microbiol. 2006, 116, 53–59. [Google Scholar] [CrossRef]
- Behbahani, M.; Sayedipour, S.; Pourazar, A.; Shanehsazzadeh, M. In vitro anti-HIV-1 activities of kaempferol and kaempferol-7-Oglucoside isolated from Securigera securidaca. Res. Pharm. Sci. 2014, 9, 463–469. [Google Scholar] [PubMed]
- Özçelik, B.; Kartal, M.; Orhan, I. Cytotoxicity, antiviral and antimicrobial activities of alkaloids, flavonoids, and phenolic acids. Pharm. Biol. 2011, 49, 396–402. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.-L.; Guo, Y.-S.; Wang, C.-H.; Li, G.-Q.; Xu, J.-J.; Chung, H.Y.; Ye, W.-C.; Li, Y.-L.; Wang, G.-C. Phenolic compounds from Origanum vulgare and their antioxidant and antiviral activities. Food Chem. 2014, 152, 300–306. [Google Scholar] [CrossRef]
- Ishikawa, T.; Watanabe, T.; Tanigawa, H.; Saito, T.; Kotake, K.-I.; Ohashi, Y.; Ishii, H. Nitrosation of phenolic substrates under mildly basic conditions: Selective preparation of p-quinone monooximes and their antiviral activities. J. Org. Chem. 1996, 61, 2774–2779. [Google Scholar] [CrossRef]
- Zhang, Y.-B.; Wu, P.; Zhang, X.-L.; Xia, C.; Li, G.-Q.; Ye, W.-C.; Wang, G.-C.; Li, Y.-L. Phenolic compounds from the flowers of Bombax malabaricum and their antioxidant and antiviral activities. Molecules 2015, 20, 19947–19957. [Google Scholar] [CrossRef]
- Nile, S.H.; Kim, D.H.; Nile, A.; Park, G.S.; Gansukh, E.; Kai, G. Probing the effect of quercetin 3-glucoside from Dianthus superbus L. against influenza virus infection- In vitro and in silico biochemical and toxicological screening. Food Chem. Toxicol. 2020, 135, 110985. [Google Scholar] [CrossRef]
- Takeda, Y.; Okuyama, Y.; Nakano, H.; Yaoita, Y.; Machida, K.; Ogawa, H.; Imai, K. Antiviral Activities of Hibiscus sabdariffa L. tea extract against human influenza a virus rely largely on acidic pH but partially on a low-pH-independent mechanism. Food Environ. Virol. 2019, 12, 9–19. [Google Scholar] [CrossRef] [PubMed]
- You, H.-L.; Huang, C.-C.; Chen, C.-J.; Chang, C.-C.; Liao, P.-L.; Huang, S.-T. Anti-pandemic influenza A (H1N1) virus potential of catechin and gallic acid. J. Chin. Med Assoc. 2018, 81, 458–468. [Google Scholar] [CrossRef] [PubMed]
- Nicolaou, K.C. Organic synthesis: The art and science of replicating the molecules of living nature and creating others like them in the laboratory. Proc. R. Soc. 2014, 479, 20130690. [Google Scholar] [CrossRef] [PubMed]
- Crane, E.A.; Gademann, K. Capturing biological activity in natural products fragments by chemical synthesis. Angew. Chem. Int. Ed. 2016, 55, 3882–3902. [Google Scholar] [CrossRef]
- Newman, D.J.; Cragg, G.M. Natural products as source of new drugs over the 30 years from 1981 to 2010. J. Nat. Prod. 2012, 75, 311–335. [Google Scholar] [CrossRef]
- Yi, L.; Li, Z.; Yuan, K.; Qu, X.; Chen, J.; Wang, G.; Zhang, H.; Luo, H.; Zhu, L.; Jiang, P.; et al. Small molecules blocking the entry of severe acute respiratory syndrome coronavirus into host cells. J. Virol. 2004, 78, 11334–11339. [Google Scholar] [CrossRef]
- Lin, C.-W.; Tsai, F.-J.; Tsai, C.-H.; Lai, C.-C.; Wan, L.; Ho, T.-Y.; Hsieh, C.-C.; Chao, P.-D.L. Anti-SARS coronavirus 3C-like protease effects of Isatis indigotica root and plant-derived phenolic compounds. Antivir. Res. 2005, 68, 36–42. [Google Scholar] [CrossRef]
- Ho, T.-Y.; Wu, S.-L.; Chen, J.-C.; Li, C.-C.; Hsiang, C.-Y. Emodin blocks the SARS coronavirus spike protein and angiotensin-converting enzyme 2 interaction. Antivir. Res. 2007, 74, 92–101. [Google Scholar] [CrossRef]
- Park, J.-Y.; Kim, J.H.; Kwon, J.M.; Kwon, H.-J.; Jeong, H.J.; Kim, Y.M.; Kim, D.; Lee, W.S.; Ryu, Y.B. Dieckol, a SARS-CoV 3CLpro inhibitor, isolated from the edible brown algae Ecklonia cava. Bioorganic Med. Chem. 2013, 21, 3730–3737. [Google Scholar] [CrossRef]
- Park, J.-Y.; Ko, J.A.; Kim, D.W.; Kim, Y.M.; Kwon, H.J.; Jeong, H.J.; Kim, C.Y.; Park, K.H.; Lee, W.S.; Ryu, Y.B. Chalcones isolated from Angelica keiskei inhibit cysteine proteases of SARS-CoV. J. Enzym. Inhib. Med. Chem. 2016, 31, 23–30. [Google Scholar] [CrossRef]
- Park, J.-Y.; Yuk, H.J.; Ryu, H.W.; Lim, S.H.; Kim, K.S.; Park, K.H.; Ryu, Y.B.; Lee, W.S. Evaluation of polyphenols from Broussonetia papyrifera as coronavirus protease inhibitors. J. Enzym. Inhib. Med. Chem. 2017, 32, 504–512. [Google Scholar] [CrossRef]
- Park, J.-Y.; Kim, J.H.; Kim, Y.M.; Jeong, H.J.; Kim, D.W.; Park, K.H.; Kwon, H.; Park, S.; Lee, W.S.; Ryu, Y.B. Tanshinones as selective and slow-binding inhibitors for SARS-CoV cysteine proteases. Bioorganic Med. Chem. 2012, 20, 5928–5935. [Google Scholar] [CrossRef] [PubMed]
- Bahbah, E.I.; Negida, A.; Nabet, M.S. Purposing saikosaponins for the treatment of COVID-19. Med. Hypotheses 2020, 140, 109782. [Google Scholar] [CrossRef] [PubMed]
- Luo, W.; Su, X.; Gong, S.; Qin, Y.; Liu, W.; Li, J.; Yu, H.; Xu, Q. Anti-SARS coronavirus 3C-like protease effects of Rheum palmatum L. extracts. Biosci. Trends 2009, 3, 124–126. [Google Scholar] [PubMed]
- Kim, D.W.; Seo, K.H.; Curtis-Long, M.J.; Oh, K.Y.; Oh, J.W.; Cho, J.K.; Lee, K.H.; Park, K.H. Phenolic phytochemical displaying SARS-CoV papain-like protease inhibition from the seeds of Psoralea corylifolia. J. Enzym. Med. Chem. 2014, 29, 59–63. [Google Scholar] [CrossRef]
- Schwarz, S.; Sauter, D.; Wang, K.; Zhang, R.; Sun, B.; Karioti, A.; Bilia, A.R.; Efferth, T.; Schwarz, W. Kaempferol derivatives as antiviral drugs against the 3a channel protein of coronavirus. Planta Medica 2014, 80, 177–182. [Google Scholar] [CrossRef]
- Cheng, P.-W.; Ng, L.-T.; Chiang, L.-C.; Lin, C.-C. Antiviral effects of saikosaponins on human coronavirus 229E in vitro. Clin. Exp. Pharmacol. Physiol. 2006, 33, 612–616. [Google Scholar] [CrossRef]
- Zhang, T.; Chen, D. Anticomplementary principles of a Chinese multiherb remedy for the treatment and prevention of SARS. J. Ethnopharmacol. 2008, 117, 351–361. [Google Scholar] [CrossRef]
- Karioti, A.; Bilia, A.R.; Skaltsa, H. Quercus ilex L.: A rich source of polyacylated flavonoid glycosides. Food Chem. 2010, 123, 131–142. [Google Scholar] [CrossRef]
- Orfali, R.; Rateb, M.; Hassan, H.; Alonazi, M.; Gomaa, M.; Mahrous, N.; GabAllah, M.; Kandeil, A.; Perveen, S.; Abdelmohsen, U.; et al. Sinapic Acid Suppresses SARS CoV-2 Replication by Targeting Its Envelope Protein. Antibiotics 2021, 10, 420. [Google Scholar] [CrossRef]
- Zannella, C.; Giugliano, R.; Chianese, A.; Buonocore, C.; Vitale, G.A.; Sanna, G.; Sarno, F.; Manzin, A.; Nebbioso, A.; Termolino, P.; et al. Antiviral Activity of Vitis vinifera Leaf Extract against SARS-CoV-2 and HSV-1. Viruses 2021, 13, 1263. [Google Scholar] [CrossRef] [PubMed]
- Boroduske, A.; Jekabsons, K.; Riekstina, U.; Muceniece, R.; Rostoks, N.; Ilva, N. Wild Sambucus nigra L. from north-east edge of the species range: A valuable germplasm with inhibitory capacity against SARS-CoV2 S-protein RBD and hACE2 binding in vitro. Ind. Crops Prod. 2021, 165, 113438. [Google Scholar] [CrossRef] [PubMed]
- Signer, J.; Jonsdottir, H.R.; Albrich, W.C.; Strasser, M.; Züst, R.; Ryter, S.; Ackerman-Gäumann, R.; Lenz, N.; Siegrist, D.; Suter, A.; et al. In vitro virucidal activity of Echinaforce, an Echinacea purpurea preparation, against coronaviruses, including common cold coronavirus 229E and SARS-CoV-2. Virol. J. 2020, 17, 136. [Google Scholar] [CrossRef] [PubMed]
- Cinatl, J.; Morgenstern, B.; Bauer, G.; Chandra, P.; Rabenau, H.; Doerr, H. Glycyrrhizin, and active component of liquorice roots, and replication of SARS-associated coronavirus. Lancet 2003, 361, 2045–2046. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Ye, F.; Sun, Q.; Liang, H.; Li, C.; Li, S.; Lu, R.; Huang, B.; Tna, W.; Lai, L. Scutellaria baicalensis extract and baicalein inhibit replication of SARS-CoV-2 and its 3C-like protease in vitro. J. Enzym. Inhib. Med. Chem. 2021, 36, 497–503. [Google Scholar] [CrossRef] [PubMed]
- Ohishi, T.; Hishiki, T.; Baig, M.S.; Rajpoot, S.; Saqib, U.; Takasaki, T.; Hara, Y. Epigallocatechin gallate (EGCG) attenuates severe acute respiratory coronavirus disease 2 (SARS-CoV-2) infection by blocking the interaction of SARS-CoV-2 spike protein receptor-binding domain to human angiotensin-converting enzyme 2. PLoS ONE 2022, 17, e0271112. [Google Scholar] [CrossRef]
- Tran, H.T.T.; Gigl, M.; Le, N.P.K.; Dawid, C.; Lamy, E. In vitro effect of Taraxacum officinale lear aqueous extract on the interaction between ACE2 cell surface receptor and SARS-CoV-2 spike protein D614 and four mutants. Pharmaceuticals 2021, 14, 1055. [Google Scholar] [CrossRef]
- Touret, F.; Gilles, M.; Barral, K.; Nougairède, A.; van Helden, J.; Decroly, E.; de Lamballerie, X.; Coutard, B. In vitro screening of a FDA approved chemical library reveals potential inhibitors of SARS-CoV-2 replication. Sci. Rep. 2020, 10, 13093. [Google Scholar]
- Brown, A.S.; Ackerley, D.F.; Calcott, M.J. High-throughput screening for inhibitors of the SARS-CoV-2 protease using a FRET-biosensor. Molecules 2020, 25, 4666. [Google Scholar] [CrossRef]
- Jin, Z.; Du, X.; Xu, Y.; Deng, Y.; Liu, M.; Zhao, Y.; Zhang, B.; Li, X.; Zhang, L.; Peng, C.; et al. Structure of Mpro from SARS-CoV-2 and discovery of its inhibitors. Nature 2020, 582, 289–293. [Google Scholar] [CrossRef] [PubMed]
- Coelho, C.; Gallo, G.; Campos, C.B.; Hardy, L.; Würtele, M. Biochemical screening for SARS-CoV-2 main protease inhibitors. PLoS ONE 2020, 15, e0240079. [Google Scholar] [CrossRef] [PubMed]
- Xu, T.; Zheng, W.; Huang, R. High-throughput screening assays for SARS-CoV-2 drug development: Current status and future directions. Drug Discov. Today 2021, 26, 2439–2444. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Zhang, Y.; Qiao, W.; Zhang, J.; Qi, Z. Baricitinib, a drug with potential effect to prevent SARS-CoV-2 from entering target cells and control cytokine storm induced by COVID-19. Int. Immunopharmacol. 2020, 86, 106749. [Google Scholar] [CrossRef]
- Zhao, J.; Guo, S.; Yi, D.; Li, Q.; Ma, L.; Zhang, Y.; Wang, J.; Li, X.; Guo, F.; Lin, R.; et al. A cell-based assay to discover inhibitors of SARS-CoV-2 RNA dependent RNA polymerase. Antivir. Res. 2021, 190, 105078. [Google Scholar] [CrossRef]
- Nie, J.; Li, Q.; Wu, J.; Zhao, C.; Hao, H.; Liu, H.; Zhang, L.; Nie, L.; Qin, H.; Wang, M.; et al. Establishment and validation of a pseudovirus neutralization assay for SARS-CoV-2. Emerg. Microbes Infect. 2020, 9, 680–686. [Google Scholar] [CrossRef]
- Govinda, K.C.; Giovanni, B.; Srijan, V.; Mahmudulla, H.; Jayme, H.; Jeremy, Y. A machine learning platform to estimate anti-SARS-CoV-2 activities. Nat. Mach. Intell. 2021, 3, 527–535. [Google Scholar]
- Lan, J.; Ge, J.; Yu, J.; Shan, S.; Zhou, H.; Fan, S.; Zhang, Q.; Shi, X.; Wang, Q.; Zhang, L.; et al. Structure of the SARS-CoV-2 spike receptor-binding domain bound to the ACE2 receptor. Nature 2020, 581, 215–220. [Google Scholar] [CrossRef] [PubMed]
- Wrapp, D.; Wang, N.; Corbett, K.S.; Goldsmith, J.A.; Hsieh, C.-L.; Abiona, O.; Graham, B.S.; McLellan, J.S. Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation. Science 2020, 367, 1260–1263. [Google Scholar] [CrossRef]
- Xiu, S.; Dick, A.; Ju, H.; Mirzaie, S.; Abdi, F.; Cocklin, S.; Zhan, P.; Liu, X. Inhibitors of SARS-CoV-2 entry: Current and future opportunities. J. Med. Chem. 2020, 63, 12256–12274. [Google Scholar] [CrossRef]
- Lundin, A.; Dijkman, R.; Bergström, T.; Kann, N.; Adamiak, B.; Hannoun, C.; Kindler, E.; Jónsdóttir, H.R.; Muth, D.; Kint, J.; et al. Targeting membrane-bound viral RNA synthesis reveals potent inhibition of diverse coronaviruses including the middle east respiratory syndrome virus. PLoS Pathol. 2014, 10, e1004166. [Google Scholar] [CrossRef]
- Gurevich, E.V.; Gurevich, V.V. Therapeutic potential of small molecules and engineered proteins. In Arrestins-Pharmacology and Therapeutic Potential; Springer: Berlin/Heidelberg, Germany, 2014; pp. 1–12. [Google Scholar]
- Ngo, H.X.; Garneau-Tsodikova, S. What are the drugs of the future? MedChemComm 2018, 9, 757–758. [Google Scholar] [CrossRef] [PubMed]
- Milewska, A.; Ciejka, J.; Kaminski, K.; Karewicz, A.; Bielska, D.; Zeglen, S.; Wojciech, K.; Nowakowska, M.; Potempa, J.; Bosch, B.J.; et al. Novel polymeric inhibitors of HCoV-NL63. Antivir. Res. 2013, 97, 112–121. [Google Scholar] [CrossRef] [PubMed]
- Mitsuki, Y.Y.; Ohnishi, K.; Takagi, H.; Oshima, M.; Yamamoto, T.; Mizukoshi, F.; Terahara, K.; Kobayashi, K.; Yamamoto, N.; Yamaoka, S.; et al. A single amino acid substitution in the S1 and S2 Spike protein domains determines the neutralization escape phenotype of SARS-CoV. Microbes Infect. 2008, 10, 908–915. [Google Scholar] [CrossRef] [PubMed]
- Wahedi, H.M.; Ahmad, S.; Abbasi, S.W. Stilbene-based natural compounds as promising drug candidates against COVID-19. J. Biomol. Struct. Dyn. 2020, 39, 3225–3234. [Google Scholar] [CrossRef] [PubMed]
- Ortega-Rivera, O.A.; Shukla, S.; Shin, M.D.; Chen, A.; Beiss, V.; Moreno-Gonzalez, M.A.; Zheng, Y.; Clark, A.E.; Carlin, A.F.; Pokorski, J.K.; et al. Cowpea mosaic virus nanoparticle vaccine candidate displaying peptide epitopes can neutralize the severe acute respiratory syndrome coronavirus. ACS Infect. Dis. 2021, 7, 3096–3110. [Google Scholar] [CrossRef]
- Patrick, R.; Garad, R.; Snell, T.; Enticott, J.; Meadows, G. Australians report climate change as a bigger concern than COVID-19. J. Clim. Chang. Health 2021, 3, 100032. [Google Scholar] [CrossRef]
- Beyer, R.M.; Manica, A.; Mora, C. Shifts in global bat diversity suggest a possible role in climate change in the emergence of SARS-CoV-1 and SARS-CoV-2. Sci. Total Environ. 2021, 767, 145413. [Google Scholar] [CrossRef]
- Galeotti, C.; Bayry, J. Autoimmune and inflammatory diseases following COVID-19. Nat. Rev. Rheumatol. 2020, 16, 413–414. [Google Scholar] [CrossRef]
- Toscano, G.; Palmerini, F.; Ravaglia, S.; Ruiz, L.; Invernizzi, P.; Cuzzoni, M.G.; Baldanti, F.; Daturi, R.; Postorino, P.; Cavallini, A.; et al. Guillain-Barré syndrome associated with SARS-CoV-2. N. Engl. J. Med. 2020, 382, 2574–2576. [Google Scholar] [CrossRef]
- Zhou, Y.; Han, T.; Chen, J.; Hou, C.; Hua, L.; He, S.; Guo, Y.; Zhang, S.; Wang, Y.; Yuan, J.; et al. Clinical and autoimmune characteristics of severe and critical cases of COVID-19. Clin. Transl. Sci. 2020, 13, 1077–1086. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).