Application of Domain- and Genotype-Specific Models to Infer Post-Transcriptional Regulation of Segmentation Gene Expression in Drosophila
Abstract
1. Introduction
2. Materials and Methods
2.1. Model and Data
2.2. Model Fitting and Hypotheses
2.3. Parameter Optimization
3. Results
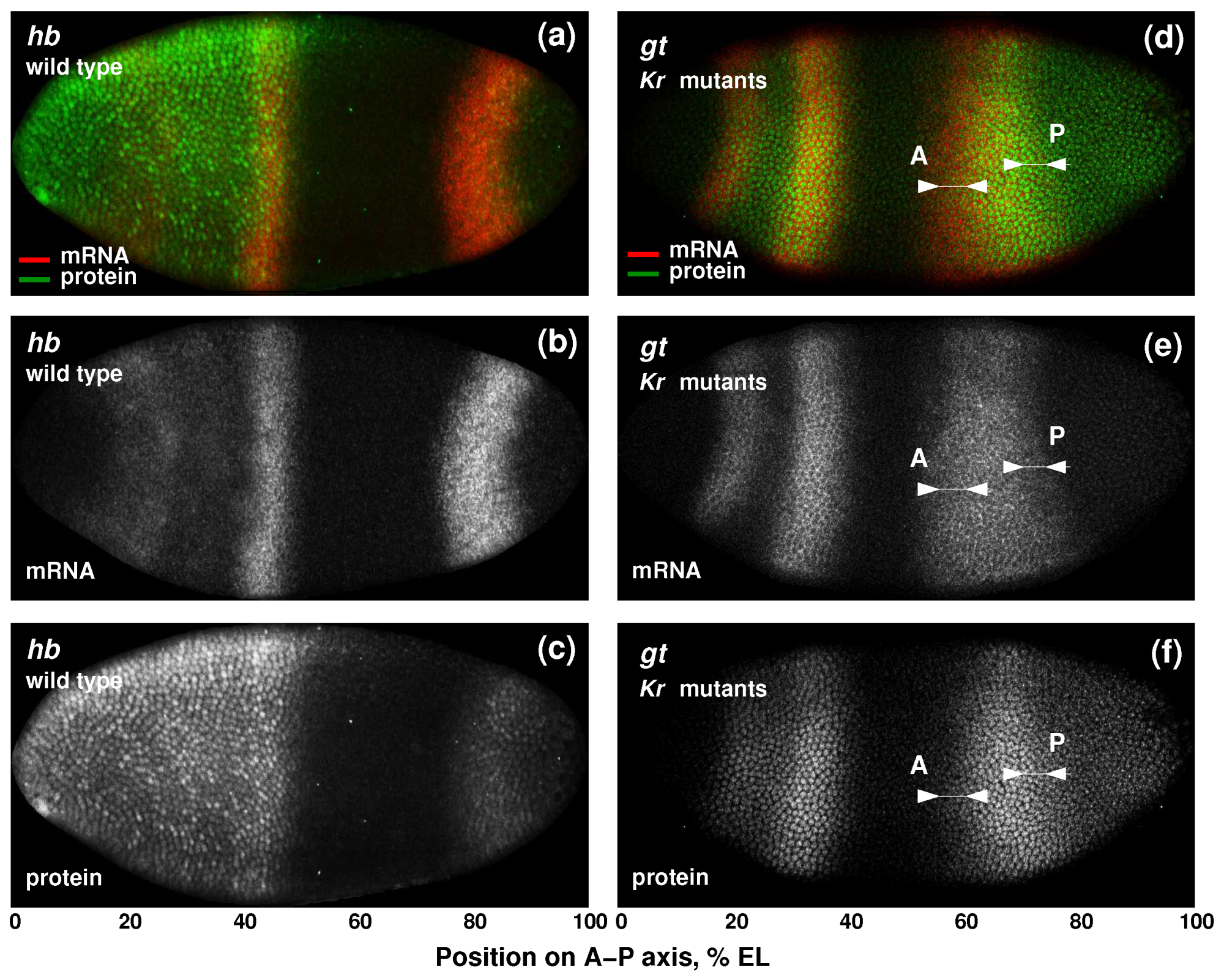
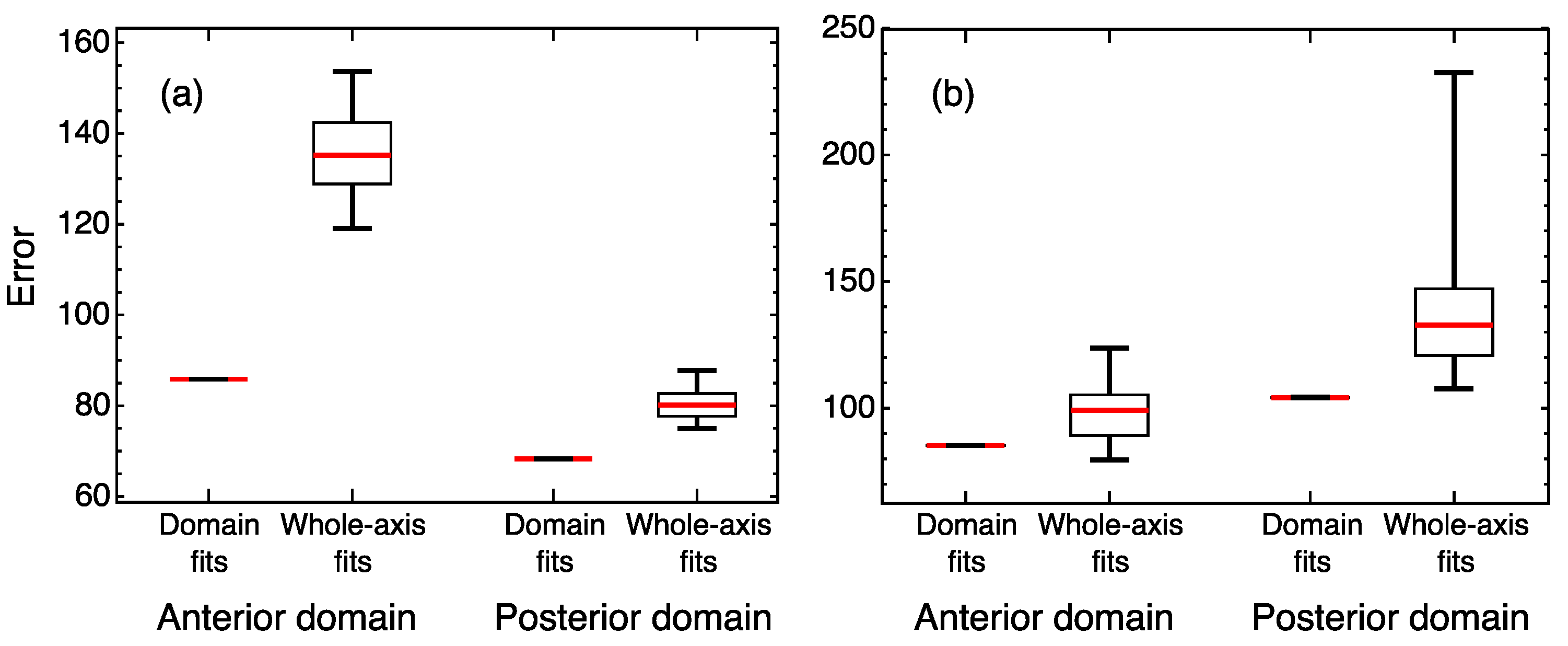
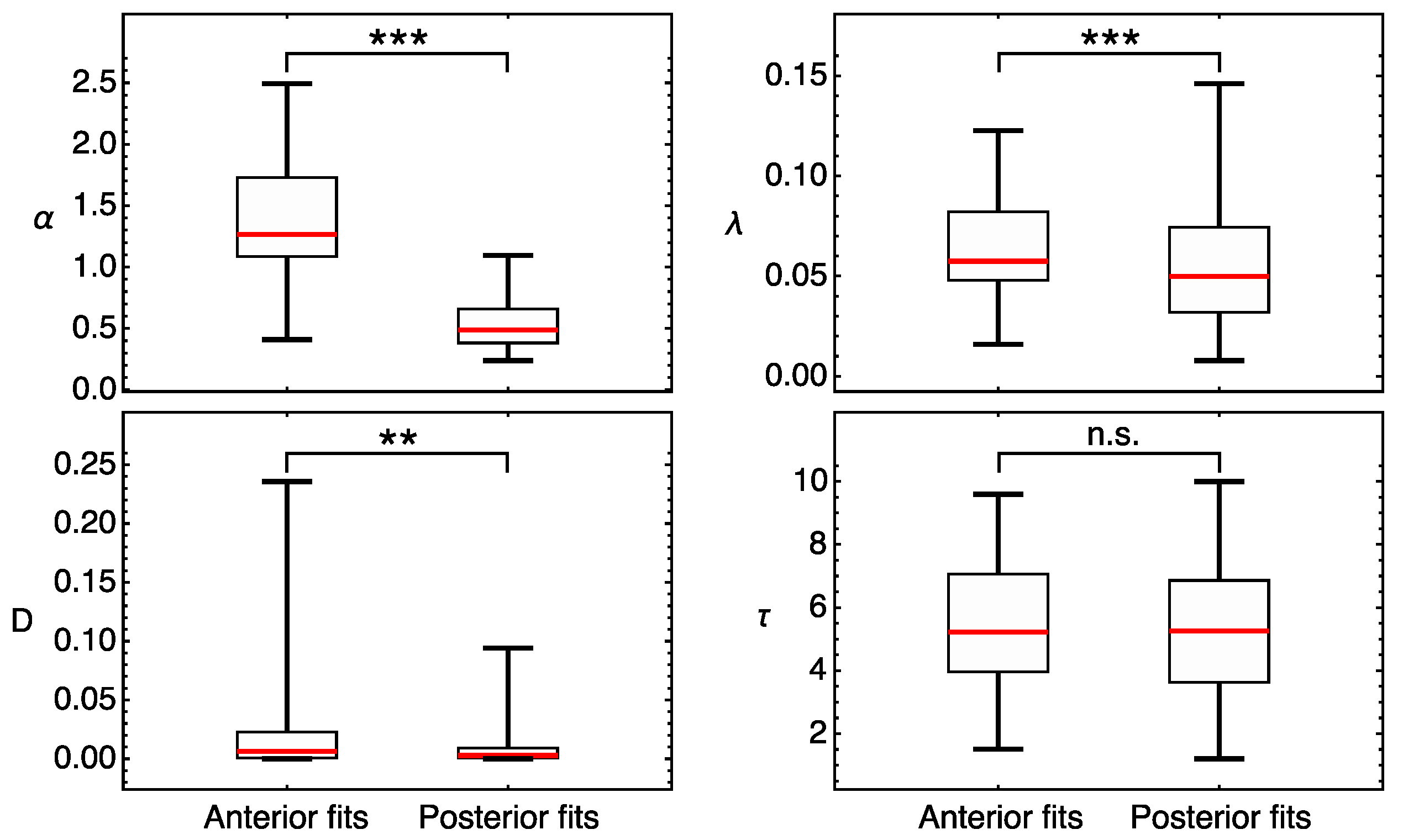
3.1. Position-Specific Models for Hunchback Expression in Wild Type Embryos
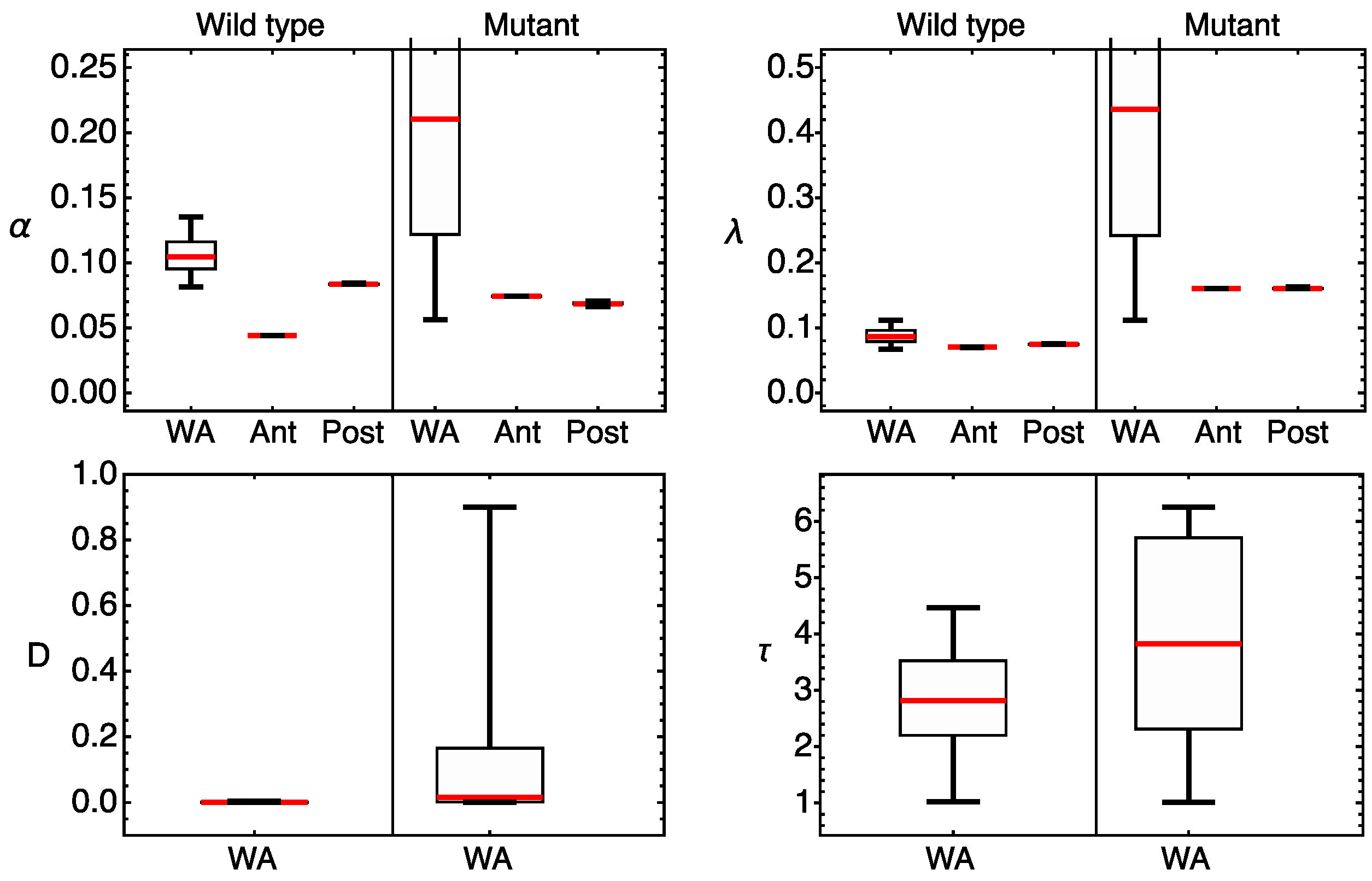
3.2. Modeling Gt Protein Dynamics in Wild-Type Embryos and Kruppel Mutants
3.3. Model Fails to Reproduce Full Dynamics of Even-Skipped Expression
4. Discussion
4.1. Model Predicts Position-Specific Regulation of Hb Protein Production
4.2. Genotype-Specific Modeling of Gt Protein Dynamics
4.3. Dynamic Sculpting of Eve Pattern: Modeling Failures and Perspectives
4.4. Limitations of the Modeling Approach
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Appendix A
References
- Schwanhäusser, B.; Busse, D.; Li, N.; Dittmar, G.; Schuchhardt, J.; Wolf, J.; Chen, W.; Selbach, M. Global quantification of mammalian gene expression control. Nature 2011, 473, 337–342. [Google Scholar] [CrossRef]
- Li, J.J.; Bickel, P.J.; Biggin, M.D. System wide analyses have underestimated protein abundances and the importance of transcription in mammals. PeerJ 2014, 2, e270. [Google Scholar] [CrossRef] [PubMed]
- Koussounadis, A.; Langdon, S.P.; Um, I.H.; Harrison, D.J.; Smith, V.A. Relationship between differentially expressed mRNA and mRNA-protein correlations in a xenograft model system. Sci. Rep. 2015, 5, 10775. [Google Scholar] [CrossRef] [PubMed]
- Cheng, Z.; Teo, G.; Krueger, S.; Rock, T.M.; Koh, H.W.; Choi, H.; Vogel, C. Differential dynamics of the mammalian mRNA and protein expression response to misfolding stress. Mol. Syst. Biol. 2016, 12, 855. [Google Scholar] [CrossRef]
- Liu, Y.; Beyer, A.; Aebersold, R. On the Dependency of Cellular Protein Levels on mRNA Abundance. Cell 2016, 165, 535–550. [Google Scholar] [CrossRef]
- Alli Shaik, A.; Wee, S.; Li, R.H.; Li, Z.; Carney, T.J.; Mathavan, S.; Gunaratne, J. Functional mapping of the zebrafish early embryo proteome and transcriptome. J. Proteome Res. 2014, 13, 5536–5550. [Google Scholar] [CrossRef] [PubMed]
- Grün, D.; Kirchner, M.; Thierfelder, N.; Stoeckius, M.; Selbach, M.; Rajewsky, N. Conservation of mRNA and protein expression during development of C. Elegans. Cell Rep. 2014, 6, 565–577. [Google Scholar] [CrossRef] [PubMed]
- Casas-Vila, N.; Bluhm, A.; Sayols, S.; Dinges, N.; Dejung, M.; Altenhein, T.; Kappei, D.; Altenhein, B.; Roignant, J.Y.; Butter, F. The developmental proteome of Drosophila Melanogaster. Genome Res. 2017, 27, 1273–1285. [Google Scholar] [CrossRef]
- Gao, Y.; Liu, X.; Tang, B.; Li, C.; Kou, Z.; Li, L.; Liu, W.; Wu, Y.; Kou, X.; Li, J.; et al. Protein Expression Landscape of Mouse Embryos during Pre-implantation Development. Cell Rep. 2017, 21, 3957–3969. [Google Scholar] [CrossRef]
- Becker, K.; Bluhm, A.; Casas-Vila, N.; Dinges, N.; Dejung, M.; Sayols, S.; Kreutz, C.; Roignant, J.Y.; Butter, F.; Legewie, S. Quantifying post-transcriptional regulation in the development of Drosophila Melanogaster. Nat. Commun. 2018, 9, 4970. [Google Scholar] [CrossRef]
- Smits, A.H.; Lindeboom, R.G.; Perino, M.; van Heeringen, S.J.; Veenstra, G.J.; Vermeulen, M. Global absolute quantification reveals tight regulation of protein expression in single Xenopus Eggs. Nucleic Acids Res. 2014, 42, 9880–9891. [Google Scholar] [CrossRef]
- Vogel, C.; Marcotte, E.M. Insights into the regulation of protein abundance from proteomic and transcriptomic analyses. Nat. Rev. Genet. 2012, 13, 227–232. [Google Scholar] [CrossRef]
- Mair, F.; Erickson, J.R.; Voillet, V.; Simoni, Y.; Bi, T.; Tyznik, A.J.; Martin, J.; Gottardo, R.; Newell, E.W.; Prlic, M. A Targeted Multi-omic Analysis Approach Measures Protein Expression and Low-Abundance Transcripts on the Single-Cell Level. Cell Rep. 2020, 31, 107499. [Google Scholar] [CrossRef]
- Akam, M. The molecular basis for metameric pattern in the Drosophila Embryo. Development 1987, 101, 1–22. [Google Scholar] [CrossRef] [PubMed]
- Jaeger, J. The gap gene network. Cell. Mol. Life Sci. CMLS 2011, 68, 243–274. [Google Scholar] [CrossRef]
- Jaeger, J.; Reinitz, J. Drosophila Blastoderm Patterning. Curr. Opin. Genet. Dev. 2012, 22, 533–541. [Google Scholar] [CrossRef] [PubMed]
- Frasch, M.; Hoey, T.; Rushlow, C.; Doyle, H.; Levine, M. Characterization and localization of the Even-Skipped Protein Drosophila. EMBO J. 1987, 6, 749–759. [Google Scholar] [CrossRef] [PubMed]
- Surkova, S.; Kosman, D.; Kozlov, K.N.; Myasnikova, E.; Samsonova, A.A.; Spirov, A.; Vanario-Alonso, C.E.; Samsonova, M.G.; Reinitz, J. Characterization of the Drosophila Segm. Determ. Morphome. Dev. Biol. 2008, 313, 844–862. [Google Scholar] [CrossRef]
- Loncar, D.; Singer, S.J. Cell membrane formation during the cellularization of the syncytial blastoderm of Drosophila. Proc. Natl. Acad. Sci. USA 1995, 92, 2199–2203. [Google Scholar] [CrossRef]
- Jaeger, J.; Surkova, S.; Blagov, M.; Janssens, H.; Kosman, D.; Kozlov, K.N.; Myasnikova, E.; Vanario-Alonso, C.E.; Samsonova, M.G.; Sharp, D.H.; et al. Dynamic control of positional information in the early Drosophila Embryo. Nature 2004, 430, 368–371. [Google Scholar] [CrossRef]
- Jaeger, J.; Blagov, M.; Kosman, D.; Kozlov, K.N.; Myasnikova, E.; Surkova, S.; Vanario-Alonso, C.E.; Samsonova, M.G.; Sharp, D.H.; Reinitz, J. Dynamical analysis of regulatory interactions in the gap gene system of Drosophila Melanogaster. Genetics 2004, 167, 1721–1737. [Google Scholar] [CrossRef] [PubMed]
- Perkins, T.J.; Jaeger, J.; Reinitz, J.; Glass, L. Reverse engineering the gap gene network of Drosophila Melanogaster. PLoS Comput. Biol. 2006, 2, e51. [Google Scholar] [CrossRef]
- Surkova, S.; Spirov, A.V.; Gursky, V.V.; Janssens, H.; Kim, A.R.; Radulescu, O.; Vanario-Alonso, C.E.; Sharp, D.H.; Samsonova, M.G.; Reinitz, J. Canalization of gene expression in the Drosophila Blastoderm Gap Gene Cross Regul. PLoS Biol. 2009, 7, e1000049. [Google Scholar] [CrossRef]
- Surkova, S.; Spirov, A.V.; Gursky, V.V.; Janssens, H.; Kim, A.R.; Radulescu, O.; Vanario-Alonso, C.E.; Sharp, D.H.; Samsonova, M.G.; Reinitz, J. Canalization of gene expression and domain shifts in the Drosophila Blastoderm by Dynamical Attractors. PLoS Comput. Biol. 2009, 5, e1000303. [Google Scholar] [CrossRef]
- Crombach, A.; Wotton, K.R.; Cicin-Sain, D.; Ashyraliyev, M.; Jaeger, J. Efficient reverse-engineering of a developmental gene regulatory network. PLoS Comput. Biol. 2012, 8, e1002589. [Google Scholar] [CrossRef] [PubMed]
- Kozlov, K.N.; Surkova, S.; Myasnikova, E.; Reinitz, J.; Samsonova, M.G. Modeling of gap gene expression in Drosophila Kruppel Mutants. PLoS Comput. Biol. 2012, 8, e1002635. [Google Scholar] [CrossRef]
- Clark, E. Dynamic patterning by the Drosophila Pair-Rule Netw. Reconciles Long-Germ Short-Germ Segmentation. PLoS Biol. 2017, 15, e2002439. [Google Scholar] [CrossRef]
- Becker, K.; Balsa-Canto, E.; Cicin-Sain, D.; Hoermann, A.; Janssens, H.; Banga, J.R.; Jaeger, J. Reverse-Engineering Post-Transcriptional Regulation of Gap Genes in Drosophila Melanogaster. PLoS Comput. Biol. 2013, 9, e1003281. [Google Scholar] [CrossRef]
- Wharton, R.P.; Struhl, G. RNA regulatory elements mediate control of Drosophila Body Pattern Posterior Morphogen Nanos. Cell 1991, 67, 955–967. [Google Scholar] [CrossRef]
- Rivera-Pomar, R.; Niessing, D.; Schmidt-Ott, U.; Gehring, W.; Jäckle, H. RNA binding and translational suppression by Bicoid. Nature 1996, 379, 746–749. [Google Scholar] [CrossRef]
- Cho, P.F.; Gamberi, C.; Cho-Park, Y.A.; Cho-Park, I.B.; Lasko, P.; Sonenberg, N. Cap-dependent translational inhibition establishes two opposing morphogen gradients in Drosophila Embryos. Curr. Biol. 2006, 16, 2035–2041. [Google Scholar] [CrossRef] [PubMed]
- Lasko, P. Posttranscriptional regulation in Drosophila Oocytes Early Embryos. Wiley Interdiscip. Rev. RNA 2011, 2, 408–416. [Google Scholar] [CrossRef]
- Surkova, S.; Sokolkova, A.; Kozlov, K.; Nuzhdin, S.V.; Samsonova, M. Quantitative analysis reveals genotype- and domain- specific differences between mRNA and protein expression of segmentation genes in Drosophila. Dev. Biol. 2019, 448, 48–58. [Google Scholar] [CrossRef] [PubMed]
- Chowdhury, T.A.; Koceja, C.; Eisa-Beygi, S.; Kleinstiver, B.P.; Kumar, S.N.; Lin, C.W.; Li, K.; Prabhudesai, S.; Joung, J.K.; Ramchandran, R. Temporal and Spatial Post-Transcriptional Regulation of Zebrafish Tie1 MRNA Long Noncoding RNA During Brain Vascular Assembly. Arter. Thromb Vasc Biol. 2018, 38, 1562–1575. [Google Scholar] [CrossRef]
- Maier, T.; Schmidt, A.; Güell, M.; Kühner, S.; Gavin, A.C.; Aebersold, R.; Serrano, L. Quantification of mRNA and protein and integration with protein turnover in a bacterium. Mol. Syst. Biol. 2011, 7, 511. [Google Scholar] [CrossRef]
- Jovanovic, M.; Rooney, M.; Mertins, P.; Przybylski, D.; Chevrier, N.; Satija, R.; Rodriguez, E.H.; Fields, A.P.; Schwartz, S.; Raychowdhury, R.; et al. Dynamic profiling of the protein life cycle in response to pathogens. Science 2015, 347, 1259038. [Google Scholar] [CrossRef]
- Eldon, E.D.; Pirrotta, V. Interactions of the Drosophila gap gene giant with maternal and zygotic pattern-forming genes. Development 1991, 111, 367–378. [Google Scholar] [CrossRef] [PubMed]
- Kraut, R.; Levine, M. Spatial regulation of the gap gene Giant Drosophila development. Development 1991, 111, 601–609. [Google Scholar] [CrossRef] [PubMed]
- Surkova, S.; Golubkova, E.; Panok, L.; Mamon, L.; Reinitz, J.; Samsonova, M. Quantitative dynamics and increased variability of segmentation gene expression in the Drosophila Krüppel Knirps Mutants. Dev. Biol. 2013, 376, 99–112. [Google Scholar] [CrossRef]
- Sonoda, J.; Wharton, R. Recruitment of Nanos to Hunchback MRNA Pumilio. Genes Dev. 1999, 13, 2704–2712. [Google Scholar] [CrossRef]
- Chagnovich, D.; Lehmann, R. Poly(A)-independent regulation of maternal hunchback translation in the Drosophila Embryo. Proc. Natl. Acad. Sci. USA 2001, 98, 11359–11364. [Google Scholar] [CrossRef]
- Vinter, D.J.; Hoppe, C.; Minchington, T.G.; Sutcliffe, C.; Ashe, H.L. Dynamics of hunchback translation in real-time and at single-mRNA resolution in the Drosophila Embryo. Development 2021, 148, dev196121. [Google Scholar] [CrossRef] [PubMed]
- Wieschaus, E.; Nüsslein-Volhard, C.; Kluding, H. Krüppel, a gene whose activity is required early in the zygotic genome for normal embryonic segmentation. Dev. Biol. 1984, 104, 172–186. [Google Scholar] [CrossRef]
- Choi, H.M.; Beck, V.A.; Pierce, N.A. Next-generation in situ hybridization chain reaction: Higher gain, lower cost, greater durability. ACS Nano 2014, 8, 4284–4294. [Google Scholar] [CrossRef]
- Nagaso, H.; Murata, T.; Day, N.; Yokoyama, K.K. Simultaneous detection of RNA and protein by in situ hybridization and immunological staining. J. Histochem. Cytochem. 2001, 49, 1177–1182. [Google Scholar] [CrossRef] [PubMed]
- Surkova, S.; Myasnikova, E.; Janssens, H.; Kozlov, K.N.; Samsonova, A.A.; Reinitz, J.; Samsonova, M. Pipeline for acquisition of quantitative data on segmentation gene expression from confocal images. Fly 2008, 2, 58–66. [Google Scholar] [CrossRef]
- Wu, X.; Vasisht, V.; Kosman, D.; Reinitz, J.; Small, S. Thoracic patterning by the Drosophila Gap Gene Hunchback. Dev. Biol. 2001, 237, 79–92. [Google Scholar] [CrossRef]
- Perry, M.W.; Bothma, J.P.; Luu, R.D.; Levine, M. Precision of Hunchback Expression in the Drosophila Embryo. Curr. Biol. 2012, 22, 2247–2252. [Google Scholar] [CrossRef]
- Mohler, J.; Eldon, E.D.; Pirrotta, V. A novel spatial transcription pattern associated with the segmentation gene, giant, of Drosophila. EMBO J. 1989, 8, 1539–1548. [Google Scholar] [CrossRef]
- Janssens, H.; Hou, S.; Jaeger, J.; Kim, A.R.; Myasnikova, E.; Sharp, D.; Reinitz, J. Quantitative and predictive model of transcriptional control of the Drosophila melanogaster even skipped gene. Nat. Genet. 2006, 38, 1159–1165. [Google Scholar] [CrossRef]
- Liu, Y.; Barr, K.; Reinitz, J. Fully interpretable deep learning model of transcriptional control. Bioinformatics 2020, 36 (Suppl. 1), i499–i507. [Google Scholar] [CrossRef] [PubMed]
- Lasko, P. mRNA localization and translational control in Drosophila Oogenesis. Cold Spring Harb. Perspect. Biol. 2012, 4, a012294. [Google Scholar] [CrossRef] [PubMed]
- McKee, A.E.; Minet, E.; Stern, C.; Riahi, S.; Stiles, C.D.; Silver, P.A. A genome-wide in situ hybridization map of RNA-binding proteins reveals anatomically restricted expression in the developing mouse brain. BMC Dev. Biol. 2005, 5, 14. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Forinash, K.D.; McGivern, J.; Fritz, B.; Dorey, K.; Sheets, M.D. Spatially restricted translation of the xCR1 mRNA in Xenopus Embryos. Mol. Cell. Biol. 2009, 29, 3791–3802. [Google Scholar] [CrossRef]
- Kaschula, R.; Pinho, S.; Alonso, C.R. MicroRNA-dependent regulation of Hox gene expression sculpts fine-grain morphological patterns in a Drosophila Appendage. Development 2018, 145, dev161133. [Google Scholar] [CrossRef]
- Chua, B.A.; Van Der Werf, I.; Jamieson, C.; Signer, R. Post-Transcriptional Regulation of Homeostatic, Stressed, and Malignant Stem Cells. Cell Stem Cell 2020, 26, 138–159. [Google Scholar] [CrossRef] [PubMed]
- Pultz, M.A.; Westendorf, L.; Gale, S.D.; Hawkins, K.; Lynch, J.; Pitt, J.N.; Reeves, N.L.; Yao, J.C.; Small, S.; Desplan, C.; et al. A major role for zygotic Hunchback Patterning Nasonia Embryo. Development 2005, 132, 3705–3715. [Google Scholar] [CrossRef]
- Perry, M.W.; Boettiger, A.N.; Levine, M. Multiple enhancers ensure precision of gap gene-expression patterns in the Drosophila Embryo. Proc. Natl. Acad. Sci. USA 2011, 108, 13570–13575. [Google Scholar] [CrossRef] [PubMed]
- Menon, K.P.; Andrews, S.; Murthy, M.; Gavis, E.R.; Zinn, K. The translational repressors Nanos and Pumilio have divergent effects on presynaptic terminal growth and postsynaptic glutamate receptor subunit composition. J. Neurosci. 2009, 29, 5558–5572. [Google Scholar] [CrossRef]
- Miles, W.O.; Korenjak, M.; Griffiths, L.M.; Dyer, M.A.; Provero, P.; Dyson, N.J. Post-transcriptional gene expression control by NANOS is up-regulated and functionally important in pRb-deficient cells. EMBO J. 2014, 33, 2201–2215. [Google Scholar] [CrossRef]
- Olesnicky, E.C.; Bhogal, B.; Gavis, E.R. Combinatorial use of translational co-factors for cell type-specific regulation during neuronal morphogenesis in Drosophila. Dev. Biol. 2012, 365, 208–218. [Google Scholar] [CrossRef]
- Tanenbaum, M.E.; Gilbert, L.A.; Qi, L.S.; Weissman, J.S.; Vale, R.D. A protein-tagging system for signal amplification in gene expression and fluorescence imaging. Cell 2014, 159, 635–646. [Google Scholar] [CrossRef] [PubMed]
- Pichon, X.; Bastide, A.; Safieddine, A.; Chouaib, R.; Samacoits, A.; Basyuk, E.; Peter, M.; Mueller, F.; Bertrand, E. Visualization of single endogenous polysomes reveals the dynamics of translation in live human cells. J. Cell Biol. 2016, 214, 769–781. [Google Scholar] [CrossRef]
- Dufourt, J.; Bellec, M.; Trullo, A.; Dejean, M.; De Rossi, S.; Favard, C.; Lagha, M. Imaging translation dynamics in live embryos reveals spatial heterogeneities. Science 2021, 372, 840–844. [Google Scholar] [CrossRef]
- Berrocal, A.; Lammers, N.C.; Garcia, H.G.; Eisen, M.B. Kinetic sculpting of the seven stripes of the Drosophila even-skipped gene. eLife 2020, 9, e61635. [Google Scholar] [CrossRef] [PubMed]
- Buchler, N.E.; Gerland, U.; Hwa, T. Nonlinear protein degradation and the function of genetic circuits. Proc. Natl. Acad. Sci. USA 2005, 102, 9559–9564. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Hansen, D.; Eck, E.; Kim, Y.J.; Turner, M.; Alamos, S.; Garcia, H. Real-time single-cell characterization of the eukaryotic transcription cycle reveals correlations between RNA initiation, elongation, and cleavage. PLoS Comput. Biol. 2021, 17, e1008999. [Google Scholar] [CrossRef]



Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Duk, M.A.; Gursky, V.V.; Samsonova, M.G.; Surkova, S.Y. Application of Domain- and Genotype-Specific Models to Infer Post-Transcriptional Regulation of Segmentation Gene Expression in Drosophila. Life 2021, 11, 1232. https://doi.org/10.3390/life11111232
Duk MA, Gursky VV, Samsonova MG, Surkova SY. Application of Domain- and Genotype-Specific Models to Infer Post-Transcriptional Regulation of Segmentation Gene Expression in Drosophila. Life. 2021; 11(11):1232. https://doi.org/10.3390/life11111232
Chicago/Turabian StyleDuk, Maria A., Vitaly V. Gursky, Maria G. Samsonova, and Svetlana Yu. Surkova. 2021. "Application of Domain- and Genotype-Specific Models to Infer Post-Transcriptional Regulation of Segmentation Gene Expression in Drosophila" Life 11, no. 11: 1232. https://doi.org/10.3390/life11111232
APA StyleDuk, M. A., Gursky, V. V., Samsonova, M. G., & Surkova, S. Y. (2021). Application of Domain- and Genotype-Specific Models to Infer Post-Transcriptional Regulation of Segmentation Gene Expression in Drosophila. Life, 11(11), 1232. https://doi.org/10.3390/life11111232






