Development of a Simple Kinetic Mathematical Model of Aggregation of Particles or Clustering of Receptors
Abstract
:1. Introduction
2. Modeling Approaches
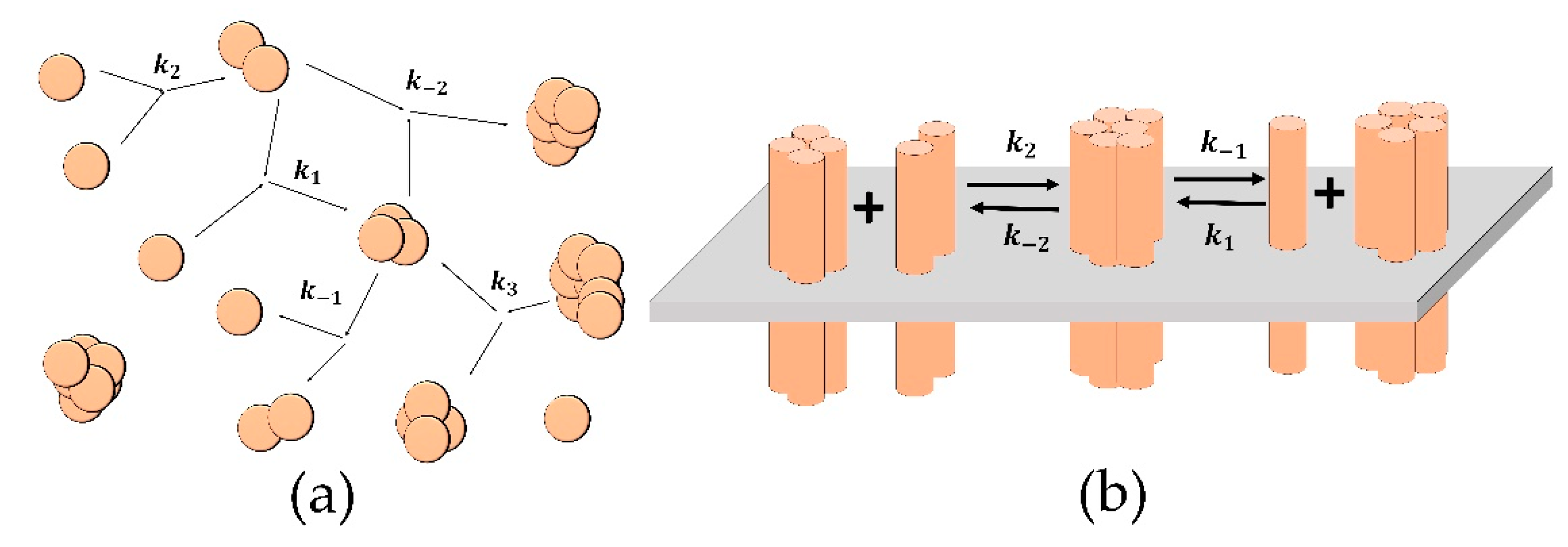
2.1. Aggregation Model
2.1.1. Model Equations
- The mechanism of fragmentation is assumed to be independent of the aggregate size.
2.1.2. Model Parameters
2.1.3. Platelet Aggregation Experiments
2.2. Clustering Model
2.2.1. Model Equations
2.2.2. Model Parameters
2.3. “2-Equation” Model
2.4. Methods for Parameter Estimation, Model Solution, and Comparison of the Models
3. Results
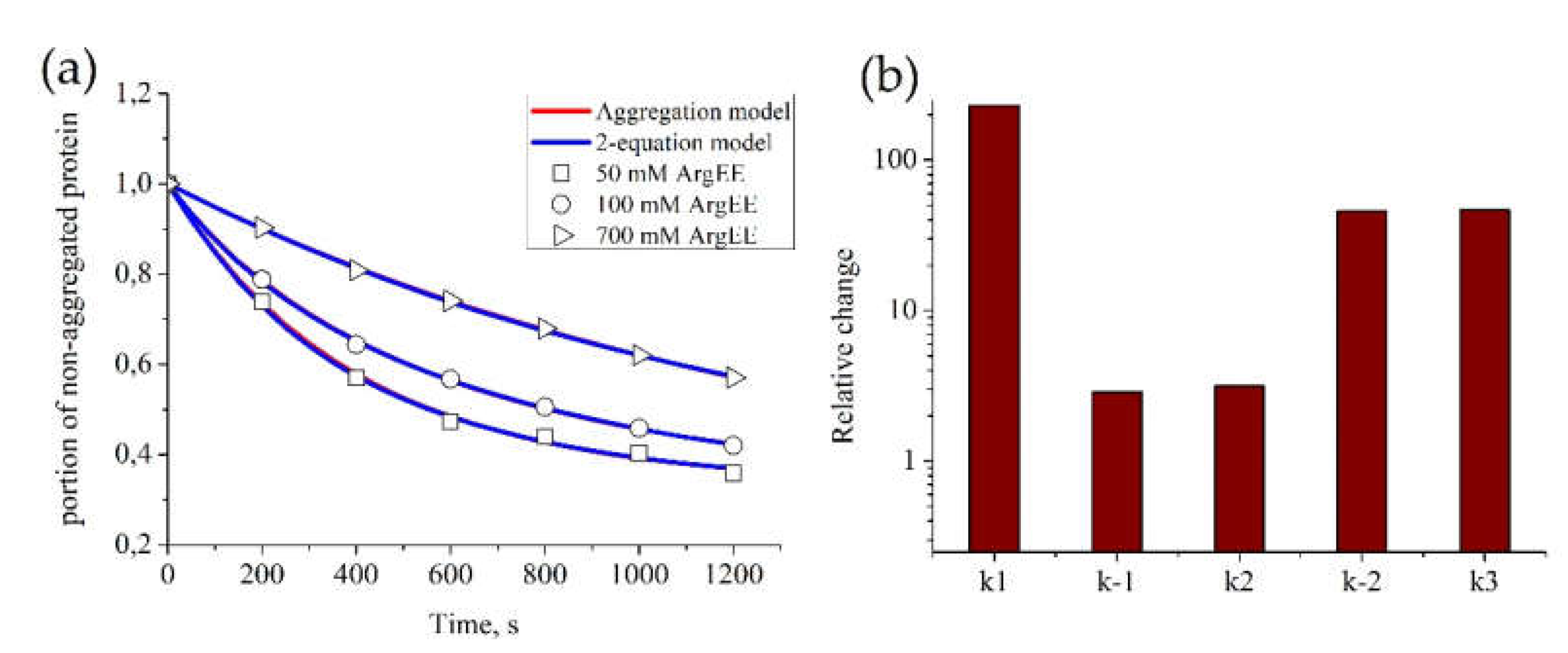
3.1. Description of Protein Aggregation Data
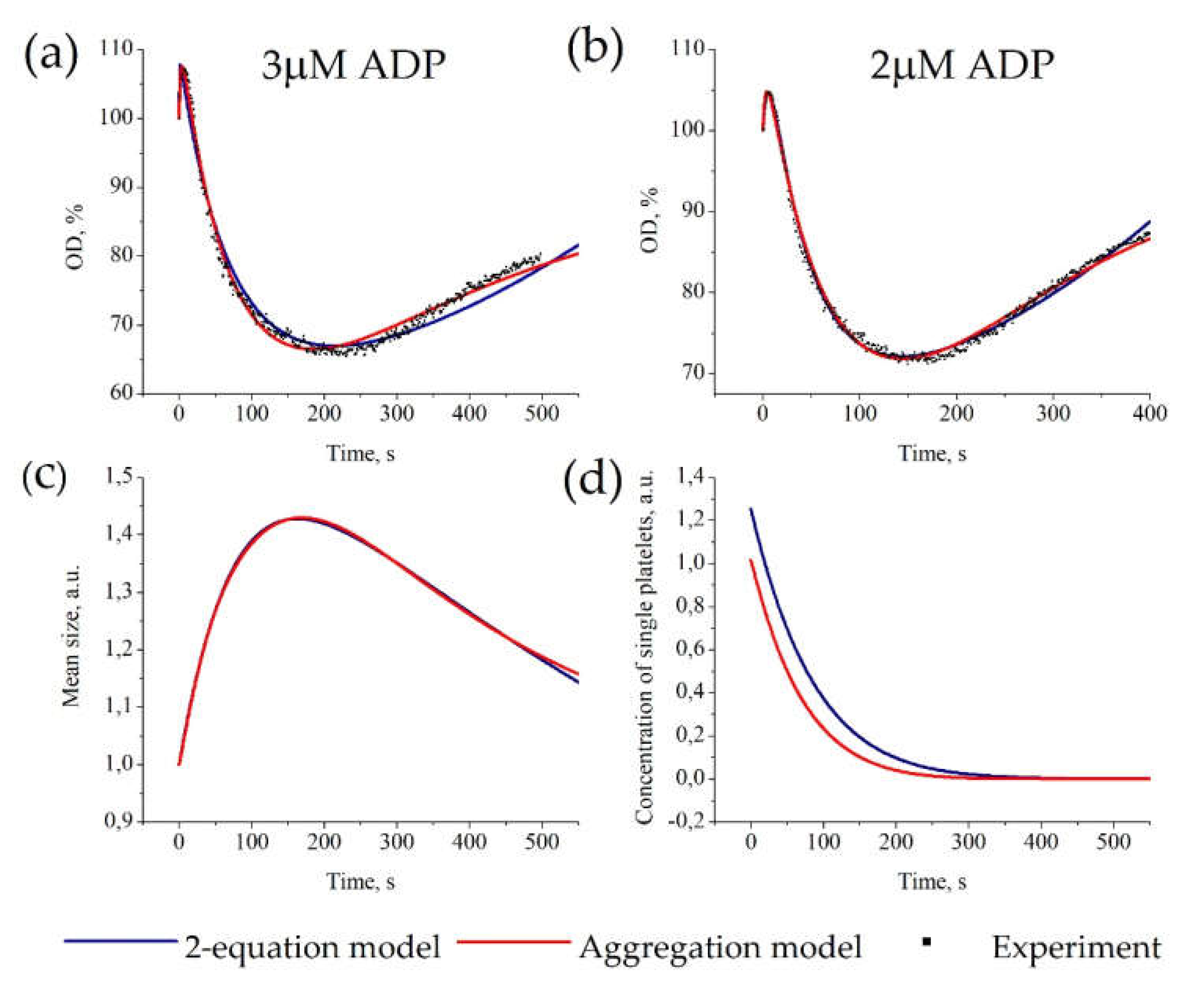
3.2. Description of Platelet Aggregation Data
3.3. Additional Restrictions on Parameter Values for “Aggregation Model”
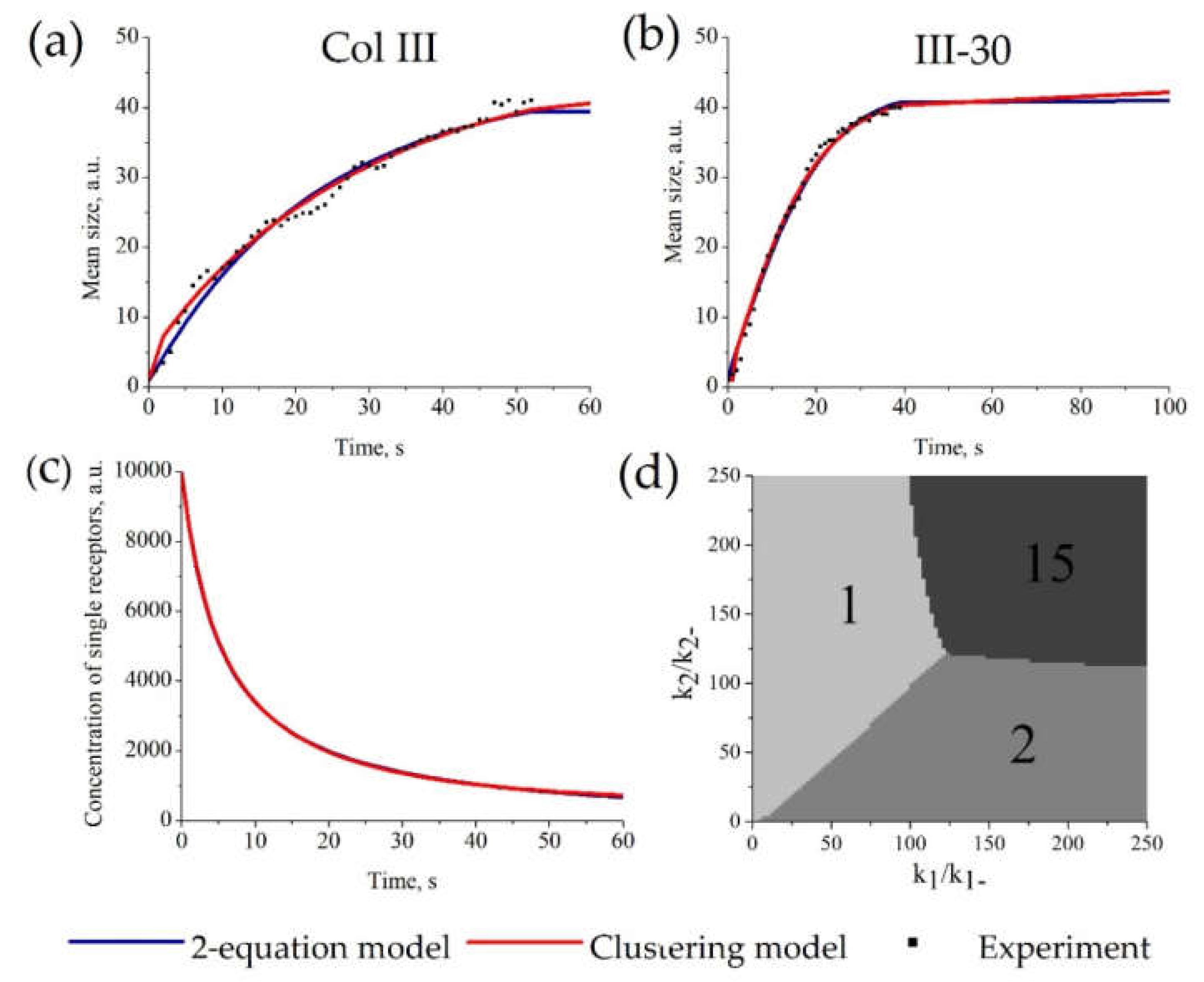
3.4. Description of Receptor Clustering Data
3.5. Additional Restrictions on Parameter Values for “Clustering Model”
3.6. Features of Receptor Clustering Process Revealed by the Mathematical Models
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Alarcon, B.; Lythe, G.; Ferez, M.; Molina-Paris, C.; Castro, M.; van Santen, H.M. Receptor Pre-Clustering and T cell Responses: Insights into Molecular Mechanisms. Front. Immunol. 2014, 5, 1–11. [Google Scholar]
- Detmers, P.A.; Wright, S.D.; Olsen, E.; Kimball, B.; Cohn, Z.A. Aggregation of complement receptors on human neutrophils in the absence of ligand. J. Cell Biol. 1987, 105, 1137–1145. [Google Scholar] [CrossRef] [Green Version]
- Head, B.P.; Patel, H.H.; Insel, P.A. Interaction of membrane/lipid rafts with the cytoskeleton: Impact on signaling and function: Membrane/lipid rafts, mediators of cytoskeletal arrangement and cell signaling. Biochim. Biophys. Acta Biomembr. 2014, 1838, 532–545. [Google Scholar] [CrossRef] [Green Version]
- Shen, Z.; Liu, S.; Li, X.; Wan, Z.; Mao, Y.; Chen, C.; Liu, W. Conformational change within the extracellular domain of B cell receptor in B cell activation upon antigen binding. eLife 2019, 8, e42271. [Google Scholar] [CrossRef]
- Li, L.; Hu, J.; Shi, X.; Shao, Y.; Song, F. Lipid rafts enhance the binding constant of membrane-anchored receptors and ligands. Soft Matter 2017, 13, 4294–4304. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bray, D.; Levin, M.D.; Morton-Firth, C.J. Receptor clustering as a cellular mechanism to control sensitivity. Nature 1998, 393, 85–88. [Google Scholar] [CrossRef] [PubMed]
- Hartman, N.C.; Groves, J.T. Signaling clusters in the cell membrane. Curr. Opin. Cell Biol. 2011, 23, 370–376. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tian, T.; Harding, A.; Inder, K.; Plowman, S.; Parton, R.G.; Hancock, J.F. Plasma membrane nanoswitches generate high-fidelity Ras signal transduction. Nat. Cell Biol. 2007, 9, 905–914. [Google Scholar] [CrossRef]
- Welf, E.S.; Naik, U.P.; Ogunnaike, B.A. A Spatial Model for Integrin Clustering as a Result of Feedback between Integrin Activation and Integrin Binding. Biophys. J. 2012, 103, 1379–1389. [Google Scholar] [CrossRef] [Green Version]
- Caré, B.R.; Soula, H.A. Impact of receptor clustering on ligand binding. BMC Syst. Biol. 2011, 5, 1–13. [Google Scholar] [CrossRef] [Green Version]
- Gopalakrishnan, M.; Forsten-Williams, K.; Nugent, M.A.; Täuber, U.C. Effects of receptor clustering on ligand dissociation kinetics: Theory and simulations. Biophys. J. 2005, 89, 3686–3700. [Google Scholar] [CrossRef] [Green Version]
- Mugler, A.; Bailey, A.G.; Takahashi, K.; ten Wolde, P.R. Membrane clustering and the role of rebinding in biochemical signaling. Biophys. J. 2012, 102, 1069–1078. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shea, L.D.; Linderman, J.J. Compartmentalization of Receptors and Enzymes Affects Activation for a Collision Coupling Mechanism. J. Theor. Biol. 1998, 191, 249–258. [Google Scholar] [CrossRef]
- Fallahi-Sichani, M.; Linderman, J.J. Lipid Raft-Mediated Regulation of G-Protein Coupled Receptor Signaling by Ligands which Influence Receptor Dimerization: A Computational Study. PLoS ONE 2009, 4, e6604. [Google Scholar] [CrossRef] [PubMed]
- Gardina, P.J.; Manson, M.D. Attractant Signaling by an Aspartate Chemoreceptor Dimer with a Single Cytoplasmic Domain. Science 1996, 274, 425–426. [Google Scholar] [CrossRef]
- Mello, B.A.; Shaw, L.; Tu, Y. Effects of receptor interaction in bacterial chemotaxis. Biophys. J. 2004, 87, 1578–1595. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- WANANT, S.; QUON, M.J. Insulin Receptor Binding Kinetics: Modeling and Simulation Studies. J. Theor. Biol. 2000, 205, 355–364. [Google Scholar] [CrossRef]
- Kong, S.-M.; Liu, H.; Xue, Y.-H.; Liu, X.-L.; Jia, X.-X.; Cui, F.-C. Polymerization-induced polymer aggregation or polymer aggregation-enhanced polymerization? A computer simulation study. Phys. Chem. Chem. Phys. 2018, 20, 24379–24388. [Google Scholar] [CrossRef] [PubMed]
- May, R.C. The Arp2/3 complex: A central regulator of the actin cytoskeleton. Cell. Mol. Life Sci. CMLS 2001, 58, 1607–1626. [Google Scholar] [CrossRef]
- Choquet, D.; Triller, A. The role of receptor diffusion in the organization of the postsynaptic membrane. Nat. Rev. Neurosci. 2003, 4, 251–265. [Google Scholar] [CrossRef]
- Kholodenko, B.N.; Hoek, J.B.; Westerhoff, H.V. Why cytoplasmic signalling proteins should be recruited to cell membranes. Trends Cell Biol. 2000, 10, 173–178. [Google Scholar] [CrossRef]
- Arosio, P.; Rima, S.; Lattuada, M.; Morbidelli, M. Population Balance Modeling of Antibodies Aggregation Kinetics. J. Phys. Chem. B 2012, 116, 7066–7075. [Google Scholar] [CrossRef] [PubMed]
- Fornari, S.; Schäfer, A.; Kuhl, E.; Goriely, A. Spatially-extended nucleation-aggregation-fragmentation models for the dynamics of prion-like neurodegenerative protein-spreading in the brain and its connectome. J. Theor. Biol. 2019, 486, 110102. [Google Scholar] [CrossRef] [PubMed]
- Kier, L.B.; Cheng, C.-K.; Nelson, J.D. Models of Solute Aggregation Using Cellular Automata. Chem. Biodivers. 2009, 6, 396–401. [Google Scholar] [CrossRef] [PubMed]
- Lahiri, S.; Wang, Y.; Esposito, M.; Lacoste, D. Kinetics and thermodynamics of reversible polymerization in closed systems. New J. Phys. 2015, 17, 085008. [Google Scholar] [CrossRef] [Green Version]
- Przybycien, T.M.; Bailey, J.E. Aggregation kinetics in salt-induced protein precipitation. AIChE J. 1989, 35, 1779–1790. [Google Scholar] [CrossRef]
- Richardson, G.; Cummings, L.J.; Harris, H.J.; O’Shea, P. Toward a mathematical model of the assembly and disassembly of membrane microdomains: Comparison with experimental models. Biophys. J. 2007, 92, 4145–4156. [Google Scholar] [CrossRef] [Green Version]
- Zhu, H.; Pang, P.Y.H.; Sun, Y.; Dhar, P. Asynchronous adaptive time step in quantitative cellular automata modeling. BMC Bioinform. 2004, 5, 1–7. [Google Scholar] [CrossRef] [Green Version]
- Mukhopadhyay, D.; De, R. Aggregation dynamics of active cells on non-adhesive substrate. Phys. Biol. 2019, 16, 046006. [Google Scholar] [CrossRef] [Green Version]
- Stilck, J. Lattice models for confined polymers. Braz. J. Phys. 1998, 28. [Google Scholar] [CrossRef]
- Wang, H.; Wingreen, N.S.; Mukhopadhyay, R. Self-organized periodicity of protein clusters in growing bacteria. Phys. Rev. Lett. 2008, 101, 1–4. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zidar, M.; Kuzman, D.; Ravnik, M. Characterisation of protein aggregation with the Smoluchowski coagulation approach for use in biopharmaceuticals. Soft Matter 2018, 14, 6001–6012. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Filkova, A.A.; Martyanov, A.A.; Garzon Dasgupta, A.K.; Panteleev, M.A.; Sveshnikova, A.N. Quantitative dynamics of reversible platelet aggregation: Mathematical modelling and experiments. Sci. Rep. 2019, 9, 1–9. [Google Scholar] [CrossRef]
- Smoluchowski, M. Attempt for a mathematical theory of kinetic coagulation of colloid solutions. Z. Phys. Chem. 1917, 92, 129–168. [Google Scholar]
- Kinney, M.A.; Sargent, C.Y.; McDevitt, T.C. The multiparametric effects of hydrodynamic environments on stem cell culture. Tissue Eng. Part. B Rev. 2011, 17, 249–262. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Matthews, L.S.; Shotorban, B.; Hyde, T.W. Cosmic Dust Aggregation with Stochastic Charging. Astrophys. J. 2013, 776, 103. [Google Scholar] [CrossRef] [Green Version]
- Philo, J.; Arakawa, T. Mechanisms of Protein Aggregation. Curr. Pharm. Biotechnol. 2009, 10, 348–351. [Google Scholar] [CrossRef]
- Borzova, V.A.; Markossian, K.A.; Kleymenov, S.Y.; Kurganov, B.I. A change in the aggregation pathway of bovine serum albumin in the presence of arginine and its derivatives. Sci. Rep. 2017, 7, 1–12. [Google Scholar] [CrossRef]
- Koltai, K.; Kesmarky, G.; Feher, G.; Tibold, A.; Toth, K. Platelet aggregometry testing: Molecular mechanisms, techniques and clinical implications. Int. J. Mol. Sci. 2017, 18, 1803. [Google Scholar] [CrossRef]
- Filkova, A.; Garzon Dasgupta, A.K.; Sveshnikova, A. The reversible platelet aggregation is caused by an inhibition of iPLA2-dependent TXA2 synthesis in the presence of extracellular calcium. In FEBS Open Bio, Proceedings of 44th FEBS Congress Krakow, Poland, 6–11 July 2019; Wiley: Hoboken, NJ, USA, 2019; Volume 9, p. 133. [Google Scholar]
- Garzon Dasgupta, A.K.; Filkova, A.; Panteleev, M.; Sveshnikova, A. Strong platelet activation in response to ADP in large platelet aggregates. In Proceedings of the ECTH 2019 Abstract Book, Glasgow, UK, 2–4 October 2019; MCI Group: Geneva, Switzerland, 2019; Volume 1, p. 259. [Google Scholar]
- Beer Bestimmung der Absorption des rothen Lichts in farbigen Flüssigkeiten. Ann. Phys. Chem. 1852, 162, 78–88. [CrossRef] [Green Version]
- Feltham, D.L.; Garside, J. A mathematical model of crystallization in an emulsion. J. Chem. Phys. 2005, 122, 174910. [Google Scholar] [CrossRef] [PubMed]
- Ma, Y.; Pandzic, E.; Nicovich, P.R.; Yamamoto, Y.; Kwiatek, J.; Pageon, S.V.; Benda, A.; Rossy, J.; Gaus, K. An intermolecular FRET sensor detects the dynamics of T cell receptor clustering. Nat. Commun. 2017, 8, 1–11. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Morrison, I.E.; Anderson, C.M.; Georgiou, G.N.; Stevenson, G.V.; Cherry, R.J. Analysis of receptor clustering on cell surfaces by imaging fluorescent particles. Biophys. J. 1994, 67, 1280–1290. [Google Scholar] [CrossRef] [Green Version]
- Bene, L.; Fulwyler, M.J.; Damjanovich, S. Detection of receptor clustering by flow cytometric fluorescence anisotropy measurements. Cytom. J. Int. Soc. Anal. Cytol. 2000, 40, 292–306. [Google Scholar] [CrossRef]
- Medda, R.; Giske, A.; Cavalcanti-Adam, E.A. Challenges in imaging cell surface receptor clusters. Opt. Lasers Eng. 2016, 76, 3–8. [Google Scholar] [CrossRef]
- Zhang, S.; Reinhard, B.M. Characterizing Large-Scale Receptor Clustering on the Single Cell Level: A Comparative Plasmon Coupling and Fluorescence Superresolution Microscopy Study. J. Phys. Chem. B 2019, 123, 5494–5505. [Google Scholar] [CrossRef] [PubMed]
- Siljander, P.R.-M.; Munnix, I.C.A.; Smethurst, P.A.; Deckmyn, H.; Lindhout, T.; Ouwehand, W.H.; Farndale, R.W.; Heemskerk, J.W.M. Platelet receptor interplay regulates collagen-induced thrombus formation in flowing human blood. Blood 2004, 103, 1333–1341. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Heemskerk, J.W.M.; Kuijpers, M.J.E.; Munnix, I.C.A.; Siljander, P.R.M. Platelet Collagen Receptors and Coagulation. A Characteristic Platelet Response as Possible Target for Antithrombotic Treatment. Trends Cardiovasc. Med. 2005, 15, 86–92. [Google Scholar] [CrossRef] [PubMed]
- Poulter, N.S.; Pollitt, A.Y.; Owen, D.M.; Gardiner, E.E.; Andrews, R.K.; Shimizu, H.; Ishikawa, D.; Bihan, D.; Farndale, R.W.; Moroi, M.; et al. Clustering of glycoprotein VI (GPVI) dimers upon adhesion to collagen as a mechanism to regulate GPVI signaling in platelets. J. Thromb. Haemost. 2017, 15, 549–564. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Petzold, L. Automatic Selection of Methods for Solving Stiff and Nonstiff Systems of Ordinary Differential Equations. SIAM J. Sci. Stat. Comput. 1983, 4, 136–148. [Google Scholar] [CrossRef]
- Petzold, L.; Hindmarsh, A. LSODA (Livermore Solver of Ordinary Differential Equations); Computing and Mathematics Research Division, Lawrence Livermore National Laboratory: Livermore, CA, USA, 1997; Volume 24. [Google Scholar]
- Kennedy, J.; Eberhart, R. Particle Swarm Optimization. In Neural Networks, Proceedings of ICNN’s—International Conference on Neural Networks, Perth, WA, Australia, 27 November–1 December 1995; Institute of Electrical and Electronics Engineers: Perth, WA, Australia, 1995; Volume 4, pp. 1942–1948. ISBN 0-7803-2768-3. [Google Scholar]
- Hooke, R.; Jeeves, T.A. “Direct search” solution of numerical and statistical problems. J. Assoc. Comput. Mach. 1961, 8, 212–229. [Google Scholar] [CrossRef]
- Marquardt, D.W. An algorithm for least squares estimation of nonlinear parameters. SIAM J. 1963, 11, 431–441. [Google Scholar] [CrossRef]
- Strawderman, R. Model. Selection and Inference: A Practical Information-Theoretic Approach; Burnham, K.P., Anderson, D.R., Eds.; Springer: Berlin/Heidelberg, Germany, 2000; Volume 95. [Google Scholar]
- Stokes, G.G. On the Effect of Internal Friction of Fluids on the Motion of Pendulums; Transactions of the Cambridge Philosophical Society: Cambridge, UK, 1851; Volume 9, pp. 8–106. [Google Scholar]
- Abbott, A.J.; Nelsestuen, G.L. The collisional limit: An important consideration for membrane-associated enzymes and receptors. FASEB J. 1988, 2, 2858–2866. [Google Scholar] [CrossRef] [PubMed]
- Steinfeld, J.I.; Francisco, J.S.; Hase, W.L. Chemical Kinetics and Dynamics; Prentice Hall: Englewood Cliffs, NJ, USA, 1989; Volume 3. [Google Scholar]
- Ghosh, P.; Vaidya, A.; Kumar, A.; Rangachari, V. Determination of critical nucleation number for a single nucleation amyloid-β aggregation model. Math. Biosci. 2016, 273, 70–79. [Google Scholar] [CrossRef] [Green Version]
- Guy, R.D.; Fogelson, A.L. Probabilistic Modeling of Platelet Aggregation: Effects of Activation Time and Receptor Occupancy. J. Theor. Biol. 2002, 219, 33–53. [Google Scholar] [CrossRef]
- Leiderman, K.; Fogelson, A.L. The influence of hindered transport on the development of platelet thrombi under flow. Bull. Math. Biol. 2013, 75, 1255–1283. [Google Scholar] [CrossRef] [PubMed]
- Burroughs, N.J.; Lazic, Z.; van der Merwe, P.A. Ligand detection and discrimination by spatial relocalization: A kinase-phosphatase segregation model of TCR activation. Biophys. J. 2006, 91, 1619–1629. [Google Scholar] [CrossRef] [Green Version]
- Martyanov, A.; Balabin, F.A.; Dunster, J.L.; Panteleev, M.A.; Gibbins, J.; Sveshnikova, A.N. Diffusional and chemical control in the tyrosine kinase network of platelet CLEC-2 signalling. bioRxiv 2019. bioRxiv:529859. [Google Scholar]
- Sveshnikova, A.N.; Balatskiy, A.V.; Demianova, A.S.; Shepelyuk, T.O.; Shakhidzhanov, S.S.; Balatskaya, M.N.; Pichugin, A.V.; Ataullakhanov, F.I.; Panteleev, M.A. Systems biology insights into the meaning of the platelet’s dual-receptor thrombin signaling. J. Thromb. Haemost. 2016, 14, 2045–2057. [Google Scholar] [CrossRef]
- Fiala, G.J.; Kaschek, D.; Blumenthal, B.; Reth, M.; Timmer, J.; Schamel, W.W.A. Pre-clustering of the B cell antigen receptor demonstrated by mathematically extended electron microscopy. Front. Immunol. 2013, 4, 427. [Google Scholar] [CrossRef] [Green Version]
- Hingant, E.; Yvinec, R. Deterministic and Stochastic Becker-Döring equations: Past and Recent Mathematical Developments. arXiv 2016, arXiv:1609.00697. [Google Scholar]
- Kim, S.; Zhen, J.; Popel, A.S.; Intaglietta, M.; Johnson, P.C. Contributions of collision rate and collision efficiency to erythrocyte aggregation in postcapillary venules at low flow rates. Am. J. Physiol. Heart Circ. Physiol. 2007, 293, H1947–H1954. [Google Scholar] [CrossRef] [PubMed] [Green Version]

| Parameter | ArgEE Concentration, mM | Pearson Correlation Coefficient | ||
|---|---|---|---|---|
| 50 | 100 | 700 | ||
| k1 | 2.3 × 10−3 | 4.3 × 10−3 | 2.3 × 10−3 | −0.69 |
| k−1 | 1.3 × 10−3 | 1.6 × 10−4 | 4.5 × 10−4 | −0.34 |
| k2 | 8.6 × 10−4 | 7.1 × 10−4 | 2.7 × 10−4 | −0.98 |
| k−2 | 0.052 | 0.022 | 1.1 × 10−3 | −0.85 |
| k3 | 0.015 | 5.3 × 10−3 | 3.4 × 10−4 | −0.80 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Garzon Dasgupta, A.K.; Martyanov, A.A.; Filkova, A.A.; Panteleev, M.A.; Sveshnikova, A.N. Development of a Simple Kinetic Mathematical Model of Aggregation of Particles or Clustering of Receptors. Life 2020, 10, 97. https://doi.org/10.3390/life10060097
Garzon Dasgupta AK, Martyanov AA, Filkova AA, Panteleev MA, Sveshnikova AN. Development of a Simple Kinetic Mathematical Model of Aggregation of Particles or Clustering of Receptors. Life. 2020; 10(6):97. https://doi.org/10.3390/life10060097
Chicago/Turabian StyleGarzon Dasgupta, Andrei K., Alexey A. Martyanov, Aleksandra A. Filkova, Mikhail A. Panteleev, and Anastasia N. Sveshnikova. 2020. "Development of a Simple Kinetic Mathematical Model of Aggregation of Particles or Clustering of Receptors" Life 10, no. 6: 97. https://doi.org/10.3390/life10060097
APA StyleGarzon Dasgupta, A. K., Martyanov, A. A., Filkova, A. A., Panteleev, M. A., & Sveshnikova, A. N. (2020). Development of a Simple Kinetic Mathematical Model of Aggregation of Particles or Clustering of Receptors. Life, 10(6), 97. https://doi.org/10.3390/life10060097






