Chromatography-Free Purification Strategies for Large Biological Macromolecular Complexes Involving Fractionated PEG Precipitation and Density Gradients
Abstract
1. Introduction
2. Pitfalls and Difficulties in the Investigation of Macromolecular Complex Function
3. Pitfalls of Chromatographic Purification Strategies for Large Macromolecular Complexes
4. Development of Chromatography-Free Purification Procedures
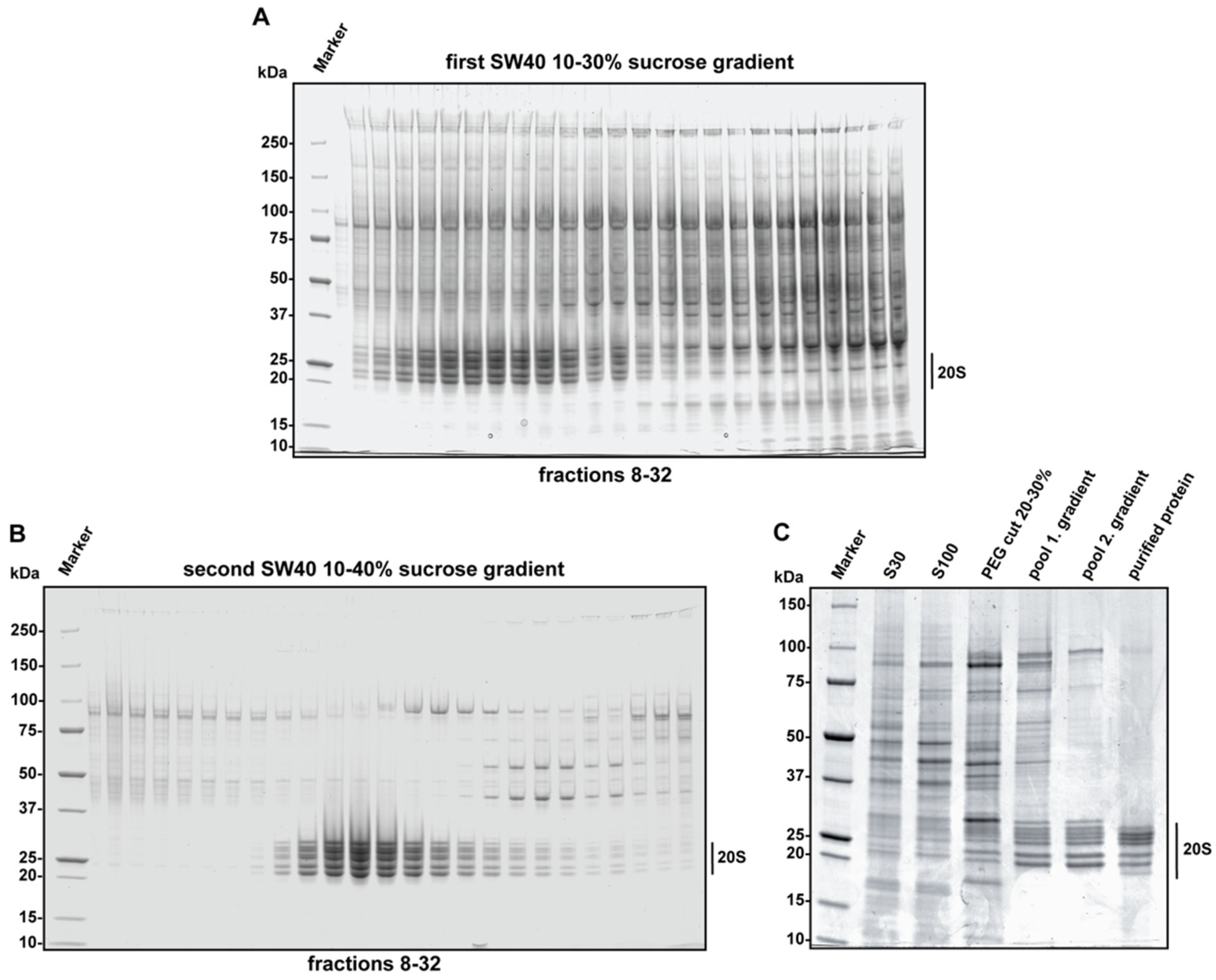
4.1. Case Study 1: Human Constitutive 20S Proteasome
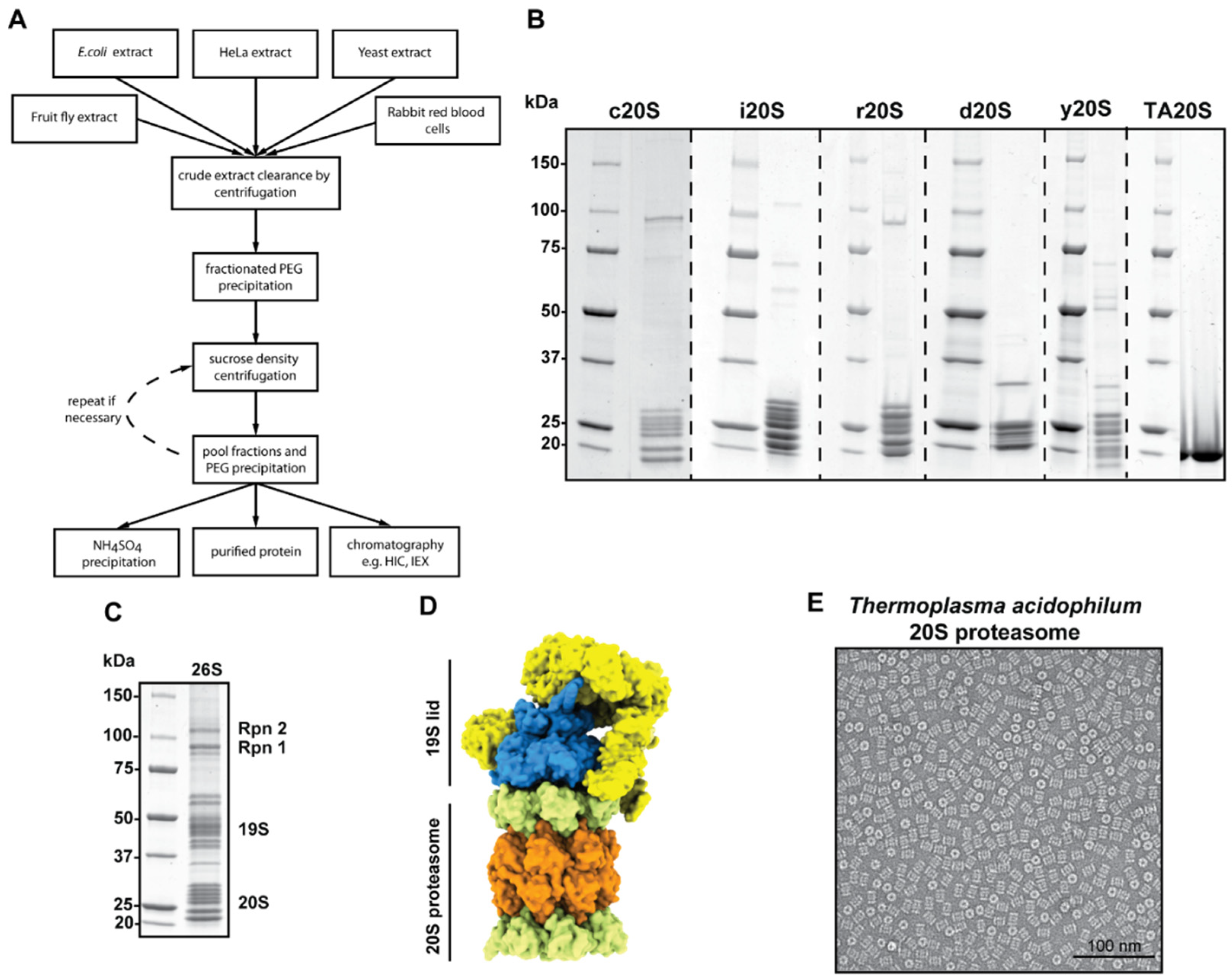
4.2. Case Study 2: 20S Proteasome from Various Organisms and Recombinant Source
4.3. Case Study 3: 26S Proteasome from HeLa Cells
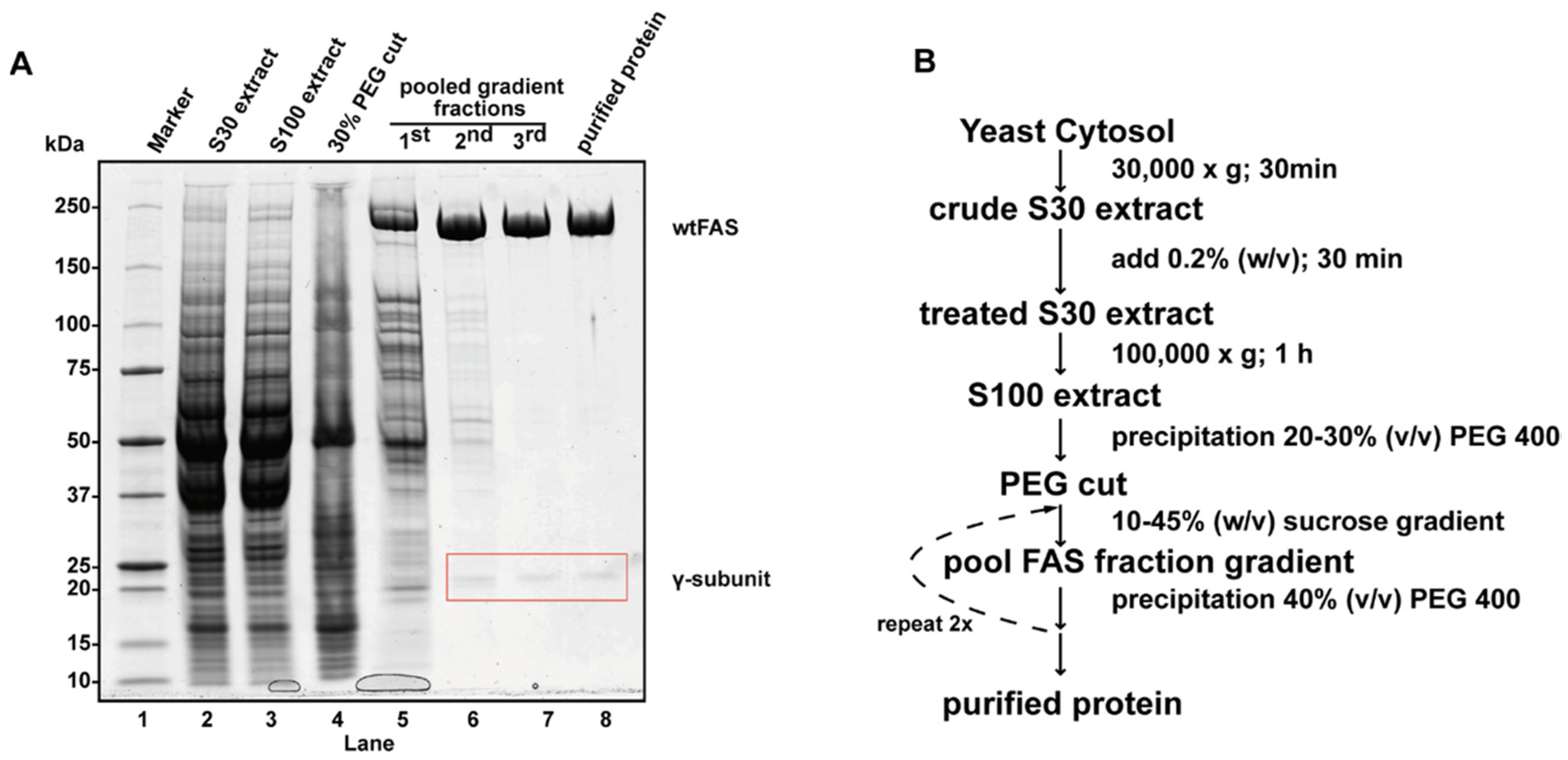
4.4. Case Study 4: Yeast Fatty Acid Synthase
4.5. Case Study 5: Combination of PEG Precipitation with Chromatographic Procedures
5. Considerations for the Incorporation into Biopharmaceutical Production Pipelines
6. Outlook and Perspectives
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Referenced Patents
References
- Hartl, F.U. Cellular Homeostasis and Aging. Annu. Rev. Biochem. 2016, 85, 1–4. [Google Scholar] [CrossRef]
- Marth, J.D. A unified vision of the building blocks of life. Nat. Cell Biol. 2008, 10, 1015–1016. [Google Scholar] [CrossRef] [PubMed]
- Menche, J.; Sharma, A.; Kitsak, M.; Ghiassian, S.D.; Vidal, M.; Loscalzo, J.; Barabasi, A.L. Disease networks. Uncovering disease-disease relationships through the incomplete interactome. Science 2015, 347, 1257601. [Google Scholar] [CrossRef] [PubMed]
- Miller, W.; Makova, K.D.; Nekrutenko, A.; Hardison, R.C. Comparative genomics. Annu. Rev. Genom. Hum. Genet. 2004, 5, 15–56. [Google Scholar] [CrossRef]
- Venter, J.C.; Adams, M.D.; Myers, E.W.; Li, P.W.; Mural, R.J.; Sutton, G.G.; Smith, H.O.; Yandell, M.; Evans, C.A.; Holt, R.A.; et al. The sequence of the human genome. Science 2001, 291, 1304–1351. [Google Scholar] [CrossRef]
- Lander, E.S.; Linton, L.M.; Birren, B.; Nusbaum, C.; Zody, M.C.; Baldwin, J.; Devon, K.; Dewar, K.; Doyle, M.; FitzHugh, W.; et al. Initial sequencing and analysis of the human genome. Nature 2001, 409, 860–921. [Google Scholar] [CrossRef]
- Wang, Z.; Gerstein, M.; Snyder, M. RNA-Seq: A revolutionary tool for transcriptomics. Nat. Rev. Genet. 2009, 10, 57–63. [Google Scholar] [CrossRef] [PubMed]
- Saliba, A.E.; Westermann, A.J.; Gorski, S.A.; Vogel, J. Single-cell RNA-seq: Advances and future challenges. Nucleic Acids Res. 2014, 42, 8845–8860. [Google Scholar] [CrossRef] [PubMed]
- Baralle, F.E.; Giudice, J. Alternative splicing as a regulator of development and tissue identity. Nat. Rev. Mol. Cell Biol. 2017, 18, 437–451. [Google Scholar] [CrossRef]
- Eisenberg, E.; Levanon, E.Y. A-to-I RNA editing—immune protector and transcriptome diversifier. Nat. Rev. Genet. 2018, 19, 473–490. [Google Scholar] [CrossRef]
- Harvey, R.F.; Smith, T.S.; Mulroney, T.; Queiroz, R.M.L.; Pizzinga, M.; Dezi, V.; Villenueva, E.; Ramakrishna, M.; Lilley, K.S.; Willis, A.E. Trans-acting translational regulatory RNA binding proteins. Wiley Interdiscip. Rev. RNA 2018, 9, e1465. [Google Scholar] [CrossRef] [PubMed]
- Forler, D.; Kocher, T.; Rode, M.; Gentzel, M.; Izaurralde, E.; Wilm, M. An efficient protein complex purification method for functional proteomics in higher eukaryotes. Nat. Biotechnol. 2003, 21, 89–92. [Google Scholar] [CrossRef] [PubMed]
- Rigaut, G.; Shevchenko, A.; Rutz, B.; Wilm, M.; Mann, M.; Seraphin, B. A generic protein purification method for protein complex characterization and proteome exploration. Nat. Biotechnol. 1999, 17, 1030–1032. [Google Scholar] [CrossRef] [PubMed]
- Puig, O.; Caspary, F.; Rigaut, G.; Rutz, B.; Bouveret, E.; Bragado-Nilsson, E.; Wilm, M.; Seraphin, B. The tandem affinity purification (TAP) method: A general procedure of protein complex purification. Methods 2001, 24, 218–229. [Google Scholar] [CrossRef] [PubMed]
- Blackstock, W.P.; Weir, M.P. Proteomics: Quantitative and physical mapping of cellular proteins. Trends Biotechnol. 1999, 17, 121–127. [Google Scholar] [CrossRef]
- Shevchenko, A.; Jensen, O.N.; Podtelejnikov, A.V.; Sagliocco, F.; Wilm, M.; Vorm, O.; Mortensen, P.; Shevchenko, A.; Boucherie, H.; Mann, M. Linking genome and proteome by mass spectrometry: Large-scale identification of yeast proteins from two dimensional gels. Proc. Natl. Acad. Sci. USA 1996, 93, 14440–14445. [Google Scholar] [CrossRef]
- Gavin, A.C.; Bosche, M.; Krause, R.; Grandi, P.; Marzioch, M.; Bauer, A.; Schultz, J.; Rick, J.M.; Michon, A.M.; Cruciat, C.M.; et al. Functional organization of the yeast proteome by systematic analysis of protein complexes. Nature 2002, 415, 141–147. [Google Scholar] [CrossRef]
- Ito, T.; Chiba, T.; Ozawa, R.; Yoshida, M.; Hattori, M.; Sakaki, Y. A comprehensive two-hybrid analysis to explore the yeast protein interactome. Proc. Natl. Acad. Sci. USA 2001, 98, 4569–4574. [Google Scholar] [CrossRef]
- LaCava, J.; Molloy, K.R.; Taylor, M.S.; Domanski, M.; Chait, B.T.; Rout, M.P. Affinity proteomics to study endogenous protein complexes: Pointers, pitfalls, preferences and perspectives. Biotechniques 2015, 58, 103–119. [Google Scholar] [CrossRef]
- Gorka, M.; Swart, C.; Siemiatkowska, B.; Martinez-Jaime, S.; Skirycz, A.; Streb, S.; Graf, A. Protein Complex Identification and quantitative complexome by CN-PAGE. Sci. Rep. 2019, 9, 11523. [Google Scholar] [CrossRef]
- Hakhverdyan, Z.; Domanski, M.; Hough, L.E.; Oroskar, A.A.; Oroskar, A.R.; Keegan, S.; Dilworth, D.J.; Molloy, K.R.; Sherman, V.; Aitchison, J.D.; et al. Rapid, optimized interactomic screening. Nat. Methods 2015, 12, 553–560. [Google Scholar] [CrossRef] [PubMed]
- Terpe, K. Overview of tag protein fusions: From molecular and biochemical fundamentals to commercial systems. Appl. Microbiol. Biotechnol. 2003, 60, 523–533. [Google Scholar] [CrossRef] [PubMed]
- Oeffinger, M. Two steps forward-one step back: Advances in affinity purification mass spectrometry of macromolecular complexes. Proteomics 2012, 12, 1591–1608. [Google Scholar] [CrossRef]
- Barends, T.R.; Foucar, L.; Botha, S.; Doak, R.B.; Shoeman, R.L.; Nass, K.; Koglin, J.E.; Williams, G.J.; Boutet, S.; Messerschmidt, M.; et al. De novo protein crystal structure determination from X-ray free-electron laser data. Nature 2014, 505, 244–247. [Google Scholar] [CrossRef] [PubMed]
- Cavalli, A.; Salvatella, X.; Dobson, C.M.; Vendruscolo, M. Protein structure determination from NMR chemical shifts. Proc. Natl. Acad. Sci. USA 2007, 104, 9615–9620. [Google Scholar] [CrossRef]
- Cheng, Y.; Grigorieff, N.; Penczek, P.A.; Walz, T. A primer to single-particle cryo-electron microscopy. Cell 2015, 161, 438–449. [Google Scholar] [CrossRef] [PubMed]
- Manjasetty, B.A.; Turnbull, A.P.; Panjikar, S.; Bussow, K.; Chance, M.R. Automated technologies and novel techniques to accelerate protein crystallography for structural genomics. Proteomics 2008, 8, 612–625. [Google Scholar] [CrossRef]
- Mueller, M.; Jenni, S.; Ban, N. Strategies for crystallization and structure determination of very large macromolecular assemblies. Curr. Opin. Struct. Biol. 2007, 17, 572–579. [Google Scholar] [CrossRef]
- Stellato, F.; Oberthur, D.; Liang, M.; Bean, R.; Gati, C.; Yefanov, O.; Barty, A.; Burkhardt, A.; Fischer, P.; Galli, L.; et al. Room-temperature macromolecular serial crystallography using synchrotron radiation. IUCrJ 2014, 1, 204–212. [Google Scholar] [CrossRef]
- Chari, A.; Haselbach, D.; Kirves, J.M.; Ohmer, J.; Paknia, E.; Fischer, N.; Ganichkin, O.; Moller, V.; Frye, J.J.; Petzold, G.; et al. ProteoPlex: Stability optimization of macromolecular complexes by sparse-matrix screening of chemical space. Nat. Methods 2015, 12, 859–865. [Google Scholar] [CrossRef]
- Magnusson, A.O.; Szekrenyi, A.; Joosten, H.J.; Finnigan, J.; Charnock, S.; Fessner, W.D. nanoDSF as screening tool for enzyme libraries and biotechnology development. FEBS J. 2019, 286, 184–204. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Yu, S.; Siedler, M.; Ihnat, P.M.; Filoti, D.I.; Lu, M.; Zuo, L. A power compensated differential scanning calorimeter for protein stability characterization. Sens. Actuators B Chem. 2018, 256, 946–952. [Google Scholar] [CrossRef]
- VanAernum, Z.L.; Busch, F.; Jones, B.J.; Jia, M.; Chen, Z.; Boyken, S.E.; Sahasrabuddhe, A.; Baker, D.; Wysocki, V.H. Rapid online buffer exchange for screening of proteins, protein complexes and cell lysates by native mass spectrometry. Nat. Protoc. 2020, 15, 1132–1157. [Google Scholar] [CrossRef]
- Johansson, P.; Wiltschi, B.; Kumari, P.; Kessler, B.; Vonrhein, C.; Vonck, J.; Oesterhelt, D.; Grininger, M. Inhibition of the fungal fatty acid synthase type I multienzyme complex. Proc. Natl. Acad. Sci. USA 2008, 105, 12803–12808. [Google Scholar] [CrossRef]
- Leggett, D.S.; Glickman, M.H.; Finley, D. Purification of proteasomes, proteasome subcomplexes, and proteasome-associated proteins from budding yeast. Methods Mol. Biol. 2005, 301, 57–70. [Google Scholar] [CrossRef]
- Myers, L.C.; Leuther, K.; Bushnell, D.A.; Gustafsson, C.M.; Kornberg, R.D. Yeast RNA polymerase II transcription reconstituted with purified proteins. Methods 1997, 12, 212–216. [Google Scholar] [CrossRef] [PubMed]
- Unverdorben, P.; Beck, F.; Sledz, P.; Schweitzer, A.; Pfeifer, G.; Plitzko, J.M.; Baumeister, W.; Forster, F. Deep classification of a large cryo-EM dataset defines the conformational landscape of the 26S proteasome. Proc. Natl. Acad. Sci. USA 2014, 111, 5544–5549. [Google Scholar] [CrossRef]
- Jungbauer, A.; Hahn, R. Chapter 22 Ion-Exchange Chromatography. In Guide to Protein Purification, 2nd ed.; Methods in Enzymology; Academic Press: Cambridge, MA, USA, 2009; pp. 349–371. [Google Scholar]
- Muhleip, A.; McComas, S.E.; Amunts, A. Structure of a mitochondrial ATP synthase with bound native cardiolipin. Elife 2019, 8, e51179. [Google Scholar] [CrossRef]
- Rackiewicz, M.; Grosse-Hovest, L.; Alpert, A.J.; Zarei, M.; Dengjel, J. Hydrophobic Interaction Chromatography for Bottom-Up Proteomics Analysis of Single Proteins and Protein Complexes. J. Proteome Res. 2017, 16, 2318–2323. [Google Scholar] [CrossRef]
- Dalvai, M.; Loehr, J.; Jacquet, K.; Huard, C.C.; Roques, C.; Herst, P.; Cote, J.; Doyon, Y. A Scalable Genome-Editing-Based Approach for Mapping Multiprotein Complexes in Human Cells. Cell Rep. 2015, 13, 621–633. [Google Scholar] [CrossRef]
- Makowska-Grzyska, M.; Kim, Y.; Maltseva, N.; Li, H.; Zhou, M.; Joachimiak, G.; Babnigg, G.; Joachimiak, A. Protein production for structural genomics using E. coli expression. Methods Mol. Biol. 2014, 1140, 89–105. [Google Scholar] [CrossRef] [PubMed]
- Jungbauer, A. Chromatographic media for bioseparation. J. Chromatogr. A 2005, 1065, 3–12. [Google Scholar] [CrossRef] [PubMed]
- Afeyan, N.B.; Gordon, N.F.; Mazsaroff, I.; Varady, L.; Fulton, S.P.; Yang, Y.B.; Regnier, F.E. Flow-through particles for the high-performance liquid chromatographic separation of biomolecules: Perfusion chromatography. J. Chromatogr. A 1990, 519, 1–29. [Google Scholar] [CrossRef]
- Yao, Y.; Czymmek, K.J.; Pazhianur, R.; Lenhoff, A.M. Three-dimensional pore structure of chromatographic adsorbents from electron tomography. Langmuir 2006, 22, 11148–11157. [Google Scholar] [CrossRef] [PubMed]
- Arrua, R.D.; Alvarez Igarzabal, C.I. Macroporous monolithic supports for affinity chromatography. J. Sep. Sci. 2011, 34, 1974–1987. [Google Scholar] [CrossRef]
- Singh, K.; Graf, B.; Linden, A.; Sautner, V.; Urlaub, H.; Tittmann, K.; Stark, H.; Chari, A. Discovery of a Regulatory Subunit of the Yeast Fatty Acid Synthase. Cell 2020, 180, 1130–1143.e20. [Google Scholar] [CrossRef]
- Masuoka, T.; Takatsu, Y. Turbulence model for flow through porous media. Int. J. Heat Mass Transf. 1996, 39, 2803–2809. [Google Scholar] [CrossRef]
- Gagnon, P. The emerging generation of chromatography tools for virus purification. BioProcess Int. 2008, 6, 24–30. [Google Scholar]
- Kirkland, J.J.; Truszkowski, F.A.; Dilks, C.H.; Engel, G.S. Superficially porous silica microspheres for fast high-performance liquid chromatography of macromolecules. J. Chromatogr. A 2000, 890, 3–13. [Google Scholar] [CrossRef]
- Gagnon, P. Technology trends in antibody purification. J. Chromatogr. A 2012, 1221, 57–70. [Google Scholar] [CrossRef]
- Jungbauer, A. Continuous downstream processing of biopharmaceuticals. Trends Biotechnol. 2013, 31, 479–492. [Google Scholar] [CrossRef]
- Belenkii, B.G. Monolithic stationary phases: Yesterday, today, and tomorrow. Russ. J. Bioorganic Chem. 2006, 32, 323–332. [Google Scholar] [CrossRef]
- Hahn, R.; Panzer, M.; Hansen, E.; Mollerup, J.; Jungbauer, A. Mass transfer properties of monoliths. Sep. Sci. Technol. 2002, 37, 1545–1565. [Google Scholar] [CrossRef]
- Jungbauer, A.; Hahn, R. Monoliths for fast bioseparation and bioconversion and their applications in biotechnology. J. Sep. Sci. 2004, 27, 767–778. [Google Scholar] [CrossRef] [PubMed]
- Lynch, K.B.; Ren, J.; Beckner, M.A.; He, C.; Liu, S. Monolith columns for liquid chromatographic separations of intact proteins: A review of recent advances and applications. Anal. Chim. Acta 2019, 1046, 48–68. [Google Scholar] [CrossRef] [PubMed]
- Trauner, A.; Bennett, M.H.; Williams, H.D. Isolation of bacterial ribosomes with monolith chromatography. PLoS ONE 2011, 6, e16273. [Google Scholar] [CrossRef] [PubMed]
- Safarik, I.; Safarikova, M. Magnetic techniques for the isolation and purification of proteins and peptides. Biomagn. Res. Technol. 2004, 2, 7. [Google Scholar] [CrossRef]
- Albertsson, P.A. Partition of proteins in liquid polymer-polymer two-phase systems. Nature 1958, 182, 709–711. [Google Scholar] [CrossRef]
- González-González, M.; Picó, G.; Lima, Á.S.; Ussemane Mussagy, C.; Pereira, J.F.B.; Fernandez-Lahore, H.M.; Asenjo, J.A.; Rito-Palomares, M. Aqueous two-phase systems in Latin America: Perspective and future trends. J. Chem. Technol. Biotechnol. 2021. [Google Scholar] [CrossRef]
- Pereira, J.F.B.; Freire, M.G.; Coutinho, J.A.P. Aqueous two-phase systems: Towards novel and more disruptive applications. Fluid Phase Equilibria 2020, 505, 112341. [Google Scholar] [CrossRef]
- Albertsson, P.A.; Philipson, L. Antigen-antibody in liquid two-phase systems: A method for studying immunological reactions. Nature 1960, 185, 38–40. [Google Scholar] [CrossRef] [PubMed]
- Polson, A.; Potgieter, G.M.; Largier, J.F.; Mears, G.E.F.; Joubert, F.J. The fractionation of protein mixtures by linear polymers of high molecular weight. Biochim. et Biophys. Acta (BBA)-Gen. Subj. 1964, 82, 463–475. [Google Scholar] [CrossRef]
- Atha, D.H.; Ingham, K.C. Mechanism of precipitation of proteins by polyethylene glycols. Analysis in terms of excluded volume. J. Biol. Chem. 1981, 256, 12108–12117. [Google Scholar] [CrossRef]
- Lee, L.L.; Lee, J.C. Thermal stability of proteins in the presence of poly(ethylene glycols). Biochemistry 1987, 26, 7813–7819. [Google Scholar] [CrossRef] [PubMed]
- Hammerling, F.; Ladd Effio, C.; Andris, S.; Kittelmann, J.; Hubbuch, J. Investigation and prediction of protein precipitation by polyethylene glycol using quantitative structure-activity relationship models. J. Biotechnol. 2017, 241, 87–97. [Google Scholar] [CrossRef] [PubMed]
- Grosshans, S.; Wang, G.; Hubbuch, J. Water on hydrophobic surfaces: Mechanistic modeling of polyethylene glycol-induced protein precipitation. Bioprocess Biosyst. Eng. 2019, 42, 513–520. [Google Scholar] [CrossRef]
- Iqbal, M.; Tao, Y.; Xie, S.; Zhu, Y.; Chen, D.; Wang, X.; Huang, L.; Peng, D.; Sattar, A.; Shabbir, M.A.; et al. Aqueous two-phase system (ATPS): An overview and advances in its applications. Biol. Proced. Online 2016, 18, 18. [Google Scholar] [CrossRef]
- Grosshans, S.; Suhm, S.; Hubbuch, J. Precipitation of complex antibody solutions: Influence of contaminant composition and cell culture medium on the precipitation behavior. Bioprocess Biosyst. Eng. 2019, 42, 1039–1051. [Google Scholar] [CrossRef]
- Kurimoto, E.; Satoh, T.; Ito, Y.; Ishihara, E.; Okamoto, K.; Yagi-Utsumi, M.; Tanaka, K.; Kato, K. Crystal structure of human proteasome assembly chaperone PAC4 involved in proteasome formation. Protein Sci. 2017, 26, 1080–1085. [Google Scholar] [CrossRef]
- Le Tallec, B.; Barrault, M.B.; Courbeyrette, R.; Guerois, R.; Marsolier-Kergoat, M.C.; Peyroche, A. 20S proteasome assembly is orchestrated by two distinct pairs of chaperones in yeast and in mammals. Mol. Cell 2007, 27, 660–674. [Google Scholar] [CrossRef]
- Toste Rego, A.; da Fonseca, P.C.A. Characterization of Fully Recombinant Human 20S and 20S-PA200 Proteasome Complexes. Mol. Cell 2019, 76, 138–147.e5. [Google Scholar] [CrossRef]
- Schrader, J.; Henneberg, F.; Mata, R.A.; Tittmann, K.; Schneider, T.R.; Stark, H.; Bourenkov, G.; Chari, A. The inhibition mechanism of human 20S proteasomes enables next-generation inhibitor design. Science 2016, 353, 594–598. [Google Scholar] [CrossRef] [PubMed]
- Ukmar-Godec, T.; Fang, P.; Ibanez de Opakua, A.; Henneberg, F.; Godec, A.; Pan, K.T.; Cima-Omori, M.S.; Chari, A.; Mandelkow, E.; Urlaub, H.; et al. Proteasomal degradation of the intrinsically disordered protein tau at single-residue resolution. Sci. Adv. 2020, 6, eaba3916. [Google Scholar] [CrossRef] [PubMed]
- Becker, P.B.; Wu, C. Cell-free system for assembly of transcriptionally repressed chromatin from Drosophila embryos. Mol. Cell. Biol. 1992, 12, 2241–2249. [Google Scholar] [CrossRef]
- Kish-Trier, E.; Hill, C.P. Structural biology of the proteasome. Annu. Rev. Biophys. 2013, 42, 29–49. [Google Scholar] [CrossRef] [PubMed]
- Mao, Y. Structure, Dynamics and Function of the 26S Proteasome. Subcell Biochem. 2021, 96, 1–151. [Google Scholar] [CrossRef]
- Finley, D. Recognition and processing of ubiquitin-protein conjugates by the proteasome. Annu. Rev. Biochem. 2009, 78, 477–513. [Google Scholar] [CrossRef]
- Marshall, R.S.; Vierstra, R.D. Dynamic Regulation of the 26S Proteasome: From Synthesis to Degradation. Front. Mol. Biosci. 2019, 6, 40. [Google Scholar] [CrossRef]
- Haselbach, D.; Schrader, J.; Lambrecht, F.; Henneberg, F.; Chari, A.; Stark, H. Long-range allosteric regulation of the human 26S proteasome by 20S proteasome-targeting cancer drugs. Nat. Commun. 2017, 8, 15578. [Google Scholar] [CrossRef]
- Huber, E.M.; Basler, M.; Schwab, R.; Heinemeyer, W.; Kirk, C.J.; Groettrup, M.; Groll, M. Immuno- and constitutive proteasome crystal structures reveal differences in substrate and inhibitor specificity. Cell 2012, 148, 727–738. [Google Scholar] [CrossRef]
- Murata, S.; Takahama, Y.; Kasahara, M.; Tanaka, K. The immunoproteasome and thymoproteasome: Functions, evolution and human disease. Nat. Immunol. 2018, 19, 923–931. [Google Scholar] [CrossRef]
- Dechavanne, V.; Vilbois, F.; Glez, L.; Antonsson, B. Purification and separation of the 20S immunoproteasome from the constitutive proteasome and identification of the subunits by LC-MS. Protein Expr. Purif. 2013, 87, 100–110. [Google Scholar] [CrossRef]
- Hammerschmidt, N.; Hobiger, S.; Jungbauer, A. Continuous polyethylene glycol precipitation of recombinant antibodies: Sequential precipitation and resolubilization. Process. Biochem. 2016, 51, 325–332. [Google Scholar] [CrossRef]
- Lohmann, L.J.; Strube, J. Accelerating Biologics Manufacturing by Modeling: Process Integration of Precipitation in mAb Downstream Processing. Processes 2020, 8, 58. [Google Scholar] [CrossRef]
- Martinez, M.; Spitali, M.; Norrant, E.L.; Bracewell, D.G. Precipitation as an Enabling Technology for the Intensification of Biopharmaceutical Manufacture. Trends Biotechnol. 2019, 37, 237–241. [Google Scholar] [CrossRef]
- Oelmeier, S.A.; Ladd-Effio, C.; Hubbuch, J. Alternative separation steps for monoclonal antibody purification: Combination of centrifugal partitioning chromatography and precipitation. J. Chromatogr. A 2013, 1319, 118–126. [Google Scholar] [CrossRef]
- Dos Santos, R.; Carvalho, A.L.; Roque, A.C. Renaissance of protein crystallization and precipitation in biopharmaceuticals purification. Biotechnol. Adv. 2017, 35, 41–50. [Google Scholar] [CrossRef]
- Geng, F.; Xie, Y.; Wang, J.; Li, S.; Jin, Y.; Ma, M. Large-scale purification of ovalbumin using polyethylene glycol precipitation and isoelectric precipitation. Poult. Sci. 2019, 98, 1545–1550. [Google Scholar] [CrossRef]
- Asakura, S.; Oosawa, F. Interaction between particles suspended in solutions of macromolecules. J. Polym. Sci. 1958, 33, 183–192. [Google Scholar] [CrossRef]
- Odijk, T. Depletion theory and the precipitation of protein by polymer. J. Phys. Chem. B 2009, 113, 3941–3946. [Google Scholar] [CrossRef]
- Polson, A. A theory for the displacement of proteins and viruses with polyethylene glycol. Prep. Biochem. 1977, 7, 129–154. [Google Scholar] [CrossRef] [PubMed]
- Sim, S.L.; He, T.; Tscheliessnig, A.; Mueller, M.; Tan, R.B.; Jungbauer, A. Protein precipitation by polyethylene glycol: A generalized model based on hydrodynamic radius. J. Biotechnol. 2012, 157, 315–319. [Google Scholar] [CrossRef] [PubMed]
- Grimm, C.; Chari, A.; Reuter, K.; Fischer, U. A crystallization screen based on alternative polymeric precipitants. Acta Cryst. D Biol. Cryst. 2010, 66, 685–697. [Google Scholar] [CrossRef]
- Maestro, B.; Velasco, I.; Castillejo, I.; Arevalo-Rodriguez, M.; Cebolla, A.; Sanz, J.M. Affinity partitioning of proteins tagged with choline-binding modules in aqueous two-phase systems. J. Chromatogr. A 2008, 1208, 189–196. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Henneberg, F.; Chari, A. Chromatography-Free Purification Strategies for Large Biological Macromolecular Complexes Involving Fractionated PEG Precipitation and Density Gradients. Life 2021, 11, 1289. https://doi.org/10.3390/life11121289
Henneberg F, Chari A. Chromatography-Free Purification Strategies for Large Biological Macromolecular Complexes Involving Fractionated PEG Precipitation and Density Gradients. Life. 2021; 11(12):1289. https://doi.org/10.3390/life11121289
Chicago/Turabian StyleHenneberg, Fabian, and Ashwin Chari. 2021. "Chromatography-Free Purification Strategies for Large Biological Macromolecular Complexes Involving Fractionated PEG Precipitation and Density Gradients" Life 11, no. 12: 1289. https://doi.org/10.3390/life11121289
APA StyleHenneberg, F., & Chari, A. (2021). Chromatography-Free Purification Strategies for Large Biological Macromolecular Complexes Involving Fractionated PEG Precipitation and Density Gradients. Life, 11(12), 1289. https://doi.org/10.3390/life11121289





