Multi-Objective Optimization Algorithm Based on Sperm Fertilization Procedure (MOSFP)
Abstract
:1. Introduction
2. State of the Art
Sperm Swarm Optimization Algorithm
- The normal pH value of a healthy female reproductive system is around 4.5–5.5 [24]. However, low pH of mucus acidic may destroy and deactivate the motility of sperm. Therefore, during the ovulation, the pH of vaginal acid or acidic is in the range of 7 to 4, which is very appropriate for sperm motility and is considered very alkaline and non-toxic to sperm [25]. The pH value of the female reproductive system is affected by the type of food consumed [26] and emotional or mood status of the female, such as happiness or sadness, etc. [27]. Based on this information, we can estimate that the value of pH will be varied in the range of 7 to 14.
- The temperature of the female reproductive system plays a significant role of determining the sperms movement direction. The scientists found that the sperm head acts like a temperature sensor to search for a warmer area (the egg location) [28]. Furthermore, the sperm head can response and sense to a temperature difference of <0.0006 °C [28]. The temperature inside the vagina can change based on women status. The average temperature inside the vagina is approximately between 35.1 and 37.4 °C [29]. However, this temperature may reach 38.5 °C in some cases due to vaginal blood pressure circulation [30].Based on this information, we can estimate that the value of temperature will be varied in the range of 35.1–38.5.
- The initial velocity of the sperm: is the velocity that acquired by each sperm after the ejaculation in the cervix zone. The sperm swarms take random positions inside the cervix and their velocity is affected by the pH value in that position. This velocity can be represented in the following rule:where:
- D: is the velocity damping factor, which is a random number between 0 and 1.
- Vi: is the sperm velocity.
- pH_Rand1: is the pH value, which is a random number in the range of (7, 14).
- Personal sperm current best solution: is the best solution that achieved so far by the sperm. As we mentioned previously, sperm head acts like a temperature sensor, which prefers to swim towards warmer temperatures (egg location) [28]. In addition, sperm moves forward to search on the guidance (higher concentrations of molecules) that produced and released by the egg, which this guidance knew as Chemo-taxis [31]. From this information, we can realize that the sperm will not swim backward to the cervix, but, will go forward towards warmer temperatures (the egg location in fallopian tubes). Based on that, this position can be achieved by comparing the sperm current position on X-axis and Y-axis with a sperm past position that is stored in the memory. The past position can be replaced by the current position if the current position is better than the past position. Personal sperm current best solution can be represented in the following equation:where:
- sb_ solution []: is the best solution that has been achieved so far, which denoted by Sperm Best (sb_ solution).
- pH_Rand2: is the pH value, which is a random number in the range of (7, 14).
- Temp_Rand1: is the area temperature, which is a random number in range (35.1, 38.5).
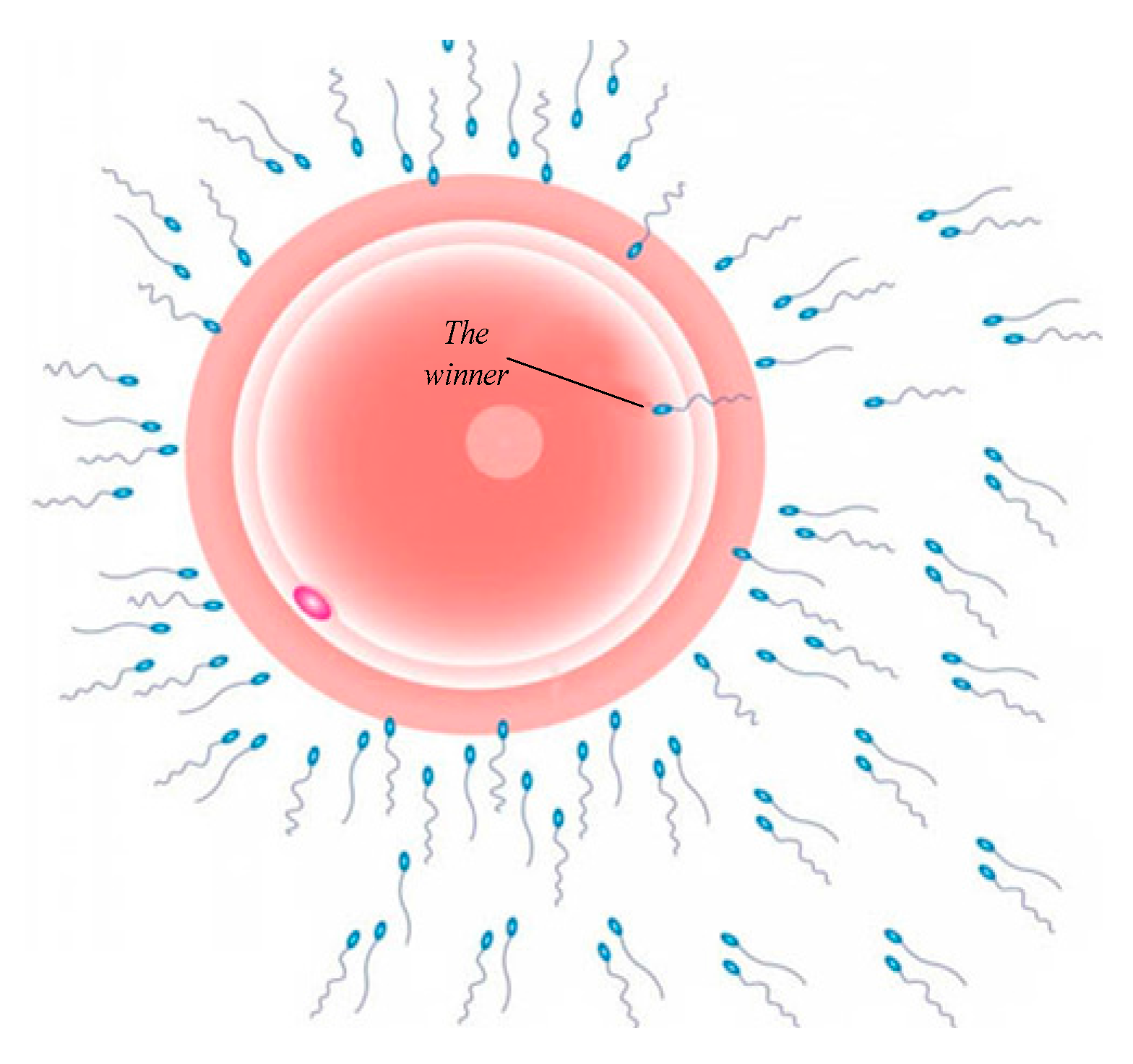
- Global best value is used to determine which sperm’s data is currently closest to the target (at the end, this sperm will represent as the winner). The value of sperm global best value is represented as the following equation:where:
- sgb_solution[]: is the best solution of any sperm has achieved so far which denoted by Sperm Global Best (sgb_solution).
- pH_Rand3: is the pH value, which is a random number in the range of (7, 14).
- Temp_Rand2: is the area temperature, which is a random number in the range of (35.1, 38.5).
- current[]: is the current best solution represented by the following equation.
where v[] is the sperm velocity which measured by companies the previous velocities in one equation as follows:
| Algorithm 1 Sperm swarm optimization (SSO) |
| 1: Begin |
| 2: Step 1: initialize positions for all sperms. |
| 3: Step 2: for i=1:number of sperm do |
| 4: Step 3: evaluate the fitness for each sperm |
| 5: If obtained fitness > sperm best solution then |
| 6: Set the current value as the sperm best solution |
| 7: End if |
| 8: End for |
| 9: Step 4: choose the sperm global best solution based on the winner. |
| 10: Step 5: for i=1:number of sperm Do |
| 11: Do the swim using velocity update rule |
| 12: Update sperm location on the search space |
| 13: End for |
| 14: Step 6: for i=1:number of sperm Do |
| 15: Apply mutation operation on the sperm value |
| 16: End for |
| 17: Step 7: while maximum iterations not achieved return to step 2 and repeat until reaching the |
| 17: maximum number of iterations. |
| 18: End procedure. |
3. Multi-Objective Optimization Algorithm Based on Sperm Fertilization Procedure (MOSFP)
- Domination: x1, may dominate a position vector, x2 (x1 < x2), if and only if and for at least one k [33].
- Pareto optimal: A vector F is defined as Pareto-optimal if no vector exists x F such that for at least one . An objective vector is called Pareto optimal if the corresponding vector is the Pareto optimal. The set of the Pareto optimal decision vectors F is denoted by [34].
- F is the Pareto optimal of a position vector if no position vector that dominates it, F. The Pareto optimal solution is non-dominated solution, which not dominated by other solutions [34].
- Pareto optimal set: is a set containing all the Pareto optimal vectors [35].
- Pareto front (PF): is defined as [35].
| Algorithm 2 Mutation |
| 1: Begin |
| 2: Step 1: for i = 0 to population size do |
| 3: If (i % 3 == 0) then |
| 4: Sperms_ mutated with a non-uniform mutation operator |
| 5: Else if (i % 3 == 1) then |
| 7: Sperms_ mutated with a uniform mutation operator |
| 8: Else |
| 9: Sperms_ without mutation |
| 10: End if |
| 11: End for |
| 12: End procedure. |
| Algorithm 3 Multi-objective optimization algorithm based on sperm fertilization procedure (MOSFP) |
| 1: Begin |
| 2: Step 1: initialize positions for all sperms. |
| 3: Step 2: initialize Winners. |
| 4: Step 2: archive the Winners in |
| 5: Step 3: crowd the winners using crowding operation. |
| 6: Step 4: define counter (i) and define number of maximum iterations (iMax). |
| 7: Step 5: do //this do is a do - while |
| 8: For <each sperm> do |
| 9: Select winner from the sperm swarm |
| 10: Update sperms positions using the predefined sperm velocity update rule (perform swim) |
| 11: Perform mutation procedure (Algorithm 1) |
| 12: Evaluate the fitness for each sperm |
| 13: Update personal sperm current best solution |
| 14: End for |
| 15: Update Set of Winners (SoW) |
| 16: Archive winner in |
| 17: Crowd the SoW using crowding operation |
| 18: Update value of counter (i) |
| 19: Step 6: While i < iMax |
| 20: Step 7: archive results in |
| 21: End procedure. |
4. Comparison Strategy
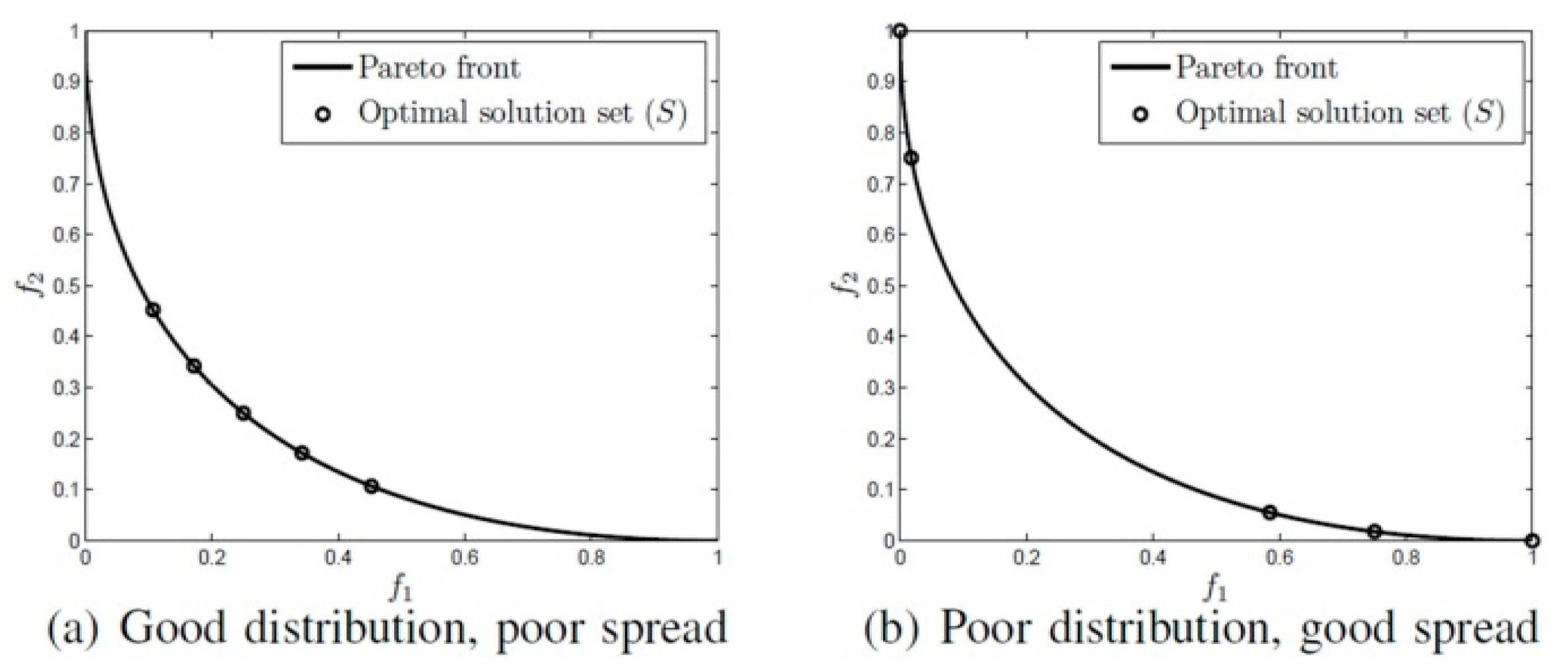
- Minimize the distance between the global Pareto front (the well-known Pareto front of any problem) and the Pareto front that produced by our approach.
- Maximize the spread of the solutions that produced by our approach, so uniform distribution of vectors can be achieved.
- Maximize the convergence of algorithm to obtain a good quality of the Pareto optimal set found.
- : is the Euclidean distance measured based on the distance between consecutive solutions,
- : is the average of
- and : are the minimum Euclidean distance measured based on the distance between solutions in S to the radical (bounding) solutions of the Pareto front (P).
- Non-dominated Sorting Genetic Algorithm (NSGA-II): is one of the most popular algorithms in MOO area. This algorithm performs a set of operators that deduced from the single objective Genetic Algorithm (GA). These operations are selection, crossover and mutation operators [9]. The pseudo-code of NSGA-II is summarized in Algorithm 4 [8].
Algorithm 4 Non-dominated sorting genetic algorithm II (NSGA-II) 1: Begin 2: Step 1: initialize Population 3: Generate random population – size M 4: Step 2: evaluate objective values 5: Step 3: assign rank (level) based on Pareto dominance-“sort” 6: Step 4: generate child population 7: Step 5: binary tournament selection 8: Step 6: recombination and mutation 9: Step 7: for i=1 to number of generations do 10: With parent and child population 11: Assign rank (level) based on Pareto – “sort” 12: Generate sets on non-dominated fronts 13: Loop (inside) by adding solutions to next generation 14: Starting from the “first” front until M individuals found 15: Determine crowding distances between points on each front 16: Select points (elitist) on the lower front (with lower rank)and are outside a crowding 16: distance 17: Create next generation 18: Binary tournament Selection 19: Recombination and mutation 20: Increment generation index 21: End for 22: End Procedure. - Strength Pareto Evolutionary Algorithm 2 (SPEA2): is a multi-objective algorithm, which is created as an improvement version of strength Pareto evolutionary algorithm (SPEA) algorithm [56]. This algorithm proposed by Zitzler et al. [57]. The procedure of this algorithm works based on dominance, which each single individual dominates or is dominated by other solutions. The nearest neighbor technique is used to guide the search. In addition, this algorithm operates based on preserving the boundary solutions by using archive truncation method [58]. Algorithm 5 summarizes the pseudo-code of this algorithm [59].
Algorithm 5 Strength Pareto evolutionary algorithm 2 (SPEA2) 1: Begin 2: Step 1: initialize population P 3: Step 2: evaluate objective functions 4: Step 3: create external archive A 5: Step 4: for i=1 to number of generations do 6: Compute fitness of individual in P and A 7: Add non-dominated individuals from P and A 8: If capacity of A is exceeded Then 9: Remove individuals from A by truncation operator 10: End If 11: Perform binary tournament selection to create mating pool 12: Perform crossover 13: Perform mutation 14: End for 15: End Procedure. - Optimized Multi-objective Particle Swarm Optimization (OMOPSO): proposed by Sierra et al. [11]. This algorithm performs a set of operators to solve MOOPs, such as crowding the global best solutions that known as leaders, performs mutation, after that, performs an archive of the leaders. The pseudo-code of OMOPSO is summarized in Algorithm 6.In order to compare our algorithm with the previously mentioned algorithms, we use two techniques, including, quantitative and qualitative tests. For the quantitative test, we adopt the IGD, SP, and measures, while for the qualitative test, we compare between the quality of Pareto fronts that is achieved by our algorithm and the Pareto fronts that achieved by the other algorithms. For these purposes, two test suites called Zitzler-Deb-Thiele (ZDT), and Walking-Fish-Group (WFG) have been used in this work.
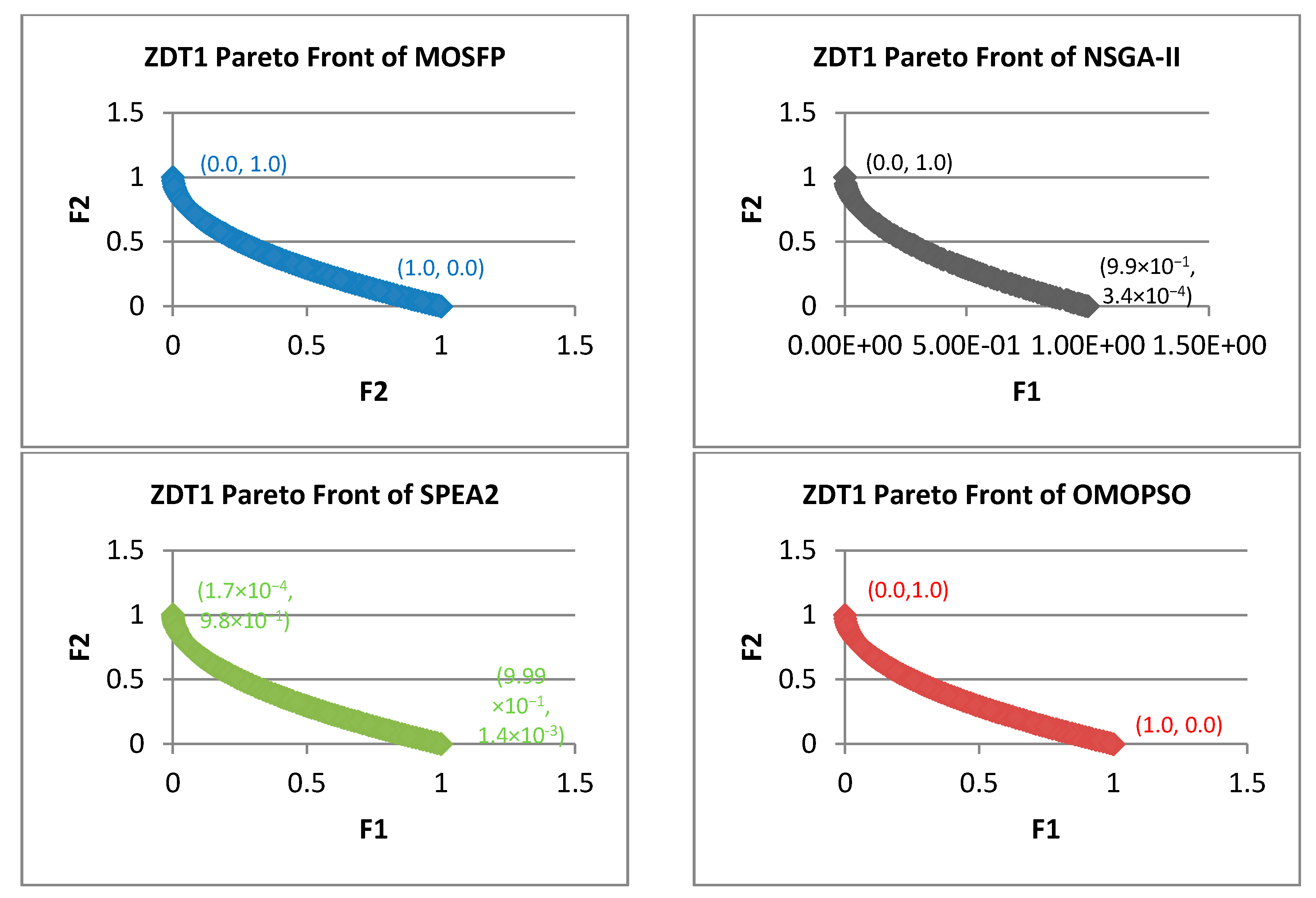
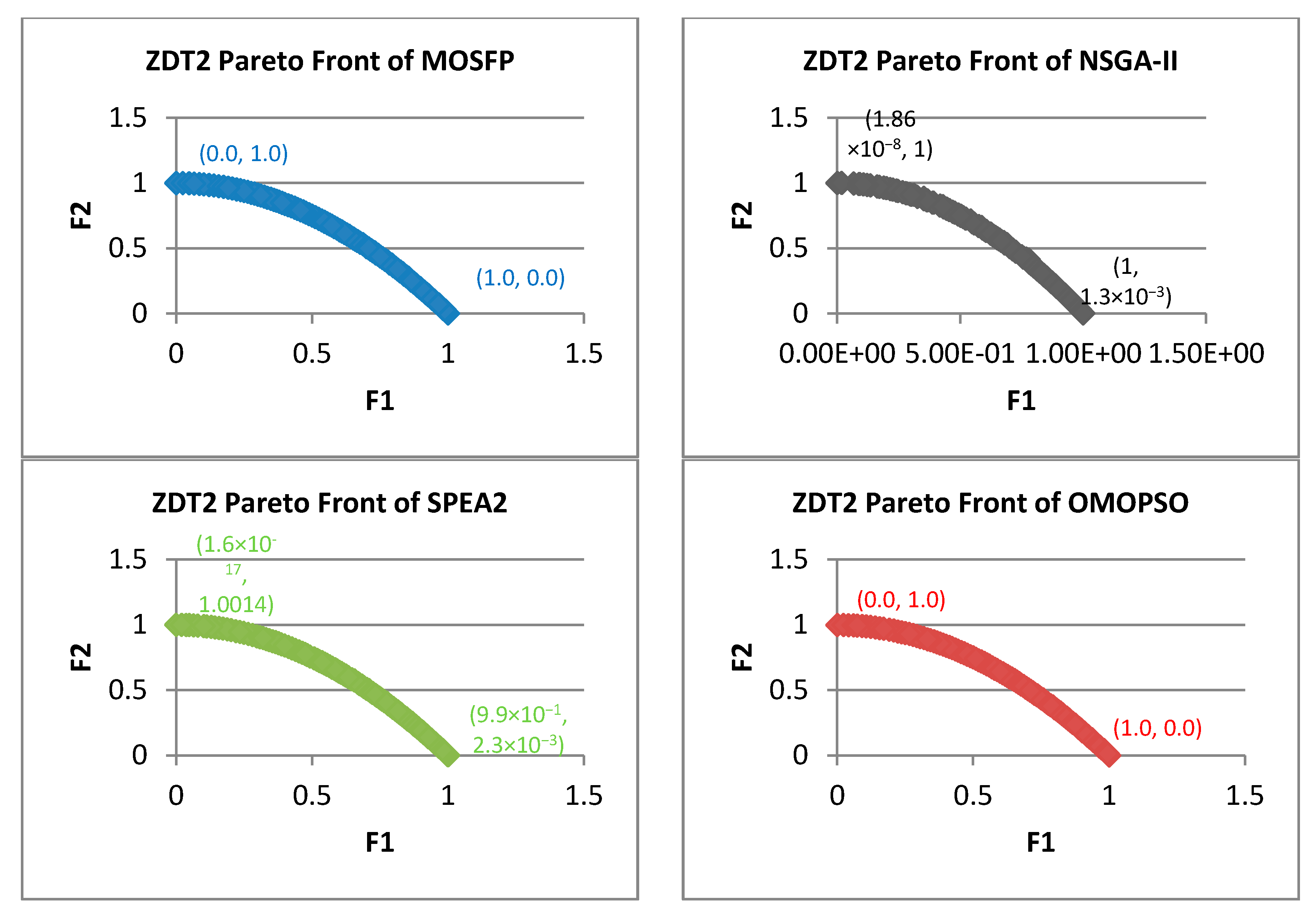
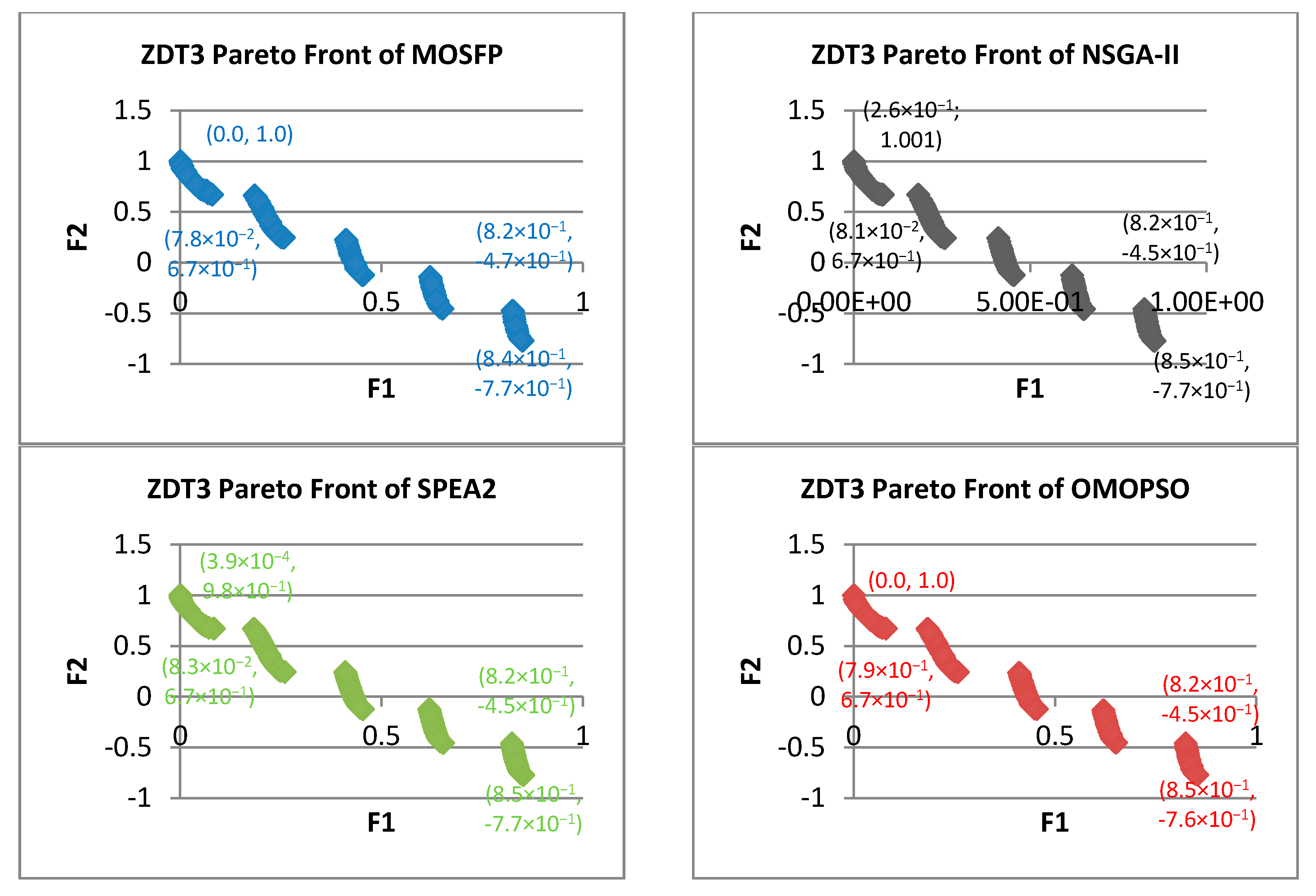
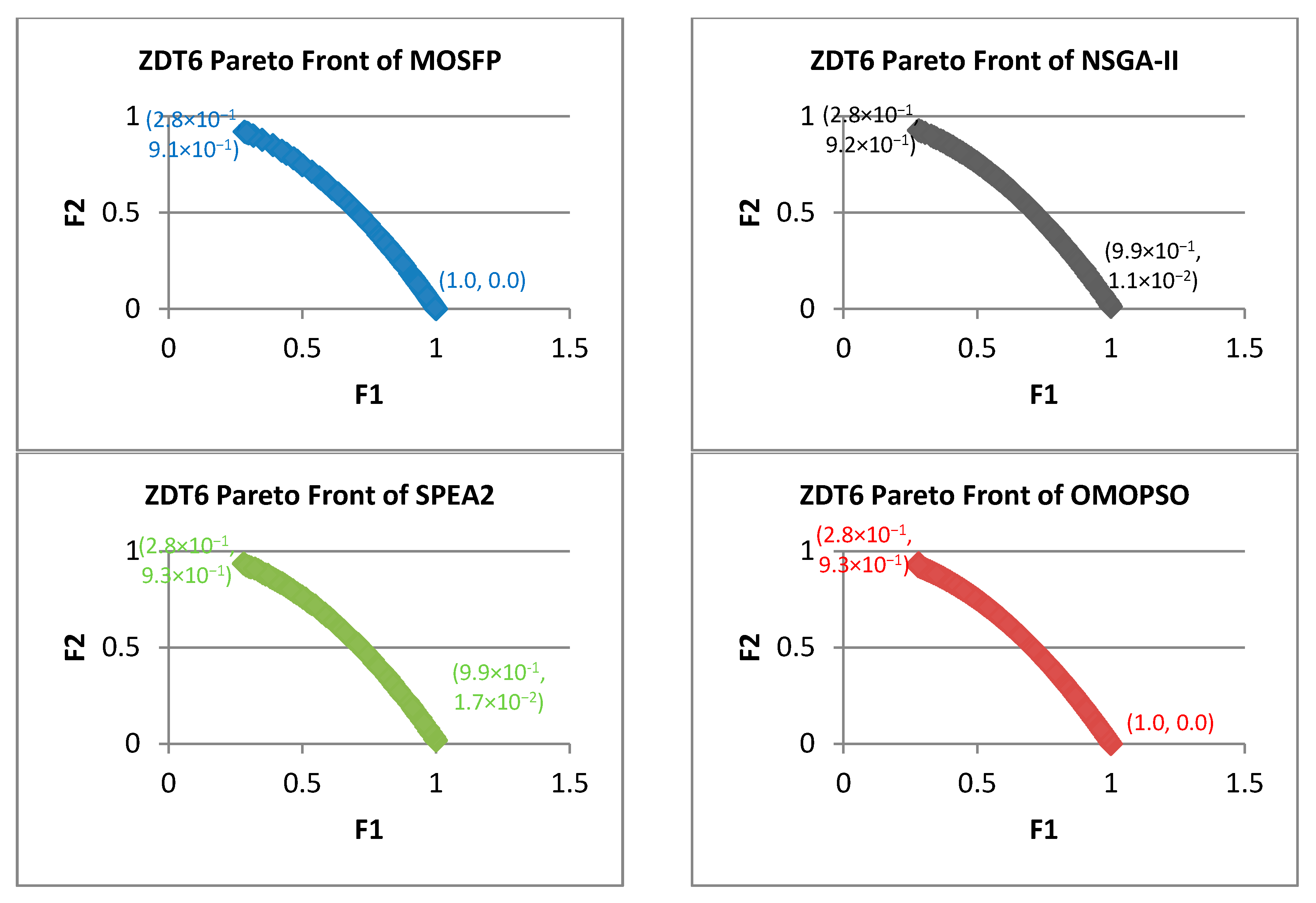
- Zitzler-Deb-Thiele (ZDT) Test Suite: this suite is proposed by Zitzler et al. [60]. ZDT suite is the most widely utilized suite of benchmark functions in evolutionary algorithms and swarm intelligence algorithms literature. The characteristics of ZDT problems are analyzed in Table 1. From the table, we can notice that the geometry of ZDT1 is convex, and the geometry of ZDT2 is concave. In addition, ZDT3 is disconnected on both the Pareto front and the Pareto optimal set, while the other functions consist of convex or concave components. The ZDT benchmark functions share many characteristics, including, how multimodality can cause a disconnected Pareto problems such as in ZDT3, and many-to-one Pareto problem, such as in ZDT6. These functions utilize only a single parameter, which means that these functions own only one variable. There are many advantages of using ZDT problem as a benchmark function for MOOAs, including, that this suite is well defined and employed in many research papers, which facilitate the comparisons with new MOOA. In addition, the Pareto optimal fronts of this suite are easy to apply and to understand [61]. For these reasons, we choose this suite to test our proposed algorithm.
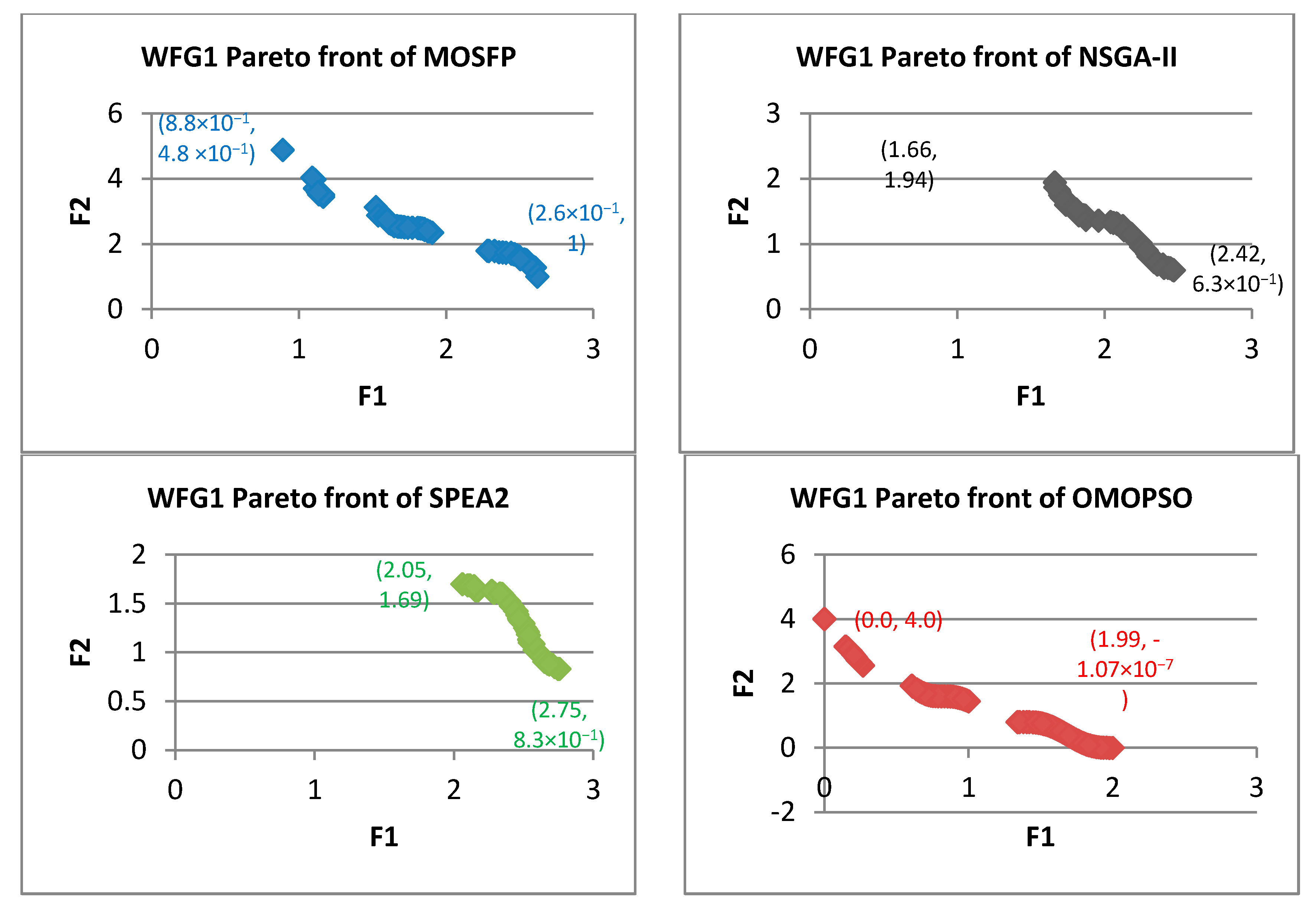
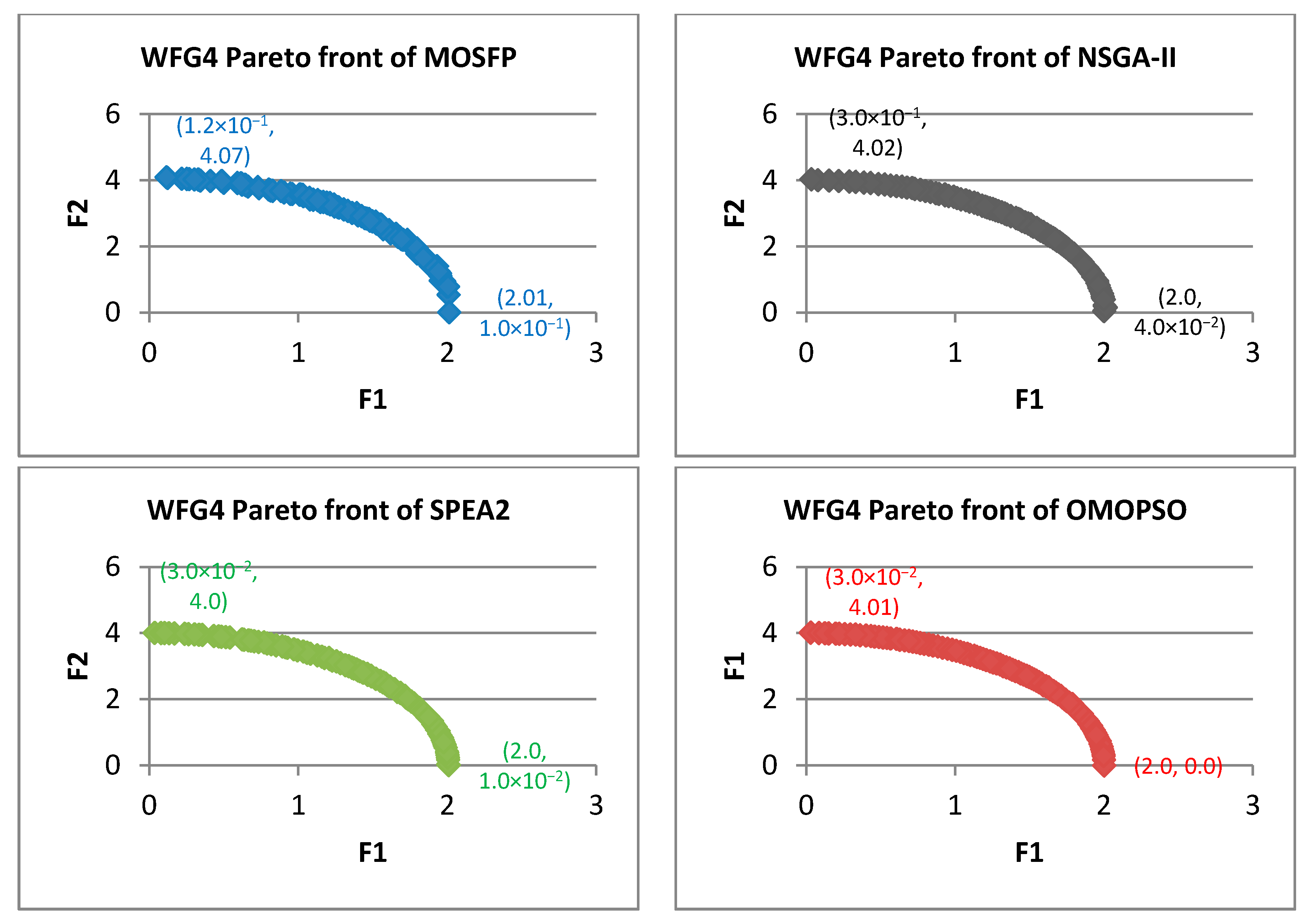
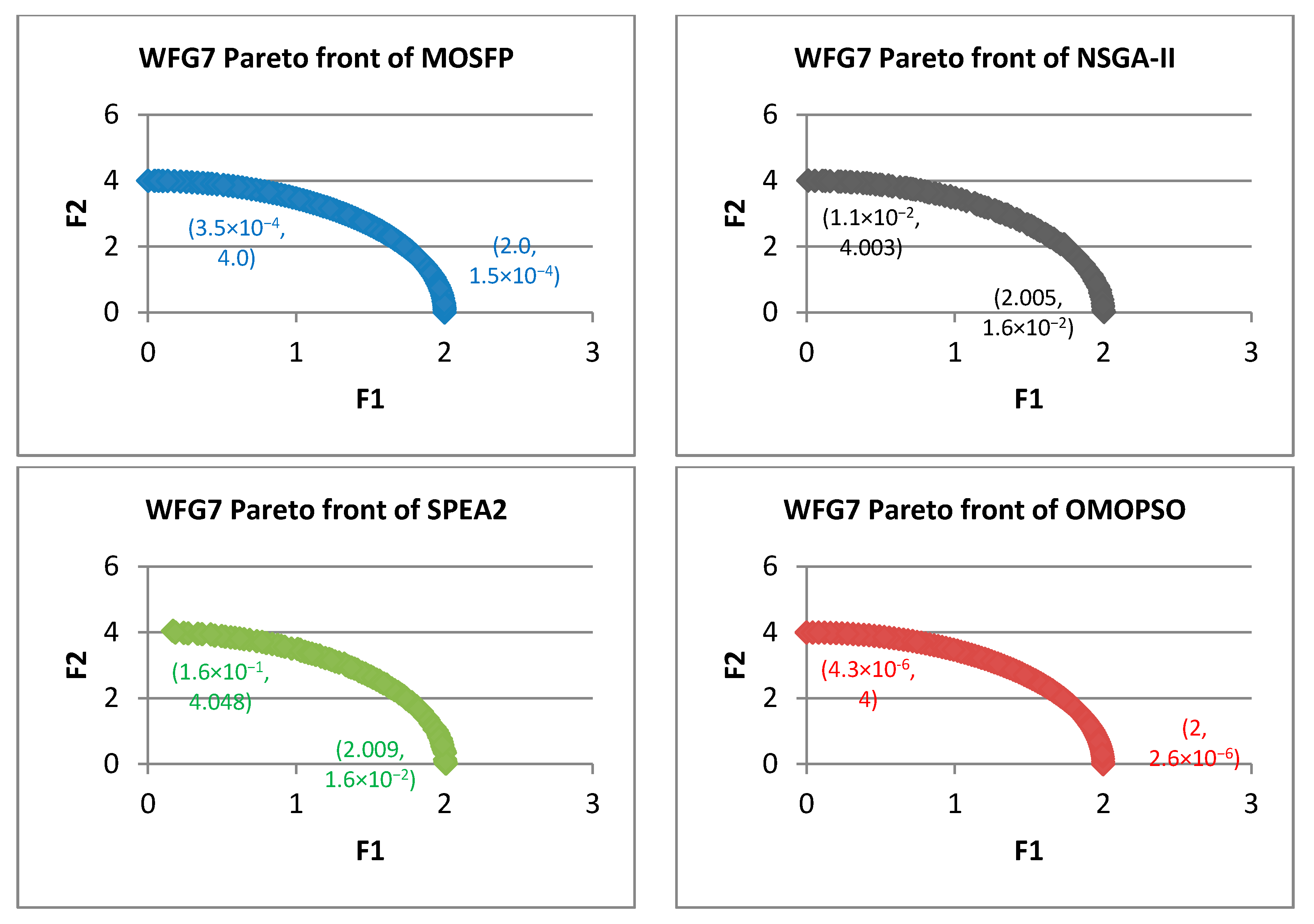
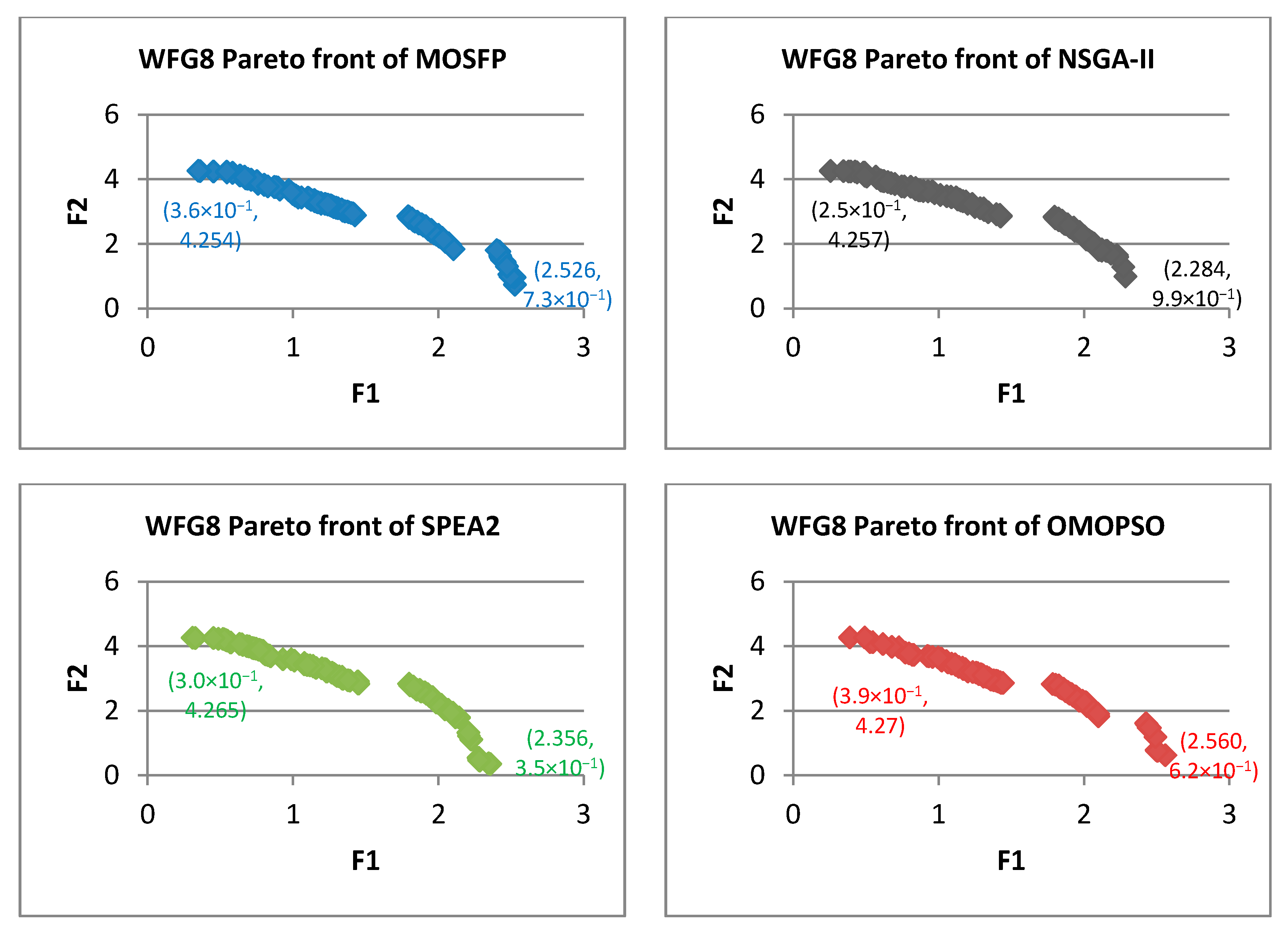
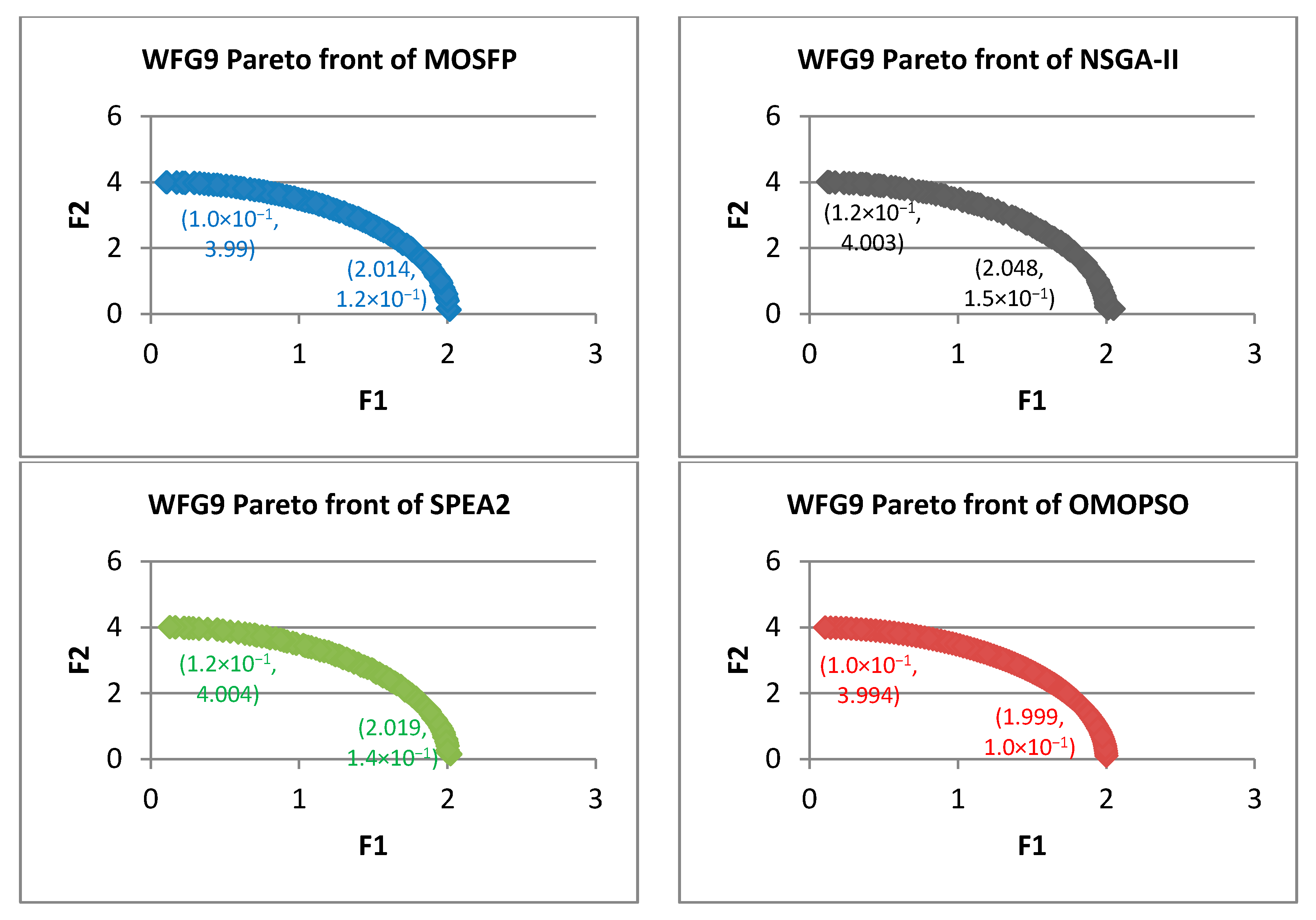
Algorithm 6 Optimized multi-objective particle swarm optimization (OMOPSO) 1: Begin 2: Step 1: initialize swarm and leaders. Send leaders to 3: Step 2: crowding(leaders), g = 0 4: Step 3: While g < max number of iterations (gmax) 5: For <each particle> do 6: Select leader. Flight. Mutation. Evaluation. Update pbest 7: End for 8: Update leaders, Send leaders to -archive 9: Crowding(leaders), g++ 10: End while 11: Step 4: Report results in -archive 12: End Procedure. - Walking-Fish-Group (WFG) Test Suite: WFG suite consists of nine problems introduced by [62]. The characteristics of these problems are outlined in Table 2 [60]. WFG1 and WFG7 are the only both unimodal and separable. WFG6 and WFG9 are the non-separable reduction problems, and they are more difficult than that of WFG3 and WFG2. WFG4 is a multimodality problem with a large “hill sizes” and it is more difficult than that of WFG9. WFG1 employs different parameters by using dissimilar weights in its variable weighted sum reduction. The parameters of WFG7 are dependent on the position and distance-related parameters, whereas WFG9 is more complex, which its distance-related parameters depend on other distance-related parameters. WFG8 is different than WFG9, wherein its distance-related parameters depend on other position and distance-related parameters. WFG5 is highly deceptive function and can be more difficult than that of WFG9.
5. Experimental Test and Results
6. Discussion
7. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
Appendix A
| No | Problem | Model | Domain |
| 1 | ZDT1 | ||
| 2 | ZDT2 | ||
| 3 | ZDT3 | ||
| 4 | ZDT6 |
| No. | Problem | Type | Model |
| 5 | WFG1 | Shape | |
| t1 | |||
| t2 | |||
| t3 | |||
| t4 | |||
| 6 | WFG2 | Shape | |
| t1 | As t1 from WFG1. (linear shift) | ||
| t2 | |||
| t3 | |||
| 7 | WFG3 | Shape | |
| t1:3 | As t1:3 from WFG2. (linear shift, non-separable reduction, and weighted sum reduction.) | ||
| 8 | WFG4 | Shape | |
| t1 | |||
| t2 | |||
| 9 | WFG5 | Shape | |
| t1 | |||
| t2 | As t2 from WFG4 (Weighted sum reduction.) | ||
| 10 | WFG6 | Shape | |
| t1 | As t1 from WFG1. (Linear shift.) | ||
| t2 | |||
| 11 | WFG7 | Shape | |
| t1 | |||
| t2 | As t1 from WFG1. (Linear shift). | ||
| t3 | As t2 from WFG4. (weighted sum reduction.) | ||
| 12 | WFG8 | Shape | |
| t1 | |||
| t2 | As t1 from WFG1. (Linear shift). | ||
| t3 | As t2 from WFG4. (weighted sum reduction.) | ||
| 13 | WFG9 | Shape | |
| t1 | |||
| t2 | |||
| t3 | As t2 from WFG6. (non-separable reduction.) |
References
- Everson, R.M.; Fieldsend, J.E.; Singh, S. Full elite sets for multi-objective Optimization. In Proceedings of the 5th International Conference Adaptive Computing Design Manufacture (ACDM 2002), Devon, UK, 16–18 April 2002; pp. 343–354. [Google Scholar] [CrossRef]
- Knowles, J.D.; Corne, D.W. Approximating the nondominated front using the Pareto archived evolution strategy. Evolut. Comput. 2000, 8, 149–172. [Google Scholar] [CrossRef] [PubMed]
- Capitanescu, F.; Marvuglia, A.; Benetto, E.; Ahmadi, A.; Tiruta-Barna, L. Linear programming-based directed local search for expensive multi-objective optimization problems: Application to drinking water production plants. Eur. J. Oper. Res. 2017, 262, 322–334. [Google Scholar] [CrossRef]
- Sakawa, M. Fuzzy Sets and Interactive Multiobjective Optimization, 1st ed.; Singah, M.G., Ed.; Springer Science & Business Media: Higashi-Hiroshima, Japan, 2013; pp. 1–308. ISBN 978-1-4899-1633-4. [Google Scholar]
- Watanabe, K.; Hashem, M.M.A. Evolutionary Computations: New Algorithms and Their Applications to Evolutionary Robots, 1st ed.; Springer: Berlin/Heidelberg, Germany, 2004; Volume 147, pp. 1–172. ISBN 978-3-540-39883-7. [Google Scholar]
- Holland, J.H. Genetic algorithms. Sci. Am. 1992, 267, 66–73. [Google Scholar] [CrossRef]
- Kennedy, J.; Eberhart, R. Particle swarm optimization. In Proceedings of the IEEE International Conference on Neural Networks, Perth, Australia, 27 November–1 December 1995; pp. 1942–1948. [Google Scholar] [CrossRef]
- Coello, C.; Lamont, G.; Van-Veldhuizen, D. Evolutionary Algorithms for Solving Multiobjective Problems, 2nd ed.; Lamont-Veldhuizen, G.B., Ed.; Springer: New York, NY, USA, 2013; pp. 1–800. ISBN 978-0-387-36797-2. [Google Scholar]
- Srinivas, N.; Deb, K. Muiltiobjective optimization using nondominated sorting in genetic algorithms. Evolut. Comput. 1994, 2, 221–248. [Google Scholar] [CrossRef]
- Coello Coello, C.A. Mopso: A proposal for multiple objective particle swarm optimization. In Proceedings of the 2002 Congress on Evolutionary Computation (CEC 2002), Honolulu, HI, USA, 12–17 May 2002; pp. 1051–1056. [Google Scholar] [CrossRef]
- Sierra, M.R.; Coello, C.C. Improving pso-based multi-objective optimization using crowding, mutation and e-dominance, Evolutionary multi-criterion optimization. In Proceedings of the International Conference on Evolutionary Multi-Criterion Optimization, Guanajuato, Mexico, 9–11 March 2005; pp. 505–519. [Google Scholar] [CrossRef]
- Venkatarayalu, N.V.; Ray, T. Single and multi-objective design of yagi-uda antennas using computational intelligence, Evolutionary Computation. In Proceedings of the IEEE Evolutionary Computation (CEC’03), Canberra, ACT, Australia, 8–12 December 2003; pp. 1237–1242. [Google Scholar] [CrossRef]
- Watanabe, S.; Hiroyasu, T.; Miki, M. Parallel evolutionary multi-criterion optimization for mobile telecommunication networks optimization. Evolutionary Methods for Design, Optimization and Control with Applications to Industrial Problems. In Proceedings of the EUROGEN 2001, Athens, Greece, 19–21 September 2001; pp. 167–172. [Google Scholar]
- Shaw, K.; Nortcliffe, A.; Thompson, M.; Love, J.; Fleming, P.; Fonseca, C. Assessing the performance of multiobjective genetic algorithms for optimization of a batch process scheduling problem. In Proceedings of the IEEE 1999 Congress on Evolutionary Computation (CEC 99), Washington, DC, USA, 6–9 July 1999; pp. 37–45. [Google Scholar] [CrossRef]
- Barone, L.; While, L.; Hingston, P. Designing crushers with a multi-objective evolutionary algorithm. In Proceedings of the 4th Annual Conference on Genetic and Evolutionary Computation, New York, NY, USA, 9–13 July 2002; pp. 995–1002. [Google Scholar] [CrossRef]
- Buche, D.; Stoll, P.; Dornberger, R.; Koumoutsakos, P. Multiobjective evolutionary algorithm for the optimization of noisy combustion processes. IEEE Trans. Syst. Man Cybern. Part C Appl. Rev. 2002, 32, 460–473. [Google Scholar] [CrossRef]
- Engrand, P. A multi-Objective Optimization Approach Based on Simulated Annealing and Its Application to Nuclear Fuel Management. Electricite de France 1998, 30, 1–8. [Google Scholar]
- Hughes, E.J. Swarm guidance using a multi-objective co-evolutionary on-line evolutionary algorithm. In Proceedings of the IEEE Congress on Evolutionary Computation (CEC2004), Portland, OR, USA, 19–23 June 2004; pp. 2357–2363. [Google Scholar] [CrossRef]
- Garcia, J.; Martin, J.R.; Gonzales, A.; Blanco, P. Hybrid heuristic and mathematical programming in oil pipelines networks. In Proceedings of the IEEE Congress on Evolutionary Computation (CEC2004), Portland, OR, USA, 19–23 June 2004; pp. 1479–1486. [Google Scholar] [CrossRef]
- Zitzler, E.; Laumanns, M.; Bleuler, S. A tutorial on evolutionary multiobjective optimization. In Metaheuristics for Multiobjective Optimization, 1st ed.; Gandibleux, X., Sevaux, M., Sörensen, K., T’kindt, V., Eds.; Springer: Berlin/Heidelberg, Germany, 2004; Volume 535, pp. 3–37. ISBN 978-3-540-20637-8. [Google Scholar]
- Shehadeh, H.A.; Idris, M.Y.I.; Ahmedy, I.; Noor, N.M. Sperm Swarm Optimization (SSO) Algorithm. Comput. Intell. Neurosci. 2017. under review. [Google Scholar]
- Tung, C.K.; Fiore, A.G.; Ardon, F.; Suarez, S.S.; Wu, M. Collective dynamics of sperm in viscoelastic fluid. APS Meet. Abstr. 2016, 2, 10–30. [Google Scholar]
- Nalepa, J.; Kawulok, M. A memetic algorithm to select training data for support vector machines. In Proceedings of the 2014 Annual Conference on Genetic and Evolutionary Computation ACM, Vancouver, BC, Canada, 12–16 July 2014; pp. 573–580. [Google Scholar] [CrossRef]
- Das Neves, J.; Bahia, M.F. Gels as vaginal drug delivery systems. Int. J. Pharm. 2006, 318, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Rodrigues, J.; Caldeira, J.; Vaidya, B. A novel intra-body sensor for vaginal temperature monitoring. Sensor 2009, 9, 2797–2808. [Google Scholar] [CrossRef] [PubMed]
- Borges, S.F.; Silva, J.G.; Teixeira, P.C. Survival and biofilm formation of Listeria monocytogenes in simulated vaginal fluid: Influence of pH and strain origin. FEMS Immunol. Med. Microbiol. 2011, 62, 315–320. [Google Scholar] [CrossRef] [PubMed]
- Edmunds, M.W.; Mayhew, M.S. Pharmacology for the Primary Care Provider-E-Book, 1st ed.; Elsevier Health Sciences: Washington, DC, USA, 2013; pp. 1–863. ISBN 978-0-323-08790-2. [Google Scholar]
- Bahat, A.; Caplan, S.R.; Eisenbach, M. Thermotaxis of human sperm cells in extraordinarily shallow temperature gradients over a wide range. PLoS ONE 2012, 7, e41915. [Google Scholar] [CrossRef] [PubMed]
- Christian Nicole. 2013. Available online: https://christiannicole72838.wordpress.com/2013/06/26/from-the-figure-for-women-to-understand-their-own-sexual-organs-gender/ (accessed on 21 September 2017).
- Health. 2011. Available online: http://health-of-people.blogspot.com/2011/03/interestingly-statistics-of-physical.html (accessed on 28 July 2017).
- Bahat, A.; Eisenbach, M. Sperm thermotaxis. Mol. Cell. Endocrinol. 2006, 252, 115–119. [Google Scholar] [CrossRef] [PubMed]
- Engelbrecht, A.P. Fundamentals of Computational Swarm Intelligence; John Wiley & Sons: Hoboken, NJ, USA, 2006; pp. 1–672. ISBN 0470091916. [Google Scholar]
- Chamaani, S.; Mirtaheri, S.A.; Teshnehlab, M.; Shooredeli, M.A. Modified multi-objective particle swarm optimization for electromagnetic absorber design. In Proceedings of the IEEE Asia-Pacific Conference on Applied Electromagnetics APACE, Kuala Lampur, Malaysia, 4–6 December 2007; pp. 1–5. [Google Scholar] [CrossRef]
- Lindroth, P.; Patriksson, M.; Strömberg, A.B. Approximating the Pareto optimal set using a reduced set of objective functions. Eur. J. Oper. Res. 2010, 207, 1519–1534. [Google Scholar] [CrossRef]
- Zheng, X.; Liu, H. A scalable coevolutionary multi-objective particle swarm optimizer. Int. J. Comput. Intell. Syst. 2010, 3, 590–600. [Google Scholar] [CrossRef]
- Lei, J.; Fu, G.; Yang, L.; Fu, D. Multi-objective optimization design of the yagi-uda antenna with an x-shape driven dipole. J. Electromagn. Waves Appl. 2007, 21, 963–972. [Google Scholar] [CrossRef]
- Lee, Y.H.; Cahill, B.J.; Porter, S.J.; Marvin, A.C. A novel evolutionary learning technique for multi-objective array antenna optimization. Prog. Electromagn. Res. 2004, 48, 125–144. [Google Scholar] [CrossRef]
- Coello, C.A.C.; Pulido, G.T.; Lechuga, M.S. Handling multiple objectives with particle swarm optimization. IEEE Trans. Evolut. Comput. 2004, 8, 256–279. [Google Scholar] [CrossRef]
- Deb, K.; Pratap, A.; Agarwal, S.; Meyarivan, T. A fast and elitist multiobjective genetic algorithm: Nsga-ii. IEEE Trans. Evolut. Comput. 2002, 6, 182–197. [Google Scholar] [CrossRef]
- Ray, T.; Liew, K. A swarm metaphor for multiobjective design optimization. Eng. Optim. 2002, 34, 141–153. [Google Scholar] [CrossRef]
- Li, X. A non-dominated sorting particle swarm optimizer for multiobjective optimization. In Proceedings of the Genetic and Evolutionary Computation—GECCO 2003, Chicago, IL, USA, 3 August 2003; Springer: Berlin, Germany, 2003; p. 198. [Google Scholar] [CrossRef]
- Laumanns, M.; Thiele, L.; Deb, K.; Zitzler, E. Combining convergence and diversity in evolutionary multiobjective optimization. Evolut. Comput. 2002, 10, 263–282. [Google Scholar] [CrossRef] [PubMed]
- Yue, X.; Guo, Z.; Yin, Y.; Liu, X. Many-objective E-dominance dynamical evolutionary algorithm based on adaptive grid. Soft Comput. 2016, 2, 1–10. [Google Scholar] [CrossRef]
- Guzek, M.; Diaz, C.O.; Pecero, J.E.; Bouvry, P.; Zomaya, A.Y. Impact of voltage levels number for energy-aware bi-objective dag scheduling for multi-processors systems. In Proceedings of the International Conference on Advances in Information Technology, Bangkok, Thailand, 6–7 December 2012; pp. 70–80. [Google Scholar] [CrossRef]
- Guzek, M.; Pecero, J.E.; Dorronsoro, B.; Bouvry, P. Multi-objective evolutionary algorithms for energy-aware scheduling on distributed computing systems. Appl. Soft Comput. 2014, 24, 432–446. [Google Scholar] [CrossRef]
- Dai, C.; Wang, Y.; Hu, L. An improved α-dominance strategy for many-objective optimization problems. Soft Comput. 2016, 20, 1105–1111. [Google Scholar] [CrossRef]
- Bezerra, L.C.; López-Ibáñez, M.; Stützle, T. An empirical assessment of the properties of inverted generational distance on multi-and many-objective optimization. In Proceedings of the International Conference on Evolutionary Multi-Criterion Optimization, Münster, Germany, 19–22 March 2017; pp. 31–45. [Google Scholar] [CrossRef]
- Yang, S.; Ong, Y.S.; Jin, Y. (Eds.) Evolutionary Computation in Dynamic and Uncertain Environments, 1st ed.; Springer: Berlin/Heidelberg, Germany, 2007; Volume 51, pp. 1–605. ISBN 3-540-49772-2. [Google Scholar]
- Monroy, R.; Arroyo-Figueroa, G.; Sucar, L.E.; Sossa, H. Micai Advances in artificial intelligence. In Proceedings of the Third Mexican International Conference on Artificial Intelligence, Mexico City, Mexico, 26–30 April 2004; pp. 622–631. [Google Scholar] [CrossRef]
- Jiang, S.; Cai, Z. A novel hybrid particle swarm optimization for multi-objective problems. In Proceedings of the International Conference on Artificial Intelligence and Computational Intelligence, Shanghai, China, 7–8 November 2009; pp. 28–37. [Google Scholar] [CrossRef]
- Jiang, S.; Ong, Y.S.; Zhang, J.; Feng, L. Consistencies and contradictions of performance metrics in multiobjective optimization. IEEE Trans. Cybern. 2014, 44, 2391–2404. [Google Scholar] [CrossRef] [PubMed]
- Riquelme, N.; Von Lücken, C.; Baran, B. Performance metrics in multi-objective optimization, Computing. In Proceedings of the IEEE Conference of Latin American (CLEI), Arequipa, Peru, 19–23 October 2015; pp. 1–11. [Google Scholar] [CrossRef]
- Hamdy, M.; Nguyen, A.-T.; Hensen, J.L. A performance comparison of multi-objective optimization algorithms for solving nearly-zero-energy-building design problems. Energy Build. 2016, 121, 57–71. [Google Scholar] [CrossRef]
- Acampora, G.; Ishibuchi, H.; Vitiello, A. A comparison of multi-objective evolutionary algorithms for the ontology meta-matching problem. In Proceedings of the IEEE Congress on Evolutionary Computation (CEC), Beijing, China, 6–11 July 2014; pp. 413–420. [Google Scholar] [CrossRef]
- Gharari, R.; Poursalehi, N.; Abbasi, M.; Aghaie, M. Implementation of strength pareto evolutionary algorithm ii in the multiobjective burnable poison placement optimization of kwu pressurized water reactor. Nucl. Eng. Technol. 2016, 48, 1126–1139. [Google Scholar] [CrossRef]
- Eckart, Z.; Marco, L.; Lothar, T. SPEA2: Improving the strength pareto evolutionary algorithm for multiobjective optimization. In Proceedings of the EUROGEN’2001 Eurogen, Evolutionary Method for Design: Optimization and Control for Industrial Problem, Barcelona, Spain, 19–21 September 2001; pp. 1–21. [Google Scholar] [CrossRef]
- Theophila, B. Study of Hardware and Software Optimizations of Spea2 on Hybrid Fpgas, 1st ed.; Lukowiak, M., Amuso, V.J., Shaaban, R., Jason, E., Eds.; Rochester Institute of Technology: Rochester, NY, USA, 2008; pp. 1–79. [Google Scholar]
- Bandyopadhyay, S.; Bhattacharya, R. On some aspects of nature-based algorithms to solve multi-objective problems. In Artificial intelligence, Evolutionary Computing and Metaheuristics; Yang, X., Ed.; Springer: Berlin/Heidelberg, Germany, 2013; Chapter 9; pp. 477–524. ISBN 978-3-642-29693-2. [Google Scholar]
- Pollack, S.L.; Hicks, H.T.; Harrison, W.J. Decision Tables Theory and Practice; Cannian, R.G., Cougar, J., Eds.; John Wiley & Sons: Hoboken, NJ, USA, 1971; pp. 1–179. ISBN 047169150X. [Google Scholar]
- Huband, S.; Hingston, P.; Barone, L.; While, L. A review of multiobjective test problems and a scalable test problem toolkit. IEEE Trans. Evolut. Comput. 2006, 10, 477–506. [Google Scholar] [CrossRef]
- Huband, S.; Barone, L.; While, R.L.; Hingston, P. A scalable multi-objective test problem toolkit. In Proceedings of the International Conference on Evolutionary Multi-Criterion Optimization, Guanajuato, Mexico, 9–11 March 2005; pp. 280–295. [Google Scholar] [CrossRef]
- Deb, K.; Thiele, L.; Laumanns, M.; Zitzler, E. Scalable multi-objective optimization test problems. In Proceedings of the IEEE Congress on Evolutionary Computation (CEC’2002), Honolulu, HI, USA, 12–17 May 2002; pp. 825–830. [Google Scholar] [CrossRef]
- Chase, N.; Rademacher, M.; Goodman, E.; Averill, R.; Sidhu, R. A benchmark study of multi-objective optimization methods. Red Cedar Technol. 2009, 6, 1–24. [Google Scholar]
- Bara’a, A.A.; Khalil, E.A.; Cosar, A. Multi-objective evolutionary routing protocol for efficient coverage in mobile sensor networks. Soft Comput. 2015, 19, 2983–2995. [Google Scholar] [CrossRef]
- Ibdah, Y.; Ding, Y. Path loss models for low-height mobiles in forest and urban. Wirel. Person. Commun. 2017, 92, 455–465. [Google Scholar] [CrossRef]
- Hamdan, M.; Yassein, M.B.; Shehadeh, H.A. Multi-objective optimization modeling of interference in home health care sensors. In Proceedings of the IEEE 11th International Conference on Innovations in Information Technology (IIT), Dubai, UAE, 1–3 November 2015; pp. 219–224. [Google Scholar] [CrossRef]




| Name | ZDT1 | ZDT2 | ZDT3 | ZDT6 | ||||
|---|---|---|---|---|---|---|---|---|
| Objective | f1 | f2 | f1 | f2 | f1 | f2 | f1 | f2 |
| R3: # Parameters | 1 | ✓ | 1 | ✓ | 1 | ✓ | 1 | ✓ |
| F2: Separability | S | S | S | S | S | S | S | S |
| F5: Modality | U | U | U | U | U | M | U | U |
| R1: No Extremal | ✗ | ✗ | ✗ | ✗ | ||||
| R2: No Medial | ✓ | ✓ | ✓ | ✓ | ||||
| R5: Diss. Domains | ✗ | ✗ | ✗ | ✗ | ||||
| R6: Diss. Ranges | ✗ | ✗ | ✓ | ✓ | ||||
| R7: Optima known | ✓ | ✓ | ✓ | ✓ | ||||
| F1: Geometry | convex | concave | disconnected | concave | ||||
| F3: Bias | - | - | - | + | ||||
| F4a: Pareto Many-to-one | - | - | - | + | ||||
| F4b: Flat Regions | - | - | - | - | ||||
| Name | Objective | F2: Separability | Modality | Bias | Geometry |
|---|---|---|---|---|---|
| WFG1 | f1:M | S | U | Polynomial, flat | Convex, Mixed |
| WFG2 | f1:M-1 fM | NS | U | − | Convex, disconnected |
| NS | M | ||||
| WFG3 | f1:M | NS | U | − | Linear, degenerate |
| WFG4 | f1:M | S | M | − | Concave |
| WFG5 | f1:M | S | D | − | Concave |
| WFG6 | f1:M | NS | U | − | Concave |
| WFG7 | f1:M | S | U | Parameter dependent | Concave |
| WFG8 | f1:M | NS | U | Parameter dependent | Concave |
| WFG9 | f1:M | NS | M, D | Parameter dependent | Concave |
| Parameters | MOSFP | OMOPSO | NSGA-II | SPEA2 |
|---|---|---|---|---|
| Population size | 100 | 100 | 100 | 100 |
| Archive size | (winner) 100 | 100 | (Elite) 100 | 100 |
| Mating pool size | - | - | - | 100 |
| Maximum generation | 5000 | 5000 | 5000 | 5000 |
| Crossover probability | - | - | 0.9 | 0.9 |
| Mutation probability | 1/d where d is the variable code size | |||
| Method | Metrics | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| SP | IGD | |||||||||||
| Best | Worst | Aver. | Med. | Best | Worst | Aver. | Med. | Best | Worst | Aver. | Med. | |
| ZDT1 | ||||||||||||
| MOSFP | 5.01 × 10−3 | 5.57 × 10−3 | 5.27 × 10−3 | 5.25 × 10−3 | 2.76 × 10−2 | 9.85 × 10−2 | 6.23 × 10−2 | 6.22 × 10−2 | 3.44 × 10−5 | 3.61 × 10−5 | 3.51 × 10−5 | 3.51 × 10−5 |
| OMOPSO | 4.92 × 10−3 | 5.40 × 10−3 | 5.15 × 10−3 | 5.15 × 10−3 | 3.29 × 10−2 | 8.74 × 10−2 | 5.54 × 10−2 | 5.52 × 10−2 | 3.39 × 10−5 | 3.60 × 10−5 | 3.48 × 10−5 | 3.48 × 10−5 |
| NSGA-II | 1.11 × 10−1 | 2.67 × 10−1 | 1.61 × 10−1 | 1.51 × 10−1 | 5.84 × 10−1 | 8.17 × 10−1 | 6.80 × 10−1 | 6.87 × 10−1 | 7.65 × 10−4 | 1.92 × 10−3 | 1.18 × 10−3 | 1.14 × 10−3 |
| SPEA2 | 1.60 × 10−1 | 4.58 × 10−1 | 2.82 × 10−1 | 2.74 × 10−1 | 8.92 × 10−1 | 6.11 × 10−1 | 7.43 × 10−1 | 7.35 × 10−1 | 1.04 × 10−3 | 2.71 × 10−3 | 1.76 × 10−3 | 1.72 × 10−3 |
| ZDT2 | ||||||||||||
| MOSFP | 5.018 × 10−3 | 5.65 × 10−3 | 5.27 × 10−3 | 5.28 × 10−3 | 2.75 × 10−2 | 9.95 × 10−2 | 5.62 × 10−2 | 5.41 × 10−2 | 3.41 × 10−5 | 3.60 × 10−5 | 3.49 × 10−5 | 3.49 × 10−5 |
| OMOPSO | 4.93 × 10−3 | 5.32 × 10−3 | 5.13 × 10−3 | 5.14 × 10−3 | 2.45 × 10−2 | 8.30 × 10−2 | 5.53 × 10−2 | 5.56 × 10−2 | 3.43 × 10−5 | 3.56 × 10−5 | 3.48 × 10−5 | 3.47 × 10−5 |
| NSGA-II | 3.46 × 10−1 | 1.457 | 6.32 × 10−1 | 5.88 × 10−1 | 6.38 × 10−1 | 1.011 | 8.33 × 10−1 | 8.32 × 10−1 | 1.54 × 10−3 | 9.44 × 10−3 | 3.10 × 10−3 | 2.75 × 10−3 |
| SPEA2 | 5.21 × 10−1 | 1.775 | 1.107 | 1.051 | 8.14 × 10−1 | 1.0162 | 9.38 × 10−1 | 9.51 × 10−1 | 2.50 × 10−3 | 1.16 × 10−2 | 6.34 × 10−3 | 5.31 × 10−3 |
| ZDT3 | ||||||||||||
| MOSFP | 3.37 × 10−3 | 5.28 × 10−3 | 4.13 × 10−3 | 4.11 × 10−3 | 6.99 × 10−1 | 7.02 × 10−1 | 7.01 × 10−1 | 7.01 × 10−1 | 2.78 × 10−5 | 3.08 × 10−5 | 2.91 × 10−5 | 2.91 × 10−5 |
| OMOPSO | 3.54 × 10−3 | 5.25 × 10−3 | 4.18 × 10−3 | 4.06 × 10−3 | 6.99 × 10−1 | 7.01 × 10−1 | 7.00 × 10−1 | 7.00 × 10−1 | 2.74 × 10−5 | 3.04 × 10−5 | 2.86 × 10−5 | 2.86 × 10−5 |
| NSGA-II | 1.51 × 10−1 | 3.36 × 10−1 | 2.10 × 10−1 | 2.01 × 10−1 | 7.82 × 10−1 | 9.39 × 10−1 | 8.63 × 10−1 | 8.62 × 10−1 | 3.96 × 10−4 | 1.40 × 10−3 | 8.73 × 10−4 | 8.87 × 10−4 |
| SPEA2 | 1.91 × 10−1 | 6.06 × 10−1 | 2.92 × 10−1 | 2.86 × 10−1 | 7.88 × 10−1 | 9.37 × 10−1 | 8.64 × 10−1 | 8.62 × 10−1 | 7.43 × 10−4 | 1.98 × 10−3 | 1.30 × 10−3 | 1.31 × 10−3 |
| ZDT6 | ||||||||||||
| MOSFP | 5.32 × 10−3 | 6.53 × 10−3 | 5.90 × 10−3 | 5.91 × 10−3 | 1.33 × 10−1 | 1.81 × 10−1 | 1.55 × 10−1 | 1.55 × 10−1 | 3.21 × 10−5 | 3.35 × 10−5 | 3.27 × 10−5 | 3.27 × 10−5 |
| OMOPSO | 4.73 × 10−3 | 9.00 × 10−3 | 5.59 × 10−3 | 5.42 × 10−3 | 5.59 × 10−2 | 2.53 × 10−1 | 1.13 × 10−1 | 1.08 × 10−1 | 3.09 × 10−5 | 3.41 × 10−5 | 3.19 × 10−5 | 3.17 × 10−5 |
| NSGA-II | 2.34 × 10−2 | 3.53 × 10−2 | 2.71 × 10−2 | 2.65 × 10−2 | 6.04 × 10−2 | 1.36 | 7.41 × 10−1 | 9.53 × 10−1 | 4.67 × 10−5 | 1.37 × 10−3 | 2.03 × 10−4 | 1.60 × 10−4 |
| SPEA2 | 1.85 × 10−2 | 3.84 × 10−1 | 6.98 × 10−2 | 6.09 × 10−2 | 9.03 × 10−1 | 1.436 | 1.332 | 1.345 | 1.99 × 10−4 | 2.29 × 10−4 | 2.04 × 10−4 | 2.03 × 10−4 |
| Method | Metrics | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| SP | IGD | |||||||||||
| Best | Worst | Aver. | Med. | Best | Worst | Aver. | Med. | Best | Worst | Aver. | Med. | |
| WFG1 | ||||||||||||
| MOSFP | 8.59× 10−1 | 1.199 | 1.092 | 1.094 | 8.01 × 10−1 | 1.057 | 9.31 × 10−1 | 9.31 × 10−1 | 2.72 × 10−3 | 4.93 × 10−3 | 4.33 × 10−3 | 4.37 × 10−3 |
| OMOPSO | 7.68 × 10−2 | 1.022 | 1.61 × 10−1 | 1.45 × 10−1 | 7.41 × 10−1 | 1.238 | 8.42 × 10−1 | 8.33 × 10−1 | 3.04 × 10−5 | 3.49 × 10−3 | 1.05 × 10−4 | 4.81 × 10−5 |
| NSGA-II | 1.441 | 2.020 | 1.740 | 1.746 | 8.63 × 10−1 | 1.030 | 9.37 × 10−1 | 9.33 × 10−1 | 3.93 × 10−3 | 6.17 × 10−3 | 5.06 × 10−3 | 5.07 × 10−3 |
| SPEA2 | 1.562 | 2.227 | 1.923 | 1.922 | 8.08 × 10−1 | 1.266 | 1.080 | 1.070 | 4.46 × 10−3 | 7.02 × 10−3 | 5.77 × 10−3 | 5.73 × 10−3 |
| WFG2 | ||||||||||||
| MOSFP | 1.14 × 10−2 | 1.92 × 10−2 | 1.42 × 10−2 | 1.39 × 10−2 | 7.57 × 10−1 | 7.96 × 10−1 | 7.77 × 10−1 | 7.74 × 10−1 | 5.07 × 10−5 | 7.37 × 10−5 | 5.91 × 10−5 | 5.90 × 10−5 |
| OMOPSO | 7.48 × 10−3 | 1.04 × 10−2 | 9.01 × 10−3 | 8.94 × 10−3 | 7.56 × 10−1 | 7.59 × 10−1 | 7.57 × 10−1 | 7.57 × 10−1 | 3.81 × 10−5 | 4.22 × 10−5 | 4.01 × 10−5 | 4.00 × 10−5 |
| NSGA-II | 1.59 × 10−2 | 8.15 × 10−1 | 4.83 × 10−1 | 7.99 × 10−1 | 7.86 × 10−1 | 1.006 | 8.59 × 10−1 | 8.50 × 10−1 | 6.21 × 10−5 | 1.77 × 10−3 | 1.06 × 10−3 | 1.73 × 10−3 |
| SPEA2 | 1.64 × 10−2 | 8.18 × 10−1 | 5.03 × 10−1 | 8.01 × 10−1 | 8.01 × 10−1 | 1.076 | 9.27 × 10−1 | 9.22 × 10−1 | 6.85 × 10−5 | 1.77 × 10−3 | 1.11 × 10−3 | 1.73 × 10−3 |
| WFG3 | ||||||||||||
| MOSFP | 1.55 × 10−2 | 1.81 × 10−2 | 1.81 × 10−2 | 1.66 × 10−2 | 2.40 × 10−2 | 6.63 × 10−2 | 4.22 × 10−2 | 4.29 × 10−2 | 4.48 × 10−5 | 4.81 × 10−5 | 4.62 × 10−5 | 4.61 × 10−5 |
| OMOPSO | 1.39 × 10−2 | 1.51 × 10−2 | 1.43 × 10−2 | 1.43 × 10−2 | 1.54 × 10−2 | 4.85 × 10−2 | 2.95 × 10−2 | 2.89 × 10−2 | 4.37 × 10−5 | 4.64 × 10−5 | 4.49 × 10−5 | 4.5 × 10−5 |
| NSGA-II | 3.37 × 10−2 | 7.05 × 10−2 | 4.60 × 10−2 | 4.43 × 10−2 | 2.62 × 10−1 | 4.10 × 10−1 | 3.44 × 10−1 | 3.44 × 10−1 | 7.06 × 10−5 | 1.35 × 10−4 | 8.55 × 10−5 | 8.34 × 10−5 |
| SPEA2 | 2.74 × 10−2 | 1.12 × 10−1 | 4.87 × 10−2 | 4.44 × 10−2 | 1.70 × 10−1 | 3.09 × 10−1 | 2.16 × 10−1 | 2.16 × 10−1 | 5.99 × 10−5 | 1.32 × 10−4 | 8.58 × 10−5 | 8.60 × 10−5 |
| WFG4 | ||||||||||||
| MOSFP | 2.87 × 10−2 | 1.91 × 10−1 | 4.71 × 10−2 | 4.16 × 10−2 | 3.10 × 10−1 | 4.38 × 10−1 | 3.72 × 10−1 | 3.74 × 10−1 | 1.13 × 10−4 | 1.55 × 10−4 | 1.25 × 10−4 | 1.24 × 10−4 |
| OMOPSO | 1.25 × 10−2 | 4.47 × 10−2 | 1.90 × 10−2 | 1.52 × 10−2 | 9.02 × 10−2 | 2.43 × 10−1 | 1.38 × 10−1 | 1.27 × 10−1 | 8.13 × 10−5 | 1.14 × 10−4 | 8.86 × 10−5 | 8.51 × 10−5 |
| NSGA-II | 6.26 × 10−2 | 1.55 × 10−1 | 1.09 × 10−1 | 1.10 × 10−1 | 4.55 × 10−1 | 6.69 × 10−1 | 5.56 × 10−1 | 5.53 × 10−1 | 2.90 × 10−4 | 4.30 × 10−4 | 3.60 × 10−4 | 3.54 × 10−4 |
| SPEA2 | 3.01 × 10−2 | 4.17 × 10−1 | 1.04 × 10−1 | 7.02 × 10−2 | 2.46 × 10−1 | 3.51 × 10−1 | 3.00 × 10−1 | 3.02 × 10−1 | 1.01 × 10−4 | 1.74 × 10−4 | 1.16 × 10−4 | 1.14 × 10−4 |
| WFG5 | ||||||||||||
| MOSFP | 4.99 × 10−2 | 6.52 × 10−2 | 5.61 × 10−2 | 5.59 × 10−2 | 9.78 × 10−2 | 1.76 × 10−1 | 1.40 × 10−1 | 1.40 × 10−1 | 2.66 × 10−4 | 4.24 × 10−4 | 3.00 × 10−4 | 2.95 × 10−4 |
| OMOPSO | 4.93 × 10−2 | 9.35 × 10−2 | 5.73 × 10−2 | 5.71 × 10−2 | 9.05 × 10−2 | 3.47 × 10−1 | 1.19 × 10−1 | 1.18 × 10−1 | 2.57 × 10−4 | 6.84 × 10−4 | 3.05 × 10−4 | 2.95 × 10−4 |
| NSGA-II | 5.30 × 10−2 | 1.33 × 10−1 | 1.04 × 10−1 | 1.04 × 10−1 | 3.15 × 10−1 | 4.55 × 10−1 | 3.88 × 10−1 | 3.85 × 10−1 | 3.14 × 10−4 | 1.15 × 10−3 | 8.11 × 10−4 | 8.26 × 10−4 |
| SPEA2 | 9.73 × 10−2 | 1.50 × 10−1 | 1.19 × 10−1 | 1.17 × 10−1 | 2.29 × 10−1 | 3.34 × 10−1 | 2.82 × 10−1 | 2.81 × 10−1 | 7.32 × 10−4 | 1.30 × 10−3 | 9.75 × 10−4 | 947 × 10−4 |
| WFG6 | ||||||||||||
| MOSFP | 1.55 × 10−2 | 2.08 × 10−2 | 1.70 × 10−2 | 1.69 × 10−2 | 8.58 × 10−2 | 1.52 × 10−1 | 1.20 × 10−1 | 1.22 × 10−1 | 4.98 × 10−5 | 5.46 × 10−5 | 5.21 × 10−5 | 5.21 × 10−5 |
| OMOPSO | 1.29 × 10−2 | 1.45 × 10−2 | 1.36 × 10−2 | 1.36 × 10−2 | 7.70 × 10−2 | 1.42 × 10−1 | 1.05 × 10−1 | 1.06 × 10−1 | 4.71 × 10−5 | 5.16 × 10−5 | 4.90 × 10−5 | 4.88 × 10−5 |
| NSGA-II | 3.49 × 10−2 | 1.30 × 10−1 | 6.10 × 10−2 | 5.75 × 10−2 | 3.19 × 10−1 | 5.17 × 10−1 | 3.92 × 10−1 | 3.86 × 10−1 | 7.59 × 10−5 | 5.32 × 10−4 | 1.64 × 10−4 | 1.45 × 10−4 |
| SPEA2 | 3.41 × 10−2 | 2.13 × 10−1 | 8.74 × 10−2 | 7.79 × 10−2 | 2.52 × 10−1 | 6.56 × 10−1 | 3.61 × 10−1 | 3.44 × 10−1 | 7.17 × 10−5 | 4.08 × 10−4 | 1.76 × 10−4 | 1.67 × 10−4 |
| WFG7 | ||||||||||||
| MOSFP | 1.47 × 10−2 | 1.89 × 10−2 | 1.57 × 10−2 | 1.56 × 10−2 | 9.38 × 10−2 | 1.44 × 10−1 | 1.20 × 10−1 | 1.20 × 10−1 | 4.94 × 10−5 | 5.38 × 10−5 | 5.14 × 10−5 | 5.14 × 10−5 |
| OMOPSO | 1.29 × 10−2 | 1.45 × 10−2 | 1.36 × 10−2 | 1.36 × 10−2 | 8.68 × 10−2 | 1.38 × 10−1 | 1.10 × 10−1 | 1.09 × 10−1 | 4.71 × 10−5 | 5.10 × 10−5 | 4.93 × 10−5 | 4.93 × 10−5 |
| NSGA-II | 2.98 × 10−2 | 7.71 × 10−2 | 4.23 × 10−2 | 3.98 × 10−2 | 3.13 × 10−1 | 4.48 × 10−1 | 3.82 × 10−1 | 3.79 × 10−1 | 6.75 × 10−5 | 8.45 × 10−5 | 7.47 × 10−5 | 7.42 × 10−5 |
| SPEA2 | 2.85 × 10−2 | 3.34 × 10−1 | 8.82 × 10−2 | 7.23 × 10−2 | 2.41 × 10−1 | 4.41 × 10−1 | 2.92 × 10−1 | 2.89 × 10−1 | 6.00 × 10−5 | 3.34 × 10−4 | 7.35 × 10−5 | 6.85 × 10−5 |
| WFG8 | ||||||||||||
| MOSFP | 3.05 × 10−1 | 4.82 × 10−1 | 3.66 × 10−1 | 3.66 × 10−1 | 6.30 × 10−1 | 1.071 | 7.54 × 10−1 | 7.34 × 10−1 | 1.73 × 10−3 | 2.18 × 10−3 | 2.02 × 10−3 | 2.02 × 10−3 |
| OMOPSO | 5.31 × 10−2 | 4.89 × 10−1 | 3.78 × 10−1 | 4.87 × 10−1 | 4.05 × 10−1 | 7.83 × 10−1 | 5.06 × 10−1 | 4.89 × 10−1 | 2.33 × 10−4 | 2.11 × 10−3 | 1.79 × 10−3 | 2.08 × 10−3 |
| NSGA-II | 1.94 × 10−1 | 7.27 × 10−1 | 4.62 × 10−1 | 4.97 × 10−1 | 6.60 × 10−1 | 9.34 × 10−1 | 7.95 × 10−1 | 7.94 × 10−1 | 1.10 × 10−3 | 2.61 × 10−3 | 2.26 × 10−3 | 2.37 × 10−3 |
| SPEA2 | 3.99 × 10−1 | 8.15 × 10−1 | 6.14 × 10−1 | 6.00 × 10−1 | 6.45 × 10−1 | 9.09 × 10−1 | 7.67 × 10−1 | 7.58 × 10−1 | 2.04 × 10−3 | 2.68 × 10−3 | 2.44 × 10−3 | 2.46 × 10−3 |
| WFG9 | ||||||||||||
| MOSFP | 3.59 × 10−2 | 1.05 × 10−1 | 9.26 × 10−2 | 9.28 × 10−2 | 1.37 × 10−1 | 2.27 × 10−1 | 1.84 × 10−1 | 1.83 × 10−1 | 8.15 × 10−5 | 9.25 × 10−5 | 8.69 × 10−5 | 8.70 × 10−5 |
| OMOPSO | 1.53 × 10−2 | 8.35 × 10−2 | 7.58 × 10−2 | 7.81 × 10−2 | 6.31 × 10−2 | 1.20 × 10−1 | 9.51 × 10−2 | 9.56 × 10−2 | 5.32 × 10−5 | 5.77 × 10−5 | 5.50 × 10−5 | 5.50 × 10−5 |
| NSGA-II | 8.76 × 10−2 | 1.76 × 10−1 | 1.09 × 10−1 | 1.04 × 10−1 | 3.05 × 10−1 | 4.39 × 10−1 | 3.63 × 10−1 | 3.65 × 10−1 | 8.17 × 10−5 | 1.19 × 10−4 | 9.87 × 10−5 | 9.81 × 10−5 |
| SPEA2 | 8.86 × 10−2 | 2.44 × 10−1 | 1.30 × 10−1 | 1.21 × 10−1 | 2.29 × 10−1 | 3.25 × 10−1 | 2.86 × 10−1 | 2.86 × 10−1 | 7.76 × 10−5 | 1.54 × 10−4 | 9.38 × 10−5 | 9.32 × 10−5 |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shehadeh, H.A.; Ldris, M.Y.I.; Ahmedy, I. Multi-Objective Optimization Algorithm Based on Sperm Fertilization Procedure (MOSFP). Symmetry 2017, 9, 241. https://doi.org/10.3390/sym9100241
Shehadeh HA, Ldris MYI, Ahmedy I. Multi-Objective Optimization Algorithm Based on Sperm Fertilization Procedure (MOSFP). Symmetry. 2017; 9(10):241. https://doi.org/10.3390/sym9100241
Chicago/Turabian StyleShehadeh, Hisham A., Mohd Yamani Idna Ldris, and Ismail Ahmedy. 2017. "Multi-Objective Optimization Algorithm Based on Sperm Fertilization Procedure (MOSFP)" Symmetry 9, no. 10: 241. https://doi.org/10.3390/sym9100241
APA StyleShehadeh, H. A., Ldris, M. Y. I., & Ahmedy, I. (2017). Multi-Objective Optimization Algorithm Based on Sperm Fertilization Procedure (MOSFP). Symmetry, 9(10), 241. https://doi.org/10.3390/sym9100241





