Analysis of the C2H2 Gene Family in Maize (Zea mays L.) under Cold Stress: Identification and Expression
Abstract
1. Introduction
2. Materials and Methods
2.1. Bioinformatics Analysis of the C2H2 Gene Family
2.2. Plant Materials and Treatments
2.3. RT-qPCR Analysis of The Candidate Genes
3. Results
3.1. Bioinformatics Analysis of the C2H2 Gene Family
3.1.1. Identification of C2H2 Members
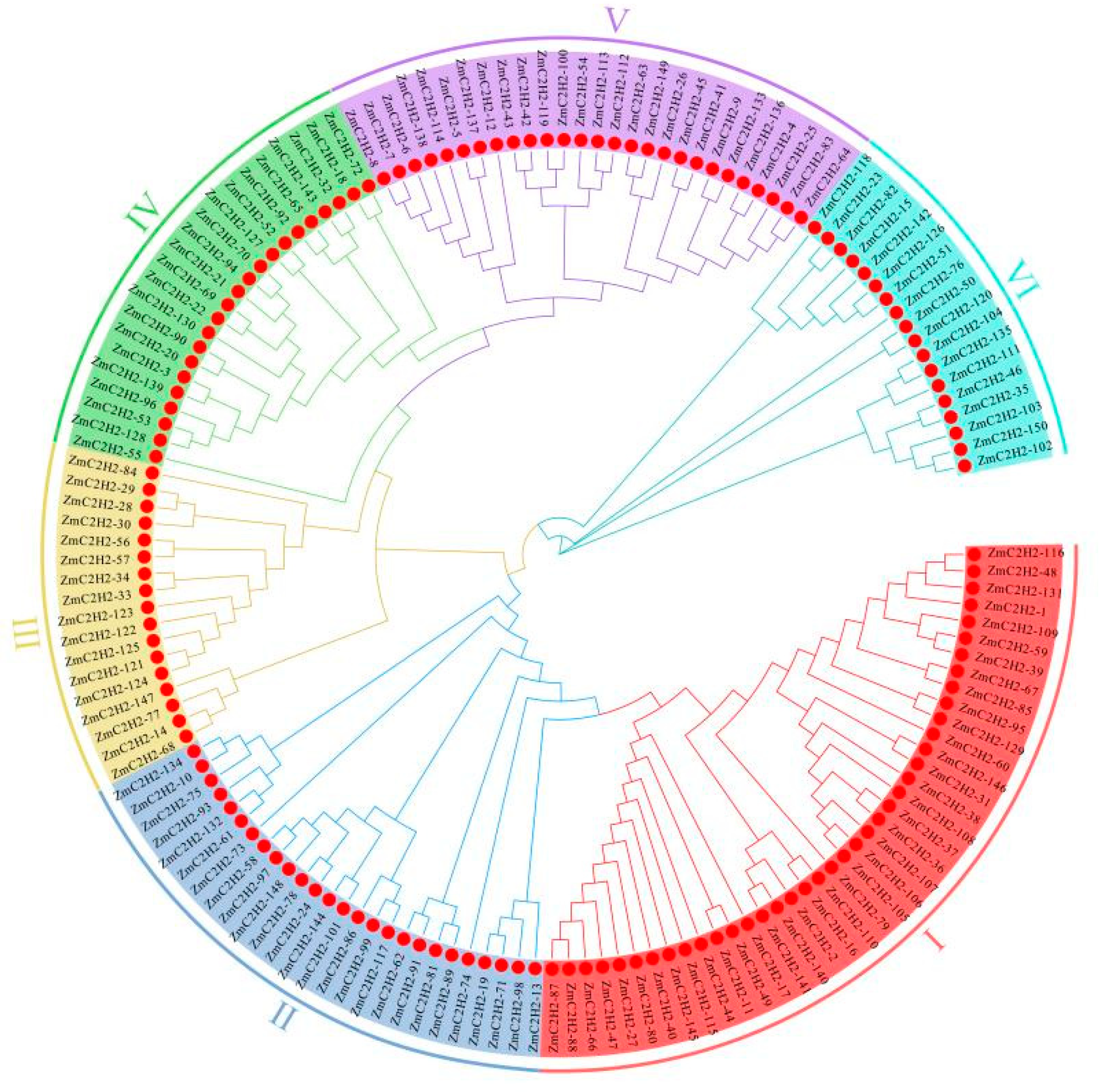
3.1.2. Evolution Analysis of the C2H2 Members
3.1.3. Motifs and Gene Structure of the ZmC2H2s
3.1.4. Cis-Acting Elements of the ZmC2H2s
3.1.5. Collinearity and Ka/Ks Analysis of the ZmC2H2s
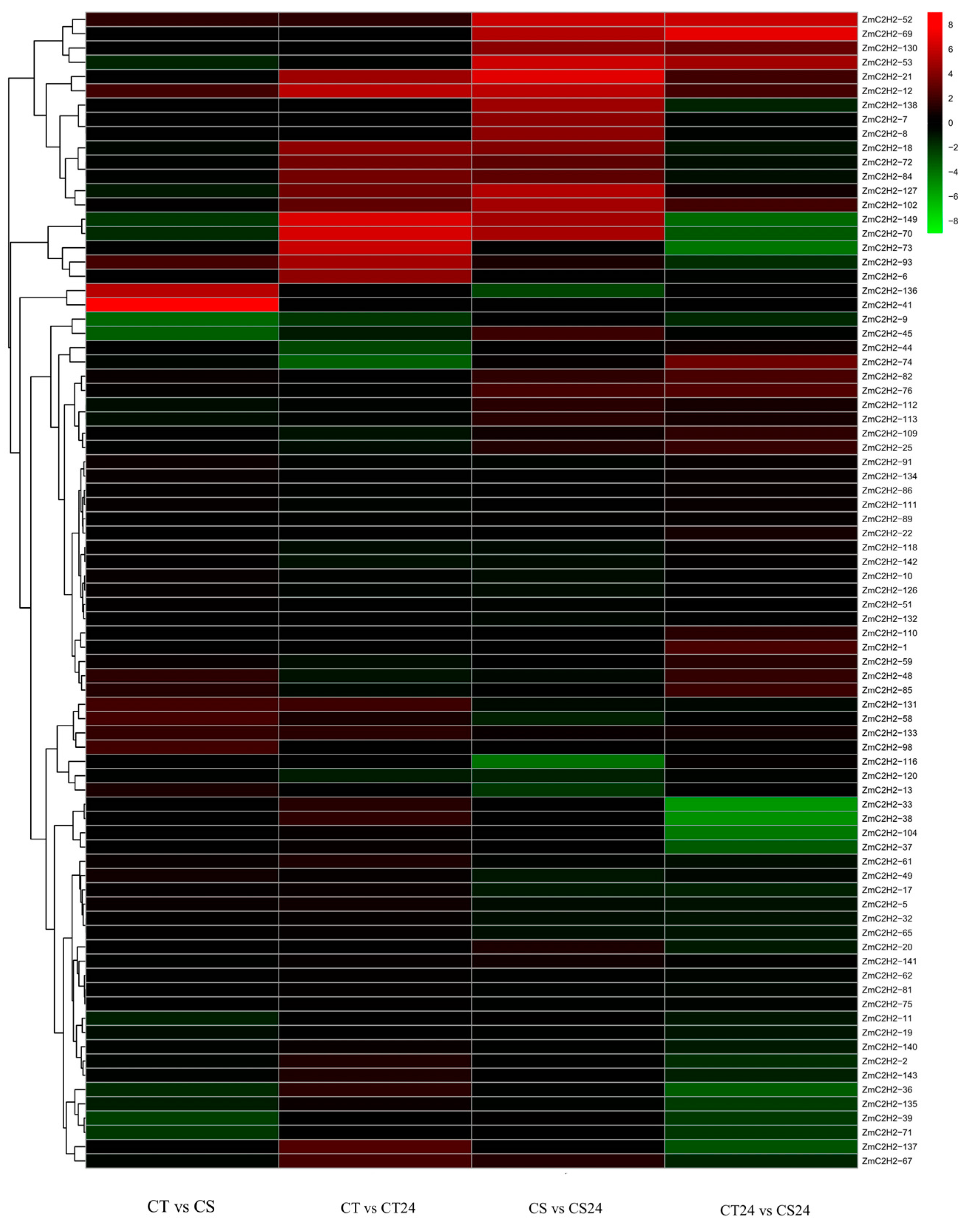
3.2. RNA-Seq Analysis of the ZmC2H2s
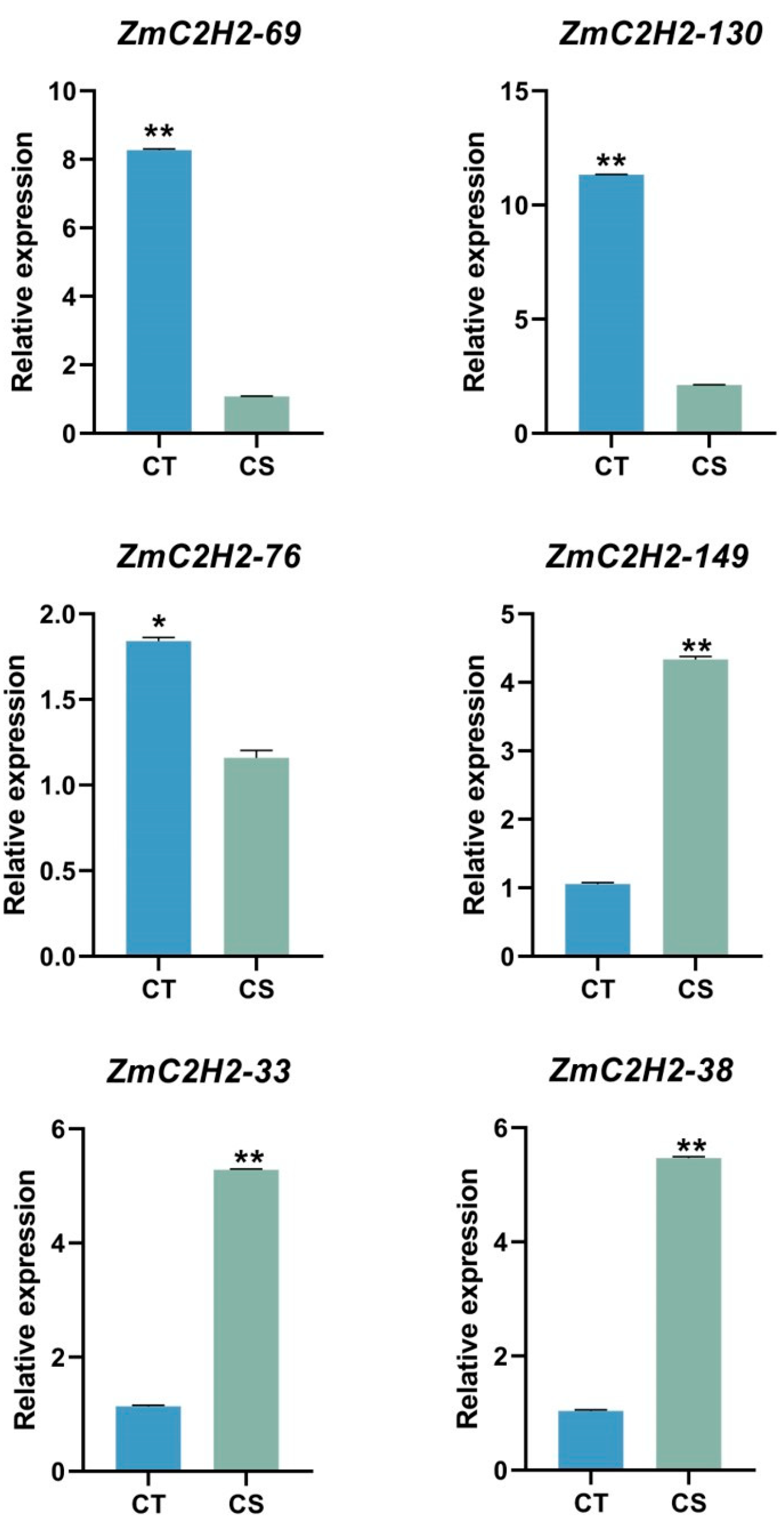
3.3. The Expression of the ZmC2H2s under Cold Stress
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Enders, T.A.; Dennis, S.S.; Oakland, J.; Callen, S.T.; Gehan, M.A.; Miller, N.D.; Spalding, E.P.; Springer, N.M.; Hirsch, C.D. Classifying cold-stress responses of inbred maize seedlings using RGB imaging. Plant Direct 2019, 3, e00104. [Google Scholar] [CrossRef] [PubMed]
- Allen, D.J.; Ort, D.R. Impacts of chilling temperatures on photosynthesis in warm-climate plants. Trends Plant Sci. 2001, 6, 36–42. [Google Scholar] [CrossRef]
- Mackeh, R.; Marr, A.K.; Fadda, A.; Kino, T. C2H2-Type Zinc Finger Proteins: Evolutionarily Old and New Partners of the Nuclear Hormone Receptors. Nucl. Recept. Signal. 2018, 15, 1550762918801071. [Google Scholar] [CrossRef] [PubMed]
- Carroll, S.B. Evo-devo and an expanding evolutionary synthesis: A genetic theory of morphological evolution. Cell 2008, 134, 25–36. [Google Scholar] [CrossRef]
- Xie, M.; Sun, J.; Gong, D.; Kong, Y. The Roles of Arabidopsis C1-2i Subclass of C2H2-type Zinc-Finger Transcription Factors. Genes 2019, 10, 653. [Google Scholar] [CrossRef]
- Han, G.; Lu, C.; Guo, J.; Qiao, Z.; Sui, N.; Qiu, N.; Wang, B. C2H2 Zinc Finger Proteins: Master Regulators of Abiotic Stress Responses in Plants. Front. Plant Sci. 2020, 11, 115. [Google Scholar] [CrossRef]
- Han, G.; Li, Y.; Qiao, Z.; Wang, C.; Zhao, Y.; Guo, J.; Chen, M.; Wang, B. Advances in the Regulation of Epidermal Cell Development by C2H2 Zinc Finger Proteins in Plants. Front. Plant Sci. 2021, 12, 754512. [Google Scholar] [CrossRef]
- Kielbowicz-Matuk, A. Involvement of plant C(2)H(2)-type zinc finger transcription factors in stress responses. Plant Sci. 2012, 185–186, 78–85. [Google Scholar] [CrossRef] [PubMed]
- Lyu, T.; Cao, J. Cys(2)/His(2) Zinc-Finger Proteins in Transcriptional Regulation of Flower Development. Int. J. Mol. Sci. 2018, 19, 2589. [Google Scholar] [CrossRef]
- Laity, J.H.; Lee, B.M.; Wright, P.E. Zinc finger proteins: New insights into structural and functional diversity. Curr. Opin. Struct. Biol. 2001, 11, 39–46. [Google Scholar] [CrossRef] [PubMed]
- Gupta, S.K.; Rai, A.K.; Kanwar, S.S.; Sharma, T.R. Comparative analysis of zinc finger proteins involved in plant disease resistance. PLoS ONE 2012, 7, e42578. [Google Scholar] [CrossRef]
- Chen, Y.; Wang, G.; Pan, J.; Wen, H.; Du, H.; Sun, J.; Zhang, K.; Lv, D.; He, H.; Cai, R.; et al. Comprehensive Genomic Analysis and Expression Profiling of the C2H2 Zinc Finger Protein Family Under Abiotic Stresses in Cucumber (Cucumis sativus L.). Genes 2020, 11, 171. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Khan, A.R.; Gan, Y. C2H2 Zinc Finger Proteins Response to Abiotic Stress in Plants. Int. J. Mol. Sci. 2022, 23, 2730. [Google Scholar] [CrossRef] [PubMed]
- Wang, K.; Ding, Y.; Cai, C.; Chen, Z.; Zhu, C. The role of C2H2 zinc finger proteins in plant responses to abiotic stresses. Physiol. Plant. 2019, 165, 690–700. [Google Scholar] [CrossRef] [PubMed]
- Takatsuji, H.; Mori, M.; Benfey, P.N.; Ren, L.; Chua, N.H. Characterization of a zinc finger DNA-binding protein expressed specifically in Petunia petals and seedlings. EMBO J. 1992, 11, 241–249. [Google Scholar] [CrossRef]
- Joseph, M.P.; Papdi, C.; Kozma-Bognar, L.; Nagy, I.; Lopez-Carbonell, M.; Rigo, G.; Koncz, C.; Szabados, L. The Arabidopsis ZINC FINGER PROTEIN3 Interferes with Abscisic Acid and Light Signaling in Seed Germination and Plant Development. Plant Physiol. 2014, 165, 1203–1220. [Google Scholar] [CrossRef]
- Lyu, T.; Hu, Z.; Liu, W.; Cao, J. Arabidopsis Cys(2)/His(2) zinc-finger protein MAZ1 is essential for intine formation and exine pattern. Biochem. Biophys. Res. Commun. 2019, 518, 299–305. [Google Scholar] [CrossRef]
- Zhuang, H.; Wang, H.L.; Zhang, T.; Zeng, X.Q.; Chen, H.; Wang, Z.W.; Zhang, J.; Zheng, H.; Tang, J.; Ling, Y.H.; et al. NONSTOP GLUMES1 Encodes a C2H2 Zinc Finger Protein That Regulates Spikelet Development in Rice. Plant Cell 2020, 32, 392–413. [Google Scholar] [CrossRef]
- Chang, J.; Yu, T.; Yang, Q.; Li, C.; Xiong, C.; Gao, S.; Xie, Q.; Zheng, F.; Li, H.; Tian, Z.; et al. Hair, encoding a single C2H2 zinc-finger protein, regulates multicellular trichome formation in tomato. Plant J. 2018, 96, 90–102. [Google Scholar] [CrossRef]
- Huang, L.; Jiang, Q.; Wu, J.; An, L.; Zhou, Z.; Wong, C.; Wu, M.; Yu, H.; Gan, Y. Zinc finger protein 5 (ZFP5) associates with ethylene signaling to regulate the phosphate and potassium deficiency-induced root hair development in Arabidopsis. Plant Mol. Biol. 2020, 102, 143–158. [Google Scholar] [CrossRef]
- Zhao, T.; Wu, T.; Zhang, J.; Wang, Z.; Pei, T.; Yang, H.; Li, J.; Xu, X. Genome-Wide Analyses of the Genetic Screening of C(2)H(2)-Type Zinc Finger Transcription Factors and Abiotic and Biotic Stress Responses in Tomato (Solanum lycopersicum) Based on RNA-Seq Data. Front. Genet. 2020, 11, 540. [Google Scholar] [CrossRef]
- Wang, Y.; Dou, D.; Wang, X.; Li, A.; Sheng, Y.; Hua, C.; Cheng, B.; Chen, X.; Zheng, X.; Wang, Y. The PsCZF1 gene encoding a C2H2 zinc finger protein is required for growth, development and pathogenesis in Phytophthora sojae. Microb. Pathog. 2009, 47, 78–86. [Google Scholar] [CrossRef]
- Vogel, J.T.; Zarka, D.G.; Van Buskirk, H.A.; Fowler, S.G.; Thomashow, M.F. Roles of the CBF2 and ZAT12 transcription factors in configuring the low temperature transcriptome of Arabidopsis. Plant J. 2005, 41, 195–211. [Google Scholar] [CrossRef]
- Kim, Y.H.; Kim, M.D.; Park, S.C.; Yang, K.S.; Jeong, J.C.; Lee, H.S.; Kwak, S.S. SCOF-1-expressing transgenic sweetpotato plants show enhanced tolerance to low-temperature stress. Plant Physiol. Biochem. 2011, 49, 1436–1441. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Guo, X.; Lei, C.; Cheng, Z.; Lin, Q.; Wang, J.; Wu, F.; Wang, J.; Wan, J. Overexpression of SlCZFP1, a Novel TFIIIA-type Zinc Finger Protein from Tomato, Confers Enhanced Cold Tolerance in Transgenic Arabidopsis and Rice. Plant Mol. Biol. Report. 2010, 29, 185–196. [Google Scholar] [CrossRef]
- Han, Y.C.; Fu, C.C. Cold-inducible MaC2H2s are associated with cold stress response of banana fruit via regulating MaICE1. Plant Cell Rep. 2019, 38, 673–680. [Google Scholar] [CrossRef]
- Yu, G.H.; Jiang, L.L.; Ma, X.F.; Xu, Z.S.; Liu, M.M.; Shan, S.G.; Cheng, X.G. A soybean C2H2-type zinc finger gene GmZF1 enhanced cold tolerance in transgenic Arabidopsis. PLoS ONE 2014, 9, e109399. [Google Scholar] [CrossRef]
- Agarwal, P.; Arora, R.; Ray, S.; Singh, A.K.; Singh, V.P.; Takatsuji, H.; Kapoor, S.; Tyagi, A.K. Genome-wide identification of C2H2 zinc-finger gene family in rice and their phylogeny and expression analysis. Plant Mol. Biol. 2007, 65, 467–485. [Google Scholar] [CrossRef] [PubMed]
- Yuan, S.; Li, X.; Li, R.; Wang, L.; Zhang, C.; Chen, L.; Hao, Q.; Zhang, X.; Chen, H.; Shan, Z.; et al. Genome-Wide Identification and Classification of Soybean C2H2 Zinc Finger Proteins and Their Expression Analysis in Legume-Rhizobium Symbiosis. Front. Microbiol. 2018, 9, 126. [Google Scholar] [CrossRef] [PubMed]
- Cui, H.; Chen, J.; Liu, M.; Zhang, H.; Zhang, S.; Liu, D.; Chen, S. Genome-Wide Analysis of C2H2 Zinc Finger Gene Family and Its Response to Cold and Drought Stress in Sorghum [Sorghum bicolor (L.) Moench]. Int. J. Mol. Sci. 2022, 23, 5571. [Google Scholar] [CrossRef] [PubMed]
- Matsuoka, Y.; Vigouroux, Y.; Goodman, M.M.; Sanchez, G.J.; Buckler, E.; Doebley, J. A single domestication for maize shown by multilocus microsatellite genotyping. Proc. Natl. Acad. Sci. USA 2002, 99, 6080–6084. [Google Scholar] [CrossRef] [PubMed]
- Piperno, D.R.; Ranere, A.J.; Holst, I.; Iriarte, J.; Dickau, R. Starch grain and phytolith evidence for early ninth millennium B.P. maize from the Central Balsas River Valley, Mexico. Proc. Natl. Acad. Sci. USA 2009, 106, 5019–5024. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Zhang, J.; Xu, Q.; Wang, D.; Di, H.; Huang, J.; Yang, X.; Wang, Z.; Zhang, L.; Dong, L.; et al. Identification of candidate tolerance genes to low-temperature during maize germination by GWAS and RNA-seqapproaches. BMC Plant Biol. 2020, 20, 333. [Google Scholar] [CrossRef] [PubMed]
- Hussain, H.A.; Hussain, S.; Khaliq, A.; Ashraf, U.; Anjum, S.A.; Men, S.; Wang, L. Chilling and Drought Stresses in Crop Plants: Implications, Cross Talk, and Potential Management Opportunities. Front. Plant Sci. 2018, 9, 393. [Google Scholar] [CrossRef]
- Farooq, M.; Aziz, T.; Wahid, A.; Lee, D.-J.; Siddique, K.H.M. Chilling tolerance in maize: Agronomic and physiological approaches. Crop Pasture Sci. 2009, 60, 501–516. [Google Scholar] [CrossRef]
- Chen, M.; Tang, J. Effects of low temperature stress on chlorophyll fluorescence characteristics of com seedlings. J. Inn. Mong. Agric. Univ. Nat. Sci. Ed. 2012, 33, 20–24. [Google Scholar]
- Yang, D.; Sun, Y.; Irfan, A.R.; Liu, X.; Jinying, L.V.; Juping, Y.U. Effect of low temperature stress on germination and physiological of maize seedling. J. Northeast. Agric. Univ. 2018, 49, 1–8+44. [Google Scholar]
- Juurakko, C.L.; diCenzo, G.C.; Walker, V.K. Cold acclimation and prospects for cold-resilient crops. Plant Stress 2021, 2, 100028. [Google Scholar] [CrossRef]
- Bietz, S.; Inhester, T.; Lauck, F.; Sommer, K.; von Behren, M.M.; Fahrrolfes, R.; Flachsenberg, F.; Meyder, A.; Nittinger, E.; Otto, T.; et al. From cheminformatics to structure-based design: Web services and desktop applications based on the NAOMI library. J. Biotechnol. 2017, 261, 207–214. [Google Scholar] [CrossRef]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.; Xia, R. TBtools: An Integrative Toolkit Developed for Interactive Analyses of Big Biological Data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular Evolutionary Genetics Analysis across Computing Platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef] [PubMed]
- Bailey, T.L.; Boden, M.; Buske, F.A.; Frith, M.; Grant, C.E.; Clementi, L.; Ren, J.; Li, W.W.; Noble, W.S. MEME SUITE: Tools for motif discovery and searching. Nucleic Acids Res. 2009, 37, W202–W208. [Google Scholar] [CrossRef]
- Hu, B.; Jin, J.; Guo, A.Y.; Zhang, H.; Luo, J.; Gao, G. GSDS 2.0: An upgraded gene feature visualization server. Bioinformatics 2015, 31, 1296–1297. [Google Scholar] [CrossRef] [PubMed]
- Lescot, M.; Dehais, P.; Thijs, G.; Marchal, K.; Moreau, Y.; Van de Peer, Y.; Rouze, P.; Rombauts, S. PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res. 2002, 30, 325–327. [Google Scholar] [CrossRef]
- Wang, Y.; Tang, H.; Debarry, J.D.; Tan, X.; Li, J.; Wang, X.; Lee, T.H.; Jin, H.; Marler, B.; Guo, H.; et al. MCScanX: A toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res. 2012, 40, e49. [Google Scholar] [CrossRef] [PubMed]
- Qu, L.J.; Zhu, Y.X. Transcription factor families in Arabidopsis: Major progress and outstanding issues for future research. Curr. Opin. Plant Biol. 2006, 9, 544–549. [Google Scholar] [CrossRef]
- Liu, Z.; Coulter, J.A.; Li, Y.; Zhang, X.; Meng, J.; Zhang, J.; Liu, Y. Genome-wide identification and analysis of the Q-type C2H2 gene family in potato (Solanum tuberosum L.). Int. J. Biol. Macromol. 2020, 153, 327–340. [Google Scholar] [CrossRef]
- Jiao, Z.; Wang, L.; Du, H.; Wang, Y.; Wang, W.; Liu, J.; Huang, J.; Huang, W.; Ge, L. Genome-wide study of C2H2 zinc finger gene family in Medicago truncatula. BMC Plant Biol. 2020, 20, 401. [Google Scholar] [CrossRef]
- Jiang, Y.; Liu, L.; Pan, Z.; Zhao, M.; Zhu, L.; Han, Y.; Li, L.; Wang, Y.; Wang, K.; Liu, S.; et al. Genome-wide analysis of the C2H2 zinc finger protein gene family and its response to salt stress in ginseng, Panax ginseng Meyer. Sci Rep. 2022, 12, 10165. [Google Scholar] [CrossRef]
- Zhang, S.; Liu, J.; Zhong, G.; Wang, B. Genome-Wide Identification and Expression Patterns of the C2H2-Zinc Finger Gene Family Related to Stress Responses and Catechins Accumulation in Camellia sinensis [L.] O. Kuntze. Int. J. Mol. Sci. 2021, 22, 4197. [Google Scholar] [CrossRef]
- Zhang, Q.; Geng, J.; Du, Y.; Zhao, Q.; Zhang, W.; Fang, Q.; Yin, Z.; Li, J.; Yuan, X.; Fan, Y.; et al. Heat shock transcription factor (Hsf) gene family in common bean (Phaseolus vulgaris): Genome-wide identification, phylogeny, evolutionary expansion and expression analyses at the sprout stage under abiotic stress. BMC Plant Biol 2022, 22, 33. [Google Scholar] [CrossRef] [PubMed]
- Mallick, B.; Kumari, M.; Pradhan, S.K.; Parmeswaran, C.; Acharya, G.C.; Naresh, P.; Das, B.; Shashankar, P. Genome-wide analysis and characterization of heat shock transcription factors (Hsfs) in common bean (Phaseolus vulgaris L.). Funct. Integr. Genom. 2022, 22, 743–756. [Google Scholar] [CrossRef] [PubMed]
- Sequeira-Mendes, J.; Gutierrez, C. Links between genome replication and chromatin landscapes. Plant J. 2015, 83, 38–51. [Google Scholar] [CrossRef]
- Archibald, J.M. Endosymbiosis and Eukaryotic Cell Evolution. Curr. Biol. 2015, 25, R911–R921. [Google Scholar] [CrossRef] [PubMed]
- Qin, Z.; Wang, Y.; Wang, Q.; Li, A.; Hou, F.; Zhang, L. Evolution Analysis of Simple Sequence Repeats in Plant Genome. PLoS ONE 2015, 10, e0144108. [Google Scholar] [CrossRef]
- Englbrecht, C.C.; Schoof, H.; Bohm, S. Conservation, diversification and expansion of C2H2 zinc finger proteins in the Arabidopsis thaliana genome. BMC Genom. 2004, 5, 39. [Google Scholar] [CrossRef]
- Faraji, S.; Rasouli, S.H.; Kazemitabar, S.K. Genome-wide exploration of C2H2 zinc finger family in durum wheat (Triticum turgidum ssp. Durum): Insights into the roles in biological processes especially stress response. Biometals 2018, 31, 1019–1042. [Google Scholar] [CrossRef]
- Liu, L.; White, M.J.; MacRae, T.H. Transcription factors and their genes in higher plants functional domains, evolution and regulation. Eur. J. Biochem. 1999, 262, 247–257. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Cao, X.; Li, J.; Niu, Q.; Mo, Y.; Xiao, L. Genome-wide characterization of C2H2 zinc-finger gene family provides insight into the mechanisms and evolution of the dehydration-rehydration responses in Physcomitrium and Arabidopsis. Front. Plant Sci. 2022, 13, 953459. [Google Scholar] [CrossRef]
- Fedotova, A.A.; Bonchuk, A.N.; Mogila, V.A.; Georgiev, P.G. C2H2 Zinc Finger Proteins: The Largest but Poorly Explored Family of Higher Eukaryotic Transcription Factors. Acta Nat. 2017, 9, 47–58. [Google Scholar] [CrossRef]
- Liu, Q.; Wang, Z.; Xu, X.; Zhang, H.; Li, C. Genome-Wide Analysis of C2H2 Zinc-Finger Family Transcription Factors and Their Responses to Abiotic Stresses in Poplar (Populus trichocarpa). PLoS ONE 2015, 10, e0134753. [Google Scholar] [CrossRef] [PubMed]
- Alam, I.; Batool, K.; Cui, D.L.; Yang, Y.Q.; Lu, Y.H. Comprehensive genomic survey, structural classification and expression analysis of C2H2 zinc finger protein gene family in Brassica rapa L. PLoS ONE 2019, 14, e0216071. [Google Scholar] [CrossRef] [PubMed]
- Ding, G.; Lorenz, P.; Kreutzer, M.; Li, Y.; Thiesen, H.J. SysZNF: The C2H2 zinc finger gene database. Nucleic Acids Res. 2009, 37, D267–D273, (Database issue). [Google Scholar] [CrossRef]
- Muthamilarasan, M.; Bonthala, V.S.; Mishra, A.K.; Khandelwal, R.; Khan, Y.; Roy, R.; Prasad, M. C2H2 type of zinc finger transcription factors in foxtail millet define response to abiotic stresses. Funct. Integr. Genom. 2014, 14, 531–543. [Google Scholar] [CrossRef] [PubMed]



Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Li, S.; Li, Y.; Cai, Q.; Li, X.; Sun, Y.; Yu, T.; Yang, J.; Zhang, J. Analysis of the C2H2 Gene Family in Maize (Zea mays L.) under Cold Stress: Identification and Expression. Life 2023, 13, 122. https://doi.org/10.3390/life13010122
Li S, Li Y, Cai Q, Li X, Sun Y, Yu T, Yang J, Zhang J. Analysis of the C2H2 Gene Family in Maize (Zea mays L.) under Cold Stress: Identification and Expression. Life. 2023; 13(1):122. https://doi.org/10.3390/life13010122
Chicago/Turabian StyleLi, Sinan, Yunlong Li, Quan Cai, Xin Li, Yan Sun, Tao Yu, Jianfei Yang, and Jianguo Zhang. 2023. "Analysis of the C2H2 Gene Family in Maize (Zea mays L.) under Cold Stress: Identification and Expression" Life 13, no. 1: 122. https://doi.org/10.3390/life13010122
APA StyleLi, S., Li, Y., Cai, Q., Li, X., Sun, Y., Yu, T., Yang, J., & Zhang, J. (2023). Analysis of the C2H2 Gene Family in Maize (Zea mays L.) under Cold Stress: Identification and Expression. Life, 13(1), 122. https://doi.org/10.3390/life13010122




