An Intelligent Technique for Initial Distribution of Genetic Algorithms
Abstract
:1. Introduction
2. The Proposed Method
- Initialization step:
- (a)
- Set as the number of chromosomes.
- (b)
- Set as the maximum number of allowed generations.
- (c)
- Initialize randomly the chromosomes in S. In most implementations of genetic algorithms, the chromosomes will be selected using some random number distribution. In the present work, the chromosomes will be selected using the sampling technique described in Section 2.3.
- (d)
- Set as the selection rate of the algorithm, with .
- (e)
- Set as the mutation rate, with .
- (f)
- Set iter = 0.
- For every chromosome : Calculate the fitness of chromosome .
- Genetic operations step:
- (a)
- Selection procedure: The chromosomes are sorted according to their fitness values. Denote as the integer part of ; chromosomes with the lowest fitness values are transferred intact to the next generation. The remaining chromosomes are substituted by offspring created in the crossover procedure. During the selection process, for each offspring, two parents are selected from the population using tournament selection.
- (b)
- Crossover procedure: For every pair of selected parents, two additional chromosomes and are produced using the following equations:where . The values are uniformly distributed random numbers, with [60].
- (c)
- Replacement procedure:
- i.
- For to , do:
- Replace using the next offspring created in the crossover procedure.
- ii.
- EndFor:
- (d)
- Mutation procedure:
- i.
- For every chromosome , do:
- For each element of , a uniformly distributed random number is drawn. The element is altered randomly if .
- ii.
- EndFor
- Termination check step:
- (a)
- Set .
- (b)
- If or the proposed stopping rule of Tsoulos [61] holds, then goto the local search step, else goto Step 2.
- Local search step: Apply a local search procedure to the chromosome of the population with the lowest fitness value, and report the obtained minimum. In the current work, the BFGS variant of Powell [62] was used as a local search procedure.
2.1. Proposed Initialization Distribution
2.2. Chromosome Rejection Rule
2.3. The Proposed Sampling Procedure
- Take random samples from the objective function using a uniform distribution.
- Calculate the k centers of the points using the k-means algorithm provided in Algorithm 1.
- Remove from the set of centers C points that are close to each other.
- Return the set of centers C as the set of chromosomes.
| Algorithm 1 The k-means algorithm. |
|
3. Experiments
3.1. Test Functions
- Bohachevsky 1 (Bf1) function:with .
- Bohachevsky 2 (Bf2) function:with .
- Branin function: with .
- CM function:where . In the experiments conducted, the value was used.
- Camel function:
- Easom function:with
- Exponential function, defined as:The values were used in the executed experiments.
- Griewank2 function:
- Griewank10 function: The function is given by the equation:with .
- Gkls function: is a function with w local minima, described in [74] with , and n is a positive integer between 2 and 100. The values and were used in the experiments conducted.
- Goldstein and Price function:with .
- Hansen function: , .
- Hartman 3 function:with and and
- Hartman 6 function:with and and
- Potential function: The molecular conformation corresponding to the global minimum of the energy of N atoms interacting via the Lennard–Jones potential [75] was used as a test function here, and it is defined by:The values were used in the experiments conducted. Also, for the experiments conducted, the values 1 were used.
- Rastrigin function:
- Rosenbrock function:The values were used in the experiments conducted.
- Shekel 5 function:with and
- Shekel 7 function:with and .
- Shekel 10 function:with and .
- Sinusoidal function:The values of and were used in the experiments conducted.
- Test2N function:The function has in the specified range, and in our experiments, we used .
- Test30N function:with , with the local minima in the search space. For our experiments, we used .
3.2. Experimental Results
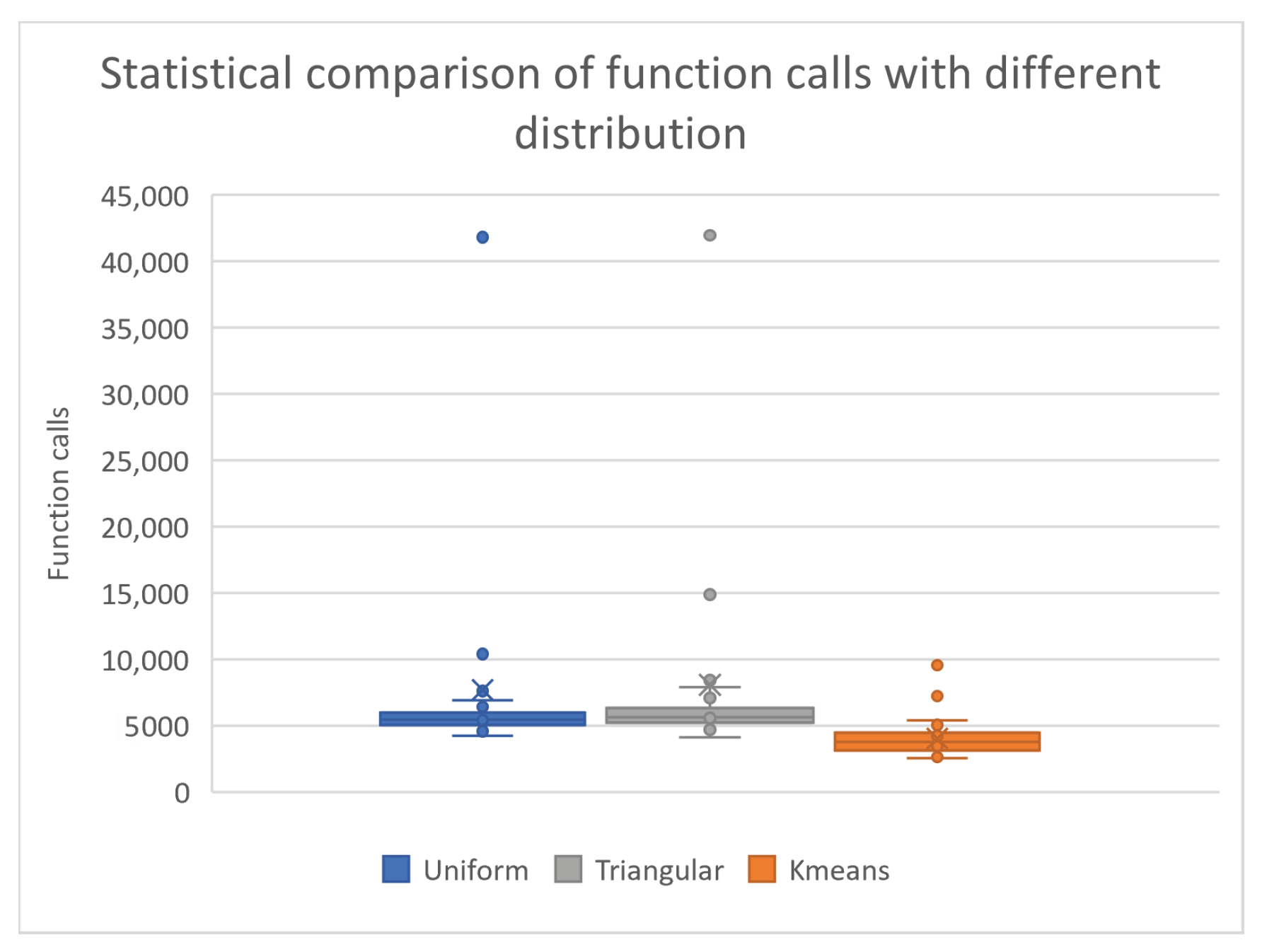
- The column UNIFORM indicates the incorporation of uniform sampling in the genetic algorithm. In this case, randomly selected chromosomes using uniform sampling were used in the genetic algorithm.
- The column TRIANGULAR defines the usage of the triangular distribution [76] for the initial samples of the genetic algorithm. For this case, randomly selected chromosomes with a triangular distribution were used in the genetic algorithm.
- The column KMEANS denotes the application of k-means sampling as proposed here in the genetic algorithm. In this case, randomly selected points were sampled from the objective function and k centers were produced using the k-means algorithm. In order to have a fair comparison between the results produced between the proposed technique and the rest, the number of centers produced by the k-means method was set to be equal to the number of chromosomes of the rest of the techniques. Ten-times the number of initial points were used to produce the centers. In addition, through the discard process of Algorithm 2, some centers were eliminated.
- The numbers in the cells represent the average number of function calls required to obtain the global minimum. The fraction in parentheses denotes the percentage where the global minimum was successfully discovered. If this fraction is absent, then the global minimum was successfully discovered in all runs.
- In every table, an additional line was added under the name TOTAL, representing the total number of function calls and, in parentheses, the average success rate in finding the global minimum.
| Algorithm 2 Chromosome rejection rule. |
|
| Parameter | Meaning | Value |
|---|---|---|
| Number of chromosomes | 200 | |
| Initial samples for k-means | 2000 | |
| k | Number of centers in k-means | 200 |
| Maximum number of allowed generations | 200 | |
| Selection rate | 0.9 | |
| Mutation rate | 0.05 | |
| Small value used in comparisons |
- High conditioned elliptic function, defined as
- CM function, defined as
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Yang, L.; Robin, D.; Sannibale, F.; Steier, C.; Wan, W. Global optimization of an accelerator lattice using multiobjective genetic algorithms. Nucl. Instruments Methods Phys. Res. Sect. A Accel. Spectrom. Detect. Assoc. Equip. 2009, 609, 50–57. [Google Scholar] [CrossRef]
- Iuliano, E. Global optimization of benchmark aerodynamic cases using physics-based surrogate models. Aerosp. Sci. Technol. 2017, 67, 273–286. [Google Scholar] [CrossRef]
- Duan, Q.; Sorooshian, S.; Gupta, V. Effective and efficient global optimization for conceptual rainfall-runoff models. Water Resour. Res. 1992, 28, 1015–1031. [Google Scholar] [CrossRef]
- Heiles, S.; Johnston, R.L. Global optimization of clusters using electronic structure methods. Int. J. Quantum Chem. 2013, 113, 2091–2109. [Google Scholar] [CrossRef]
- Shin, W.H.; Kim, J.K.; Kim, D.S.; Seok, C. GalaxyDock2: Protein–ligand docking using beta-complex and global optimization. J. Comput. Chem. 2013, 34, 2647–2656. [Google Scholar] [CrossRef]
- Liwo, A.; Lee, J.; Ripoll, D.R.; Pillardy, J.; Scheraga, H.A. Protein structure prediction by global optimization of a potential energy function. Biophysics 1999, 96, 5482–5485. [Google Scholar] [CrossRef]
- Gaing, Z.-L. Particle swarm optimization to solving the economic dispatch considering the generator constraints. IEEE Trans. Power Syst. 2003, 18, 1187–1195. [Google Scholar] [CrossRef]
- Maranas, C.D.; Androulakis, I.P.; Floudas, C.A.; Berger, A.J.; Mulvey, J.M. Solving long-term financial planning problems via global optimization. J. Econ. Dyn. Control. 1997, 21, 1405–1425. [Google Scholar] [CrossRef]
- Lee, E.K. Large-Scale Optimization-Based Classification Models in Medicine and Biology. Ann. Biomed. Eng. 2007, 35, 1095–1109. [Google Scholar] [CrossRef]
- Cherruault, Y. Global optimization in biology and medicine. Math. Comput. Model. 1994, 20, 119–132. [Google Scholar] [CrossRef]
- Wolfe, M.A. Interval methods for global optimization. Appl. Math. Comput. 1996, 75, 179–206. [Google Scholar]
- Csendes, T.; Ratz, D. Subdivision Direction Selection in Interval Methods for Global Optimization. SIAM J. Numer. Anal. 1997, 34, 922–938. [Google Scholar] [CrossRef]
- Price, W.L. Global optimization by controlled random search. J. Optim. Theory Appl. 1983, 40, 33–348. [Google Scholar] [CrossRef]
- Krivy, I.; Tvrdik, J. The controlled random search algorithm in optimizing regression models. Comput. Stat. Data Anal. 1995, 20, 229–234. [Google Scholar] [CrossRef]
- Ali, M.M.; Torn, A.; Viitanen, S. A Numerical Comparison of Some Modified Controlled Random Search Algorithms. J. Glob. Optim. 1997, 11, 377–385. [Google Scholar] [CrossRef]
- Kirkpatrick, S.; Gelatt, C.D.; Vecchi, M.P. Optimization by simulated annealing. Science 1983, 220, 671–680. [Google Scholar] [CrossRef]
- Ingber, L. Very fast simulated re-annealing. Math. Comput. Model. 1989, 12, 967–973. [Google Scholar] [CrossRef]
- Eglese, R.W. Simulated annealing: A tool for operational research. Eur. J. Oper. Res. 1990, 46, 271–281. [Google Scholar] [CrossRef]
- Storn, R.; Price, K. Differential Evolution—A Simple and Efficient Heuristic for Global Optimization over Continuous Spaces. J. Glob. Optim. 1997, 11, 341–359. [Google Scholar] [CrossRef]
- Liu, J.; Lampinen, J. A Fuzzy Adaptive Differential Evolution Algorithm. Soft Comput. 2005, 9, 448–462. [Google Scholar] [CrossRef]
- Kennedy, J.; Eberhart, R. Particle swarm optimization. In Proceedings of the ICNN’95—International Conference on Neural Networks, Perth, WA, Australia, 27 November–1 December 1995; Volume 4, pp. 1942–1948. [Google Scholar] [CrossRef]
- Poli, R.; Kennedy, J.K.; Blackwell, T. Particle swarm optimization An Overview. Swarm Intell. 2007, 1, 33–57. [Google Scholar] [CrossRef]
- Trelea, I.C. The particle swarm optimization algorithm: Convergence analysis and parameter selection. Inf. Process. Lett. 2003, 85, 317–325. [Google Scholar] [CrossRef]
- Dorigo, M.; Birattari, M.; Stutzle, T. Ant colony optimization. IEEE Comput. Intell. Mag. 2006, 1, 28–39. [Google Scholar] [CrossRef]
- Socha, K.; Dorigo, M. Ant colony optimization for continuous domains. Eur. J. Oper. Res. 2008, 185, 1155–1173. [Google Scholar] [CrossRef]
- Perez, M.; Almeida, F.; Moreno-Vega, J.M. Genetic algorithm with multistart search for the p-Hub median problem. In Proceedings of the 24th EUROMICRO Conference (Cat. No.98EX204), Vasteras, Sweden, 27 August 1998; Volume 2, pp. 702–707. [Google Scholar]
- De Oliveira, H.C.B.; Vasconcelos, G.C.; Alvarenga, G.B. A Multi-Start Simulated Annealing Algorithm for the Vehicle Routing Problem with Time Windows. In Proceedings of the 2006 Ninth Brazilian Symposium on Neural Networks (SBRN’06), Ribeirao Preto, Brazil, 23–27 October 2006; pp. 137–142. [Google Scholar]
- Liu, B.; Wang, L.; Jin, Y.H.; Tang, F.; Huang, D.X. Improved particle swarm optimization combined with chaos. Chaos Solitons Fractals 2005, 25, 1261–1271. [Google Scholar] [CrossRef]
- Shi, X.H.; Liang, Y.C.; Lee, H.P.; Lu, C.; Wang, L.M. An improved GA and a novel PSO-GA based hybrid algorithm. Inf. Process. Lett. 2005, 93, 255–261. [Google Scholar] [CrossRef]
- Garg, H. A hybrid PSO-GA algorithm for constrained optimization problems. Appl. Math. Comput. 2016, 274, 292–305. [Google Scholar] [CrossRef]
- Larson, J.; Wild, S.M. Asynchronously parallel optimization solver for finding multiple minima. Math. Program. Comput. 2018, 10, 303–332. [Google Scholar] [CrossRef]
- Bolton, H.P.J.; Schutte, J.F.; Groenwold, A.A. Multiple Parallel Local Searches in Global Optimization. In Recent Advances in Parallel Virtual Machine and Message Passing Interface, Proceedings of the EuroPVM/MPI 2000, Balatonfured, Hungary, 10–13 September 2000; Lecture Notes in Computer Science; Dongarra, J., Kacsuk, P., Podhorszki, N., Eds.; Springer: Berlin/Heidelberg, Germany, 2000; Volume 1908, p. 1908. [Google Scholar]
- Kamil, R.; Reiji, S. An Efficient GPU Implementation of a Multi-Start TSP Solver for Large Problem Instances. In Proceedings of the 14th Annual Conference Companion on Genetic and Evolutionary Computation, Philadelphia, PA, USA, 7–11 July 2012; pp. 1441–1442. [Google Scholar]
- Van Luong, T.; Melab, N.; Talbi, E.G. GPU-Based Multi-start Local Search Algorithms. In Learning and Intelligent Optimization. LION 2011; Lecture Notes in Computer Science; Coello, C.A.C., Ed.; Springer: Berlin/Heidelberg, Germany, 2011; Volume 6683. [Google Scholar] [CrossRef]
- Holland, J.H. Genetic algorithms. Sci. Am. 1992, 267, 66–73. [Google Scholar] [CrossRef]
- Stender, J. Parallel Genetic Algorithms: Theory & Applications; IOS Press: Amsterdam, The Netherlands, 1993. [Google Scholar]
- Goldberg, D. Genetic Algorithms in Search, Optimization and Machine Learning; Addison-Wesley Publishing Company: Reading, MA, USA, 1989. [Google Scholar]
- Michaelewicz, Z. Genetic Algorithms + Data Structures = Evolution Programs; Springer: Berlin/Heidelberg, Germany, 1996. [Google Scholar]
- Ansari, S.; Alnajjar, K.; Saad, M.; Abdallah, S.; Moursy, A. Automatic Digital Modulation Recognition Based on Genetic-Algorithm-Optimized Machine Learning Models. IEEE Access 2022, 10, 50265–50277. [Google Scholar] [CrossRef]
- Ji, Y.; Liu, S.; Zhou, M.; Zhao, Z.; Guo, X.; Qi, L. A machine learning and genetic algorithm-based method for predicting width deviation of hot-rolled strip in steel production systems. Inf. Sci. 2022, 589, 360–375. [Google Scholar] [CrossRef]
- Santana, Y.H.; Alonso, R.M.; Nieto, G.G.; Martens, L.; Joseph, W.; Plets, D. Indoor genetic algorithm-based 5G network planning using a machine learning model for path loss estimation. Appl. Sci. 2022, 12, 3923. [Google Scholar] [CrossRef]
- Liu, X.; Jiang, D.; Tao, B.; Jiang, G.; Sun, Y.; Kong, J.; Chen, B. Genetic algorithm-based trajectory optimization for digital twin robots. Front. Bioeng. Biotechnol. 2022, 9, 793782. [Google Scholar] [CrossRef] [PubMed]
- Nonoyama, K.; Liu, Z.; Fujiwara, T.; Alam, M.M.; Nishi, T. Energy-efficient robot configuration and motion planning using genetic algorithm and particle swarm optimization. Energies 2022, 15, 2074. [Google Scholar] [CrossRef]
- Liu, K.; Deng, B.; Shen, Q.; Yang, J.; Li, Y. Optimization based on genetic algorithms on energy conservation potential of a high speed SI engine fueled with butanol–gasoline blends. Energy Rep. 2022, 8, 69–80. [Google Scholar] [CrossRef]
- Zhou, G.; Zhu, Z.; Luo, S. Location optimization of electric vehicle charging stations: Based on cost model and genetic algorithm. Energy 2022, 247, 123437. [Google Scholar] [CrossRef]
- Chen, Q.; Hu, X. Design of intelligent control system for agricultural greenhouses based on adaptive improved genetic algorithm for multi-energy supply system. Energy Rep. 2022, 8, 12126–12138. [Google Scholar] [CrossRef]
- Min, D.; Song, Z.; Chen, H.; Wang, T.; Zhang, T. Genetic algorithm optimized neural network based fuel cell hybrid electric vehicle energy management strategy under start-stop condition. Appl. Energy 2022, 306, 118036. [Google Scholar] [CrossRef]
- Doewes, R.I.; Nair, R.; Sharma, T. Diagnosis of COVID-19 through blood sample using ensemble genetic algorithms and machine learning classifier. World J. Eng. 2022, 19, 175–182. [Google Scholar] [CrossRef]
- Choudhury, S.; Rana, M.; Chakraborty, A.; Majumder, S.; Roy, S.; RoyChowdhury, A.; Datta, S. Design of patient specific basal dental implant using Finite Element method and Artificial Neural Network technique. J. Eng. Med. 2022, 236, 1375–1387. [Google Scholar] [CrossRef]
- El-Anwar, M.I.; El-Zawahry, M.M. A three dimensional finite element study on dental implant design. J. Genet. Eng. Biotechnol. 2011, 9, 77–82. [Google Scholar] [CrossRef]
- Zheng, Q.; Zhong, J. Design of Automatic Pronunciation Error Correction System for Cochlear Implant Based on Genetic Algorithm. In Proceedings of the ICMMIA: Application of Intelligent Systems in Multi-Modal Information Analytics 2022, Online, 23 April 2022; pp. 1041–1047. [Google Scholar]
- Brahim, O.; Hamid, B.; Mohammed, N. Optimal design of inductive coupled coils for biomedical implants using metaheuristic techniques. E3S Web Conf. 2022, 351, 01063. [Google Scholar] [CrossRef]
- Tokgoz, E.; Carro, M.A. Applications of Artificial Intelligence, Machine Learning, and Deep Learning on Facial Plastic Surgeries; Springer: Cham, Switzerland, 2023; pp. 281–306. [Google Scholar]
- Wang, B.; Gomez-Aguilar, J.F.; Sabir, Z.; Raja, M.A.Z.; Xia, W.; Jahanshahi, H.; Alassafi, M.O.; Alsaadi, F. Surgery Using The Capability Of Morlet Wavelet Artificial Neural Networks. Fractals 2023, 30, 2240147. [Google Scholar] [CrossRef]
- Ahmed, M.; Seraj, R.; Islam, S.M.S. The k-means algorithm: A comprehensive survey and performance evaluation. Electronics 2020, 9, 1295. [Google Scholar] [CrossRef]
- Maaranen, H.; Miettinen, K.; Makela, M.M. Quasi-random initial population for genetic algorithms. Comput. Math. Appl. 2004, 47, 1885–1895. [Google Scholar] [CrossRef]
- Paul, P.V.; Dhavachelvan, P.; Baskaran, R. A novel population initialization technique for Genetic Algorithm. In Proceedings of the 2013 International Conference on Circuits, Power and Computing Technologies (ICCPCT), Nagercoil, India, 20–21 March 2013; pp. 1235–1238. [Google Scholar]
- Li, C.; Chu, X.; Chen, Y.; Xing, L. A knowledge-based technique for initializing a genetic algorithm. J. Intell. Fuzzy Syst. 2016, 31, 1145–1152. [Google Scholar] [CrossRef]
- Hassanat, A.B.; Prasath, V.S.; Abbadi, M.A.; Abu-Qdari, S.A.; Faris, H. An improved genetic algorithm with a new initialization mechanism based on regression techniques. Information 2018, 9, 167. [Google Scholar] [CrossRef]
- Kaelo, P.; Ali, M.M. Integrated crossover rules in real coded genetic algorithms. Eur. J. Oper. Res. 2007, 176, 60–76. [Google Scholar] [CrossRef]
- Tsoulos, I.G. Modifications of real code genetic algorithm for global optimization. Appl. Math. Comput. 2008, 203, 598–607. [Google Scholar] [CrossRef]
- Powell, M.J.D. A Tolerant Algorithm for Linearly Constrained Optimization Calculations. Math. Program. 1989, 45, 547–566. [Google Scholar] [CrossRef]
- Beyer, H.G.; Schwefel, H.P. Evolution strategies–A comprehensive introduction. Nat. Comput. 2002, 1, 3–52. [Google Scholar] [CrossRef]
- LeCun, Y.; Bengio, Y.; Hinton, G. Deep learning. Nature 2015, 521, 436–444. [Google Scholar] [CrossRef] [PubMed]
- Whitley, D. The GENITOR algorithm and selection pressure: Why rank-based allocation of reproductive trials is best. In Proceedings of the Third International Conference on Genetic Algorithms, Fairfax, VA, USA, 4–7 June 1989; pp. 116–121. [Google Scholar]
- Eiben, A.E.; Smith, J.E. Introduction to Evolutionary Computing; Springer: Berlin/Heidelberg, Germany, 2015. [Google Scholar]
- Lloyd, S. Least squares quantization in PCM. IEEE Trans. Inf. Theory 1982, 28, 129–137. [Google Scholar] [CrossRef]
- MacQueen, J.B. Some Methods for classification and Analysis of Multivariate Observations. In Proceedings of the 5th Berkeley Symposium on Mathematical Statistics and Probability, Oakland, CA, USA, 21 June–18 July 1967; pp. 281–297. [Google Scholar]
- Jain, A.K.; Murty, M.N.; Flynn, P.J. Data clustering: A review. Acm Comput. Surv. 1999, 31, 264–323. [Google Scholar] [CrossRef]
- Bishop, C.M. Pattern Recognition and Machine Learning; Springer: Berlin/Heidelberg, Germany, 2006. [Google Scholar]
- Hastie, T.; Tibshirani, R.; Friedman, J. The Elements of Statistical Learning: Data Mining, Inference, and Prediction; Springer: Berlin/Heidelberg, Germany, 2009. [Google Scholar]
- Ali, M.M.; Khompatraporn, C.; Zabinsky, Z.B. A Numerical Evaluation of Several Stochastic Algorithms on Selected Continuous Global Optimization Test Problems. J. Glob. Optim. 2005, 31, 635–672. [Google Scholar] [CrossRef]
- Floudas, C.A.; Pardalos, P.M.; Adjiman, C.; Esposoto, W.; Gümüs, Z.; Harding, S.; Klepeis, J.; Meyer, C.; Schweiger, C. Handbook of Test Problems in Local and Global Optimization; Kluwer Academic Publishers: Dordrecht, The Netherland, 1999. [Google Scholar]
- Gaviano, M.; Ksasov, D.E.; Lera, D.; Sergeyev, Y.D. Software for generation of classes of test functions with known local and global minima for global optimization. Acm Trans. Math. Softw. 2003, 29, 469–480. [Google Scholar] [CrossRef]
- Lennard-Jones, J.E. On the Determination of Molecular Fields. Proc. R. Soc. Lond. A 1924, 106, 463–477. [Google Scholar]
- Stein, W.E.; Keblis, M.F. A new method to simulate the triangular distribution. Math. Comput. Model. 2009, 49, 1143–1147. [Google Scholar] [CrossRef]
- Gropp, W.; Lusk, E.; Doss, N.; Skjellum, A. A high-performance, portable implementation of the MPI message passing interface standard. Parallel Comput. 1996, 22, 789–828. [Google Scholar] [CrossRef]
- Chandra, R.; Dagum, L.; Kohr, D.; Maydan, D.; McDonald, J.; Menon, R. Parallel Programming in OpenMP; Morgan Kaufmann Publishers Inc.: Cambridge, MA, USA, 2001. [Google Scholar]


| Problem | Uniform | Triangular | Kmeans |
|---|---|---|---|
| BF1 | 5731 | 5934 | 4478 |
| BF2 | 5648 (0.97) | 5893 | 4512 |
| BRANIN | 4680 | 4835 | 4627 |
| CM4 | 5801 | 5985 | 4431 |
| CAMEL | 4965 | 5099 | 4824 |
| EASOM | 5657 | 7089 | 4303 |
| EXP4 | 4934 | 4958 | 4539 |
| EXP8 | 5021 | 5187 | 4689 |
| EXP16 | 5063 | 5246 | 4874 |
| EXP32 | 5044 | 5244 | 5016 |
| GKLS250 | 4518 | 4710 | 4525 |
| GKLS350 | 4650 | 4833 | 4637 |
| GOLDSTEIN | 8099 | 8537 | 7906 |
| GRIEWANK2 | 5500 (0.97) | 5699 (0.97) | 4324 |
| GRIEWANK10 | 6388 (0.70) | 7482 (0.63) | 4559 |
| HANSEN | 5681 (0.93) | 6329 | 6357 |
| HARTMAN3 | 4950 | 5157 | 4998 |
| HARTMAN6 | 5288 | 5486 | 5258 |
| POTENTIAL3 | 5587 | 5806 | 5604 |
| POTENTIAL5 | 7335 | 7824 | 7450 |
| RASTRIGIN | 5703 | 5848 | 4481 |
| ROSENBROCK4 | 4241 | 4441 | 4241 |
| ROSENBROCK8 | 41,802 | 41,965 | 4523 |
| ROSENBROCK16 | 42,196 | 42,431 | 4962 |
| SHEKEL5 | 5488 (0.97) | 5193 (0.97) | 5232 (0.97) |
| SHEKEL7 | 5384 | 5711 (0.97) | 5695 (0.97) |
| SHEKEL10 | 6360 | 5989 | 6396 |
| TEST2N4 | 5000 | 5179 | 5047 |
| TEST2N5 | 5306 | 5309 | 5039 |
| TEST2N6 | 5245 | 5492 | 5107 |
| TEST2N7 | 5282 (0.93) | 5583 | 5216 |
| SINU4 | 4844 | 5046 | 4899 |
| SINU8 | 5368 | 5503 | 5509 |
| SINU16 | 6919 | 5583 | 5977 |
| TEST30N3 | 7215 | 8115 | 5270 |
| TEST30N4 | 7073 | 7455 | 6712 |
| Total | 273,966 (0.98) | 282,176 (0.985) | 186,217 (0.998) |
| Dimension | Calls (200 Uniform Samples) | Calls (200 k-Means Centers) |
|---|---|---|
| 5 | 15,637 | 4332 |
| 10 | 24,690 | 4486 |
| 15 | 39,791 | 4743 |
| 20 | 42,976 | 5194 |
| 25 | 43,617 | 7152 |
| 30 | 44,502 | 6914 |
| 35 | 45,252 | 15,065 |
| 40 | 46,567 | 13,952 |
| 45 | 47,640 | 15,193 |
| 50 | 49,393 | 22,535 |
| 55 | 50,062 | 23,692 |
| 60 | 52,293 | 25,570 |
| 65 | 52,546 | 25,678 |
| 70 | 53,346 | 28,153 |
| 75 | 54,110 | 28,328 |
| 80 | 57,209 | 29,320 |
| 85 | 60,970 | 29,371 |
| 90 | 65,319 | 32,121 |
| 95 | 68,097 | 35,721 |
| 100 | 66,803 | 35,396 |
| Total | 980,820 | 392,916 |
| Dimension | Calls (200 Uniform Samples) | Calls (200 k-Means Centers) |
|---|---|---|
| 2 | 5665 | 4718 |
| 4 | 6212 | 4431 |
| 6 | 7980 | 4390 |
| 8 | 9917 | 4449 |
| 10 | 12,076 (0.97) | 4481 |
| 12 | 14,672 | 4565 |
| 14 | 18,708 (0.87) | 4685 |
| 16 | 23,251 (0.77) | 4687 |
| 18 | 24,624 (0.77) | 4766 |
| 20 | 30,153 (0.80) | 4848 |
| 22 | 35,851 (0.77) | 15,246 (0.97) |
| 24 | 43,677 (0.93) | 7865 (0.93) |
| 26 | 41,492 (0.77) | 5627 |
| 28 | 38,017 (0.73) | 10,566 (0.97) |
| 30 | 47,538 (0.83) | 24,803 (0.90) |
| Total | 359,833 (0.84) | 110,127 (0.98) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Charilogis, V.; Tsoulos, I.G.; Stavrou, V.N. An Intelligent Technique for Initial Distribution of Genetic Algorithms. Axioms 2023, 12, 980. https://doi.org/10.3390/axioms12100980
Charilogis V, Tsoulos IG, Stavrou VN. An Intelligent Technique for Initial Distribution of Genetic Algorithms. Axioms. 2023; 12(10):980. https://doi.org/10.3390/axioms12100980
Chicago/Turabian StyleCharilogis, Vasileios, Ioannis G. Tsoulos, and V. N. Stavrou. 2023. "An Intelligent Technique for Initial Distribution of Genetic Algorithms" Axioms 12, no. 10: 980. https://doi.org/10.3390/axioms12100980
APA StyleCharilogis, V., Tsoulos, I. G., & Stavrou, V. N. (2023). An Intelligent Technique for Initial Distribution of Genetic Algorithms. Axioms, 12(10), 980. https://doi.org/10.3390/axioms12100980









