Structure, Activity and Function of the PRMT2 Protein Arginine Methyltransferase
Abstract
:1. Introduction
2. Structural features
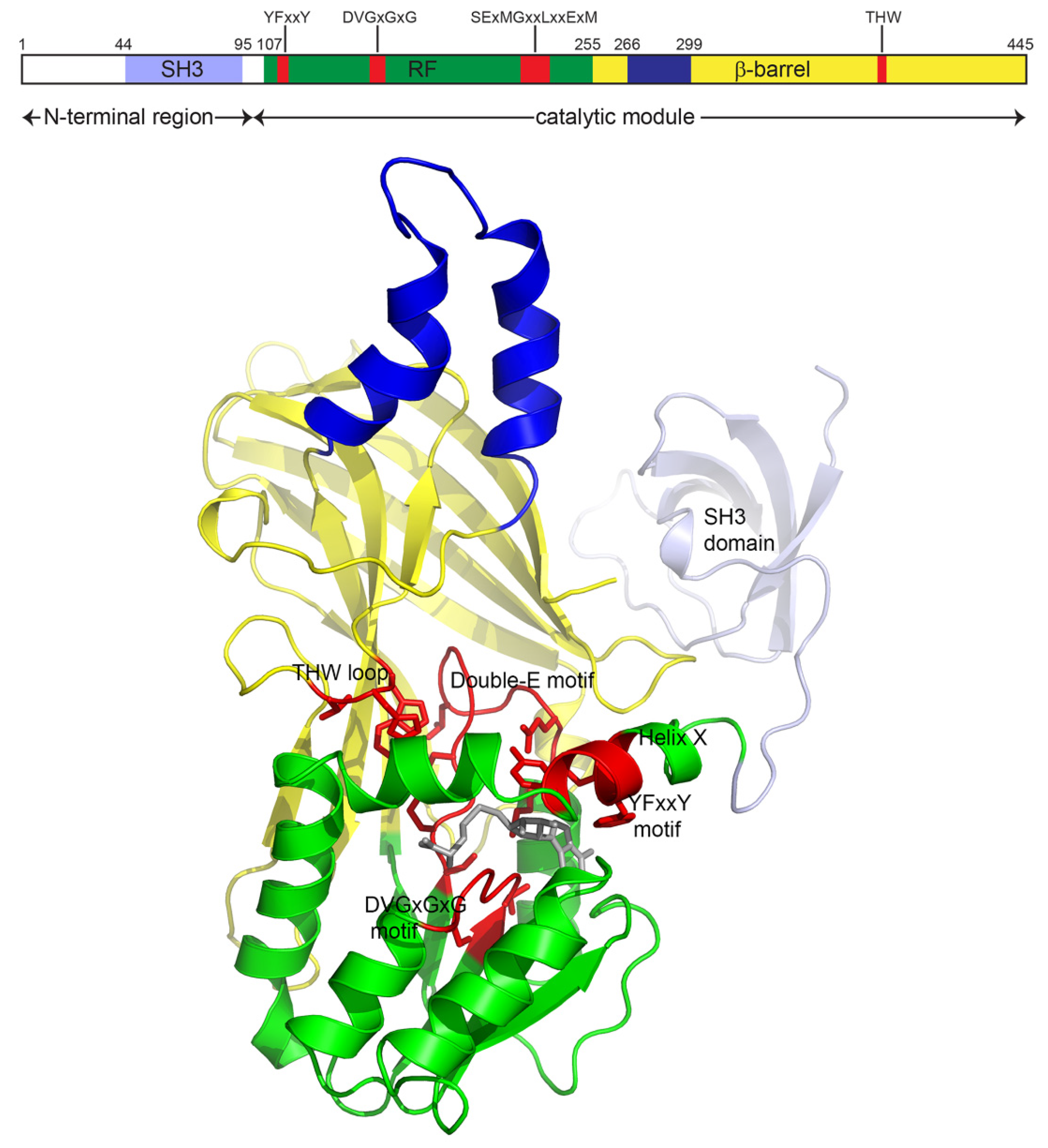
2.1. Sequence
2.2. Structure
2.3. Co-Factor Binding Site
2.4. Substrate Binding Pocket
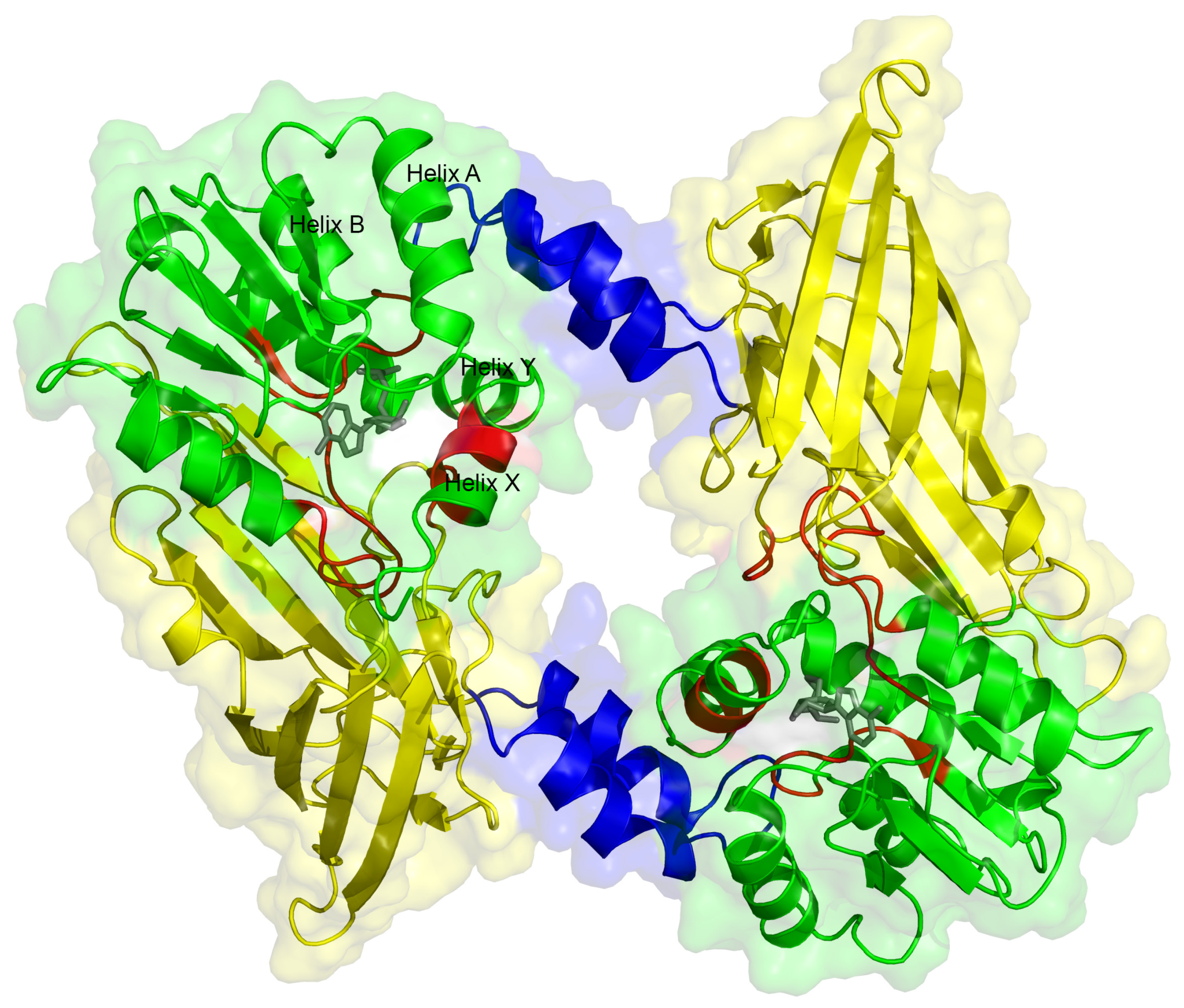
2.5. Dimerization
3. Activity
4. Interactions
4.1. Coactivation
4.2. Splicing
5. Diseases
5.1. Breast Cancer
5.2. Other Pathologies
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Acknowledgments
Conflicts of Interest
References
- Gayatri, S.; Bedford, M.T. Readers of histone methylarginine marks. Biochim. Biophys. Acta 2014, 1839, 702–710. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lorton, B.M.; Shechter, D. Cellular consequences of arginine methylation. Cell Mol. Life Sci. 2019, 76, 2933–2956. [Google Scholar] [CrossRef] [PubMed]
- Price, O.M.; Hevel, J.M. Toward Understanding Molecular Recognition between PRMTs and their Substrates. Curr. Protein Pept. Sci. 2020, 21, 713–724. [Google Scholar] [CrossRef] [PubMed]
- Wei, H.-H.; Fan, X.-J.; Hu, Y.; Tian, X.-X.; Guo, M.; Mao, M.-W.; Fang, Z.-Y.; Wu, P.; Gao, S.-X.; Peng, C.; et al. A systematic survey of PRMT interactomes reveals the key roles of arginine methylation in the global control of RNA splicing and translation. Sci. Bull. 2021, 66, 1342–1357. [Google Scholar] [CrossRef]
- Katsanis, N.; Yaspo, M.L.; Fisher, E.M. Identification and mapping of a novel human gene, HRMT1L1, homologous to the rat protein arginine N-methyltransferase 1 (PRMT1) gene. Mamm. Genome 1997, 8, 526–529. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.C.; Wang, J.D.; Chen, C.H.; Chen, Y.W.; Li, C. A novel BLAST-Based Relative Distance (BBRD) method can effectively group members of protein arginine methyltransferases and suggest their evolutionary relationship. Mol. Phylogenet. Evol. 2015, 84, 101–111. [Google Scholar] [CrossRef]
- Wang, Y.C.; Li, C. Evolutionarily conserved protein arginine methyltransferases in non-mammalian animal systems. FEBS J. 2012, 279, 932–945. [Google Scholar] [CrossRef] [Green Version]
- Kzhyshkowska, J.; Schütt, H.; Liss, M.; Kremmer, E.; Stauber, R.; Wolf, H.; Dobner, T. Heterogeneous nuclear ribonucleoprotein E1B-AP5 is methylated in its Arg-Gly-Gly (RGG) box and interacts with human arginine methyltransferase HRMT1L1. Biochem. J. 2001, 358, 305–314. [Google Scholar] [CrossRef]
- Meyer, R.; Wolf, S.S.; Obendorf, M. PRMT2, a member of the protein arginine methyltransferase family, is a coactivator of the androgen receptor. J. Steroid Biochem. Mol. Biol. 2007, 107, 1–14. [Google Scholar] [CrossRef]
- Cura, V.; Marechal, N.; Troffer-Charlier, N.; Strub, J.M.; van Haren, M.J.; Martin, N.I.; Cianferani, S.; Bonnefond, L.; Cavarelli, J. Structural studies of protein arginine methyltransferase 2 reveal its interactions with potential substrates and inhibitors. FEBS J. 2017, 284, 77–96. [Google Scholar] [CrossRef] [Green Version]
- Beitz, E. TEXshade: Shading and labeling of multiple sequence alignments using LATEX2 epsilon. Bioinformatics 2000, 16, 135–139. [Google Scholar] [CrossRef] [Green Version]
- Senior, A.W.; Evans, R.; Jumper, J.; Kirkpatrick, J.; Sifre, L.; Green, T.; Qin, C.; Zidek, A.; Nelson, A.W.R.; Bridgland, A.; et al. Improved protein structure prediction using potentials from deep learning. Nature 2020, 577, 706–710. [Google Scholar] [CrossRef]
- Zhong, J.; Cao, R.X.; Hong, T.; Yang, J.; Zu, X.Y.; Xiao, X.H.; Liu, J.H.; Wen, G.B. Identification and expression analysis of a novel transcript of the human PRMT2 gene resulted from alternative polyadenylation in breast cancer. Gene 2011, 487, 1–9. [Google Scholar] [CrossRef]
- Zhong, J.; Cao, R.X.; Zu, X.Y.; Hong, T.; Yang, J.; Liu, L.; Xiao, X.H.; Ding, W.J.; Zhao, Q.; Liu, J.H.; et al. Identification and characterization of novel spliced variants of PRMT2 in breast carcinoma. FEBS J. 2012, 279, 316–335. [Google Scholar] [CrossRef] [PubMed]
- Troffer-Charlier, N.; Cura, V.; Hassenboehler, P.; Moras, D.; Cavarelli, J. Functional insights from structures of coactivator-associated arginine methyltransferase 1 domains. EMBO J. 2007, 26, 4391–4401. [Google Scholar] [CrossRef]
- Carducci, M.; Perfetto, L.; Briganti, L.; Paoluzi, S.; Costa, S.; Zerweck, J.; Schutkowski, M.; Castagnoli, L.; Cesareni, G. The protein interaction network mediated by human SH3 domains. Biotechnol. Adv. 2012, 30, 4–15. [Google Scholar] [CrossRef] [PubMed]
- Meirson, T.; Bomze, D.; Markel, G.; Samson, A.O. kappa-helix and the helical lock and key model: A pivotal way of looking at polyproline II. Bioinformatics 2020, 36, 3726–3732. [Google Scholar] [CrossRef]
- Zhang, X.; Zhou, L.; Cheng, X. Crystal structure of the conserved core of protein arginine methyltransferase PRMT3. EMBO J. 2000, 19, 3509–3519. [Google Scholar] [CrossRef]
- Rust, H.L.; Zurita-Lopez, C.I.; Clarke, S.; Thompson, P.R. Mechanistic Studies on Transcriptional Coactivator Protein Arginine Methyltransferase 1. Biochemistry 2011, 50, 3332–3345. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Scott, H.S.; Antonarakis, S.E.; Lalioti, M.D.; Rossier, C.; Silver, P.A.; Henry, M.F. Identification and Characterization of Two Putative Human Arginine Methyltransferases (HRMT1L1 and HRMT1L2). Genomics 1998, 48, 330–340. [Google Scholar] [CrossRef]
- Frankel, A.; Brown, J.I. Evaluation of kinetic data: What the numbers tell us about PRMTs. Biochim. Biophys. Acta Proteins Proteom. 2019, 1867, 306–316. [Google Scholar] [CrossRef]
- Lakowski, T.M.; Frankel, A. Kinetic analysis of human protein arginine N-methyltransferase 2: Formation of monomethyl- and asymmetric dimethyl-arginine residues on histone H4. Biochem. J. 2009, 421, 253–261. [Google Scholar] [CrossRef]
- Blythe, S.A.; Cha, S.W.; Tadjuidje, E.; Heasman, J.; Klein, P.S. Beta-Catenin primes organizer gene expression by recruiting a histone H3 arginine 8 methyltransferase, Prmt2. Dev. Cell 2010, 19, 220–231. [Google Scholar] [CrossRef] [Green Version]
- Su, X.; Zhu, G.; Ding, X.; Lee, S.Y.; Dou, Y.; Zhu, B.; Wu, W.; Li, H. Molecular basis underlying histone H3 lysine-arginine methylation pattern readout by Spin/Ssty repeats of Spindlin1. Genes Dev. 2014, 28, 622–636. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dong, F.; Li, Q.; Yang, C.; Huo, D.; Wang, X.; Ai, C.; Kong, Y.; Sun, X.; Wang, W.; Zhou, Y.; et al. PRMT2 links histone H3R8 asymmetric dimethylation to oncogenic activation and tumorigenesis of glioblastoma. Nat. Commun. 2018, 9, 4552. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hu, G.; Yan, C.; Xie, P.; Cao, Y.; Shao, J.; Ge, J. PRMT2 accelerates tumorigenesis of hepatocellular carcinoma by activating Bcl2 via histone H3R8 methylation. Exp. Cell Res. 2020, 394, 112152. [Google Scholar] [CrossRef] [PubMed]
- Bonnefond, L.; Stojko, J.; Mailliot, J.; Troffer-Charlier, N.; Cura, V.; Wurtz, J.M.; Cianferani, S.; Cavarelli, J. Functional insights from high resolution structures of mouse protein arginine methyltransferase 6. J. Struct. Biol. 2015, 191, 175–183. [Google Scholar] [CrossRef]
- Frankel, A.; Yadav, N.; Lee, J.; Branscombe, T.L.; Clarke, S.; Bedford, M.T. The novel human protein arginine N-methyltransferase PRMT6 is a nuclear enzyme displaying unique substrate specificity. J. Biol. Chem. 2002, 277, 3537–3543. [Google Scholar] [CrossRef] [Green Version]
- Dillon, M.B.; Rust, H.L.; Thompson, P.R.; Mowen, K.A. Automethylation of protein arginine methyltransferase 8 (PRMT8) regulates activity by impeding S-adenosylmethionine sensitivity. J. Biol. Chem. 2013, 288, 27872–27880. [Google Scholar] [CrossRef] [Green Version]
- Kuhn, P.; Chumanov, R.; Wang, Y.; Ge, Y.; Burgess, R.; Xu, W. Automethylation of CARM1 allows coupling of transcription and mRNA splicing. Nucleic Acids Res. 2011, 39, 2717–2726. [Google Scholar] [CrossRef] [Green Version]
- Guo, A.; Gu, H.; Zhou, J.; Mulhern, D.; Wang, Y.; Lee, K.A.; Yang, V.; Aguiar, M.; Kornhauser, J.; Jia, X.; et al. Immunoaffinity enrichment and mass spectrometry analysis of protein methylation. Mol. Cell Proteom. 2014, 13, 372–387. [Google Scholar] [CrossRef] [Green Version]
- Larsen, S.C.; Sylvestersen, K.B.; Mund, A.; Lyon, D.; Mullari, M.; Madsen, M.V.; Daniel, J.A.; Jensen, L.J.; Nielsen, M.L. Proteome-wide analysis of arginine monomethylation reveals widespread occurrence in human cells. Sci. Signal. 2016, 9, rs9. [Google Scholar] [CrossRef]
- van Haren, M.; van Ufford, L.Q.; Moret, E.E.; Martin, N.I. Synthesis and evaluation of protein arginine N-methyltransferase inhibitors designed to simultaneously occupy both substrate binding sites. Org. Biomol. Chem. 2015, 13, 549–560. [Google Scholar] [CrossRef]
- Qi, C.; Chang, J.; Zhu, Y.; Yeldandi, A.V.; Rao, S.M.; Zhu, Y.-J. Identification of Protein Arginine Methyltransferase 2 as a Coactivator for Estrogen Receptor α. J. Biol. Chem. 2002, 277, 28624–28630. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ganesh, L.; Yoshimoto, T.; Moorthy, N.C.; Akahata, W.; Boehm, M.; Nabel, E.G.; Nabel, G.J. Protein methyltransferase 2 inhibits NF-kappaB function and promotes apoptosis. Mol. Cell. Biol. 2006, 26, 3864–3874. [Google Scholar] [CrossRef] [Green Version]
- Dalloneau, E.; Pereira, P.L.; Brault, V.; Nabel, E.G.; Herault, Y. Prmt2 regulates the lipopolysaccharide-induced responses in lungs and macrophages. J. Immunol. 2011, 187, 4826–4834. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Iwasaki, H.; Kovacic, J.C.; Olive, M.; Beers, J.K.; Yoshimoto, T.; Crook, M.F.; Tonelli, L.H.; Nabel, E.G. Disruption of protein arginine N-methyltransferase 2 regulates leptin signaling and produces leanness in vivo through loss of STAT3 methylation. Circ. Res. 2010, 107, 992–1001. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hussein, M.A.; Shrestha, E.; Ouimet, M.; Barrett, T.J.; Leone, S.; Moore, K.J.; Herault, Y.; Fisher, E.A.; Garabedian, M.J. LXR-Mediated ABCA1 Expression and Function Are Modulated by High Glucose and PRMT2. PLoS ONE 2015, 10, e0135218. [Google Scholar] [CrossRef] [Green Version]
- Li, Y.Y.; Zhou, S.H.; Chen, S.S.; Zhong, J.; Wen, G.B. PRMT2 inhibits the formation of foam cell induced by ox-LDL in RAW 264.7 macrophage involving ABCA1 mediated cholesterol efflux. Biochem. Biophys. Res. Commun. 2020, 524, 77–82. [Google Scholar] [CrossRef]
- Pak, M.L.; Lakowski, T.M.; Thomas, D.; Vhuiyan, M.I.; Husecken, K.; Frankel, A. A protein arginine N-methyltransferase 1 (PRMT1) and 2 heteromeric interaction increases PRMT1 enzymatic activity. Biochemistry 2011, 50, 8226–8240. [Google Scholar] [CrossRef]
- Yoshimoto, T.; Boehm, M.; Olive, M.; Crook, M.F.; San, H.; Langenickel, T.; Nabel, E.G. The arginine methyltransferase PRMT2 binds RB and regulates E2F function. Exp. Cell Res. 2006, 312, 2040–2053. [Google Scholar] [CrossRef]
- Hou, W.; Nemitz, S.; Schopper, S.; Nielsen, M.L.; Kessels, M.M.; Qualmann, B. Arginine Methylation by PRMT2 Controls the Functions of the Actin Nucleator Cobl. Dev. Cell 2018, 45, 262–275. [Google Scholar] [CrossRef] [Green Version]
- Ahuja, R.; Pinyol, R.; Reichenbach, N.; Custer, L.; Klingensmith, J.; Kessels, M.M.; Qualmann, B. Cordon-bleu is an actin nucleation factor and controls neuronal morphology. Cell 2007, 131, 337–350. [Google Scholar] [CrossRef] [Green Version]
- Yildirim, A.O.; Bulau, P.; Zakrzewicz, D.; Kitowska, K.E.; Weissmann, N.; Grimminger, F.; Morty, R.E.; Eickelberg, O. Increased protein arginine methylation in chronic hypoxia: Role of protein arginine methyltransferases. Am. J. Respir. Cell Mol. Biol. 2006, 35, 436–443. [Google Scholar] [CrossRef] [PubMed]
- Bedford, M.; Richard, S. Arginine methylation an emerging regulator of protein function. Mol. Cell 2005, 18, 263–272. [Google Scholar] [CrossRef] [PubMed]
- Yu, M.C. The Role of Protein Arginine Methylation in mRNP Dynamics. Mol. Biol. Int. 2011, 2011, 163827. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Blackwell, E.; Ceman, S. Arginine methylation of RNA-binding proteins regulates cell function and differentiation. Mol. Reprod. Dev. 2012, 79, 163–175. [Google Scholar] [CrossRef] [Green Version]
- Friesen, W.J.; Paushkin, S.; Wyce, A.; Massenet, S.; Pesiridis, G.S.; Duyne, G.V.; Rappsilber, J.; Mann, M.; Dreyfuss, G. The methylosome, a 20S complex containing JBP1 and pICln, produces dimethylarginine-modified Sm proteins. Mol. Cell. Biol. 2001, 21, 8289–8300. [Google Scholar] [CrossRef] [Green Version]
- Brahms, H.; Meheus, L.; de Brabandere, V.; Fischer, U.; Luhrmann, R. Symmetrical dimethylation of arginine residues in spliceosomal Sm protein B/B′ and the Sm-like protein LSm4, and their interaction with the SMN protein. RNA 2001, 7, 1531–1542. [Google Scholar] [CrossRef] [Green Version]
- Zhang, L.; Tran, N.T.; Su, H.; Wang, R.; Lu, Y.; Tang, H.; Aoyagi, S.; Guo, A.; Khodadadi-Jamayran, A.; Zhou, D.; et al. Cross-talk between PRMT1-mediated methylation and ubiquitylation on RBM15 controls RNA splicing. eLife 2015, 4. [Google Scholar] [CrossRef]
- Cheng, D.; Cote, J.; Shaaban, S.; Bedford, M.T. The arginine methyltransferase CARM1 regulates the coupling of transcription and mRNA processing. Mol. Cell 2007, 25, 71–83. [Google Scholar] [CrossRef]
- Hadjikyriacou, A.; Yang, Y.; Espejo, A.; Bedford, M.T.; Clarke, S.G. Unique Features of Human Protein Arginine Methyltransferase 9 (PRMT9) and Its Substrate RNA Splicing Factor SF3B2. J. Biol. Chem. 2015, 290, 16723–16743. [Google Scholar] [CrossRef] [Green Version]
- Sinha, R.; Allemand, E.; Zhang, Z.; Karni, R.; Myers, M.P.; Krainer, A.R. Arginine Methylation Controls the Subcellular Localization and Functions of the Oncoprotein Splicing Factor SF2/ASF. Mol. Cell. Biol. 2010, 30, 2762–2774. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Vhuiyan, M.I.; Pak, M.L.; Park, M.A.; Thomas, D.; Lakowski, T.M.; Chalfant, C.E.; Frankel, A. PRMT2 interacts with splicing factors and regulates the alternative splicing of BCL-X. J. Biochem. 2017, 162, 17–25. [Google Scholar] [CrossRef]
- Labourier, E.; Bourbon, H.-M.; Gallouzi, I.-E.; Fostier, M.; Allemand, E.; Tazi, J. Antagonism between RSF1 and SR proteins for both splice-site recognition in vitro and Drosophila development. Genes Dev. 1999, 13, 740–753. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bradley, T.; Cook, M.E.; Blanchette, M. SR proteins control a complex network of RNA-processing events. RNA 2015, 21, 75–92. [Google Scholar] [CrossRef] [Green Version]
- Zhong, J.; Cao, R.X.; Liu, J.H.; Liu, Y.B.; Wang, J.; Liu, L.P.; Chen, Y.J.; Yang, J.; Zhang, Q.H.; Wu, Y.; et al. Nuclear loss of protein arginine N-methyltransferase 2 in breast carcinoma is associated with tumor grade and overexpression of cyclin D1 protein. Oncogene 2014, 33, 5546–5558. [Google Scholar] [CrossRef] [Green Version]
- Zhong, J.; Chen, Y.J.; Chen, L.; Shen, Y.Y.; Zhang, Q.H.; Yang, J.; Cao, R.X.; Zu, X.Y.; Wen, G.B. PRMT2beta, a C-terminal splice variant of PRMT2, inhibits the growth of breast cancer cells. Oncol. Rep. 2017, 38, 1303–1311. [Google Scholar] [CrossRef]
- Ho, M.C.; Wilczek, C.; Bonanno, J.B.; Xing, L.; Seznec, J.; Matsui, T.; Carter, L.G.; Onikubo, T.; Kumar, P.R.; Chan, M.K.; et al. Structure of the arginine methyltransferase PRMT5-MEP50 reveals a mechanism for substrate specificity. PLoS ONE 2013, 8, e57008. [Google Scholar] [CrossRef]
- Shen, Y.; Zhong, J.; Liu, J.; Liu, K.; Zhao, J.; Xu, T.; Zeng, T.; Li, Z.; Chen, Y.; Ding, W.; et al. Protein arginine N-methyltransferase 2 reverses tamoxifen resistance in breast cancer cells through suppression of ER-alpha36. Oncol. Rep. 2018, 39, 2604–2612. [Google Scholar] [CrossRef]
- Bednarz-Misa, I.; Fleszar, M.G.; Fortuna, P.; Lewandowski, L.; Mierzchala-Pasierb, M.; Diakowska, D.; Krzystek-Korpacka, M. Altered L-Arginine Metabolic Pathways in Gastric Cancer: Potential Therapeutic Targets and Biomarkers. Biomolecules 2021, 11, 1086. [Google Scholar] [CrossRef]
- Zeng, S.Y.; Luo, J.F.; Quan, H.Y.; Xiao, Y.B.; Liu, Y.H.; Lu, H.Q.; Qin, X.P. Protein Arginine Methyltransferase 2 Inhibits Angiotensin II-Induced Proliferation and Inflammation in Vascular Smooth Muscle Cells. BioMed Res. Int. 2018, 2018, 1547452. [Google Scholar] [CrossRef] [Green Version]



Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cura, V.; Cavarelli, J. Structure, Activity and Function of the PRMT2 Protein Arginine Methyltransferase. Life 2021, 11, 1263. https://doi.org/10.3390/life11111263
Cura V, Cavarelli J. Structure, Activity and Function of the PRMT2 Protein Arginine Methyltransferase. Life. 2021; 11(11):1263. https://doi.org/10.3390/life11111263
Chicago/Turabian StyleCura, Vincent, and Jean Cavarelli. 2021. "Structure, Activity and Function of the PRMT2 Protein Arginine Methyltransferase" Life 11, no. 11: 1263. https://doi.org/10.3390/life11111263






