Endolithic Fungal Species Markers for Harshest Conditions in the McMurdo Dry Valleys, Antarctica
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Area
2.2. Metabarcoding Sequencing
2.3. Bioinformatics
2.4. Downstream Analysis
3. Results
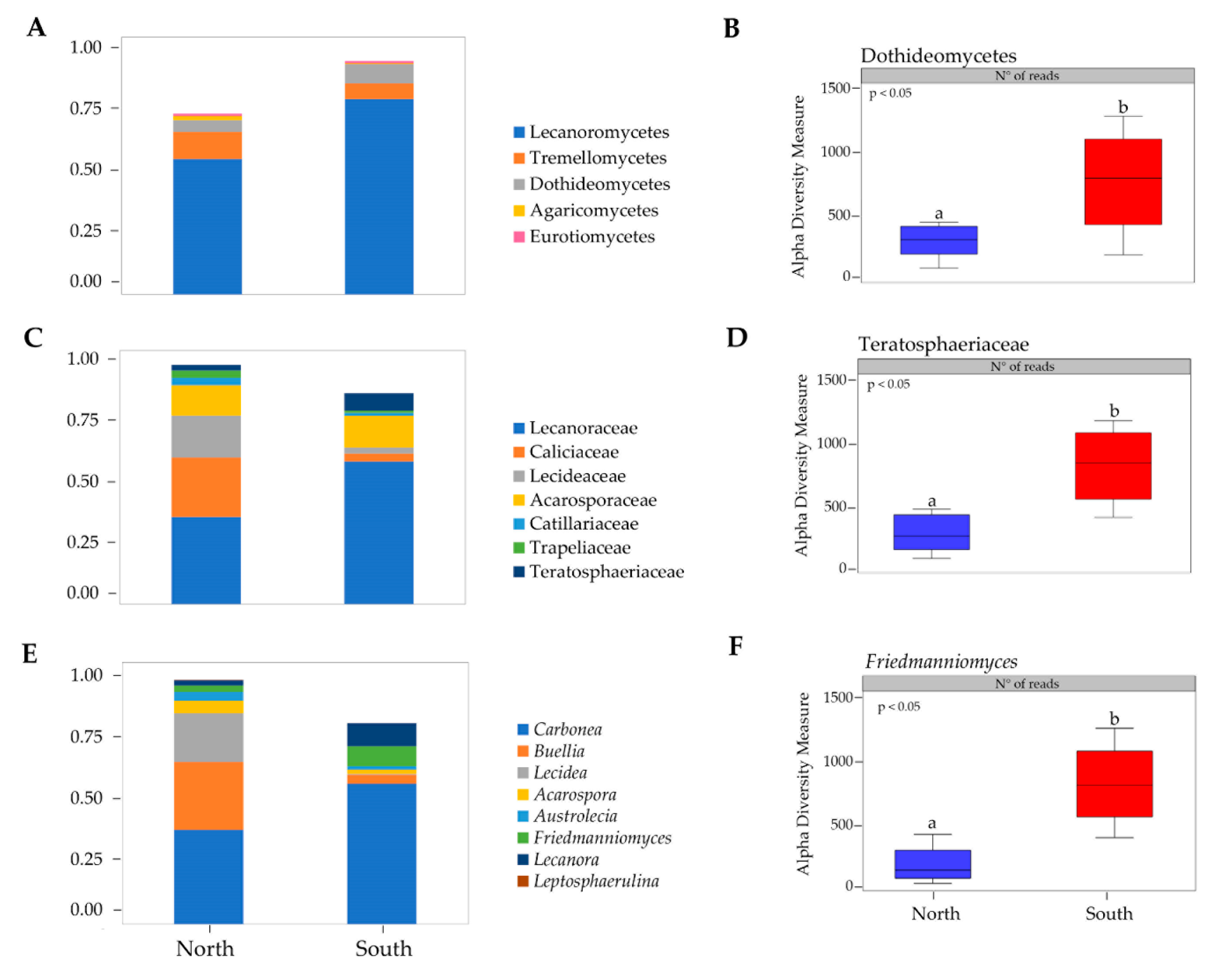
3.1. Taxonomy and Biodiversity Analysis
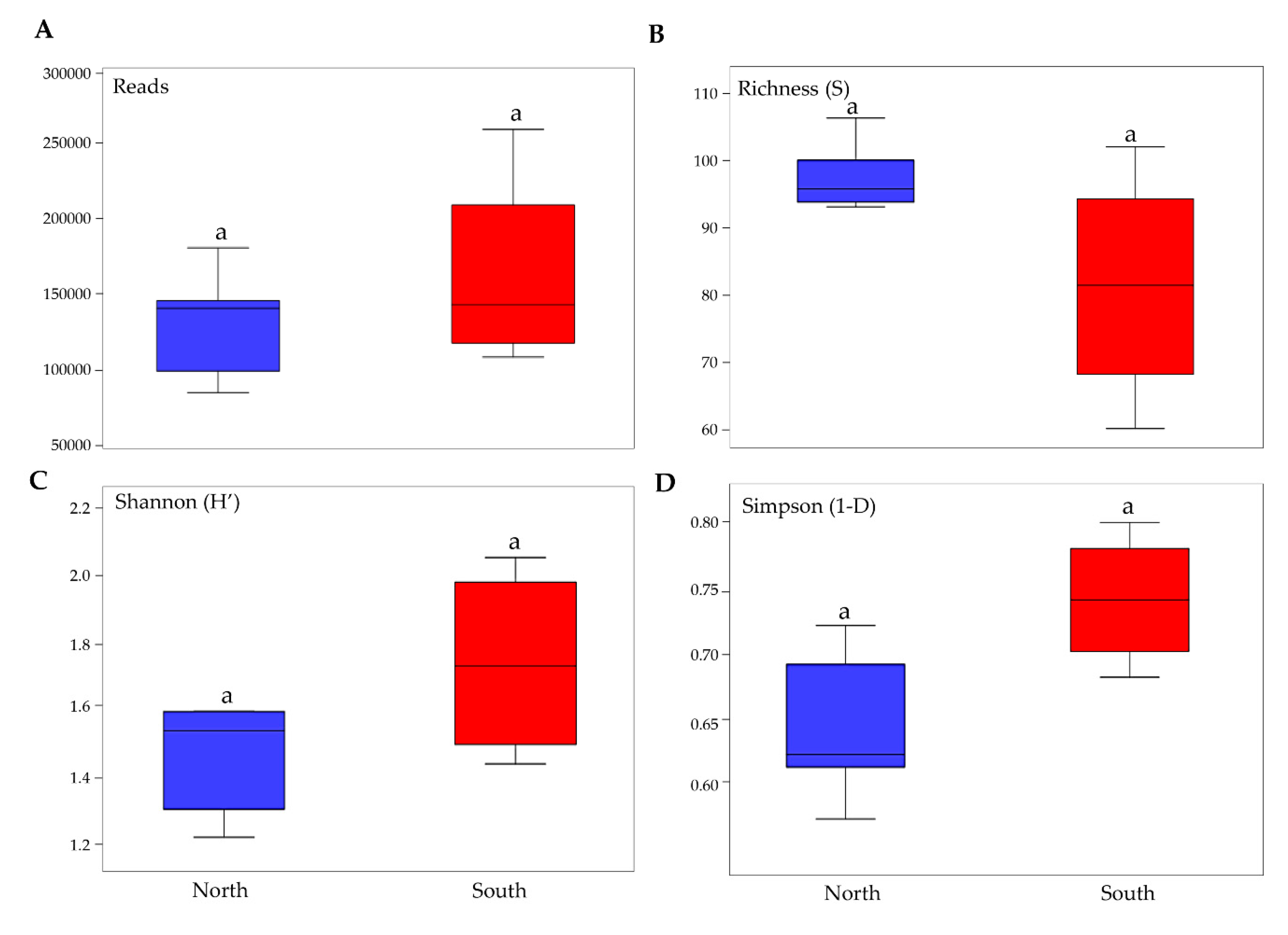
3.2. Biodiversity and Community Composition Patterns
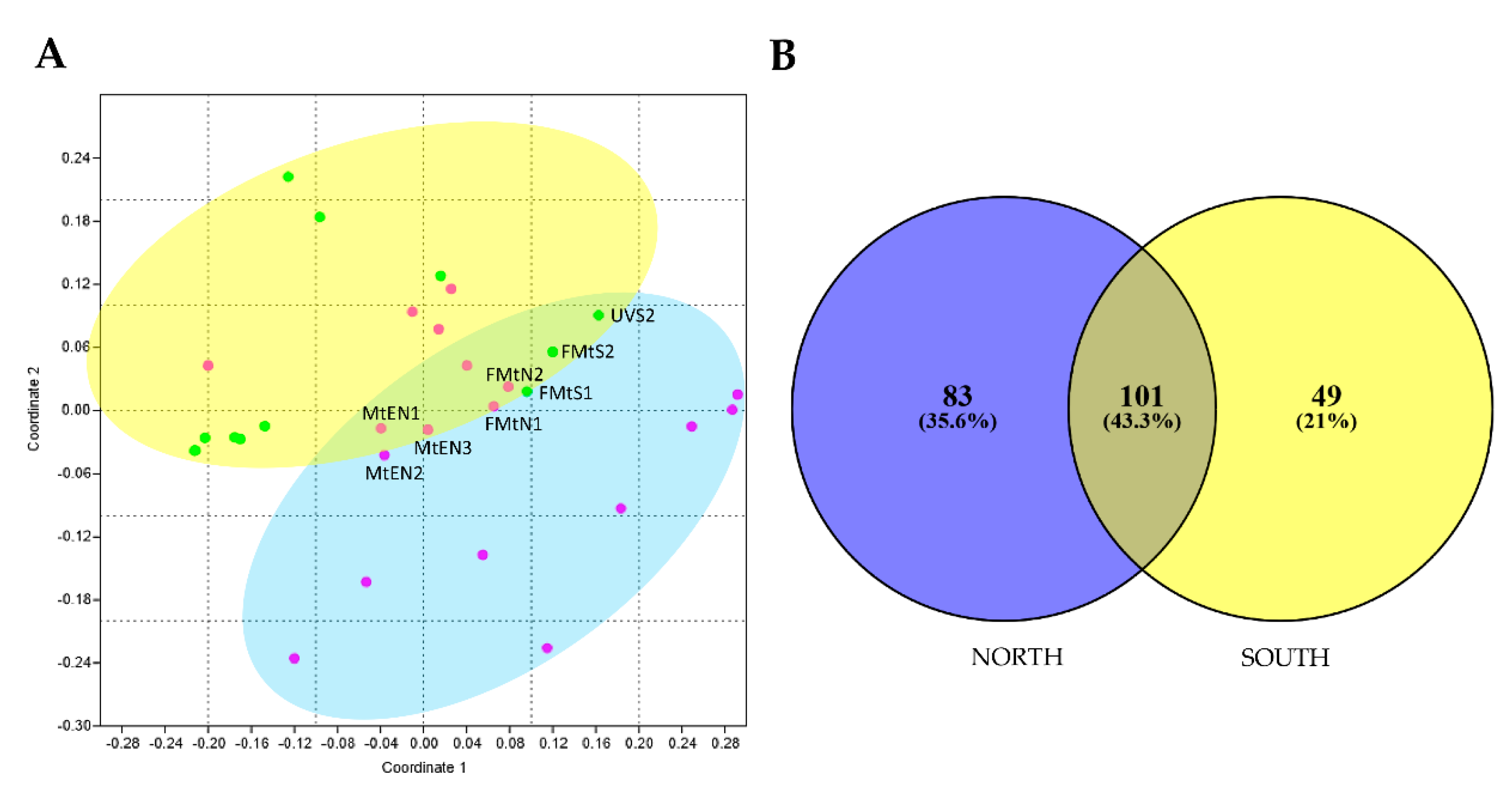
3.3. Unique and Indicator Species across North and South Sun Exposure
4. Discussion
Supplementary Materials
Author Contributions
Acknowledgments
Conflicts of Interest
References
- SCAR. SCAR Bulletin 155. Polar Rec (Gr Brit) 2004, 40, 371–382. [Google Scholar] [CrossRef]
- Doran, P.T.; McKay, C.P.; Clow, G.D.; Dana, G.L.; Fountain, A.G.; Nylen, T.; Lyons, W.B. Valley floor climate observations from the McMurdo Dry Valleys, Antarctica, 1986–2000. J. Geophys. Res. Atmos. 2002, 107, D24. [Google Scholar] [CrossRef]
- Horowitz, N.H.; Cameron, R.E.; Hubbard, J.S. Microbiology of the dry valleys of Antarctica. Science 1972, 176, 242–245. [Google Scholar] [CrossRef] [PubMed]
- Wynn-Williams, D.D.; Edwards, H.G.M. Proximal analysis of regolith habitats and protective biomolecules in situ by laser Raman spectroscopy: Overview of terrestrial Antarctic habitats and Mars analogs. Icarus 2000, 144, 486–503. [Google Scholar] [CrossRef]
- Onofri, S.; Selbmann, L.; Zucconi, L.; Pagano, S. Antarctic microfungi as models for exobiology. Planet. Space Sci. 2004, 52, 229–237. [Google Scholar] [CrossRef]
- Nienow, J.A.; Friedmann, E.I. Terrestrial lithophytic (rock) communities. In Antarctic Microbiology; Friedmann, E.I., Ed.; Wiley-Liss: New York, NY, USA, 1993; pp. 343–412. [Google Scholar]
- Friedmann, E.I.; Koriem, A.M. Life on Mars: How it disappeared (if it was ever there). Adv. Space Res. 1989, 9, 167–172. [Google Scholar] [CrossRef]
- Kappen, L. Plant activity under snow and ice, with particular reference to lichens. Arctic 1993, 46, 297–302. [Google Scholar] [CrossRef]
- Wierzchos, J.; Ascaso, C. Life, decay and fossilisation of endolithic microorganisms from the Ross Desert, Antarctica. Polar Biol. 2001, 24, 863–868. [Google Scholar] [CrossRef]
- Friedmann, E.I. Endolithic microorganisms in the Antarctic cold desert. Science 1982, 215, 1045–1053. [Google Scholar] [CrossRef]
- De la Torre, J.R.; Goebel, B.M.; Friedmann, E.; Pace, N.R. Microbial diversity of cryptoendolithic communities from the McMurdo Dry Valleys. Antarctica. Appl. Environ. Microbiol. 2003, 69, 3858–3867. [Google Scholar] [CrossRef]
- Friedmann, E.I.; Ocampo, R. Endolithic blue-green algae in dry valleys-primary producers in Antarctic desert ecosystem. Science 1976, 193, 1247–1249. [Google Scholar] [CrossRef]
- Selbmann, L.; Onofri, S.; Coleine, C.; Buzzini, P.; Canini, F.; Zucconi, L. Effect of environmental parameters on biodiversity of the fungal component in the lithic Antarctic communities. Extremophiles 2017, 21, 1069–1080. [Google Scholar] [CrossRef]
- Coleine, C.; Stajich, J.E.; Zucconi, L.; Onofri, S.; Pombubpa, N.; Egidi, E.; Frank, A.; Buzzini, P.; Selbmann, L. Antarctic cryptoendolithic fungal communities are highly adapted and dominated by Lecanoromycetes and Dothideomycetes. Front. Microbiol. 2018, 9, 1392. [Google Scholar] [CrossRef]
- Steig, E.J.; Schneider, D.P.; Rutherford, S.D.; Mann, M.E.; Comiso, J.C.; Shindell, D.T. Warming of the Antarctic ice-sheet surface since the 1957 International Geophysical Year. Nature 2009, 457, 459–462. [Google Scholar] [CrossRef]
- Hogg, I.D.; Wall, D.H. Global change and Antarctic terrestrial biodiversity. Polar Biol. 2011, 34, 1625–1627. [Google Scholar] [CrossRef]
- Archer, S.D.; de los Ríos, A.; Lee, K.C.; Niederberger, T.S.; Cary, S.C.; Coyne, K.J.; Douglas, S.; Lacap-Bugler, D.C.; Pointing, S.B. Endolithic microbial diversity in sandstone and granite from the McMurdo Dry Valleys, Antarctica. Polar Biol. 2017, 40, 997–1006. [Google Scholar] [CrossRef]
- Coleine, C.; Zucconi, L.; Onofri, S.; Pombubpa, N.; Stajich, J.E.; Selbmann, L. Sun exposure shapes functional grouping of fungi cryptoendolithic Antarctic communities. Life 2018, 8, 19. [Google Scholar] [CrossRef]
- Gonçalves, V.N.; Alves, I.M.S.; de Oliveira, F.S.; Schaefer, C.E.G.R.; Turbay, C.V.G.; Rosa, C.A.; Rosa, L.H. Rock-Inhabiting Fungi in Antarctica: New Frontiers of the Edge of Life. In Fungi of Antarctica; Springer: Cham, Switzerland, 2019; pp. 99–126. [Google Scholar] [CrossRef]
- Smith, D.P.; Peay, K.G. Sequence depth, not PCR replication, improves ecological inference from next generation DNA sequencing. PLoS ONE 2014, 9, 90234. [Google Scholar] [CrossRef]
- Palmer, J.M.; Jusino, M.A.; Banik, M.T.; Lindner, D.L. Non-biological synthetic spike-in controls and the AMPtk software pipeline improve mycobiome data. PeerJ 2018, 6, e4925. [Google Scholar] [CrossRef]
- Edgar, R.C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 2010, 26, 2460–2461. [Google Scholar] [CrossRef]
- Edgar, R.C.; Flyvbjerg, H. Error filtering, pair assembly and error correction for next-generation sequencing reads. Bioinformatics 2015, 31, 3476–3482. [Google Scholar] [CrossRef]
- Lindahl, B.D.; Nilsson, R.H.; Tedersoo, L.; Abarenkov, K.; Carlsen, T.; Kjøller, R.; Kõljalg, U.; Pennanen, T.; Rosendahl, S.; Stenlid, J.; et al. Fungal community analysis by high-throughput sequencing of amplified markers—A user’s guide. New Phytol. 2013, 199, 288–299. [Google Scholar] [CrossRef]
- Shannon, C.E.; Weaver, W. The Mathematical Theory of Communication; University of Illinois Press: Urbana, IL, USA, 1963. [Google Scholar]
- Simpson, E.H. Measurement of diversity. Nature 1949, 163, 688. [Google Scholar] [CrossRef]
- Baldrian, P.; Vetrovsky, T.; Cajthaml, T.; Dobiášová, P.; Petránková, M.; Šnajdr, J.; Eichlerová, I. Estimation of fungal biomass in forest litter and soil. Fungal Ecol. 2013, 6, 1–11. [Google Scholar] [CrossRef]
- Oksanen, J.; Blanchet, F.G.; Kindt, R.; Legendre, P.; Minchin, P.R.; O’hara, R.B.; Simpson, G.L.; Solymos, P.; Stevens, M.H.; Wagner, H.; et al. Package ‘vegan’. Community Ecol. Package 2013, 2, 1–295. [Google Scholar]
- Hammer, Ø.; Harper, D.A.T.; Ryan, P.D. PAST-Palaeontological Statistics. 2001. Available online: http://www.uv.es/~pardomv/pe/2001_1/past/pastprog/past.pdf (accessed on 1 February 2020).
- R Labs for Community Ecologists. Available online: http://ecology.msu.montana.edu/labdsv/R/ (accessed on 5 February 2020).
- Selbmann, L.; de Hoog, G.S.; Mazzaglia, A.; Friedmann, E.I.; Onofri, S. Fungi at the edge of life: Cryptoendolithic black fungi from Antarctic deserts. Stud. Mycol. 2005, 51, 1–32. [Google Scholar]
- Selbmann, L.; De Hoog, G.S.; Zucconi, L.; Isola, D.; Ruisi, S.; van den Ende, A.G.; Ruibal, C.; De Leo, F.; Onofri, S. Drought meets acid: Three new genera in a dothidealean clade of extremotolerant fungi. Stud. Mycol. 2008, 61, 1–20. [Google Scholar] [CrossRef]
- Selbmann, L.; Isola, D.; Egidi, E.; Zucconi, L.; Gueidan, C.; de Hoog, G.S.; Onofri, S. Mountain tips as reservoirs for new rock-fungal entities: Saxomyces gen. nov. and four new species from the Alps. Fungal Divers. 2014, 65, 167–182. [Google Scholar] [CrossRef]
- Selbmann, L.; Onofri, S.; Zucconi, L.; Isola, D.; Rottigni, M.; Ghiglione, C.; Piazza, P.; Alvaro, M.C.; Schiaparelli, S. Distributional records of Antarctic fungi based on strains preserved in the Culture Collection of Fungi from Extreme Environments (CCFEE) Mycological Section associated with the Italian National Antarctic Museum (MNA). MycoKeys 2015, 10, 57. [Google Scholar] [CrossRef]
- Egidi, E.; de Hoog, G.S.; Isola, D.; Onofri, S.; Quaedvlieg, W.; de Vries, M.; Verkley, G.J.M.; Stielow, J.B.; Zucconi, L.; Selbmann, L. Phylogeny and taxonomy of meristematic rock-inhabiting black fungi in the Dothidemycetes based on multi-locus phylogenies. Fungal Divers 2014, 65, 127–165. [Google Scholar] [CrossRef]
- Abdel-Hafez, S.I.I. Studies on soil mycoflora of desert soils in Saudi Arabia. Mycopathologia 1994, 80, 3e8. [Google Scholar]
- Adams, B.J.; Bardgett, R.D.; Ayres, E.; Wall, D.H.; Aislabie, J.; Bamforth, S.; Bargagli, R.; Cary, C.; Cavacini, P.; Convey, P. Diversity and distribution of Victoria Land biota. Soil Biol. Biochem. 2006, 38, 3003–3018. [Google Scholar] [CrossRef]
- Gunde-Cimerman, N.; Zalar, P.; de Hoog, S.; Plemenitaš, A. Hypersaline waters in salterns–natural ecological niches for halophilic black yeasts. FEMS Microbiol. Ecol. 2000, 32, 235–240. [Google Scholar] [CrossRef]
- Gunde-Cimerman, N.; Sonjak, S.; Zalar, P.; Frisvad, J.C.; Diderichsen, B.; Plemenitaš, A. Extremophilic fungi in arctic ice: A relationship between adaptation to low temperature and water activity. Phys. Chem. Earth Parts 2003, 28, 1273–1278. [Google Scholar] [CrossRef]
- Sterflinger, K.; Tesei, D.; Zakharova, K. Fungi in hot and cold deserts with particular reference to microcolonial fungi. Fungal Ecol. 2012, 5, 453–462. [Google Scholar] [CrossRef]
- Ruibal, C.; Selbmann, L.; Avci, S.; Martin-Sanchez, P.; Gorbushina, A. Roof-Inhabiting Cousins of Rock-Inhabiting Fungi: Novel Melanized Microcolonial Fungal Species from Photocatalytically Reactive Subaerial Surfaces. Life 2018, 8, 30. [Google Scholar] [CrossRef]
- Gorbushina, A.A.; Kotlova, E.R.; Sherstneva, O.A. Cellular responses of microcolonial rock fungi to long-term desiccation and subsequent rehydration. Stud. Mycol. 2018, 61, 91–97. [Google Scholar] [CrossRef]
- Dadachova, E.; Casadevall, A. Ionizing radiation: How fungi cope, adapt, and exploit with the help of melanin. Curr. Opin. Microbiol. 2008, 11, 525–531. [Google Scholar] [CrossRef]
- Onofri, S.; Barreca, D.; Selbmann, L.; Isola, D.; Rabbow, E.; Horneck, G.; de Vera, J.P.P.; Hatton, J.; Zucconi, L. Resistance of Antarctic black fungi and cryptoendolithic communities to simulated space and Martian conditions. Stud. Mycol. 2008, 61, 99–109. [Google Scholar] [CrossRef]
- Onofri, S.; de la Torre, R.; de Vera, J.P.; Ott, S.; Zucconi, L.; Selbmann, L.; de la Torre, R.; Ott, S.; Rabbow, E.; Iñigo, F.J.S.; et al. Survival of rock-colonizing organisms after 1.5 Years in outer space. Astrobiology 2012, 12, 508–516. [Google Scholar] [CrossRef]
- Onofri, S.; de Vera, J.P.; Zucconi, L.; Selbmann, L.; Scalzi, G.; Venkateswaran, K.J.; Rabbow, E.; de la Torre, R.; Horneck, G. Survival of Antarctic cryptoendolithic fungi in simulated Martian conditions on-board the International Space Station. Astrobiology 2015, 15, 1052–1059. [Google Scholar] [CrossRef] [PubMed]
- Onofri, S.; Selbmann, L.; Pacelli, C.; Zucconi, L.; Rabbow, E.; de Vera, J.P. Survival, DNA, and Ultrastructural Integrity of a Cryptoendolithic Antarctic Fungus in Mars and Lunar Rock Analogs Exposed Outside the International Space Station. Astrobiology 2019, 19, 170–182. [Google Scholar] [CrossRef] [PubMed]
- Selbmann, L.; Isola, D.; Zucconi, L.; Onofri, S. Resistance to UV-B induced DNA damage in extreme-tolerant cryptoendolithic Antarctic fungi: Detection by PCR assays. Fungal Biol. 2011, 115, 937–944. [Google Scholar] [CrossRef] [PubMed]
- Selbmann, L.; Pacelli, C.; Zucconi, L.; Dadachova, E.; Moeller, R.; de Vera, J.P.; Onofri, S. Resistance of an Antarctic cryptoendolithic black fungus to radiation gives new insights of astrobiological relevance. Fungal Biol. 2018, 122, 546–554. [Google Scholar] [CrossRef]
- Selbmann, L.; Egidi, E.; Isola, D.; Onofri, S.; Zucconi, L.; de Hoog, G.S.; Chinaglia, S.; Testa, L.; Tosi, S.; Balestrazzi, A.; et al. Biodiversity, evolution and adaptation of fungi in extreme environments. Plant Biosyst. 2013, 147, 237–246. [Google Scholar] [CrossRef]
- Geml, J.; Semenova, T.A.; Morgado, L.N.; Welker, J.M. Changes in composition and abundance of functional groups of arctic fungi in response to long-term summer warming. Biol. Lett. 2016, 12, 20160503. [Google Scholar] [CrossRef]
- Coleine, C.; Stajich, J.E.; Zucconi, L.; Onofri, S.; Selbmann, L. Sun exposure drives Antarctic cryptoendolithic community structure and composition. BioRxiv 2019, 676692. [Google Scholar] [CrossRef]
- Coleine, C.; Gevi, F.; Fanelli, G.; Onofri, S.; Timperio, A.M.; Selbmann, L. Metabolic responses in opposite sun-exposed Antarctic cryptoendolithic communities. BioRxiv 2019, 725663. [Google Scholar] [CrossRef]
- Shade, A.; Handelsman, J. Beyond the Venn diagram: The hunt for a core microbiome. Environ. Microbiol. 2012, 14, 4–12. [Google Scholar] [CrossRef]
- Kappen, L. Some aspects of the great success of lichens in Antarctica. Antarct. Sci. 2000, 12, 314–324. [Google Scholar] [CrossRef]
- Sancho, L.G.; Green, T.A.; Pintado, A. Slowest to fastest: Extreme range in lichen growth rates supports their use as an indicator of climate change in Antarctica. Flora 2007, 202, 667–673. [Google Scholar] [CrossRef]
- Onofri, S.; Pagano, S.; Zucconi, L.; Tosi, S. Friedmanniomyces endolithicus (Fungi, Hyphomycetes), anam-gen and sp nov, from continental Antarctica. Nova Hedwig. 1999, 68, 175–182. [Google Scholar]
- Tesei, D.; Marzban, G.; Zakharova, K.; Isola, D.; Selbmann, L.; Sterflinger, K. Alteration of protein patterns in black rock inhabiting fungi as a response to different temperatures. Fungal Biol. 2012, 116, 932–940. [Google Scholar] [CrossRef] [PubMed]
- Pacelli, C.; Bryan, R.A.; Onofri, S.; Selbmann, L.; Zucconi, L.; Shuryak, I.; Dadachova, E. Survival and redox activity of Friedmanniomyces endolithicus, an Antarctic endemic black meristematic fungus, after gamma rays exposure. Fungal Biol. 2018, 122, 1222–1227. [Google Scholar] [CrossRef] [PubMed]
- Coleine, C.; Masonjones, S.; Sterflinger, K.; Onofri, S.; Selbmann, L.; Stajich, J.E. Peculiar genomic traits in the stress-adapted cryptoendolithic endemic Antarctic fungus Friedmanniomyces endolithicus. Fungal Biol. 2020. [CrossRef]
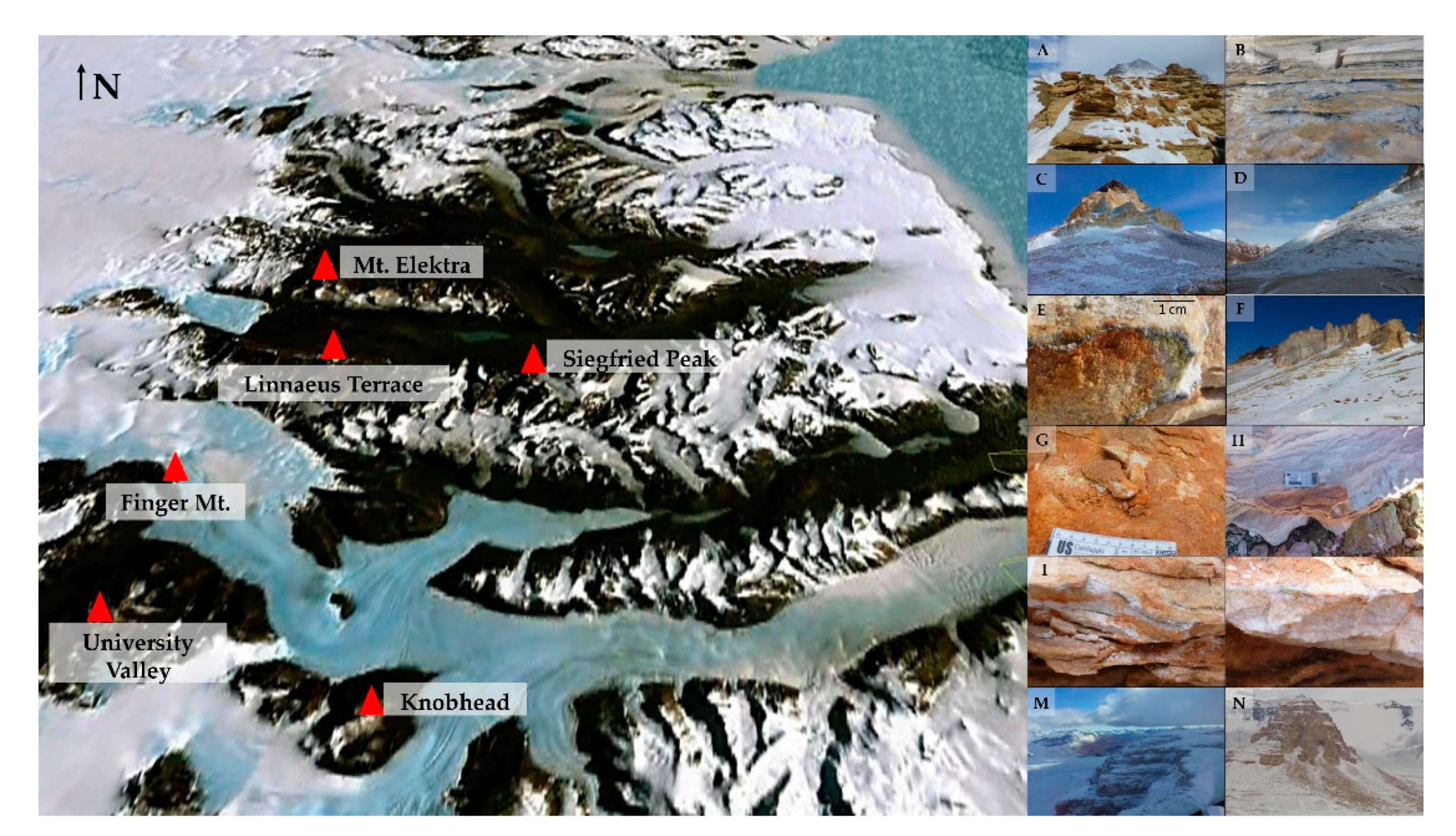

| Site | Sun Exposure | Altitude (m a.s.l.) | Relative Humidity (%) | Coordinates |
|---|---|---|---|---|
| Siegfried Peak | North | 1620 | 52 | 77°34′43.3″ S 161°47′11.7″ E |
| Siegfried Peak | South | 1620 | 54 | 77°34′39.9″ S 161°47′17.4″ E |
| Linnaeus Terrace | North | 1649 | 58 | 77°36′01.3″ S 161°05′00.5″ E |
| Linnaeus Terrace | South | 1761 | 68 | 77°37′09.9″ S 161°11′50.8″ E |
| Finger Mt. | North | 1720 | 35 | 77°45′0.9″ S 160°44′44.5″ E |
| Finger Mt. | South | 1720 | 35 | 77°45′10.3″ S 160°44′40.3″ E |
| Mt. Elektra | North | 2080 | 63 | 77°29′28.0″ S 160°54′16.4″ E |
| University Valley | North | 2090 | 18 | 77°52′27.6″ S 160°44′38.9″ E |
| University Valley | South | 2200 | 39 | 77°52′21.6″ S 160°45′20.5″ E |
| Knobhead | North | 2150 | 50 | 77°54′37.8″ S 161°34′48.8″ E |
| Knobhead | South | 2150 | 38 | 77°54′43.6″ S 161°34′39.3″ E |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Coleine, C.; Pombubpa, N.; Zucconi, L.; Onofri, S.; Stajich, J.E.; Selbmann, L. Endolithic Fungal Species Markers for Harshest Conditions in the McMurdo Dry Valleys, Antarctica. Life 2020, 10, 13. https://doi.org/10.3390/life10020013
Coleine C, Pombubpa N, Zucconi L, Onofri S, Stajich JE, Selbmann L. Endolithic Fungal Species Markers for Harshest Conditions in the McMurdo Dry Valleys, Antarctica. Life. 2020; 10(2):13. https://doi.org/10.3390/life10020013
Chicago/Turabian StyleColeine, Claudia, Nuttapon Pombubpa, Laura Zucconi, Silvano Onofri, Jason E. Stajich, and Laura Selbmann. 2020. "Endolithic Fungal Species Markers for Harshest Conditions in the McMurdo Dry Valleys, Antarctica" Life 10, no. 2: 13. https://doi.org/10.3390/life10020013
APA StyleColeine, C., Pombubpa, N., Zucconi, L., Onofri, S., Stajich, J. E., & Selbmann, L. (2020). Endolithic Fungal Species Markers for Harshest Conditions in the McMurdo Dry Valleys, Antarctica. Life, 10(2), 13. https://doi.org/10.3390/life10020013








