Rapid Transcriptional Reprogramming Triggered by Alteration of the Carbon/Nitrogen Balance Has an Impact on Energy Metabolism in Nostoc sp. PCC 7120
Abstract
1. Introduction
2. Materials and Methods
2.1. Growth and CO2 Stepdown
2.2. RNA Isolation and Transcriptomics
3. Results
3.1. Uptake and Metabolism of Carbon and Nitrogen Are Inversely Responsive to Low CO2 Conditions
3.2. Expression of Genes Encoding Photosynthetic and Respiratory Components Responds to Low CO2 Conditions
3.3. Expression of Transcription Regulators Responds to Changes in CO2 Conditions
3.4. Low CO2 Conditions Influence the Expression of Metal Homeostasis Genes
4. Discussion
4.1. Transcriptional Regulation in Response to CO2 Stepdown is Triggered by Metabolites
4.2. Altered C/N Balance Modulates the Energetic Strategy of Nostoc sp. PCC 7120
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Herrero, A.; Flores, E. Genetic responses to carbon and nitrogen availability in Anabaena. Environ. Microbiol. 2019, 21, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Burnap, R.L.; Hagemann, M.; Kaplan, A. Regulation of CO2 Concentrating Mechanism in Cyanobacteria. Life 2015, 5, 348–371. [Google Scholar] [CrossRef] [PubMed]
- Huergo, L.F.; Dixon, R. The Emergence of 2-Oxoglutarate as a Master Regulator Metabolite. Microbiol. Mol. Biol. Rev. 2015, 79, 419–435. [Google Scholar] [CrossRef]
- Zhang, C.-C.; Zhou, C.-Z.; Burnap, R.L.; Peng, L. Carbon/Nitrogen Metabolic Balance: Lessons from Cyanobacteria. Trends Plant Sci. 2018, 23, 1116–1130. [Google Scholar] [CrossRef]
- Flores, E.; Frías, J.E.; Rubio, L.M.; Herrero, A. Photosynthetic nitrate assimilation in cyanobacteria. Photosynth. Res. 2005, 83, 117–133. [Google Scholar] [CrossRef]
- Forchhammer, K.; Selim, K.A. Carbon/nitrogen homeostasis control in cyanobacteria. FEMS Microbiol. Rev. 2019, 44, 33–53. [Google Scholar] [CrossRef]
- Muro-Pastor, M.I.; Reyes, J.C.; Florencio, F.J. Cyanobacteria perceive nitrogen status by sensing intracellular 2-oxoglutarate levels. J. Biol. Chem. 2001, 276, 38320–38328. [Google Scholar] [PubMed]
- Vázquez-Bermúdez, M.F.; Herrero, A.; Flores, E. 2-Oxoglutarate increases the binding affinity of the NtcA (nitrogen control) transcription factor for the Synechococcus glnA promoter. FEBS Lett. 2002, 512, 71–74. [Google Scholar] [CrossRef]
- Li, J.-H.; Laurent, S.; Konde, V.; Bédu, S.; Zhang, C.-C. An increase in the level of 2-oxoglutarate promotes heterocyst development in the cyanobacterium Anabaena sp. strain PCC 7120. Microbiology 2003, 149, 3257–3263. [Google Scholar] [CrossRef]
- Haimovich-Dayan, M.; Lieman-Hurwitz, J.; Orf, I.; Hagemann, M.; Kaplan, A. Does 2-phosphoglycolate serve as an internal signal molecule of inorganic carbon deprivation in the cyanobacterium Synechocystis sp. PCC 6803? Environ. Microbiol. 2015, 17, 1794–1804. [Google Scholar] [CrossRef]
- Hagemann, M.; Bauwe, H. Photorespiration and the potential to improve photosynthesis. Curr. Opin. Chem. Biol. 2016, 35, 109–116. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.-L.; Postier, B.L.; Burnap, R.L. Alterations in Global Patterns of Gene Expression in Synechocystis sp. PCC 6803 in Response to Inorganic Carbon Limitation and the Inactivation of ndhR, a LysR Family Regulator. J. Biol. Chem. 2004, 279, 5739–5751. [Google Scholar] [CrossRef] [PubMed]
- Eisenhut, M.; Von Wobeser, E.A.; Jonas, L.; Schubert, H.; Ibelings, B.W.; Bauwe, H.; Matthijs, H.C.; Hagemann, M. Long-Term Response toward Inorganic Carbon Limitation in Wild Type and Glycolate Turnover Mutants of the Cyanobacterium Synechocystis sp. Strain PCC 6803. Plant Physiol. 2007, 144, 1946–1959. [Google Scholar] [CrossRef] [PubMed]
- Kaplan, A.; Hagemann, M.; Bauwe, H.; Kahlon, S.; Ogawa, T. Carbon acquisition by cyanobacteria: Mechanisms, comparative genomics and evolution. In The Cyanobacteria: Molecular Biology, Genomics and Evolution; Herrero, A., Flores, E., Eds.; Caister Academic Press: Norwich, UK, 2008; pp. 305–323. [Google Scholar]
- Klähn, S.; Orf, I.; Schwarz, D.; Matthiessen, J.K.; Kopka, J.; Hess, W.R.; Hagemann, M. Integrated Transcriptomic and Metabolomic Characterization of the Low-Carbon Response Using an ndhR Mutant of Synechocystis sp. PCC 68031. Plant Physiol. 2015, 169, 1540–1556. [Google Scholar] [CrossRef]
- Wolk, C.P.; Ernst, A.; Elhai, J. Heterocyst Metabolism and Development. In The Molecular Biology of Cyanobacteri; Springer: Dordrecht, The Netherlands, 1994; pp. 769–823. [Google Scholar] [CrossRef]
- Herrero, A.; Muro-Pastor, A.M.; Valladares, A.; Flores, E. Cellular differentiation and the NtcA transcription factor in filamentous cyanobacteria. FEMS Microbiol. Rev. 2004, 28, 469–487. [Google Scholar] [CrossRef]
- Picossi, S.; Flores, E.; Herrero, A. The LysR-type transcription factor PacR is a global regulator of photosynthetic carbon assimilation inAnabaena. Environ. Microbiol. 2015, 17, 3341–3351. [Google Scholar] [CrossRef]
- Allahverdiyeva, Y.; Vainonen, J.P.; Vorontsova, N.; Keränen, M.; Carmel, D.; Aro, E.-M. Dynamic Changes in the Proteome of Synechocystis 6803 in Response to CO2 Limitation Revealed by Quantitative Proteomics. J. Proteome Res. 2010, 9, 5896–5912. [Google Scholar] [CrossRef]
- Schwarz, D.; Nodop, A.; Hüge, J.; Purfürst, S.; Forchhammer, K.; Michel, K.-P.; Bauwe, H.; Kopka, J.; Hagemann, M. Metabolic and Transcriptomic Phenotyping of Inorganic Carbon Acclimation in the Cyanobacterium Synechococcus elongatus PCC 7942. Plant Physiol. 2011, 155, 1640–1655. [Google Scholar] [CrossRef]
- Price, G.D.; Woodger, F.J.; Badger, M.R.; Howitt, S.M.; Tucker, L. Identification of a SulP-type bicarbonate transporter in marine cyanobacteria. Proc. Natl. Acad. Sci. USA 2004, 101, 18228–18233. [Google Scholar] [CrossRef]
- McGinn, P.J.; Price, G.D.; Maleszka, R.; Badger, M.R. Inorganic Carbon Limitation and Light Control the Expression of Transcripts Related to the CO2-Concentrating Mechanism in the Cyanobacterium Synechocystis sp. Strain PCC6803. Plant Physiol. 2003, 132, 218–229. [Google Scholar] [CrossRef]
- Rippka, R.; Stanier, R.Y.; Deruelles, J.; Herdman, M.; Waterbury, J.B. Generic Assignments, Strain Histories and Properties of Pure Cultures of Cyanobacteria. Microbiology 1979, 111, 1–61. [Google Scholar] [CrossRef]
- Walter, J.; Lynch, F.; Battchikova, N.; Aro, E.-M.; Gollan, P.J. Calcium impacts carbon and nitrogen balance in the filamentous cyanobacterium Anabaena sp. PCC 7120. J. Exp. Bot. 2016, 67, 3997–4008. [Google Scholar] [CrossRef] [PubMed]
- Omata, T.; Price, G.D.; Badger, M.R.; Okamura, M.; Gohta, S.; Ogawa, T. Identification of an ATP-binding cassette transporter involved in bicarbonate uptake in the cyanobacterium Synechococcus sp. strain PCC 7942. Proc. Natl. Acad. Sci. USA 1999, 96, 13571–13576. [Google Scholar] [CrossRef] [PubMed]
- Shibata, M.; Katoh, H.; Sonoda, M.; Ohkawa, H.; Shimoyama, M.; Fukuzawa, H.; Kaplan, A.; Ogawa, T. Genes Essential to Sodium-dependent Bicarbonate Transport in Cyanobacteria: Function and phylogenetic analysis. J. Biol. Chem. 2002, 277, 18658–18664. [Google Scholar] [CrossRef] [PubMed]
- Blanco-Rivero, A.; Leganés, F.; Fernández-Valiente, E.; Calle, P.; Fernández-Piñas, F. mrpA, a gene with roles in resistance to Na+ and adaptation to alkaline pH in the cyanobacterium Anabaena sp. PCC7120. Microbiology 2005, 151, 1671–1682. [Google Scholar] [CrossRef] [PubMed]
- Fukaya, F.; Promden, W.; Hibino, T.; Tanaka, Y.; Nakamura, T.; Takabe, T. An Mrp-like cluster in the halotolerant cyanobacterium Aphanothece halophytica functions as a Na+/H+ antiporter. Appl. Environ. Microbiol. 2009, 75, 6626–6629. [Google Scholar] [CrossRef]
- Allahverdiyeva, Y.; Aro, E.-M.; Nixon, P.J. Structure and Physiological Function of NDH-1 Complexes in Cyanobacteria. In Bioenergetic Processes of Cyanobacteria; Springer: Dordrecht, The Netherlands, 2011; pp. 445–467. [Google Scholar] [CrossRef]
- Kaneko, T.; Nakamura, Y.; Wolk, C.P.; Kuritz, T.; Sasamoto, S.; Watanabe, A.; Iriguchi, M.; Ishikawa, A.; Kawashima, K.; Kimura, T.; et al. Complete Genomic Sequence of the Filamentous Nitrogen-fixing Cyanobacterium Anabaena sp. Strain PCC 7120. DNA Res. 2001, 8, 205–213. [Google Scholar] [CrossRef]
- Frías, J.E.; Flores, E.; Herrero, A. Nitrate assimilation gene cluster from the heterocyst-forming cyanobacterium Anabaena sp. strain PCC 7120. J. Bacteriol. 1997, 179, 477–486. [Google Scholar] [CrossRef]
- Frías, J.E.; Flores, E. Negative Regulation of Expression of the Nitrate Assimilation nirA Operon in the Heterocyst-Forming Cyanobacterium Anabaena sp. Strain PCC 7120. J. Bacteriol. 2010, 192, 2769–2778. [Google Scholar] [CrossRef]
- Flores, E.; Picossi, S.; Valladares, A.; Herrero, A. Transcriptional regulation of development in heterocyst-forming cyanobacteria. Biochim. Biophys. Acta Bioenergy 2019, 1862, 673–684. [Google Scholar] [CrossRef]
- Kumar, K.; Mella-Herrera, R.A.; Golden, J.W. Cyanobacterial Heterocysts. Cold Spring Harb. Perspect. Biol. 2010, 2, a000315. [Google Scholar] [CrossRef] [PubMed]
- Maldener, I.; Muro-Pastor, A.M. Cyanobacterial Heterocysts. In Encyclopedia of Life Sciences; John Wiley & Sons, Ltd.: Chichester, UK, 2010. [Google Scholar] [CrossRef]
- Aro, E.-M.; Virgin, I.; Andersson, B. Photoinhibition of Photosystem II. Inactivation, protein damage and turnover. Biochim. Biophys. Acta Bioenergy 1993, 1143, 113–134. [Google Scholar] [CrossRef]
- Komenda, J.; Hassan, H.A.; Diner, B.A.; Debus, R.J.; Barber, J.; Nixon, P.J. Degradation of the Photosystem II D1 and D2 proteins in different strains of the cyanobacterium Synechocystis PCC 6803 varying with respect to the type and level of psbA transcript. Plant Mol. Biol. 2000, 42, 635–645. [Google Scholar] [CrossRef]
- Silva, P.; Thompson, E.P.; Bailey, S.; Kruse, O.; Mullineaux, C.W.; Robinson, C.; Mann, N.H.; Nixon, P.J. FtsH Is Involved in the Early Stages of Repair of Photosystem II in Synechocystis sp PCC 6803. Plant Cell 2003, 15, 2152–2164. [Google Scholar] [CrossRef] [PubMed]
- Komenda, J.; Barker, M.; Kuviková, S.; De Vries, R.; Mullineaux, C.W.; Tichý, M.; Nixon, P.J. The FtsH Protease slr0228 Is Important for Quality Control of Photosystem II in the Thylakoid Membrane of Synechocystis sp. PCC 6803. J. Biol. Chem. 2006, 281, 1145–1151. [Google Scholar] [CrossRef] [PubMed]
- Eisenhut, M.; Georg, J.; Klähn, S.; Sakurai, I.; Mustila, H.; Zhang, P.; Hess, W.R.; Aro, E.-M. The Antisense RNA As1_flv4 in the Cyanobacterium Synechocystis sp. PCC 6803 Prevents Premature Expression of the flv4-2 Operon upon Shift in Inorganic Carbon Supply. J. Biol. Chem. 2012, 287, 33153–33162. [Google Scholar] [CrossRef]
- Bersanini, L.; Battchikova, N.; Jokel, M.; Rehman, A.; Vass, I.; Allahverdiyeva, Y.; Aro, E.M. Flavodiiron protein Flv2/Flv4-related photoprotective mechanism dissipates excitation pressure of PSII in cooperation with phycobilisomes in cyanobacteria. Plant Physiol. 2014, 164, 805–818. [Google Scholar] [CrossRef]
- Gwizdala, M.; Wilson, A.; Kirilovsky, D. In Vitro Reconstitution of the Cyanobacterial Photoprotective Mechanism Mediated by the Orange Carotenoid Protein in Synechocystis PCC 6803. Plant Cell 2011, 23, 2631–2643. [Google Scholar] [CrossRef]
- López-Igual, R.; Wilson, A.; Leverenz, R.L.; Melnicki, M.R.; De Carbon, C.B.; Sutter, M.; Turmo, A.; Perreau, F.; Kerfeld, C.A.; Kirilovsky, D. Different Functions of the Paralogs to the N-Terminal Domain of the Orange Carotenoid Protein in the Cyanobacterium Anabaena sp. PCC 7120. Plant Physiol. 2016, 171, 1852–1866. [Google Scholar] [CrossRef]
- Kufryk, G.; Hernandez-Prieto, M.A.; Kieselbach, T.; Miranda, H.; Vermaas, W.; Funk, C. Association of small CAB-like proteins (SCPs) of Synechocystis sp. PCC 6803 with Photosystem II. Photosynth. Res. 2008, 95, 135–145. [Google Scholar] [CrossRef]
- Lea-smith, D.J.; Bombelli, P.; Vasudevan, R.; Howe, C.J. Photosynthetic, Respiratory and extracellular electron transport pathways in cyanobacteria. Biochim. Biophys. Acta Bioenergy 2016, 1857, 247–255. [Google Scholar] [CrossRef] [PubMed]
- González, A.; Fillat, M.F. Overexpression, immunodetection, and site-directed mutagenesis of Anabaena sp. PCC 7120 flavodoxin: A comprehensive laboratory practice on molecular biology. Biochem. Mol. Biol. Educ. 2018, 46, 493–501. [Google Scholar] [CrossRef]
- Laudenbach, D.E.; Straus, N.A. Characterization of a cyanobacterial iron stress-induced gene similar to psbC. J. Bacteriol. 1988, 170, 5018–5026. [Google Scholar] [CrossRef] [PubMed]
- Torrado, A.; Ramírez-Moncayo, C.; Navarro, J.A.; Mariscal, V.; Molina-Heredia, F.P. Cytochrome c6 is the main respiratory and photosynthetic soluble electron donor in heterocysts of the cyanobacterium Anabaena sp. PCC 7120. Biochim. Biophys. Acta Bioenergy 2019, 1860, 60–68. [Google Scholar] [CrossRef] [PubMed]
- Ermakova, M.; Allahverdiyeva, Y.; Allahverdiyeva, Y.; Aro, E.-M.; Allahverdiyeva, Y. Novel heterocyst-specific flavodiiron proteins in Anabaena sp. PCC 7120. FEBS Lett. 2013, 587, 82–87. [Google Scholar] [CrossRef]
- Swanson, R.V.; De Lorimier, R.; Glazer, A.N. Genes encoding the phycobilisome rod substructure are clustered on the Anabaena chromosome: Characterization of the phycoerythrocyanin operon. J. Bacteriol. 1992, 174, 2640–2647. [Google Scholar] [CrossRef][Green Version]
- Ducret, A.; Sidler, W.; Wehrli, E.; Frank, G.; Zuber, H. Isolation, Characterization and Electron Microscopy Analysis of A Hemidiscoidal Phycobilisome Type from the Cyanobacterium Anabaena sp. PCC 7120. JBIC J. Biol. Inorg. Chem. 1996, 236, 1010–1024. [Google Scholar] [CrossRef]
- Khanna, N.; Lindblad, P. Cyanobacterial Hydrogenases and Hydrogen Metabolism Revisited: Recent Progress and Future Prospects. Int. J. Mol. Sci. 2015, 16, 10537–10561. [Google Scholar] [CrossRef]
- Appel, J.; Phunpruch, S.; Steinmüller, K.; Schulz, R. The bidirectional hydrogenase of Synechocystis sp. PCC 6803 works as an electron valve during photosynthesis. Arch. Microbiol. 2000, 173, 333–338. [Google Scholar] [CrossRef]
- Carrieri, D.; Wawrousek, K.; Eckert, C.; Yu, J.; Maness, P. The role of the bidirectional hydrogenase in cyanobacteria. Bioresour. Technol. 2011, 102, 8368–8377. [Google Scholar] [CrossRef]
- Gutekunst, K.; Chen, X.; Schreiber, K.; Kaspar, U.; Makam, S.; Appel, J. The Bidirectional NiFe-hydrogenase in Synechocystis sp. PCC 6803 Is Reduced by Flavodoxin and Ferredoxin and Is Essential under Mixotrophic, Nitrate-limiting Conditions. J. Biol. Chem. 2014, 289, 1930–1937. [Google Scholar] [CrossRef] [PubMed]
- Omata, T.; Gohta, S.; Takahashi, Y.; Harano, Y.; Maeda, S.-I. Involvement of a CbbR Homolog in Low CO2-Induced Activation of the Bicarbonate Transporter Operon in Cyanobacteria. J. Bacteriol. 2001, 183, 1891–1898. [Google Scholar] [CrossRef] [PubMed]
- López-Igual, R.; Picossi, S.; López-Garrido, J.; Flores, E.; Herrero, A. N and C control of ABC-type bicarbonate transporter Cmp and its LysR-type transcriptional regulator CmpR in a heterocyst-forming cyanobacterium, Anabaena sp. Environ. Microbiol. 2012, 14, 1035–1048. [Google Scholar] [CrossRef] [PubMed]
- Brahamsha, B.; Haselkorn, R. Identification of multiple RNA polymerase sigma factor homologs in the cyanobacterium Anabaena sp. strain PCC 7120: Cloning, expression, and inactivation of the sigB and sigC genes. J. Bacteriol. 1992, 174, 7273–7282. [Google Scholar] [CrossRef]
- Caslake, L.F.; Gruber, T.M.; Bryant, N.A. Expression of two alternative sigma factors of Synechococcus sp. strain PCC 7002 is modulated by carbon and nitrogen stress. Microbiology 1997, 143, 3807–3818. [Google Scholar] [CrossRef]
- Muro-Pastor, A.M.; Herrero, A.; Flores, E. Nitrogen-Regulated Group 2 Sigma Factor from Synechocystis sp. Strain PCC 6803 Involved in Survival under Nitrogen Stress. J. Bacteriol. 2001, 183, 1090–1095. [Google Scholar] [CrossRef]
- Tuominen, I.; Pollari, M.; Tyystjärvi, E.; Tyystjärvi, T. The SigB σ factor mediates high-temperature responses in the cyanobacterium Synechocystis sp. PCC6803. FEBS Lett. 2006, 580, 319–323. [Google Scholar] [CrossRef]
- Hakkila, K.; Antal, T.; Gunnelius, L.; Kurkela, J.; Matthijs, H.C.; Tyystjärvi, E.; Tyystjärvi, T. Group 2 Sigma Factor Mutant sigCDE of the Cyanobacterium Synechocystis sp. PCC 6803 Reveals Functionality of Both Carotenoids and Flavodiiron Proteins in Photoprotection of Photosystem II. Plant Cell Physiol. 2013, 54, 1780–1790. [Google Scholar] [CrossRef]
- Summerfield, T.C.; Nagarajan, S.; Sherman, L.A. Gene expression under low-oxygen conditions in the cyanobacterium Synechocystis sp. PCC 6803 demonstrates Hik31-dependent and -independent responses. Microbiology 2011, 157, 301–312. [Google Scholar] [CrossRef]
- Nagarajan, S.; Sherman, D.M.; Shaw, I.; Sherman, L.A. Functions of the Duplicated hik31 Operons in Central Metabolism and Responses to Light, Dark, and Carbon Sources in Synechocystis sp. Strain PCC 6803. J. Bacteriol. 2011, 194, 448–459. [Google Scholar] [CrossRef]
- Jiang, Y.-L.; Wang, X.-P.; Sun, H.; Han, S.-J.; Li, W.-F.; Cui, N.; Lin, G.-M.; Zhang, J.-Y.; Cheng, W.; Cao, D.-D.; et al. Coordinating carbon and nitrogen metabolic signaling through the cyanobacterial global repressor NdhR. Proc. Natl. Acad. Sci. USA 2017, 115, 403–408. [Google Scholar] [CrossRef]
- Frías, J.E.; Flores, E.; Herrero, A. Activation of the Anabaena nir operon promoter requires both NtcA (CAP family) and NtcB (LysR family) transcription factors. Mol. Microbiol. 2000, 38, 613–625. [Google Scholar] [CrossRef] [PubMed]
- Liang, J.; Scappino, L.; Haselkorn, R. The patB gene product, required for growth of the cyanobacterium Anabaena sp. strain PCC 7120 under nitrogen-limiting conditions, contains ferredoxin and helix-turn-helix domains. J. Bacteriol. 1993, 175, 1697–1704. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Hebbar, P.B.; Curtis, S.E. Characterization of devH, a Gene Encoding a Putative DNA Binding Protein Required for Heterocyst Function in Anabaena sp. Strain PCC 7120. J. Bacteriol. 2000, 182, 3572–3581. [Google Scholar] [CrossRef] [PubMed]
- Jones, K.M.; Buikema, W.J.; Haselkorn, R. Heterocyst-Specific Expression of patB, a Gene Required for Nitrogen Fixation in Anabaena sp. Strain PCC 7120. J. Bacteriol. 2003, 185, 2306–2314. [Google Scholar] [CrossRef]
- Ehira, S.; Ohmori, M. NrrA, a nitrogen-responsive response regulator facilitates heterocyst development in the cyanobacterium Anabaena sp. strain PCC 7120. Mol. Microbiol. 2006, 59, 1692–1703. [Google Scholar] [CrossRef]
- Picossi, S.; Flores, E.; Herrero, A. ChIP analysis unravels an exceptionally wide distribution of DNA binding sites for the NtcA transcription factor in a heterocyst-forming cyanobacterium. BMC Genom. 2014, 15, 22. [Google Scholar] [CrossRef]
- Tsujimoto, R.; Kamiya, N.; Fujita, Y. Identification of a cis-acting element in nitrogen fixation genes recognized by CnfR in the nonheterocystous nitrogen-fixing cyanobacterium Leptolyngbya boryana. Mol. Microbiol. 2016, 101, 411–424. [Google Scholar] [CrossRef]
- Ramírez, M.E.; Hebbar, P.B.; Zhou, R.; Wolk, C.P.; Curtis, S.E. Anabaena sp. Strain PCC 7120 Gene devH Is Required for Synthesis of the Heterocyst Glycolipid Layer. J. Bacteriol. 2005, 187, 2326–2331. [Google Scholar] [CrossRef]
- Ehira, S.; Ohmori, M. NrrA directly regulates expression of the fraF gene and antisense RNAs for fraE in the heterocyst-forming cyanobacterium Anabaena sp. strain PCC 7120. Microbiology 2014, 160, 844–850. [Google Scholar] [CrossRef]
- Katoh, H.; Hagino, N.; Grossman, A.R.; Ogawa, T. Genes Essential to Iron Transport in the Cyanobacterium Synechocystis sp. Strain PCC 6803. J. Bacteriol. 2001, 183, 2779–2784. [Google Scholar] [CrossRef] [PubMed]
- Fresenborg, L.S.; Graf, J.; Schätzle, H.; Schleiff, E. Iron homeostasis of cyanobacteria: Advancements in siderophores and metal transporters. In Advances in Cyanobacterial Biology; Academic Press: Cambridge, MA, USA, 2020. [Google Scholar] [CrossRef]
- Shen, G.; Balasubramanian, R.; Wang, T.; Wu, Y.; Hoffart, L.M.; Krebs, C.; Bryant, N.A.; Golbeck, J.H. SufR Coordinates Two [4Fe-4S]2+, 1+ Clusters and Functions as a Transcriptional Repressor of the sufBCDS Operon and an Autoregulator of sufR in Cyanobacteria. J. Biol. Chem. 2007, 282, 31909–31919. [Google Scholar] [CrossRef] [PubMed]
- Shcolnick, S.; Summerfield, T.C.; Reytman, L.; Sherman, L.A.; Keren, N. The Mechanism of Iron Homeostasis in the Unicellular Cyanobacterium Synechocystis sp. PCC 6803 and Its Relationship to Oxidative Stress. Plant Physiol. 2009, 150, 2045–2056. [Google Scholar] [CrossRef]
- Vuorijoki, L.; Tiwari, A.; Kallio, P.T.; Aro, E.-M. Inactivation of iron-sulfur cluster biogenesis regulator SufR in Synechocystis sp. PCC 6803 induces unique iron-dependent protein-level responses. Biochim. Biophys. Acta Gen. Subj. 2017, 1861, 1085–1098. [Google Scholar] [CrossRef]
- Garcia-Dominguez, M.; López-Maury, L.; Florencio, F.J.; Reyes, J.C.; Braunstein, M.; Brown, A.M.; Kurtz, S.; Jacobs, W.R. A Gene Cluster Involved in Metal Homeostasis in the Cyanobacterium Synechocystis sp. Strain PCC 6803. J. Bacteriol. 2000, 182, 1507–1514. [Google Scholar] [CrossRef]
- Huertas, M.; López-Maury, L.; Giner-Lamia, J.; Riego, A.M.S.; Florencio, F.J. Metals in Cyanobacteria: Analysis of the Copper, Nickel, Cobalt and Arsenic Homeostasis Mechanisms. Life 2014, 4, 865–886. [Google Scholar] [CrossRef]
- Daley, S.M.E.; Kappell, A.D.; Carrick, M.J.; Burnap, R.L. Regulation of the Cyanobacterial CO2-Concentrating Mechanism Involves Internal Sensing of NADP+ and α-Ketogutarate Levels by Transcription Factor CcmR. PLoS ONE 2012, 7, e41286. [Google Scholar] [CrossRef]
- Eisenhut, M.; Huege, J.; Schwarz, D.; Bauwe, H.; Kopka, J.; Hagemann, M. Metabolome Phenotyping of Inorganic Carbon Limitation in Cells of the Wild Type and Photorespiratory Mutants of the Cyanobacterium Synechocystis sp. Strain PCC 6803. Plant Physiol. 2008, 148, 2109–2120. [Google Scholar] [CrossRef]
- Figge, R.M.; Cassier-Chauvat, C.; Chauvat, F.; Cerff, R. Characterization and analysis of an NAD(P)H dehydrogenase transcriptional regulator critical for the survival of cyanobacteria facing inorganic carbon starvation and osmotic stress. Mol. Microbiol. 2001, 39, 455–469. [Google Scholar] [CrossRef]
- Nishimura, T.; Takahashi, Y.; Yamaguchi, O.; Suzuki, H.; Maeda, S.-I.; Omata, T. Mechanism of low CO2-induced activation of the cmp bicarbonate transporter operon by a LysR family protein in the cyanobacterium Synechococcus elongatus strain PCC 7942. Mol. Microbiol. 2008, 68, 98–109. [Google Scholar] [CrossRef]
- Takahashi, Y.; Yamaguchi, O.; Omata, T. Roles of CmpR, a LysR family transcriptional regulator, in acclimation of the cyanobacterium Synechococcus sp. strain PCC 7942 to low-CO2 and high-light conditions. Mol. Microbiol. 2004, 52, 837–845. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, H.; Kitamura, M.; Nakano, Y.; Katayama, M.; Takahashi, Y.; Kondo, T.; Manabe, K.; Omata, T.; Kutsuna, S. CmpR is Important for Circadian Phasing and Cell Growth. Plant Cell Physiol. 2012, 53, 1561–1569. [Google Scholar] [CrossRef] [PubMed]
- Zhang, P.; Sicora, C.I.; Vorontsova, N.; Allahverdiyeva, Y.; Allahverdiyeva, Y.; Nixon, P.J.; Aro, E.-M. FtsH protease is required for induction of inorganic carbon acquisition complexes in Synechocystis sp. PCC 6803. Mol. Microbiol. 2007, 65, 728–740. [Google Scholar] [CrossRef] [PubMed]
- Domain, F.; Houot, L.; Chauvat, F.; Cassier-Chauvat, C. Function and regulation of the cyanobacterial genes lexA, recA and ruvB: LexA is critical to the survival of cells facing inorganic carbon starvation. Mol. Microbiol. 2004, 53, 65–80. [Google Scholar] [CrossRef] [PubMed]
- Lieman-Hurwitz, J.; Haimovich, M.; Shalev-Malul, G.; Ishii, A.; Hihara, Y.; Gaathon, A.; Lebendiker, M.; Kaplan, A. A cyanobacterial AbrB-like protein affects the apparent photosynthetic affinity for CO2 by modulating low-CO2-induced gene expression. Environ. Microbiol. 2009, 11, 927–936. [Google Scholar] [CrossRef]
- Kaniya, Y.; Kizawa, A.; Miyagi, A.; Kawai-Yamada, M.; Uchimiya, H.; Kaneko, Y.; Nishiyama, Y.; Hihara, Y. Deletion of the Transcriptional Regulator cyAbrB2 Deregulates Primary Carbon Metabolism in Synechocystis sp. PCC 68031[W]. Plant Physiol. 2013, 162, 1153–1163. [Google Scholar] [CrossRef]
- Orf, I.; Schwarz, D.; Kaplan, A.; Kopka, J.; Hess, W.R.; Hagemann, M.; Klähn, S. CyAbrB2 Contributes to the Transcriptional Regulation of Low CO2 Acclimation in Synechocystis sp. PCC 6803. Plant Cell Physiol. 2016, 57, 2232–2243. [Google Scholar] [CrossRef][Green Version]
- Sjöholm, J.; Oliveira, P.; Lindblad, P. Transcription and Regulation of the Bidirectional Hydrogenase in the Cyanobacterium Nostoc sp. Strain PCC 7120. Appl. Environ. Microbiol. 2007, 73, 5435–5446. [Google Scholar] [CrossRef]
- Oliveira, P.; Lindblad, P. Transcriptional regulation of the cyanobacterial bidirectional Hox-hydrogenase. Dalton Trans. 2009, 9990–9996. [Google Scholar] [CrossRef]
- Ishii, A.; Hihara, Y. An AbrB-Like Transcriptional Regulator, Sll0822, Is Essential for the Activation of Nitrogen-Regulated Genes in Synechocystis sp. PCC 6803. Plant Physiol. 2008, 148, 660–670. [Google Scholar] [CrossRef]
- Agervald, Å.; Baebprasert, W.; Zhang, X.; Incharoensakdi, A.; Lindblad, P.; Stensjö, K. The CyAbrB transcription factor CalA regulates the iron superoxide dismutase in Nostoc sp. strain PCC 7120. Environ. Microbiol. 2010, 12, 2826–2837. [Google Scholar] [CrossRef] [PubMed]
- Higo, A.; Nishiyama, E.; Nakamura, K.; Hihara, Y.; Ehira, S. cyAbrB Transcriptional Regulators as Safety Devices To Inhibit Heterocyst Differentiation in Anabaena sp. Strain PCC 7120. J. Bacteriol. 2019, 201. [Google Scholar] [CrossRef] [PubMed]
- Tanigawa, R.; Shirokane, M.; Maeda, S.-I.; Omata, T.; Tanaka, K.; Takahashi, H. Transcriptional activation of NtcA-dependent promoters of Synechococcus sp. PCC 7942 by 2-oxoglutarate in vitro. Proc. Natl. Acad. Sci. USA 2002, 99, 4251–4255. [Google Scholar] [CrossRef]
- Zhao, M.-X.; Jiang, Y.-L.; He, Y.-X.; Chen, Y.-F.; Teng, Y.-B.; Chen, Y.; Zhang, C.-C.; Zhou, C.-Z. Structural basis for the allosteric control of the global transcription factor NtcA by the nitrogen starvation signal 2-oxoglutarate. Proc. Natl. Acad. Sci. USA 2010, 107, 12487–12492. [Google Scholar] [CrossRef]
- Espinosa, J.; Forchhammer, K.; Burillo, S.; Contreras, A. Interaction network in cyanobacterial nitrogen regulation: PipX, a protein that interacts in a 2-oxoglutarate dependent manner with PII and NtcA. Mol. Microbiol. 2006, 61, 457–469. [Google Scholar] [CrossRef] [PubMed]
- Ohashi, Y.; Shi, W.; Takatani, N.; Aichi, M.; Maeda, S.-I.; Watanabe, S.; Yoshikawa, H.; Omata, T. Regulation of nitrate assimilation in cyanobacteria. J. Exp. Bot. 2011, 62, 1411–1424. [Google Scholar] [CrossRef]
- Su, Z.; Olman, V.; Mao, F.; Xu, Y. Comparative genomics analysis of NtcA regulons in cyanobacteria: Regulation of nitrogen assimilation and its coupling to photosynthesis. Nucleic Acids Res. 2005, 33, 5156–5171. [Google Scholar] [CrossRef]
- Mitschke, J.; Vioque, A.; Haas, F.; Hess, W.R.; Muro-Pastor, A.M. Dynamics of transcriptional start site selection during nitrogen stress-induced cell differentiation in Anabaena sp. PCC7120. Proc. Natl. Acad. Sci. USA 2011, 108, 20130–20135. [Google Scholar] [CrossRef]
- Galmozzi, C.V.; Saelices, L.; Florencio, F.J.; Muro-Pastor, M.I. Posttranscriptional Regulation of Glutamine Synthetase in the Filamentous Cyanobacterium Anabaena sp. PCC 7120: Differential Expression between Vegetative Cells and Heterocysts. J. Bacteriol. 2010, 192, 4701–4711. [Google Scholar] [CrossRef]
- Kang, R.-J.; Shi, D.-J.; Cong, W.; Cai, Z.-L.; Ouyang, F. Regulation of CO2 on heterocyst differentiation and nitrate uptake in the cyanobacterium Anabaena sp. PCC 7120. J. Appl. Microbiol. 2005, 98, 693–698. [Google Scholar] [CrossRef]
- Flores, E.; Arévalo, S.; Burnat, M. Cyanophycin and arginine metabolism in cyanobacteria. Algal Res. 2019, 42, 101577. [Google Scholar] [CrossRef]
- Lopez-Gomollon, S.; Hernández, J.A.; Pellicer, S.; Angarica, V.E.; Peleato, M.L.; Fillat, M.F. Cross-talk Between Iron and Nitrogen Regulatory Networks in Anabaena (Nostoc) sp. PCC 7120: Identification of Overlapping Genes in FurA and NtcA Regulons. J. Mol. Biol. 2007, 374, 267–281. [Google Scholar] [CrossRef] [PubMed]
- Raven, J.A.; Evans, M.C.W.; Korb, R.E. The role of trace metals in photosynthetic electron transport in O2-evolving organisms. Photosynth. Res. 1999, 60, 111–150. [Google Scholar] [CrossRef]
- Gupta, J.K.; Rai, P.; Jain, K.K.; Srivastava, S. Overexpression of bicarbonate transporters in the marine cyanobacterium Synechococcus sp. PCC 7002 increases growth rate and glycogen accumulation. Biotechnol. Biofuels 2020, 13, 17. [Google Scholar] [CrossRef] [PubMed]
- Cano, M.; Holland, S.C.; Artier, J.; Burnap, R.L.; Ghirardi, M.; Morgan, J.A.; Yu, J. Glycogen Synthesis and Metabolite Overflow Contribute to Energy Balancing in Cyanobacteria. Cell Rep. 2018, 23, 667–672. [Google Scholar] [CrossRef]
- Schwarz, D.; Orf, I.; Kopka, J.; Hagemann, M. Effects of Inorganic Carbon Limitation on the Metabolome of the Synechocystis sp. PCC 6803 Mutant Defective in glnB Encoding the Central Regulator PII of Cyanobacterial C/N Acclimation. Metabolites 2014, 4, 232–247. [Google Scholar] [CrossRef]
- Vermaas, W.F. Photosynthesis and Respiration in Cyanobacteria. In Encyclopedia of Life Sciences; John Wiley & Sons, Ltd.: Chichester, UK, 2001. [Google Scholar] [CrossRef]
- Welkie, D.G.; Rubin, B.E.; Diamond, S.; Hood, R.D.; Savage, D.F.; Golden, S.S. A Hard Day’s Night: Cyanobacteria in Diel Cycles. Trends Microbiol. 2019, 27, 231–242. [Google Scholar] [CrossRef]
- Makowka, A.; Nichelmann, L.; Schulze, D.; Spengler, K.; Wittmann, C.; Forchhammer, K.; Gutekunst, K. Glycolytic Shunts Replenish the Calvin–Benson–Bassham Cycle as Anaplerotic Reactions in Cyanobacteria. Mol. Plant 2020, 13, 471–482. [Google Scholar] [CrossRef]
- Huege, J.; Goetze, J.; Schwarz, D.; Bauwe, H.; Hagemann, M.; Kopka, J. Modulation of the Major Paths of Carbon in Photorespiratory Mutants of Synechocystis. PLoS ONE 2011, 6, e16278. [Google Scholar] [CrossRef]
- Tang, J.K.-H.; Tang, Y.J.; Blankenship, R.E. Carbon Metabolic Pathways in Phototrophic Bacteria and Their Broader Evolutionary Implications. Front. Microbiol. 2011, 2, 165. [Google Scholar] [CrossRef]
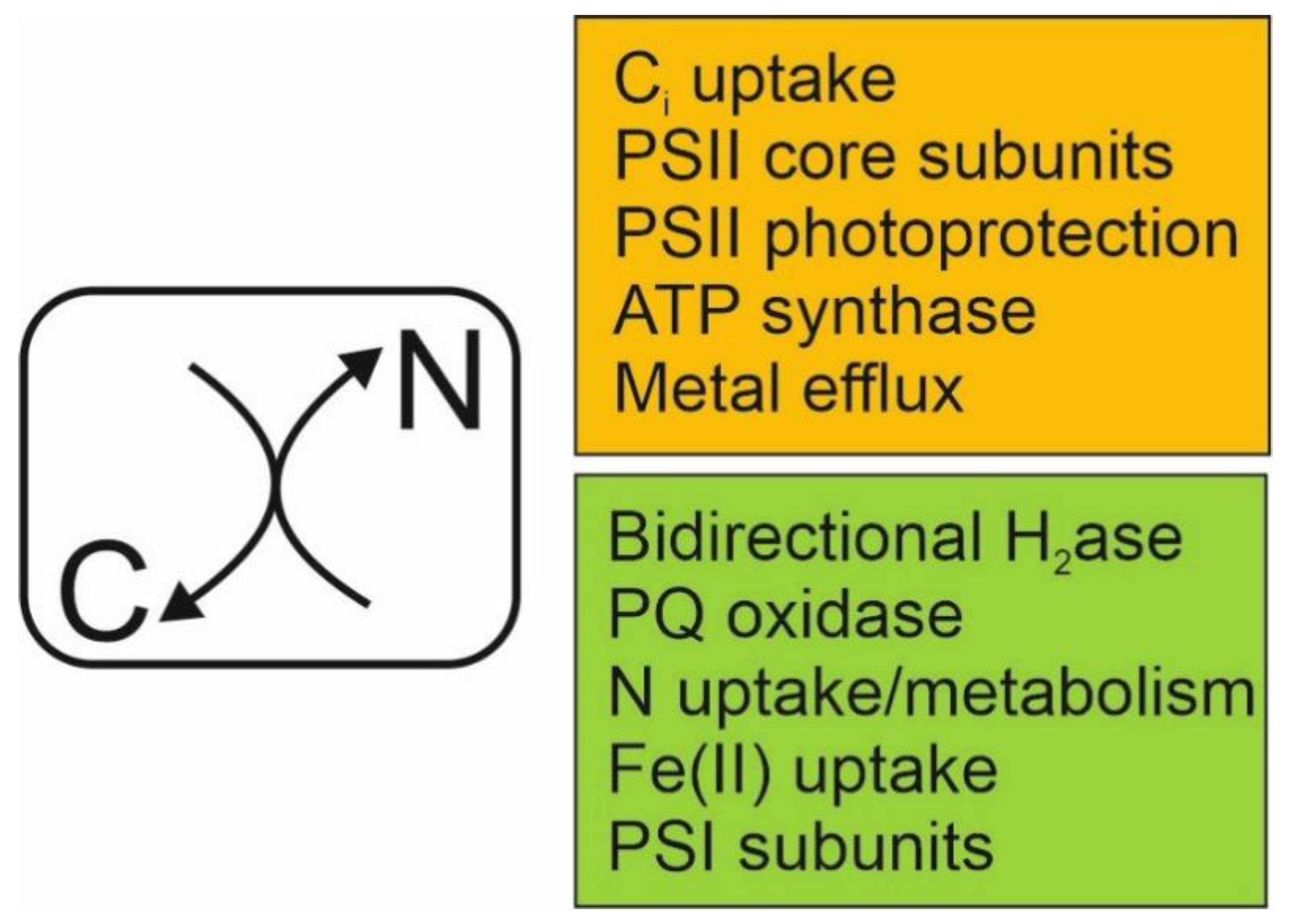
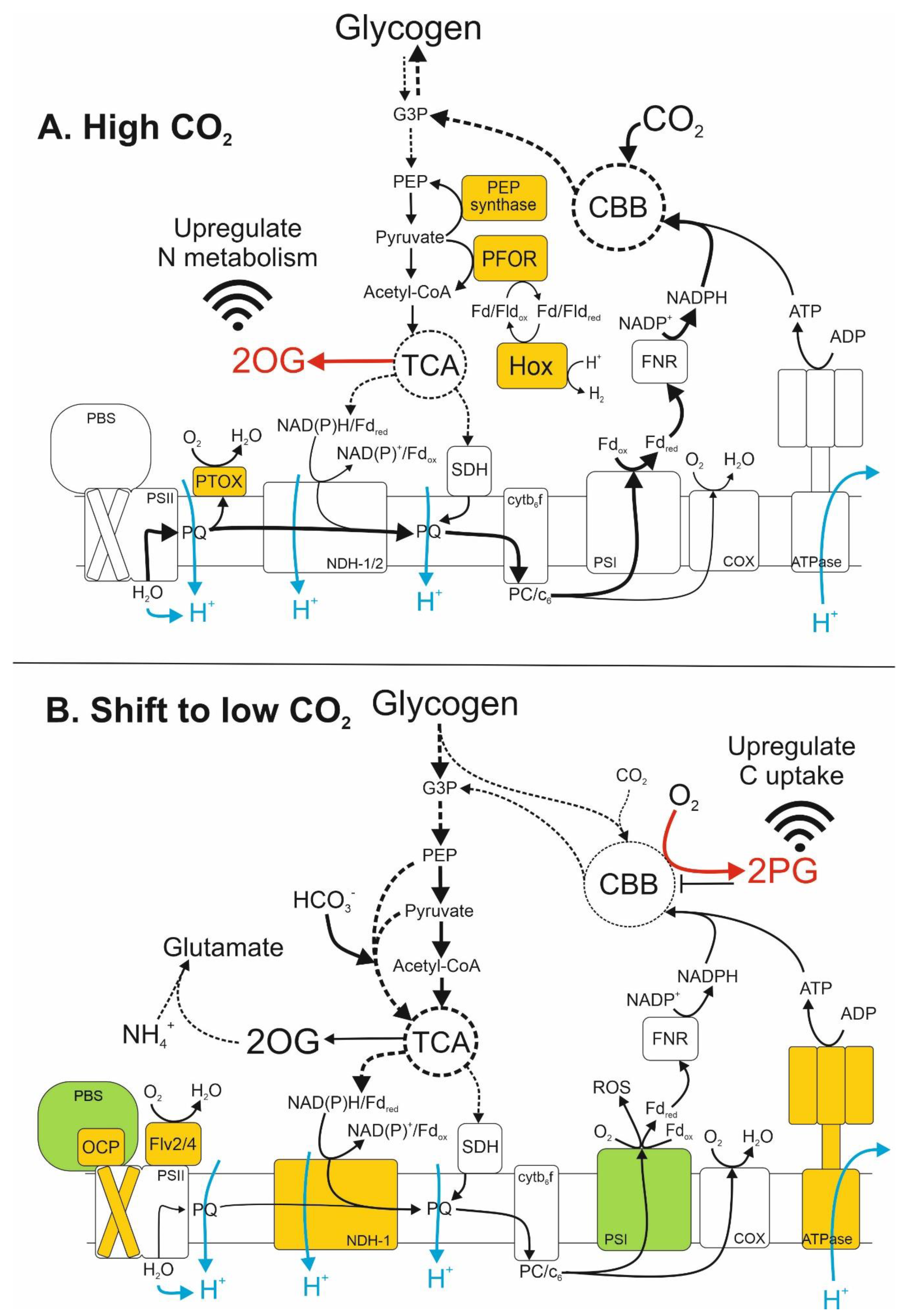
| Name 1 | Gene ID 1 | Description | Process | Fold Change 2 | p-Value 3 |
|---|---|---|---|---|---|
| sbt operon | all2133–all2134 | Na+-dependent bicarbonate permease, PII-like regulatory protein | Bicarbonate import | 78.8 | <0.001 |
| cmp operon | alr2877–alr2880 | ATP-dependent bicarbonate uptake subunits | 36.4 | <0.001 | |
| bicA operon | all1303–all1304 | Na+-dependent bicarbonate permease, Na+:H+ antiporter | 8.7 | <0.001 | |
| mrp operon | all1837–all1843 | Na+:H+ antiporter subunits | Na+ extrusion, pH regulation | 5.4 | <0.001 |
| ndhF3 | alr4156 | NDH–1MS subunit 5 | CO2 uptake | 2.5 | 0.002 |
| ndhD3 | alr4157 | NDH–1MS subunit 4 | 2.0 | <0.001 | |
| cupA | alr4158 | NDH–1MS CO2 uptake subunit | 5.3 | <0.001 | |
| nir operon | alr0607–alr0612 | Nitrate/nitrite reductase, ATP-dependent nitrate permease | Nitrate/nitrite import and metabolism | −3.5 | 0.017 |
| nirB | all0605 | Nitrate-dependent expression of nir cluster | −2.9 | <0.001 | |
| nifB | all1517 | Fe–Mo cofactor biosynthesis subunit | N2 fixation, heterocyst development and function | −2.0 | 0.002 |
| nifH2 | alr0874 | Fe–S cluster-binding nitrogenase reductase | −3.2 | 0.024 | |
| hgd, hgl clusters | all5341–alr5359 | Heterocyst glycolipid layer biosynthesis | −2.6 | 0.004 | |
| dev operon | alr3710–alr3712 | ATP-binding subunit, membrane transport subunits | −2.1 | 0.004 | |
| alr1713 | alr1713 | Similar to Mo-dependent nitrogenase, C-terminus | Unknown | −5.6 | 0.002 |
| asr1714 | asr1714 | Uncharacterised protein | −5.8 | <0.001 |
| Name 1 | Gene ID 1 | Description | Process | Fold Change 2 | p Value 3 |
|---|---|---|---|---|---|
| psbAII | alr3727 | Photosystem II D1 protein | PSII electron transport | 6.9 | <0.001 |
| psbAIII | alr4592 | Photosystem II D1 protein | 1.9 | 0.003 | |
| psbAIV | all3572 | Photosystem II D1 protein | 3.7 | <0.001 | |
| psbD | alr4548 | Photosystem II D2 protein | 3.4 | <0.001 | |
| psaA | alr5154 | Photosystem I core protein A1 | PSI electron transport | −1.9 | 0.035 |
| psaB1 | alr5155 | Photosystem I core protein A2 | −1.9 | 0.038 | |
| psaB2 | alr5314 | Photosystem I core protein A2 | −2.2 | 0.031 | |
| psaC | asr3463 | Photosystem I Fe-S subunit | −2.2 | <0.001 | |
| psaD | all0329 | Photosystem I reaction centre subunit 2 | −2.7 | <0.001 | |
| psaE | asr4319 | Photosystem I subunit E | −2.2 | 0.001 | |
| psaI | asl3849 | Photosystem I subunit I | −2.5 | 0.012 | |
| pasK | asr4775 | Photosystem I subunit K | −3.1 | 0.004 | |
| psaM | asr4657 | Photosystem I subunit M | −2.3 | 0.005 | |
| flv2 | all4444 | Flavodiiron protein | Other photosynthetic/respiratory electron transport | 23.7 | <0.001 |
| all4445 | all4445 | Unknown protein | 31.8 | <0.001 | |
| flv4 | all4446 | Flavodiiron protein | 16.7 | <0.001 | |
| isiB | alr2405 | Flavodoxin | 2.9 | 0.003 | |
| cytA | alr4251 | Cytochrome c6 | −2.2 | 0.011 | |
| flv1B-flv3B | all0177–all0178 | Flavodiiron protein (heterocyst-specific) | −2.0 | 0.004 | |
| ptox | all2096 | Alternative plastoquinone oxidase | −65.0 | <0.001 | |
| ftsH | alr1261 | FtsH protease | PSII turnover | 2.3 | <0.001 |
| ftsH2 | all3642 | FtsH protease | 2.4 | <0.001 | |
| pec operon | alr0523–alr0527 | Phycoerythrocyanin synthesis | Metabolism/binding of light-harvesting pigments | −3.1 | 0.015 |
| chlL, chlN operon | all5076–all5078 | Protochlorophyllide reductase, ATP-binding protein | −3.3 | <0.001 | |
| chlG | all4480 | Chlorophyll synthase 33 kDa subunit | 2.2 | 0.003 | |
| hemH | alr4616 | Ferrochelatase | 8.6 | <0.001 | |
| ocp | all3149 | Orange carotenoid-binding protein | 23.9 | <0.001 | |
| ocp-like | all4941 | Orange carotenoid protein-like | 3.1 | 0.010 | |
| asl3726 | asl3726 | CAB/ELIP/HLIP superfamily | 9.8 | <0.001 | |
| asr5262 | asr5262 | CAB/ELIP/HLIP superfamily | 8.6 | 0.005 | |
| atpase cluster | all0004–all0010 | ATP synthase subunits | ATP synthesis | 2.7 | <0.001 |
| ndh-1 operon | alr0223–alr0227 | NDH-1 complex subunits | Electron and proton transport | 2.3 | 0.005 |
| ndh-1 operon | all3840–all3842 | 2.1 | 0.001 | ||
| ndhB | all4883 | 2.5 | 0.003 | ||
| ndhN | alr4216 | 1.9 | 0.001 | ||
| alr1004 | alr1004 | Alanine-glyoxylate transaminase | Glycolate metabolim | −2.7 | <0.001 |
| ndbA | all1553 | NDH-2 NAD(P)H:PQ reductase | Respiration | −2.1 | <0.001 |
| hupS | all0688 | Uptake hydrogenase, small subunit | H2 uptake/evolution | −2.5 | 0.002 |
| nifJ/PFOR | alr1911 | Pyruvate-ferredoxin/flavodoxin oxidoreductase | −42.4 | <0.001 | |
| hox clusters | alr0750–all0752alr0760–alr0766 | Bidirectional hydrogenase subunits, assembly and regulation | −73.9 | 0.002 | |
| ppsA | all0635 | Phophoenolpyruvate synthase | Glycolysis | −107.4 | <0.001 |
| Name | Gene ID | Description | Process | Fold Change 1 | p Value 2 |
|---|---|---|---|---|---|
| cmpR | all0862 | LysR-type transcriptional regulator | Regulates cmp cluster | 3.1 | <0.001 |
| sigB | all7615 | Group 2 sigma factor | Response to stress | 4.6 | <0.001 |
| sigB3 | all7608 | Group 2 sigma factor | 3.8 | <0.001 | |
| C-hik31 operon | all7583–all7584 | Two-component sensor His kinase, response regulator | Regulation of central metabolism in response to glucose, low O2 | 2.8 | <0.001 |
| P-hik31 operon | alr1170–alr1171 | 1.6 | 0.001 | ||
| all7523 | all7523 | TetR family regulator | Unknown | 2.4 | 0.046 |
| putative ndhR orthologue | all4986 | LysR-type transcriptional regulator | Repression of CCM expression | −146.2 | <0.001 |
| ntcB | all0602 | LysR-type transcriptional regulator | Co-activation of nir operon | −2.4 | 0.002 |
| devH | alr3952 | CRP family transcriptional regulator | Het glycolipid biosynthesis | −2.2 | 0.006 |
| patB/cnfR | all2512 | Heterocyst patterning | Heterocyst development | −2.4 | 0.001 |
| nrrA | all4312 | OmpR family regulator | −2.1 | <0.001 |
| Name | Gene ID | Description | Process | Fold Change 1 | p Value 2 |
|---|---|---|---|---|---|
| Fe(II) transport operon | alr2118–asr2120 | Ferrous iron transporter subunits | Periplasmic iron import | −42.6 | <0.001 |
| suf operon | alr2492–alr2496 | ATPase, iron and sulphur transfer | Fe–S cluster assembly, transfer | −3.8 | <0.001 |
| Metal efflux cluster | all7606–all7611 | Proton extrusion, cation efflux | Metal cation efflux, cellular metal homeostasis | 3.1 | <0.001 |
| Metal efflux cluster | all7616–all7619 | Cadmium/nickel/zinc/cobalt efflux system | 3.9 | <0.001 | |
| Metal efflux cluster | all7629–all7633 | Cadmium/nickel/zinc/cobalt efflux system | 5.0 | <0.001 | |
| Cu2+ efflux cluster | alr7634–alr7636 | Putative copper efflux | 5.1 | 0.002 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gollan, P.J.; Muth-Pawlak, D.; Aro, E.-M. Rapid Transcriptional Reprogramming Triggered by Alteration of the Carbon/Nitrogen Balance Has an Impact on Energy Metabolism in Nostoc sp. PCC 7120. Life 2020, 10, 297. https://doi.org/10.3390/life10110297
Gollan PJ, Muth-Pawlak D, Aro E-M. Rapid Transcriptional Reprogramming Triggered by Alteration of the Carbon/Nitrogen Balance Has an Impact on Energy Metabolism in Nostoc sp. PCC 7120. Life. 2020; 10(11):297. https://doi.org/10.3390/life10110297
Chicago/Turabian StyleGollan, Peter J., Dorota Muth-Pawlak, and Eva-Mari Aro. 2020. "Rapid Transcriptional Reprogramming Triggered by Alteration of the Carbon/Nitrogen Balance Has an Impact on Energy Metabolism in Nostoc sp. PCC 7120" Life 10, no. 11: 297. https://doi.org/10.3390/life10110297
APA StyleGollan, P. J., Muth-Pawlak, D., & Aro, E.-M. (2020). Rapid Transcriptional Reprogramming Triggered by Alteration of the Carbon/Nitrogen Balance Has an Impact on Energy Metabolism in Nostoc sp. PCC 7120. Life, 10(11), 297. https://doi.org/10.3390/life10110297





