Rapid, Sensitive and Simultaneous Detection of Two Wheat RNA Viruses Using Reverse Transcription Recombinase Polymerase Amplification (RT-RPA)
Abstract
:1. Introduction
2. Materials and Methods
2.1. Virus and Plant Material
2.2. Total RNA Extraction and cDNA Synthesis
2.3. PCR Assays
2.4. Primer Design
2.5. Recombinant Plasmids and Viruses
2.6. RPA Reaction
2.7. RT-RPA Reaction Optimization
2.8. Sensitivity Assay
2.9. Specificity Assay
2.10. Applicability Assay
3. Results
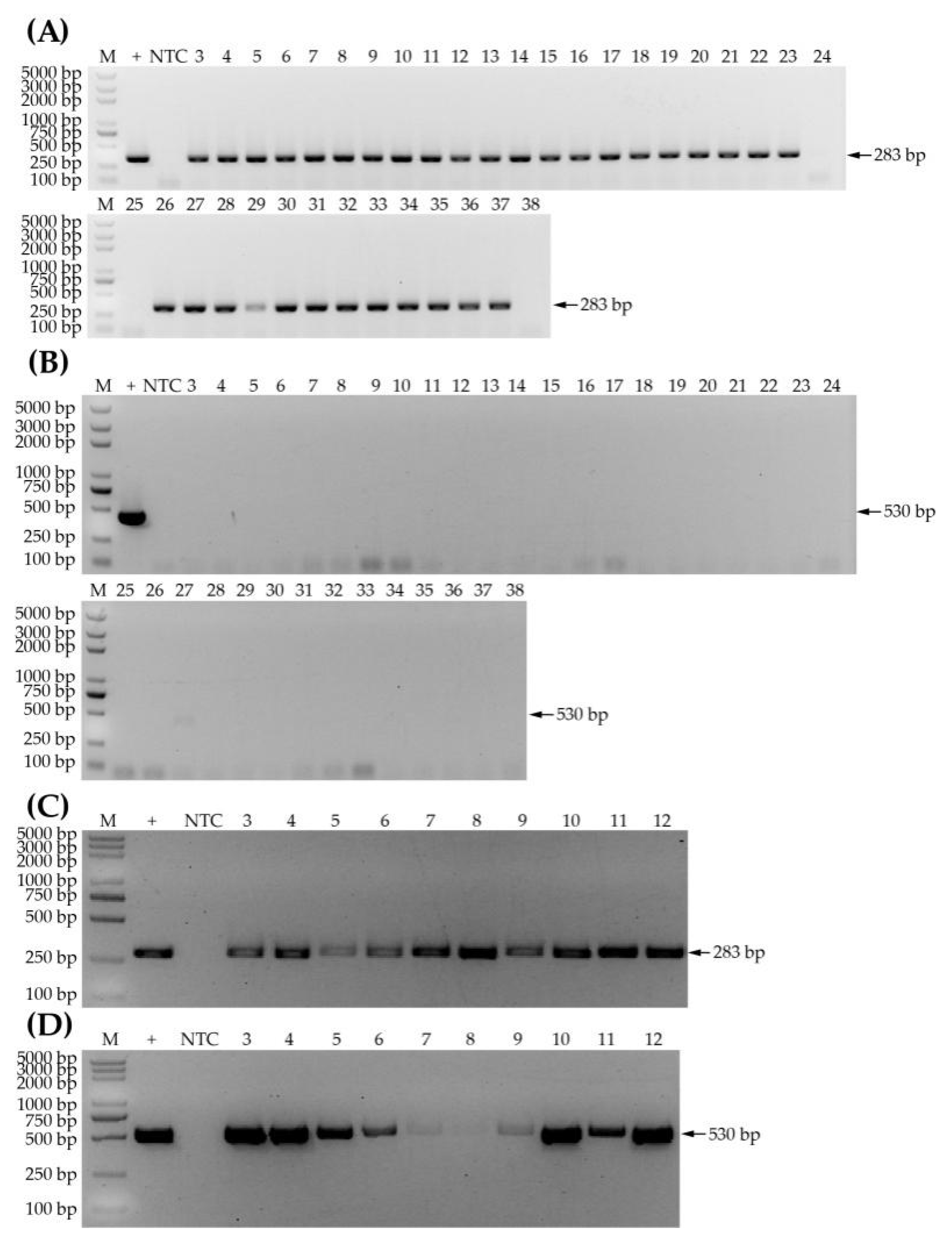
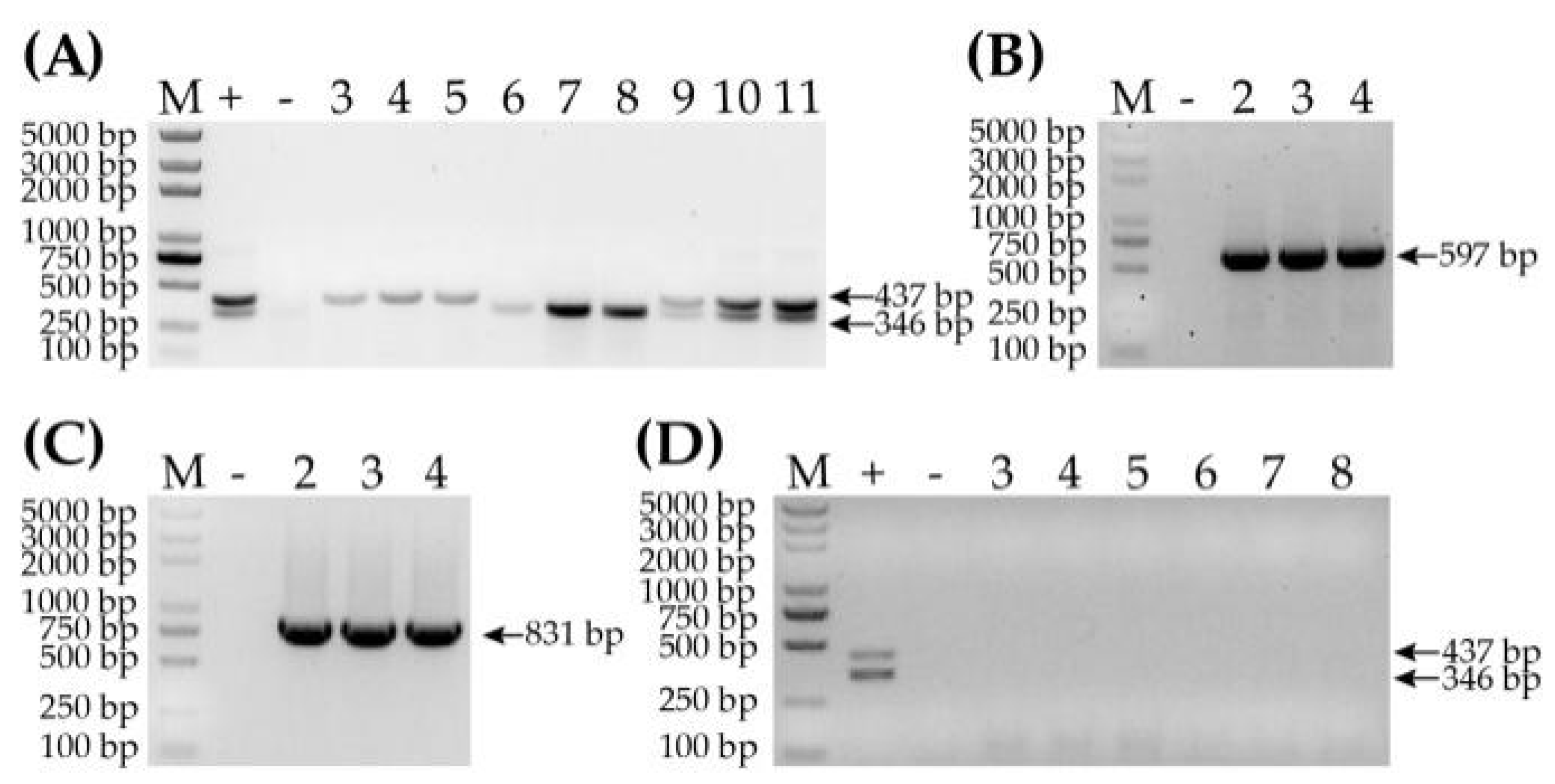
3.1. Samples Screened by PCR-CP
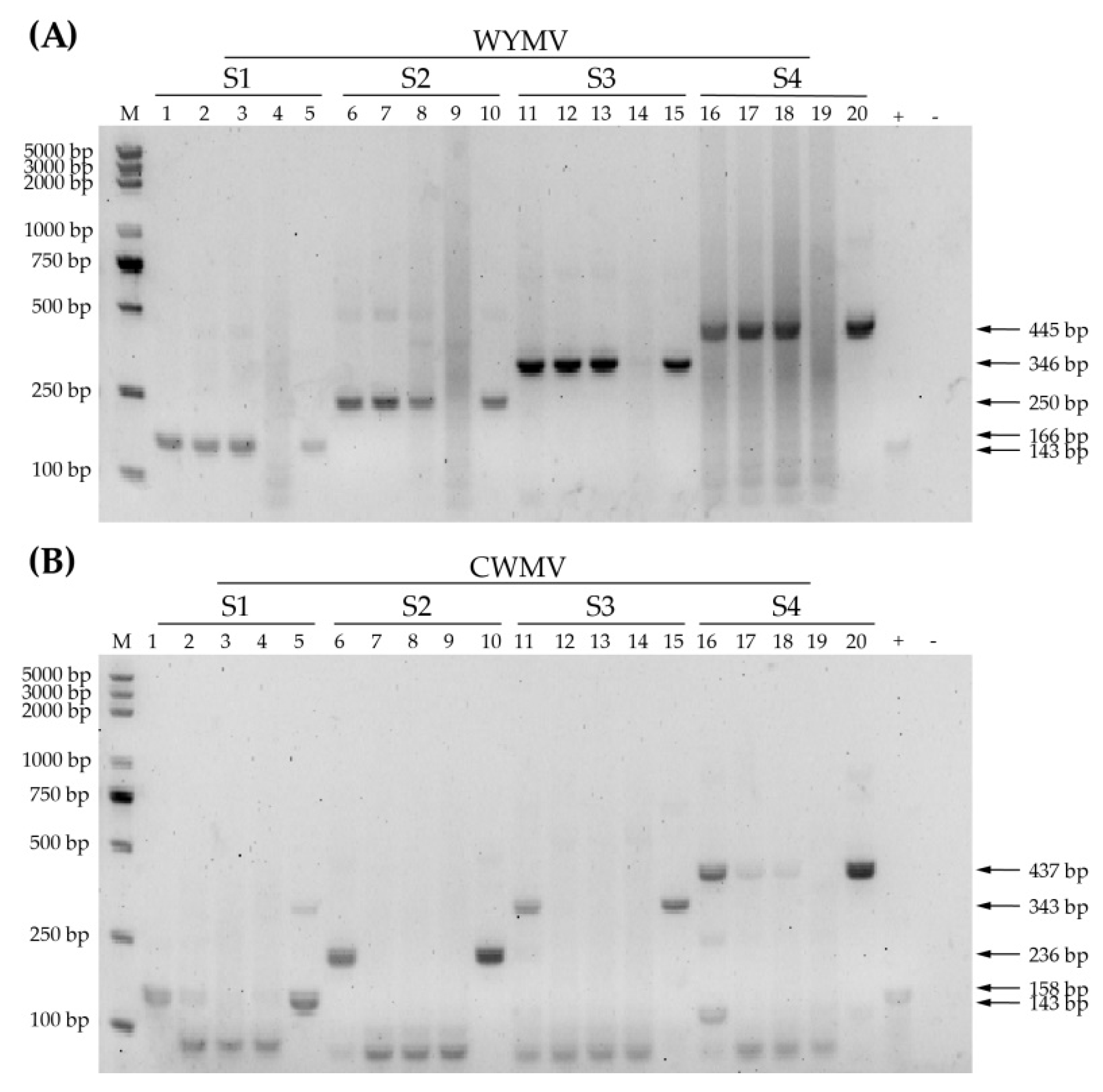
3.2. Selection of RPA Primers
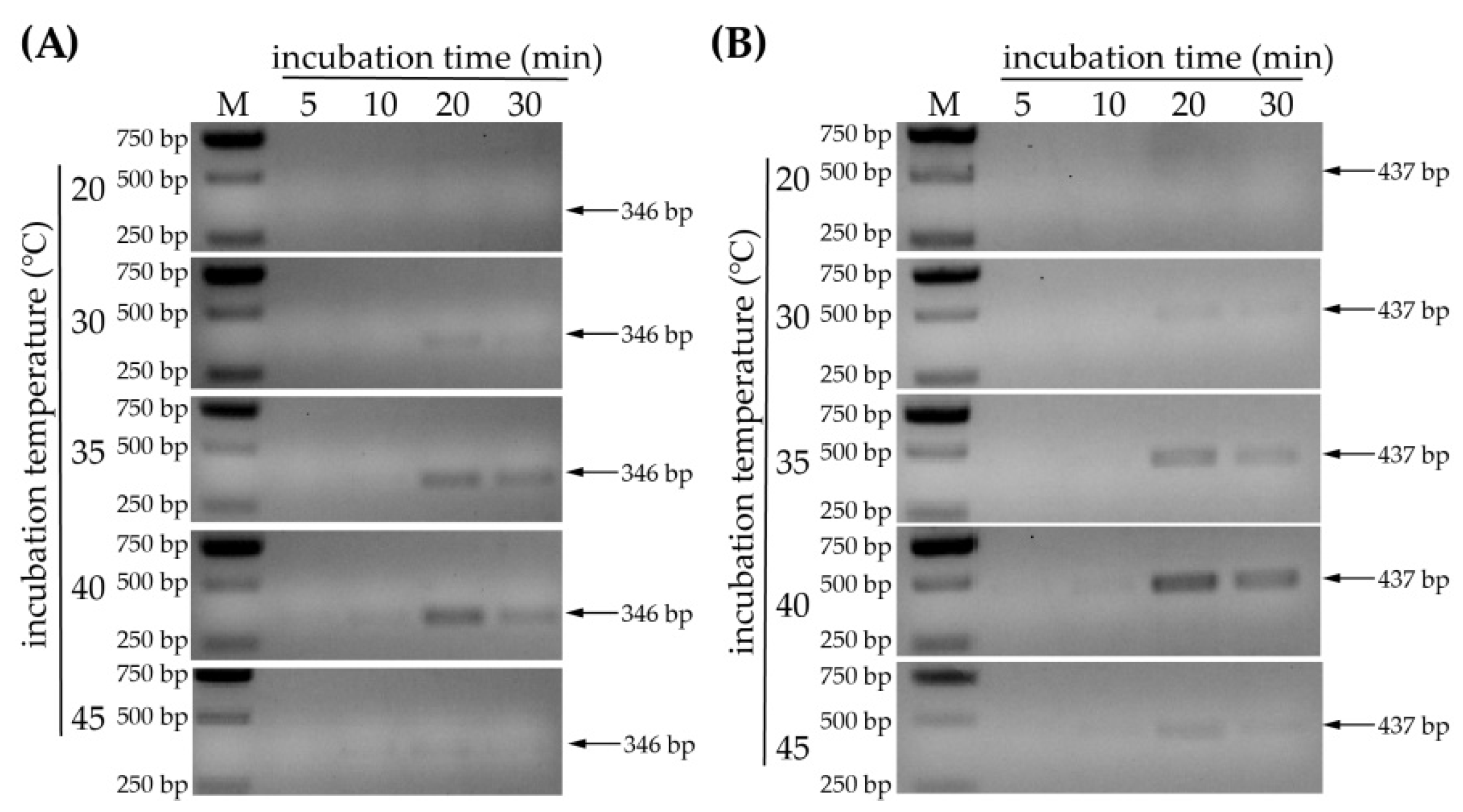
3.3. Optimization of Incubation Temperature and Time
3.4. Validation of Incubation Conditions under Simulated Field Conditions
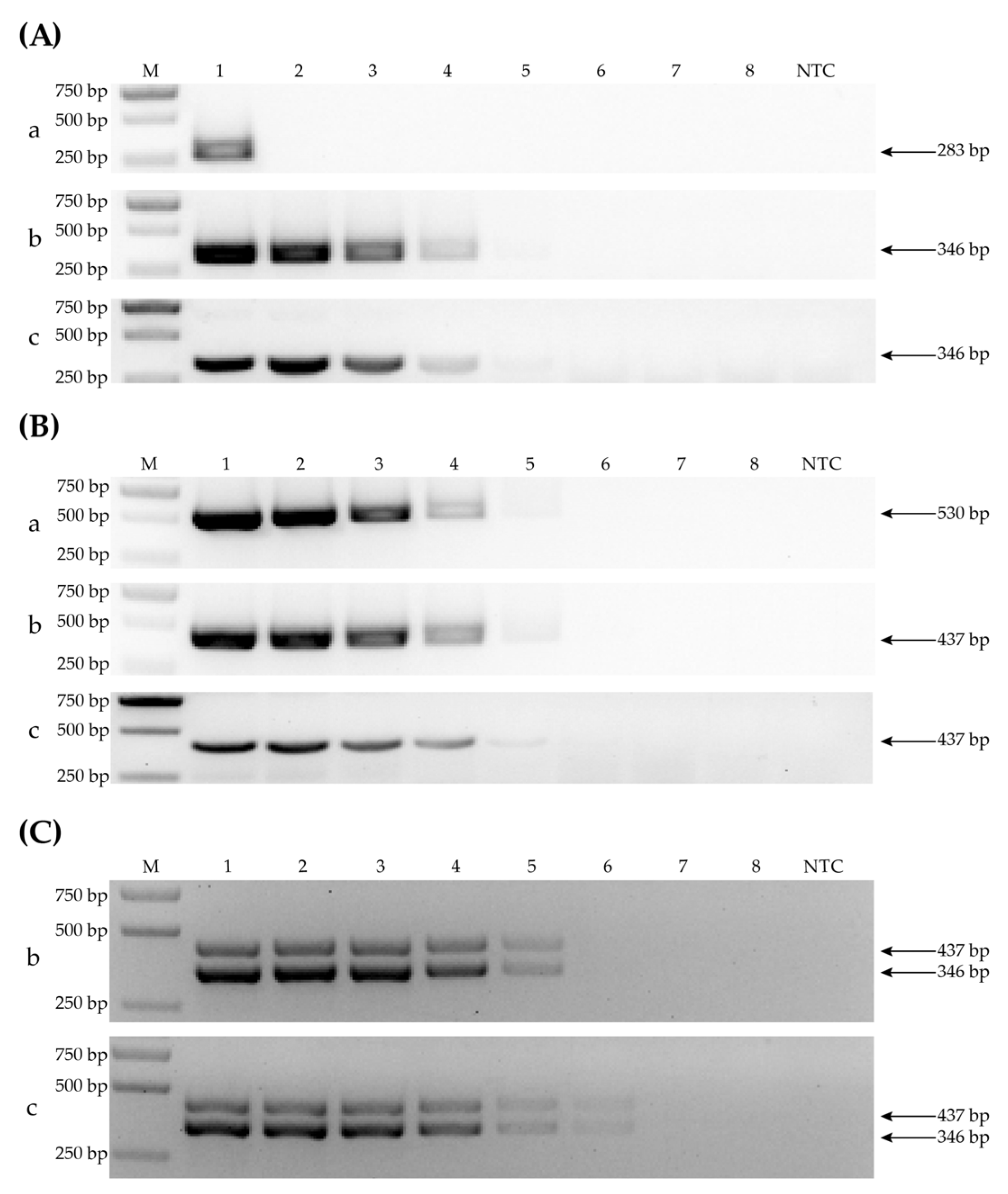
3.5. Sensitivity Determination by Comparing PCR-CP, PCR-RPA and RT-RPA
3.6. Specificity Determination of RT-RPA Method
3.7. Applicability of the RT-RPA Method
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Yamashita, Y.; Souma, C.; Ogura, R.; Suzuki, T. A single QTL on chromosome 6DS derived from a winter wheat cultivar ‘OW104′ confers resistance to wheat yellow mosaic virus. Breed. Sci. 2020, 70, 373–378. [Google Scholar] [CrossRef]
- Zhang, F.; Liu, S.; Zhang, T.; Ye, Z.; Han, X.; Zhong, K.; Yang, J.; Chen, J.; Liu, P. Construction and biological characterization of an infectious full-length cDNA clone of a Chinese isolate of wheat yellow mosaic virus. Virology 2021, 556, 101–109. [Google Scholar] [CrossRef] [PubMed]
- Adams, M.J. Responses of some American, European and Japanese wheat cultivars to soil-borne wheat viruses in China. J. Integr. Agric. 2002, 1, 78–87. [Google Scholar]
- Han, C.; Li, D.; Xing, Y.; Zhu, K.; Tian, Z.; Cai, Z.; Yu, J.; Liu, Y. Wheat yellow mosaic virus widely occurring in wheat (Triticum aestivum) in China. Plant Dis. 2000, 84, 627–630. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, J. Occurrence of fungally transmitted wheat mosaic viruses in China. Ann. Appl. Biol. 2010, 123, 55–61. [Google Scholar] [CrossRef]
- Guo, L.M.; He, J.; Li, J.; Chen, J.P.; Zhang, H.M. Chinese wheat mosaic virus: A long-term threat to wheat in China. J. Integr. Agric. 2019, 18, 821–829. [Google Scholar] [CrossRef] [Green Version]
- Kanyuka, K.; Ward, E.; Adams, M.J. Polymyxa graminis and the cereal viruses it transmits: A research challenge. Mol. Plant Pathol. 2003, 4, 393–406. [Google Scholar] [CrossRef] [Green Version]
- Diao, A.; Chen, J.; Ye, R.; Zheng, T.; Yu, S.; Antoniw, J.F.; Adams, M.J. Complete sequence and genome properties of Chinese wheat mosaic virus, a new furovirus from China. J. Gen. Virol. 1999, 80, 1141–1145. [Google Scholar] [CrossRef]
- Ohki, T.; Sasaya, T.; Maoka, T. Cylindrical inclusion protein of wheat yellow mosaic virus is involved in differential infection of wheat cultivars. Phytopathology 2019, 109, 1475–1480. [Google Scholar] [CrossRef]
- Atreya, C.D. Application of genome sequence information in potyvirus taxonomy: An overview. Arch. Virol. Suppl. 1992, 5, 17–23. [Google Scholar] [CrossRef]
- Kim, N.Y.; Oh, J.; Lee, S.H.; Kim, H.; Moon, J.S.; Jeong, R.D. Rapid and specific detection of by reverse transcription-recombinase polymerase amplification. Plant Pathol. J. 2018, 34, 575–579. [Google Scholar] [CrossRef] [PubMed]
- Kim, N.K.; Kim, S.M.; Jeong, R.D. Reverse transcription recombinase polymerase amplification assay for rapid and sensitive detection of barley yellow dwarf virus in oat. Plant Pathol. J. 2020, 36, 497–502. [Google Scholar] [CrossRef] [PubMed]
- Zhao, C.; Sun, F.; Li, X.; Lan, Y.; Du, L.; Zhou, T.; Zhou, Y. Reverse transcription-recombinase polymerase amplification combined with lateral flow strip for detection of rice black-streaked dwarf virus in plants. J. Virol. Methods 2019, 263, 96–100. [Google Scholar] [CrossRef]
- Babu, B.; Ochoa-Corona, F.M.; Paret, M.L. Recombinase polymerase amplification applied to plant virus detection and potential implications. Anal. Biochem. 2018, 546, 72–77. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Zhang, F.; Xie, L.; Song, X.J.; Li, J.; Chen, J.P.; Zhang, H.M. Functional identification of two minor capsid proteins from Chinese wheat mosaic virus using its infectious full-length cDNA clones. J. Gen. Virol. 2016, 97, 2441–2450. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, T.; Murai, M.N.; Hayashi, T.; Nasuda, S.; Yoshimura, Y.; Komatsuda, T. Resistance to wheat yellow mosaic virus in Madsen wheat is controlled by two major complementary QTLs. Theor. Appl. Genet. 2015, 128, 1569–1578. [Google Scholar] [CrossRef]
- Yang, J.; Zhang, T.; Li, J.; Wu, N.; Wu, G.; Yang, J.; Chen, X.; He, L.; Chen, J. Chinese wheat mosaic virus-derived vsiRNA-20 can regulate virus infection in wheat through inhibition of vacuolar- (H+)-PPase induced cell death. New Phytol. 2020, 226, 205–220. [Google Scholar] [CrossRef] [Green Version]
- Zhang, T.; Liu, P.; Zhong, K.; Zhang, F.; Xu, M.; He, L.; Jin, P.; Chen, J.; Yang, J. Wheat yellow mosaic virus NIb interacting with host light induced protein (LIP) facilitates its infection through perturbing the abscisic acid pathway in wheat. Biology 2019, 8, 80. [Google Scholar] [CrossRef] [Green Version]
- Fukuta, S.; Tamura, M.; Maejima, H.; Takahashi, R.; Kuwayama, S.; Tsuji, T.; Yoshida, T.; Itoh, K.; Hashizume, H.; Nakajima, Y.; et al. Differential detection of wheat yellow mosaic virus, Japanese soil-borne wheat mosaic virus and Chinese wheat mosaic virus by reverse transcription loop-mediated isothermal amplification reaction. J. Virol. Methods 2013, 189, 348–354. [Google Scholar] [CrossRef]
- Donchenko, E.K.; Pechnikova, E.V.; Mishyna, M.Y.; Manukhova, T.I.; Sokolova, O.S.; Nikitin, N.A.; Atabekov, J.G.; Karpova, O.V. Structure and properties of virions and virus-like particles derived from the coat protein of alternanthera mosaic virus. PLoS ONE 2017, 12, e0183824. [Google Scholar] [CrossRef] [Green Version]
- Kogovšek, P.; Kladnik, A.; Mlakar, J.; Znidarič, M.T.; Dermastia, M.; Ravnikar, M.; Pompe-Novak, M. Distribution of potato virus Y in potato plant organs, tissues, and cells. Phytopathology 2011, 101, 1292–1300. [Google Scholar] [CrossRef] [PubMed]
- Prendeville, H.R.; Ye, X.; Morris, T.J.; Pilson, D. Virus infections in wild plant populations are both frequent and often unapparent. Am. J. Bot. 2012, 99, 1033–1042. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Arif, M.; Ali, M.; Rehman, A.; Fahim, M. Detection of potato mop-top virus in soils and potato tubers using bait-plant bioassay, ELISA and RT-PCR. J. Virol. Methods 2014, 195, 221–227. [Google Scholar] [CrossRef] [PubMed]
- Long, H.; Peng, J.; Xuan, C.; Ye, Z.T.; Li, Z.K.; Peng, L.; Ping, C.J.; Jian, Y. Comparative proteomic analysis of Nicotiana benthamiana plants under Chinese wheat mosaic virus infection. BMC Plant Biol. 2021, 21, 51. [Google Scholar] [CrossRef]
- Poojari, S.; Alabi, O.J.; Okubara, P.A.; Naidu, R.A. SYBR(®) green-based real-time quantitative reverse-transcription PCR for detection and discrimination of grapevine viruses. J. Virol. Methods 2016, 235, 112–118. [Google Scholar] [CrossRef] [Green Version]
- Zhang, P.; Liu, Y.; Liu, W.; Massart, S.; Wang, X. Simultaneous detection of wheat dwarf virus, northern cereal mosaic virus, barley yellow striate mosaic virus and rice black-streaked dwarf virus in wheat by multiplex RT-PCR. J. Virol. Methods 2017, 249, 170–174. [Google Scholar] [CrossRef]
- Martinelli, F.; Scalenghe, R.; Davino, S.; Panno, S.; Scuderi, G.; Ruisi, P.; Villa, P.; Stroppiana, D.; Boschetti, M.; Goulart, L.R.; et al. Advanced methods of plant disease detection. A review. Agron. Sustain. Dev. 2015, 35, 1–25. [Google Scholar] [CrossRef] [Green Version]
- Katsarou, K.; Bardani, E.; Kallemi, P.; Kalantidis, K. Viral detection: Past, present, and future. Bioessays 2019, 41, e1900049. [Google Scholar] [CrossRef]
- Piepenburg, O.; Williams, C.H.; Stemple, D.L.; Armes, N.A. DNA detection using recombination proteins. PLoS Biol. 2006, 4, 1115–1121. [Google Scholar] [CrossRef]
- Zang, L.; Qiao, N.; Sun, X.; Zhang, X.; Zhao, D.; Li, J.; Zhu, X. Reverse transcription recombinase polymerase amplification assay for rapid detection of the cucurbit chlorotic yellows virus. J. Virol. Methods 2022, 300, 114388. [Google Scholar] [CrossRef]
- Jeong, H.W.; Go, S.M.; Jeong, R.D. Rapid and specific detection of apple chlorotic leaf spot virus in pear by reverse-transcription recombinase polymerase amplification. Acta Virol. 2021, 65, 237–241. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Chen, R.; Nie, X.; Zhong, Z.; Li, C.; Li, K.; Huang, W.; Fu, X.; Liu, J.; Nie, B. Rapid and sensitive detection of potato virus Y by isothermal reverse transcription-recombinase polymerase amplification assay in potato. Mol. Cell. Probes 2020, 50, 101505. [Google Scholar] [CrossRef] [PubMed]
- Jiao, Y.; Xu, C.; Li, J.; Gu, Y.; Xia, C.; Xie, Q.; Xie, Y.; An, M.; Xia, Z.; Wu, Y. Characterization and a RT-RPA assay for rapid detection of chilli veinal mottle virus (ChiVMV) in tobacco. Virol. J. 2020, 17, 33. [Google Scholar] [CrossRef] [PubMed]
- Lobato, I.M.; O’Sullivan, C.K. Recombinase polymerase amplification: Basics, applications and recent advances. Trends Anal. Chem. 2018, 98, 19–35. [Google Scholar] [CrossRef]
- Wang, R.; Zhang, F.; Wang, L.; Qian, W.; Qian, C.; Wu, J.; Ying, Y. Instant, visual, and instrument-free method for on-site screening of GTS 40-3-2 soybean based on body-heat triggered recombinase polymerase amplification. Anal. Chem. 2017, 89, 4413–4418. [Google Scholar] [CrossRef]
- Fan, X.; Li, L.; Zhao, Y.; Liu, Y.; Liu, C.; Wang, Q.; Dong, Y.; Wang, S.; Chi, T.; Song, F.; et al. Clinical validation of two recombinase-based isothermal amplification assays (RPA/RAA) for the rapid detection of african swine fever virus. Front. Microbiol. 2020, 11, 1696. [Google Scholar] [CrossRef]
- Aman, R.; Mahas, A.; Marsic, T.; Hassan, N.; Mahfouz, M.M. Efficient, rapid, and sensitive detection of plant RNA viruses with one-pot RT-RPA-CRISPR/Cas12a assay. Front. Microbiol. 2020, 11, 610872. [Google Scholar] [CrossRef]
- Koch, G.G.; Landis, J.R. The measurement of observer agreement for categorical data. Biometrics 1977, 33, 159–174. [Google Scholar] [CrossRef] [Green Version]
- Jiao, J.; Kong, K.K.; Han, J.M.; Song, S.W.; Bai, T.H.; Song, C.H.; Wang, M.M.; Yan, Z.L.; Zhang, H.T.; Zhang, R.P.; et al. Field detection of multiple RNA viruses/viroids in apple using a CRISPR/Cas12a-based visual assay. Plant Biotechnol. J. 2021, 19, 394–405. [Google Scholar] [CrossRef]
- Li, J.; Macdonald, J.; von Stetten, F. Review: A comprehensive summary of a decade development of the recombinase polymerase amplification. Analyst 2018, 144, 31–67. [Google Scholar] [CrossRef]

| Primer Name | Sequence 5’-3’ | Set | Targeted Region | Expected Product Length (bp) |
|---|---|---|---|---|
| WYMV-RPA-1-F | TTGACTTCTTTGTCCCCCGCCCGTGGATG | WYMV-S1 | WYMV coat protein | 166 |
| WYMV-RPA-1-R | TGTCCATCAGTATCCAGAACCCTGCGGTTG | |||
| WYMV-RPA-2-F | TGAATGGTGGCTTGAGACGGATCCTGCGAG | WYMV-S2 | WYMV coat protein | 250 |
| WYMV-RPA-2-R | AGGTTGGTTGTGTCGGAGGTGAGCATGGTAT | |||
| WYMV-RPA-3-F | CTTCAGACAAACACGCAGAAAACCAGACCA | WYMV-S3 | WYMV coat protein | 346 |
| WYMV-RPA-3-R | TTGGTTGTGTCGGAGGTGAGCATGGTATTGT | |||
| WYMV-RPA-4-F | ACCAGTCTTGGCATCACCACGGATGAGGCA | WYMV-S4 | WYMV coat protein | 445 |
| WYMV-RPA-4-R | GGTTGGTTGTCTTGCGGAGGTTGGTTGTGT | |||
| CWMV-RPA-1-F | CTTTTGCGACCTTGGCGGAGAGGGGGTCAGT | CWMV-S1 | CWMV coat protein | 158 |
| CWMV-RPA-1-R | CAGTCTGCCCTTGTTCTTCTGTTTCTATCT | |||
| CWMV-RPA-2-F | TAATTTCTTCTCACGGAGTAAACGCTTCGGT | CWMV-S2 | CWMV coat protein | 236 |
| CWMV-RPA-2-R | TAGATATAGCCAACGATTGATCAGTCTGCCCTT | |||
| CWMV-RPA-3-F | TGGCAGAGTACGAGGACGAGTGTGTTGTCT | CWMV-S3 | CWMV coat protein | 343 |
| CWMV-RPA-3-R | AACTAACATAAGTCATCAGTTCGCCCAAAGCAT | |||
| CWMV-RPA-4-F | AAGTTGAGACATGGCAGAGTACGAGGACGA | CWMV-S4 | CWMV coat protein | 437 |
| CWMV-RPA-4-R | TCAACTCGAACCTTCCCACTTAAGATTATACT |
| Sample | PCR | RPA | Sample | PCR | RPA | ||||
|---|---|---|---|---|---|---|---|---|---|
| WYMV | CWMV | WYMV | CWMV | WYMV | CWMV | WYMV | CWMV | ||
| Henan-1 | + | + | + | - | Anhui-1 | + | - | + | - |
| Henan-2 | + | - | + | - | Anhui-2 | + | - | + | - |
| Henan-3 | + | - | + | - | Anhui-3 | + | - | + | - |
| Henan-4 | + | - | + | - | Anhui-4 | + | - | + | - |
| Henan-5 | + | - | + | - | Anhui-5 | + | - | + | - |
| Henan-6 | + | - | + | - | Anhui-6 | + | + | + | + |
| Henan-7 | - | - | - | - | Anhui-7 | + | + | + | + |
| Henan-8 | - | - | - | - | Anhui-8 | + | + | + | + |
| Henan-9 | - | - | - | - | Anhui-9 | - | - | - | - |
| Henan-10 | + | - | + | - | Anhui-10 | - | - | - | - |
| Henan-11 | + | - | + | - | Anhui-11 | - | - | - | - |
| Henan-12 | + | - | + | - | Anhui-12 | - | - | - | - |
| Henan-13 | + | - | + | - | Anhui-13 | - | - | - | - |
| Henan-14 | + | - | + | - | Anhui-14 | - | - | - | - |
| Henan-15 | + | - | + | - | Anhui-15 | - | - | - | - |
| Henan-16 | + | - | + | - | Anhui-16 | - | - | - | - |
| Henan-17 | - | - | - | - | Anhui-17 | - | - | - | - |
| Henan-18 | - | - | - | - | Shandong-1 | + | + | + | + |
| Henan-19 | - | - | - | - | Shandong-2 | + | - | + | - |
| Henan-20 | - | - | - | - | Shandong-3 | + | + | + | - |
| Henan-21 | - | - | - | - | Shandong-4 | - | - | - | - |
| Henan-22 | - | - | - | - | Shandong-5 | - | - | - | - |
| Henan-23 | - | - | - | - | Shandong-6 | + | + | + | + |
| Henan-24 | - | - | - | - | Shandong-7 | + | + | + | + |
| Henan-25 | - | - | - | - | Shandong-8 | + | + | + | + |
| Henan-26 | + | - | + | - | Shandong-9 | + | + | + | + |
| Henan-27 | + | - | + | - | Shandong-10 | + | + | + | - |
| Henan-28 | + | - | + | - | Shandong-11 | + | + | + | + |
| Henan-29 | + | - | + | - | Shandong-12 | + | + | + | + |
| Henan-30 | + | - | + | - | Shandong-13 | + | + | + | + |
| Henan-31 | + | - | + | - | Shandong-14 | + | + | + | + |
| Henan-32 | + | - | + | - | Shandong-15 | + | - | + | - |
| Henan-33 | + | - | + | - | Shandong-16 | + | - | + | - |
| Henan-34 | + | - | + | - | Shandong-17 | + | - | + | - |
| Henan-35 | + | - | + | - | Shandong-18 | + | + | + | + |
| Henan-36 | + | - | + | - | Shandong-19 | + | + | + | + |
| Henan-37 | + | - | + | - | Shandong-20 | + | + | + | + |
| Henan-38 | + | - | + | - | Shandong-21 | + | + | + | + |
| Henan-39 | + | - | + | - | Shandong-22 | + | + | + | + |
| Henan-40 | + | - | + | - | Shandong-23 | + | + | + | + |
| Henan-41 | + | - | + | - | Shandong-24 | + | + | + | + |
| Henan-42 | + | - | + | - | Shandong-25 | + | + | + | + |
| Shanxi-1 | + | - | + | - | Shandong-26 | + | + | + | + |
| Shanxi-2 | - | - | - | - | Shandong-27 | + | + | + | + |
| Shanxi-3 | - | - | + | - | Shandong-28 | + | - | + | - |
| Shanxi-4 | - | - | - | - | Shandong-29 | + | - | + | - |
| Shanxi-5 | + | - | + | - | Shandong-30 | + | - | + | - |
| Shanxi-6 | + | - | + | - | Shandong-31 | + | - | + | - |
| Shanxi-7 | - | - | + | - | Jiangsu-1 | - | - | + | - |
| Shanxi-8 | + | - | + | - | Jiangsu-2 | + | - | + | - |
| Shanxi-9 | + | - | + | - | Jiangsu-3 | + | - | + | - |
| Shanxi-10 | + | - | + | - | Jiangsu-4 | + | - | + | - |
| Shanxi-11 | + | - | + | - | Jiangsu-5 | + | - | + | - |
| Shanxi-12 | + | - | + | - | Jiangsu-6 | + | - | + | - |
| Shanxi-13 | + | - | + | - | Jiangsu-7 | + | - | + | - |
| PCR-CPa | Performance Characteristics (%) | |||||
| Positive | Negative | Total | Sensitivity | Specificity | ||
| RPA- WYMVc | Positive | 82 | 3 | 85 | 100.0% (95.6~100.0%, 95% CI) | 89.3% (71.8~97.7%, 95% CI) |
| Negative | 0 | 25 | 25 | |||
| 82 | 28 | 110 | ||||
| Agreement Kappa value: 0.926 (0.843~1.000, 95% CI) | ||||||
| PCR-CPb | Performance Characteristics (%) | |||||
| Positive | Negative | Total | Sensitivity | Specificity | ||
| RPA- CWMVd | Positive | 22 | 0 | 22 | 88.0%(68.8~97.5%, 95% CI) | 100% (95.8~100.0%, 95% CI) |
| Negative | 3 | 81 | 84 | |||
| 25 | 81 | 106 | ||||
| Agreement Kappa value: 0.919 (0.828~1.000, 95% CI) | ||||||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chen, Z.; Zhang, T.; Lei, J.; Wang, Z.; Liu, P.; Zhong, K.; Chen, J.; Liu, J. Rapid, Sensitive and Simultaneous Detection of Two Wheat RNA Viruses Using Reverse Transcription Recombinase Polymerase Amplification (RT-RPA). Life 2022, 12, 1952. https://doi.org/10.3390/life12121952
Chen Z, Zhang T, Lei J, Wang Z, Liu P, Zhong K, Chen J, Liu J. Rapid, Sensitive and Simultaneous Detection of Two Wheat RNA Viruses Using Reverse Transcription Recombinase Polymerase Amplification (RT-RPA). Life. 2022; 12(12):1952. https://doi.org/10.3390/life12121952
Chicago/Turabian StyleChen, Zhiqing, Tianye Zhang, Jiajia Lei, Ziqiong Wang, Peng Liu, Kaili Zhong, Jianping Chen, and Jiaqian Liu. 2022. "Rapid, Sensitive and Simultaneous Detection of Two Wheat RNA Viruses Using Reverse Transcription Recombinase Polymerase Amplification (RT-RPA)" Life 12, no. 12: 1952. https://doi.org/10.3390/life12121952
APA StyleChen, Z., Zhang, T., Lei, J., Wang, Z., Liu, P., Zhong, K., Chen, J., & Liu, J. (2022). Rapid, Sensitive and Simultaneous Detection of Two Wheat RNA Viruses Using Reverse Transcription Recombinase Polymerase Amplification (RT-RPA). Life, 12(12), 1952. https://doi.org/10.3390/life12121952






