Kernel Partial Least Square Regression with High Resistance to Multiple Outliers and Bad Leverage Points on Near-Infrared Spectral Data Analysis
Abstract
1. Introduction
2. Kernel Partial Least Square Regression
3. Proposed Methods
3.1. Kernel Partial Robust GM6-Estimator (KPRGM6)
3.2. Kernel Partial Robust Modified GM6-Estimator
3.2.1. Reweighted Least Squares Based on Least Median of Squares
3.2.2. Diagnostic Robust Generalized Potential Based on Index Set Equality
3.2.3. Fast Modified Generalized Studentized Residuals
- (i)
- Observation is classified as a regular observation: If and .
- (ii)
- Observation is classified as vertical outliers: If and .
- (iii)
- Observation is classified as GLPs: If and .
- (iv)
- Observation is classified as BLPs: If and .
3.2.4. Proposed Algorithm in Kernel Partial Robust Modified GM6-Estimator
- Step 1:
- Compute the kernel Gram matrix in Equation (7) of the cross dot products among all mapped input data points.
- Step 2:
- Centralize the kernel Gram matrix as in Equation (9).
- Step 3:
- Uses identity matrix as the initial weight on the centralized kernel matrix to obtain the weighted and output vector , .
- Step 4:
- Regress on to obtain the weight , then apply the normalization and rename it as .
- Step 5:
- Continue the steps until convergence and number of PLS are determined. Here .
- Step 6:
- The new calculated latent variables denoted as the matrix of are used as the new input space.
- Step 7:
- Calculate the residual based on the initial LTS estimator using new latent variables .
- Step 8:
- Calculate the scale estimate of the residuals in Step 7.
- Step 9:
- Calculate the standardized residuals using and .
- Step 10:
- Compute the proposed improvement of initial weight in Equation (10) using and cut-off point , .
- Step 11:
- Calculate the bounded influence function for BLPs using standardized residuals and improvised initial weight , ..
- Step 12:
- Apply the weighted least squares (WLS) iteratively to obtain the parameter estimates of as in Equation (10).
- Step 13:
- Calculate the new residual from WLS and repeat Steps (8–12) until convergence.
4. Results and Discussions
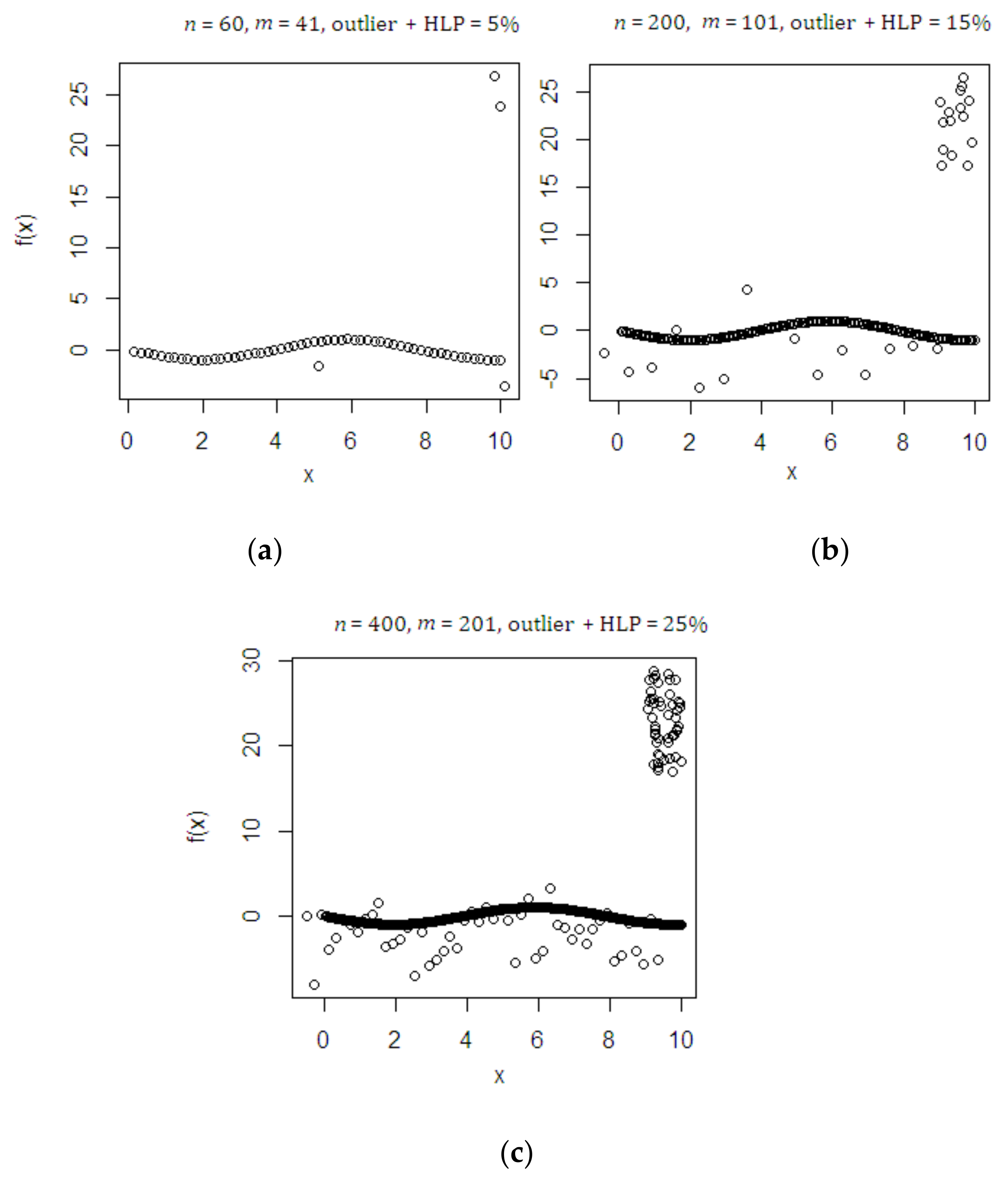
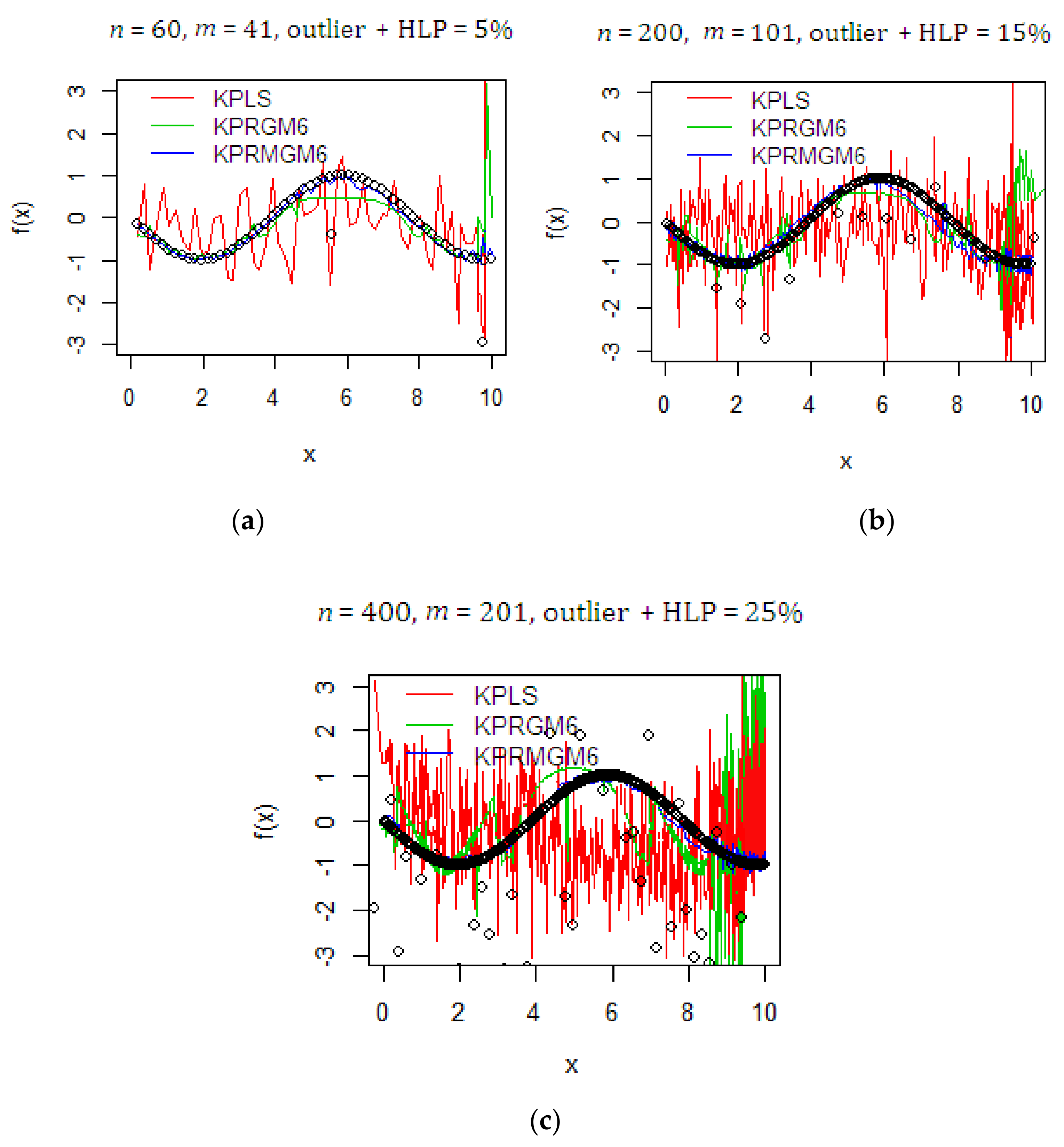
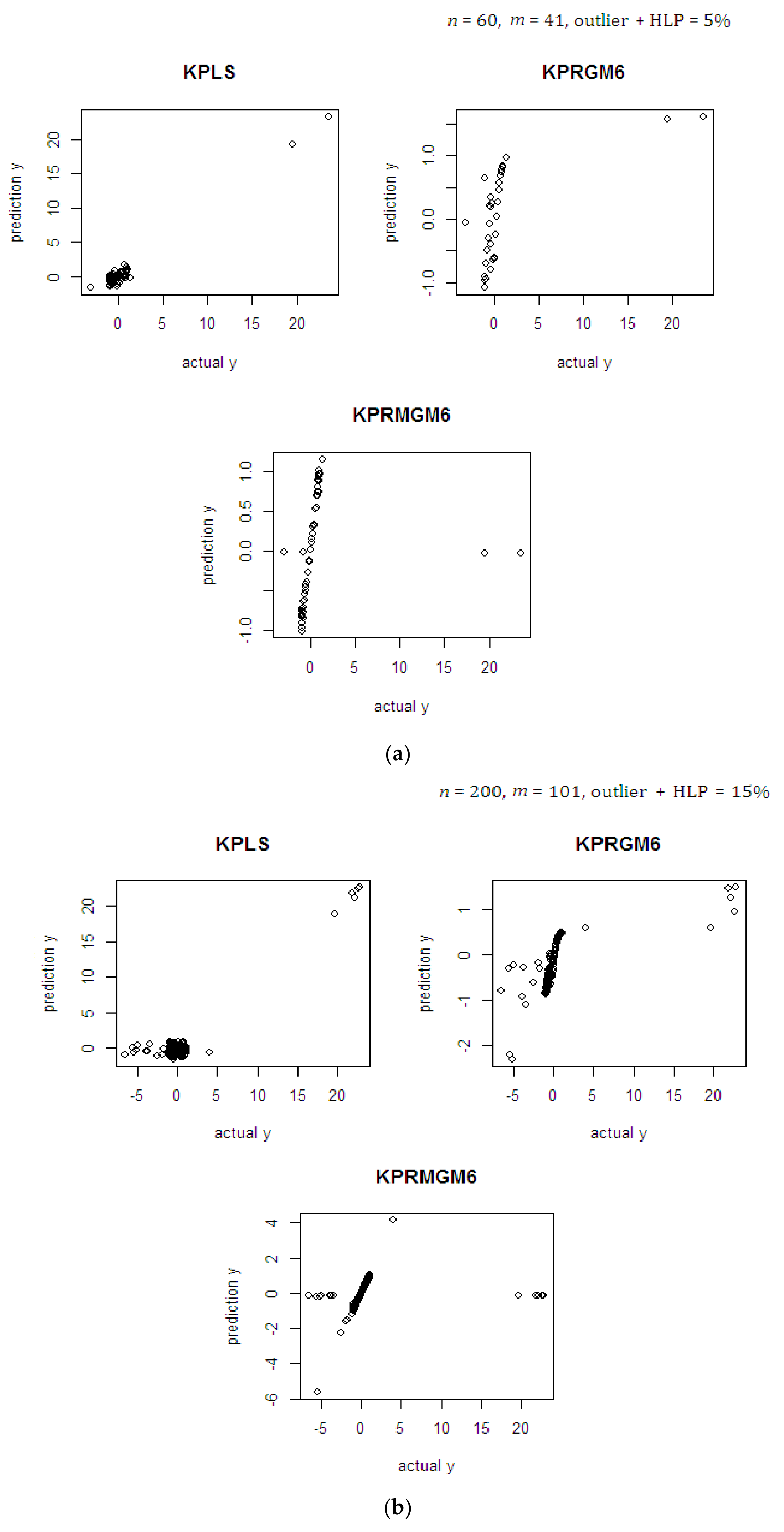
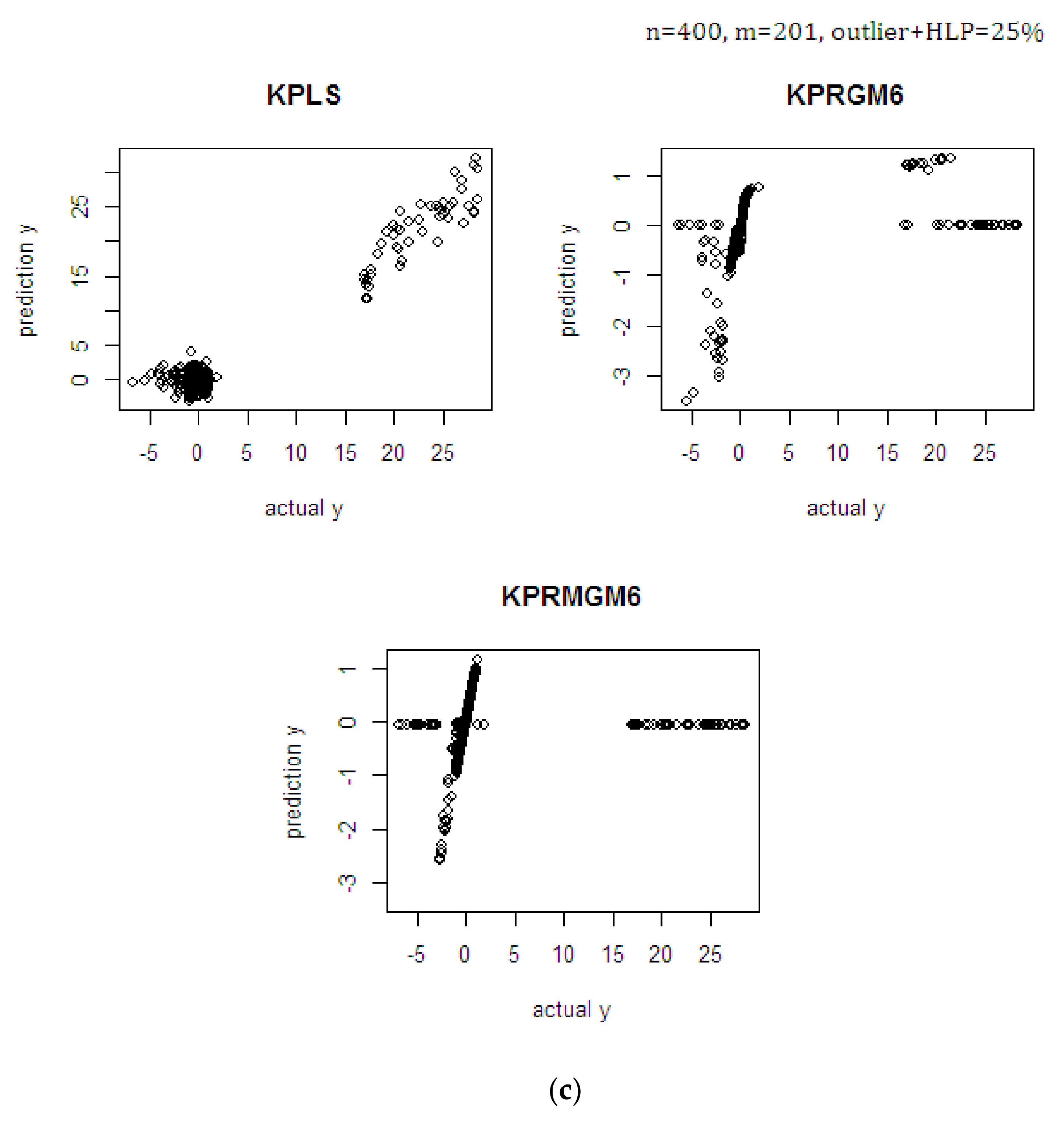
4.1. Monte Carlo Simulation Study
4.2. NIR Spectral Data
4.2.1. Oil to Dry Mesocarp
4.2.2. Oil to Wet Mesocarp
4.2.3. Fat Fatty Acids
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
List of Abbreviations
| Abbreviations | Full Form |
| ASD | Analytical Spectral Devices |
| BLPs | Bad Leverage Points |
| CP | Cut-off Point |
| CSV | Comma-Separated Values |
| DL | Deep Learning |
| DRGP | Diagnostic Robust Generalized Potential |
| FFA | Fat Fatty Acid |
| FMGT | Fast Modified Generalized Studentized |
| GLPs | Good Leverage Points |
| HLPs | High Leverage Points |
| ISE | Index Set Equality |
| KPLS | Kernel Partial Least Square |
| KPRGM6 | Kernel Partial Robust GM6-Estimator |
| KPRMGM6 | Kernel Partial Robust Modified GM6-Estimator |
| KPRMGM6 | Kernel Partial Robust M-Estimator |
| LMS | Least Median of Squares |
| LTS | Least Trimmed of Squares |
| MCD | Minimizing Covariance Determinant |
| MGT | Modified Generalized Studentized |
| ML | Machine Learning |
| MVE | Minimum Volume Ellipsoid |
| NIPALS | Nonlinear Iterative Partial Least Squares |
| NIR | Near-Infrared |
| ODM | Oil to Dry Mesocarp |
| OWM | Oil to Wet Mesocarp |
| PLSR | Partial Least Square Regression |
| RKHS | Reproducing Kernel Hilbert Spaces |
| RLS | Reweighted Least Squares |
| RMD | Robust Mahalanobis Distance |
| RMSE | Root Mean Square Error |
| SE | Standard Error |
| WLS | Weighted Least Squares |
Appendix A
| Outliers and HLPs | n | m | Methods | RMSE | R2 | SE |
|---|---|---|---|---|---|---|
| With outliers and HLPs (5%) | 60 | 41 | KPLS | 2.933 | 0.523 | 2.957 |
| KPRGM6 | 0.458 | 0.706 | 0.462 | |||
| KPRMGM6 | 0.140 | 0.921 | 0.140 | |||
| 60 | 101 | KPLS | 2.985 | 0.515 | 3.010 | |
| KPRGM6 | 0.477 | 0.640 | 0.481 | |||
| KPRMGM6 | 0.142 | 0.912 | 0.142 | |||
| 60 | 201 | KPLS | 3.051 | 0.522 | 3.077 | |
| KPRGM6 | 0.500 | 0.658 | 0.504 | |||
| KPRMGM6 | 0.098 | 0.932 | 0.101 | |||
| 200 | 41 | KPLS | 2.778 | 0.438 | 2.785 | |
| KPRGM6 | 0.422 | 0.663 | 0.423 | |||
| KPRMGM6 | 0.120 | 0.910 | 0.120 | |||
| 200 | 101 | KPLS | 2.701 | 0.430 | 2.707 | |
| KPRGM6 | 0.393 | 0.688 | 0.394 | |||
| KPRMGM6 | 0.123 | 0.909 | 0.124 | |||
| 200 | 201 | KPLS | 2.762 | 0.429 | 2.769 | |
| KPRGM6 | 0.391 | 0.652 | 0.392 | |||
| KPRMGM6 | 0.163 | 0.895 | 0.163 | |||
| 400 | 41 | KPLS | 2.830 | 0.418 | 2.834 | |
| KPRGM6 | 0.383 | 0.702 | 0.384 | |||
| KPRMGM6 | 0.198 | 0.814 | 0.200 | |||
| 400 | 101 | KPLS | 2.772 | 0.421 | 2.775 | |
| KPRGM6 | 0.415 | 0.674 | 0.416 | |||
| KPRMGM6 | 0.122 | 0.910 | 0.122 | |||
| 400 | 201 | KPLS | 2.855 | 0.427 | 2.859 | |
| KPRGM6 | 0.352 | 0.712 | 0.353 | |||
| KPRMGM6 | 0.104 | 0.959 | 0.104 | |||
| With outliers and HLPs (15%) | 60 | 41 | KPLS | 3.456 | 0.560 | 3.485 |
| KPRGM6 | 0.559 | 0.620 | 0.563 | |||
| KPRMGM6 | 0.187 | 0.859 | 0.189 | |||
| 60 | 101 | KPLS | 3.459 | 0.564 | 3.488 | |
| KPRGM6 | 0.553 | 0.700 | 0.558 | |||
| KPRMGM6 | 0.187 | 0.860 | 0.189 | |||
| 60 | 201 | KPLS | 3.484 | 0.580 | 3.513 | |
| KPRGM6 | 0.504 | 0.664 | 0.508 | |||
| KPRMGM6 | 0.125 | 0.930 | 0.126 | |||
| 200 | 41 | KPLS | 3.533 | 0.592 | 3.542 | |
| KPRGM6 | 0.572 | 0.736 | 0.573 | |||
| KPRMGM6 | 0.214 | 0.841 | 0.215 | |||
| 200 | 101 | KPLS | 3.359 | 0.597 | 3.368 | |
| KPRGM6 | 0.532 | 0.672 | 0.533 | |||
| KPRMGM6 | 0.139 | 0.898 | 0.139 | |||
| 200 | 201 | KPLS | 3.330 | 0.602 | 3.338 | |
| KPRGM6 | 0.493 | 0.707 | 0.495 | |||
| KPRMGM6 | 0.171 | 0.872 | 0.172 | |||
| 400 | 41 | KPLS | 3.505 | 0.589 | 3.510 | |
| KPRGM6 | 0.589 | 0.662 | 0.590 | |||
| KPRMGM6 | 0.217 | 0.815 | 0.218 | |||
| 400 | 101 | KPLS | 3.529 | 0.619 | 3.534 | |
| KPRGM6 | 0.515 | 0.645 | 0.515 | |||
| KPRMGM6 | 0.195 | 0.807 | 0.196 | |||
| 400 | 201 | KPLS | 3.405 | 0.639 | 3.409 | |
| KPRGM6 | 0.525 | 0.658 | 0.526 | |||
| KPRMGM6 | 0.138 | 0.851 | 0.139 | |||
| With outliers and HLPs (25%) | 60 | 41 | KPLS | 3.585 | 0.676 | 3.615 |
| KPRGM6 | 0.749 | 0.696 | 0.755 | |||
| KPRMGM6 | 0.131 | 0.899 | 0.132 | |||
| 60 | 101 | KPLS | 3.535 | 0.642 | 3.565 | |
| KPRGM6 | 0.680 | 0.670 | 0.686 | |||
| KPRMGM6 | 0.248 | 0.872 | 0.250 | |||
| 60 | 201 | KPLS | 3.471 | 0.687 | 3.501 | |
| KPRGM6 | 0.464 | 0.694 | 0.468 | |||
| KPRMGM6 | 0.154 | 0.843 | 0.155 | |||
| 200 | 41 | KPLS | 3.672 | 0.586 | 3.681 | |
| KPRGM6 | 0.653 | 0.640 | 0.655 | |||
| KPRMGM6 | 0.240 | 0.749 | 0.241 | |||
| 200 | 101 | KPLS | 3.779 | 0.680 | 3.789 | |
| KPRGM6 | 0.616 | 0.754 | 0.618 | |||
| KPRMGM6 | 0.136 | 0.896 | 0.137 | |||
| 200 | 201 | KPLS | 3.722 | 0.681 | 3.731 | |
| KPRGM6 | 0.591 | 0.717 | 0.592 | |||
| KPRMGM6 | 0.252 | 0.855 | 0.253 | |||
| 400 | 41 | KPLS | 3.646 | 0.633 | 3.651 | |
| KPRGM6 | 0.641 | 0.657 | 0.642 | |||
| KPRMGM6 | 0.248 | 0.755 | 0.249 | |||
| 400 | 101 | KPLS | 3.616 | 0.686 | 3.621 | |
| KPRGM6 | 0.578 | 0.694 | 0.579 | |||
| KPRMGM6 | 0.236 | 0.771 | 0.236 | |||
| 400 | 201 | KPLS | 3.679 | 0.684 | 3.684 | |
| KPRGM6 | 0.559 | 0.720 | 0.559 | |||
| KPRMGM6 | 0.224 | 0.785 | 0.225 |
References
- Midi, H.; Norazan, M.; Imon, A.H.M. The performance of diagnostic-robust generalized potentials for the identification of multiple high leverage points in linear regression. J. Appl. Stat. 2009, 36, 507–520. [Google Scholar]
- Bagheri, A.; Midi, H. Diagnostic plot for the identification of high leverage collinearity-influential observations. Sort Stat. Oper. Res. Trans. 2015, 39, 51–70. [Google Scholar]
- Alguraibawi, M.; Midi, H.; Imon, A.H.M. A new robust diagnostic plot for classifying good and bad high leverage points in a multiple linear regression model. Math. Probl. Eng. 2015, 2015, 1–12. [Google Scholar] [CrossRef]
- Atkinson, A.C. Fast very robust methods for the detection of multiple outliers. J. Am. Stat. Assoc. 1994, 89, 1329–1339. [Google Scholar] [CrossRef]
- Imon, A.H.M. Identifying multiple high leverage points in linear regression. J. Stat. Stud. 2002, 3, 207–218. [Google Scholar]
- Serneels, S.; Croux, C.; Filzmoser, P.; Van Espen, P.J. Partial robust M-regression. Chemom. Intell. Lab. Syst. 2005, 79, 55–64. [Google Scholar] [CrossRef]
- Jia, R.D.; Mao, Z.Z.; Chang, Y.Q.; Zhang, S.N. Kernel partial robust M-regression as a flexible robust nonlinear modeling technique. Chemom. Intell. Lab. Syst. 2010, 100, 91–98. [Google Scholar] [CrossRef]
- Wold, H. Multivariate Analysis; Krishnaiah, P.R., Ed.; Academic Press: New York, NY, USA, 1973; Volume 3, pp. 383–407. [Google Scholar]
- Rosipal, R. Nonlinear partial least squares an overview. In Chemoinformatics and Advanced Machine Learning Perspectives: Complex Computational Methods and Collaborative Techniques; IGI Global: Hershey, PA, USA, 2011; pp. 169–189. [Google Scholar]
- Yang, H.; Griffiths, P.R.; Tate, J.D. Comparison of partial least squares regression and multi-layer neural networks for quantification of nonlinear systems and application to gas phase Fourier transform infrared spectra. Anal. Chim. Acta 2003, 489, 125–136. [Google Scholar] [CrossRef]
- Balabin, R.M.; Safieva, R.Z.; Lomakina, E.I. Comparison of linear and nonlinear calibration models based on near infrared (NIR) spectroscopy data for gasoline properties prediction. Chemom. Intell. Lab. Syst. 2007, 88, 183–188. [Google Scholar] [CrossRef]
- Schölkopf, B.; Smola, A.; Müller, K.R. Nonlinear component analysis as a kernel eigenvalue problem. Neural Comput. 1998, 10, 1299–1319. [Google Scholar] [CrossRef]
- Rosipal, R.; Trejo, L.J. Kernel partial least squares regression in reproducing kernel hilbert space. J. Mach. Learn. Res. 2001, 2, 97–123. [Google Scholar]
- Bennett, K.P.; Embrechts, M.J. An optimization perspective on kernel partial least squares regression. Nato Sci. Ser. Sub Ser. Iii Comput. Syst. Sci. 2003, 190, 227–250. [Google Scholar]
- Sindhwani, V.; Minh, H.Q.; Lozano, A.C. Scalable matrix-valued kernel learning for high-dimensional nonlinear multivariate regression and Granger Causality. In Proceedings of the Twenty-Ninth Conference on Uncertainty in Artificial Intelligence, Bellevue, WA, USA, 11–15 August 2013; pp. 586–595. [Google Scholar]
- Ma, X.; Zhang, Y.; Cao, H.; Zhang, S.; Zhou, Y. Nonlinear regression with high-dimensional space mapping for blood component spectral quantitative analysis. J. Spectrosc. 2018, 1–8. [Google Scholar] [CrossRef]
- Aronszajn, N. Theory of reproducing kernels. Trans. Am. Math. Soc. 1950, 68, 337–404. [Google Scholar] [CrossRef]
- Preda, C. Regression models for functional data by reproducing kernel Hilbert spaces methods. J. Stat. Plan. Inference 2007, 137, 829–840. [Google Scholar] [CrossRef]
- Coakley, C.W.; Hettmansperger, T.P. A bounded influence, high breakdown, efficient regression estimator. J. Am. Stat. Assoc. 1993, 88, 872–880. [Google Scholar] [CrossRef]
- Rousseeuw, P.J. Regression techniques with high breakdown point. Inst. Math. Stat. Bull. 1983, 12, 155. [Google Scholar]
- Rousseeuw, P.J. Multivariate estimation with high breakdown point. In Mathematical Statistics and Applications; Grossmann, W., Pflug, G., Vincze, I., Wertz, W., Eds.; Cengage Learning: Belmont, CA, USA, 1985; Volume 37, pp. 283–297. [Google Scholar]
- Rousseeuw, P.J.; Leroy, A.M. Robust Regression and Outlier Detection. In Wiley Series in Probability and Mathematical Statistics: Applied Probability and Statistics; Wiley: New York, NY, USA, 1987. [Google Scholar]
- Rousseeuw, P.J. Least median of squares regression. J. Am. Stat. Assoc. 1984, 79, 871–880. [Google Scholar] [CrossRef]
- Midi, H. Robust Estimation of a Linearized Nonlinear Regression Model with Heteroscedastic Errors: A Simulation Study. Pertanika J. Sci. Technol. 1998, 6, 23–35. [Google Scholar]
- De Haan, J.; Sturm, J.-E. No Need to Run Millions of Regressions. Available at SSRN 246453 2000, 1–12. Available online: https://papers.ssrn.com/sol3/papers.cfm?abstract_id=246453 (accessed on 13 October 2020).
- Midi, H.; Hendi, H.T.; Arasan, J.; Uraibi, H. Fast and Robust Diagnostic Technique for the Detection of High Leverage Points. Pertanika J. Sci. Technol. 2020, 28, 1203–1220. [Google Scholar] [CrossRef]
- Silalahi, D.D.; Midi, H.; Arasan, J.; Mustafa, M.S.; Caliman, J.P. Kernel partial diagnostic robust potential to handle high-dimensional and irregular data space on near infrared spectral data. Heliyon 2020, 6, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Lim, H.A.; Midi, H. Diagnostic Robust Generalized Potential Based on Index Set Equality (DRGP (ISE)) for the identification of high leverage points in linear model. Comput. Stat. 2016, 31, 859–877. [Google Scholar] [CrossRef]
- Minasny, B.; McBratney, A. Why you don’t need to use RPD. Pedometron 2013, 33, 14–15. [Google Scholar]
- Rännar, S.; Lindgren, F.; Geladi, P.; Wold, S. A PLS kernel algorithm for data sets with many variables and fewer objects. Part 1: Theory and algorithm. J. Chemom. 1994, 8, 111–125. [Google Scholar] [CrossRef]
- Wold, H. Soft modelling by latent variables: The non-linear iterative partial least squares (NIPALS) approach. J. Appl. Probab. 1975, 12, 117–142. [Google Scholar] [CrossRef]
- Cummins, D.J.; Andrews, C.W. Iteratively reweighted partial least squares: A performance analysis by Monte Carlo simulation. J. Chemom. 1995, 9, 489–507. [Google Scholar] [CrossRef]
- Huber, P.J. Robust regression: Asymptotics, conjectures and Monte Carlo. Ann. Stat. 1973, 1, 799–821. [Google Scholar] [CrossRef]
- Rousseeuw, P.J.; Driessen, K.V. A fast algorithm for the minimum covariance determinant estimator. Technometrics 1999, 41, 212–223. [Google Scholar] [CrossRef]
- Rousseeuw, P.J.; Croux, C. Alternatives to the median absolute deviation. J. Am. Stat. Assoc. 1993, 88, 1273–1283. [Google Scholar] [CrossRef]
- Stuart, B. Infrared Spectroscopy: Fundamentals and Applications; Wiley: Toronto, ON, Canada, 2004; pp. 167–185. [Google Scholar]
- Silalahi, D.D.; Midi, H.; Arasan, J.; Mustafa, M.S.; Caliman, J.P. Robust Wavelength Selection Using Filter-Wrapper Method and Input Scaling on Near Infrared Spectral Data. Sensors 2020, 20, 5001. [Google Scholar] [CrossRef] [PubMed]
- Siew, W.L.; Tan, Y.A.; Tang, T.S. Methods of Test for Palm Oil and Palm Oil Products: Compiled; Lin, S.W., Sue, T.T., Ai, T.Y., Eds.; Palm Oil Research Institute of Malaysia: Selangor, Malaysia, 1995. [Google Scholar]
- Rao, V.; Soh, A.C.; Corley, R.H.V.; Lee, C.H.; Rajanaidu, N. Critical Reexamination of the Method of Bunch Quality Analysis in Oil Palm Breeding; PORIM Occasional Paper; FAO: Rome, Italy, 1983; Available online: https://agris.fao.org/agris-search/search.do?recordID=US201302543052 (accessed on 13 October 2020).
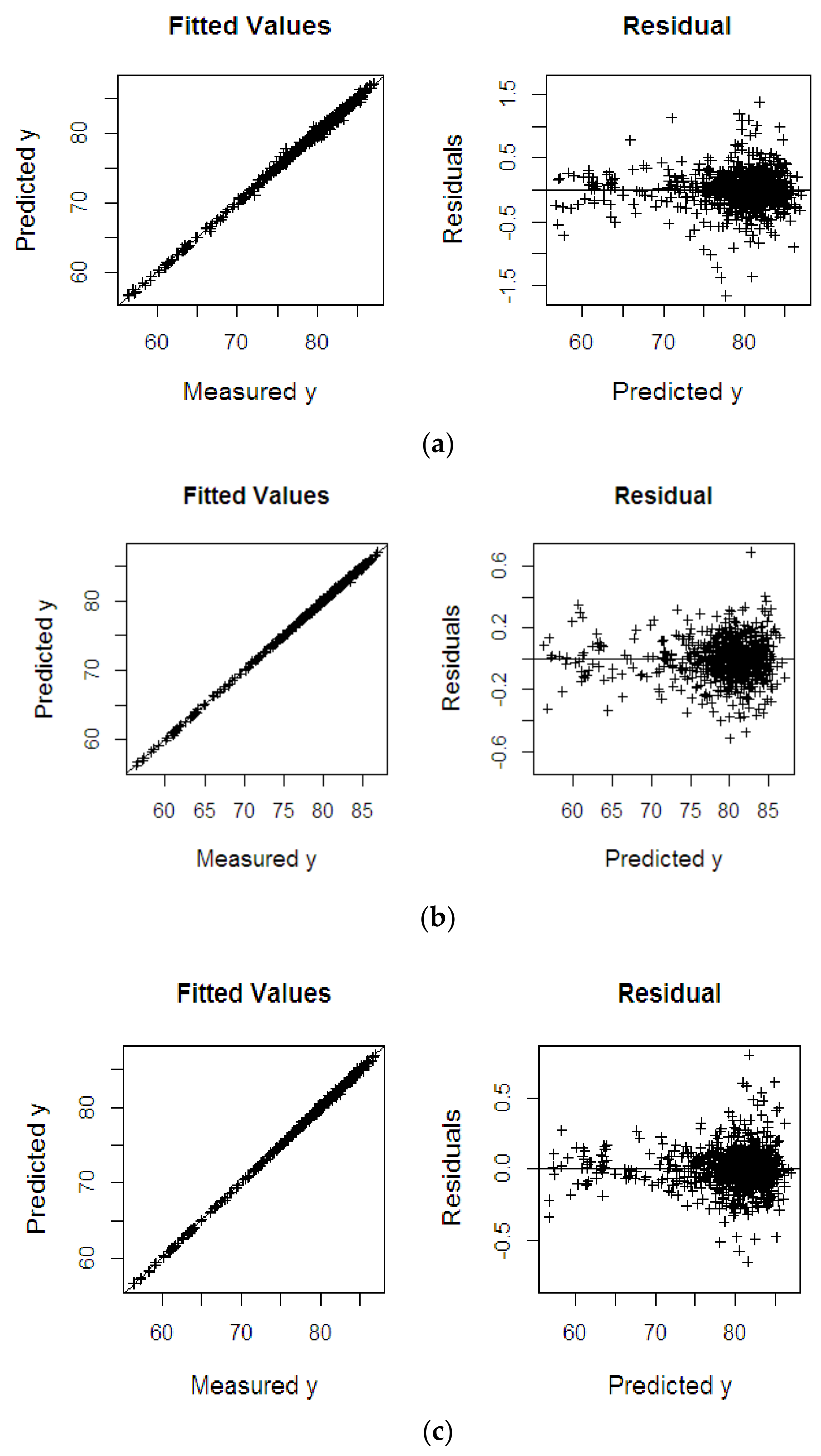
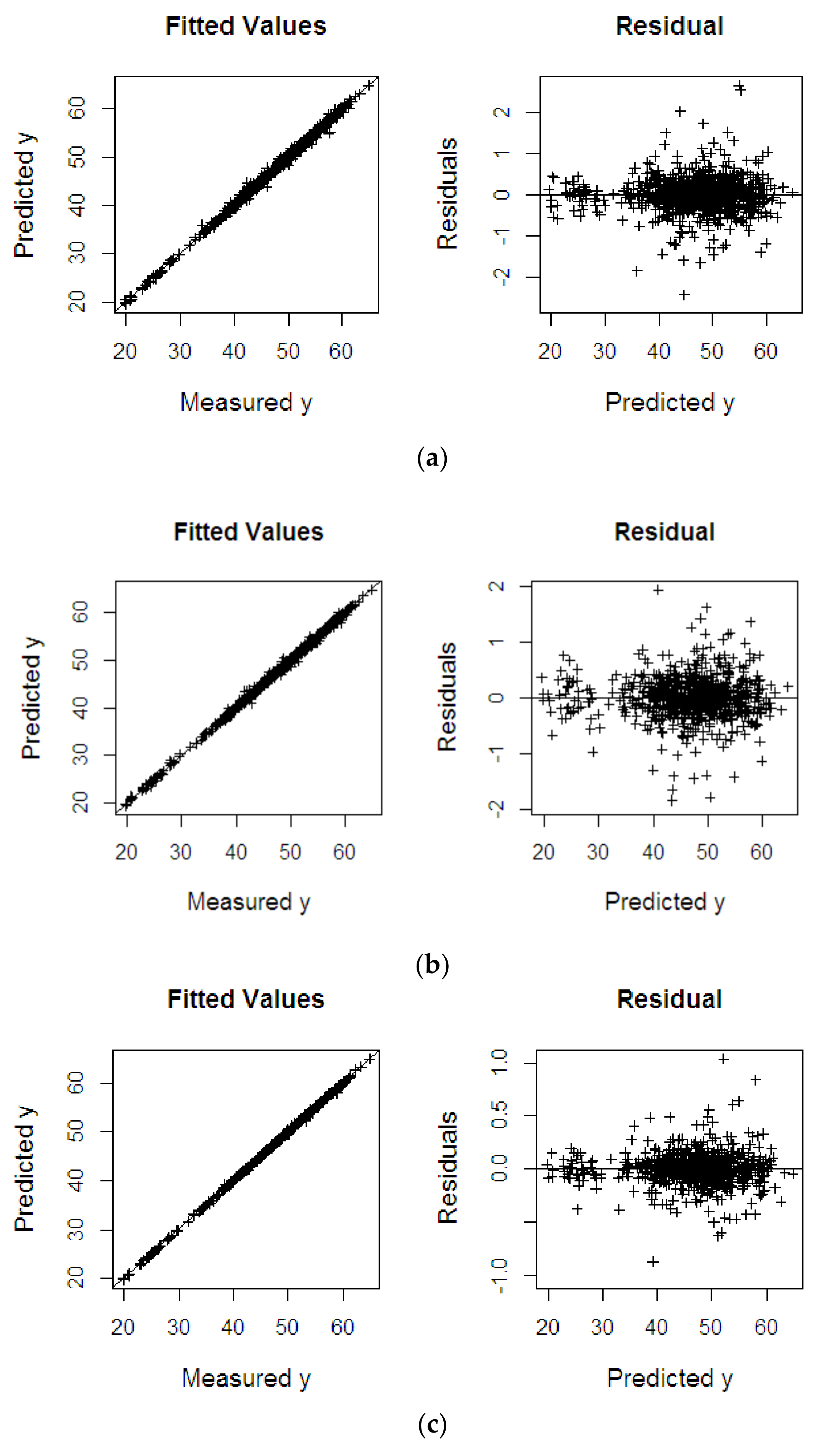
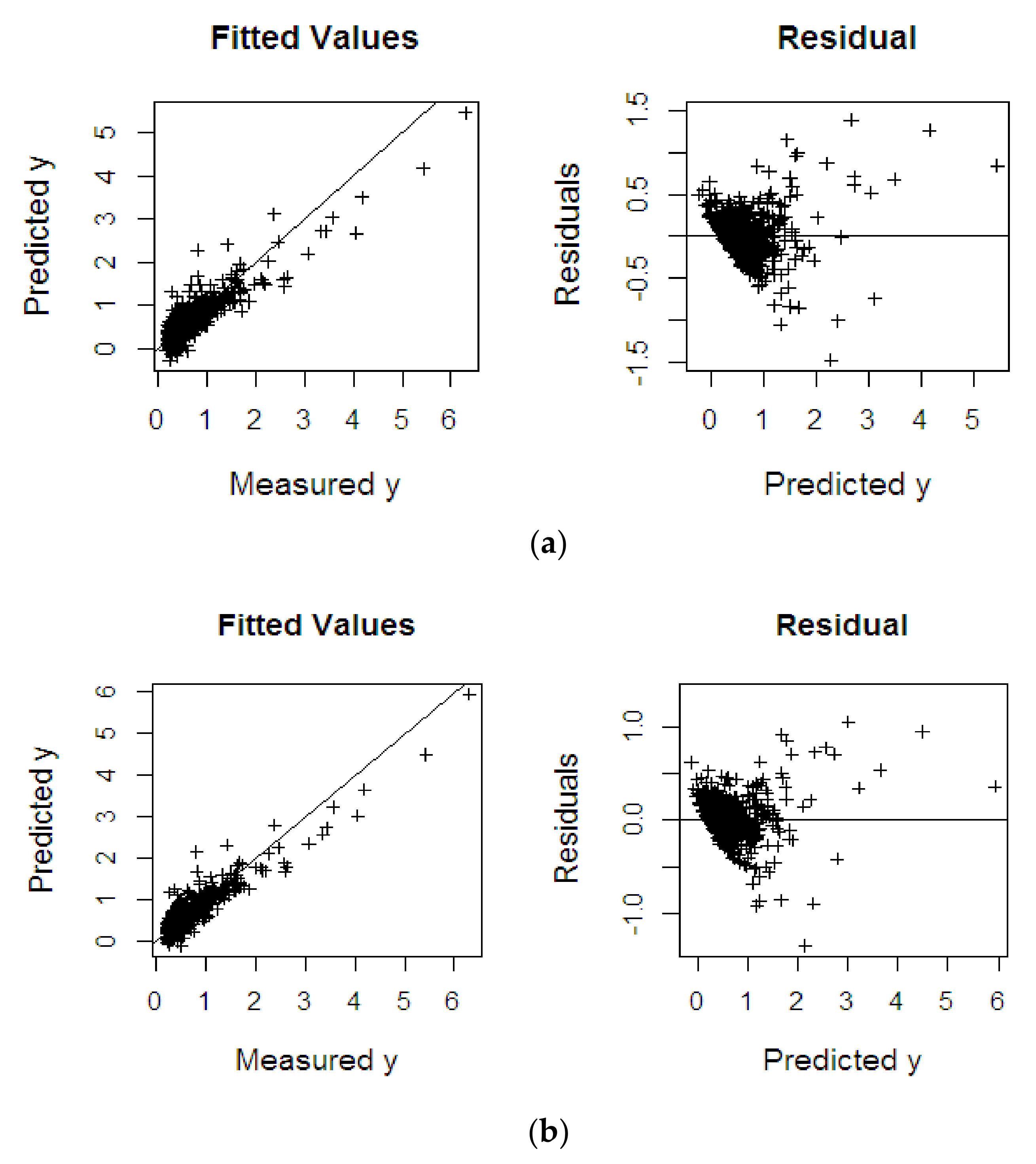
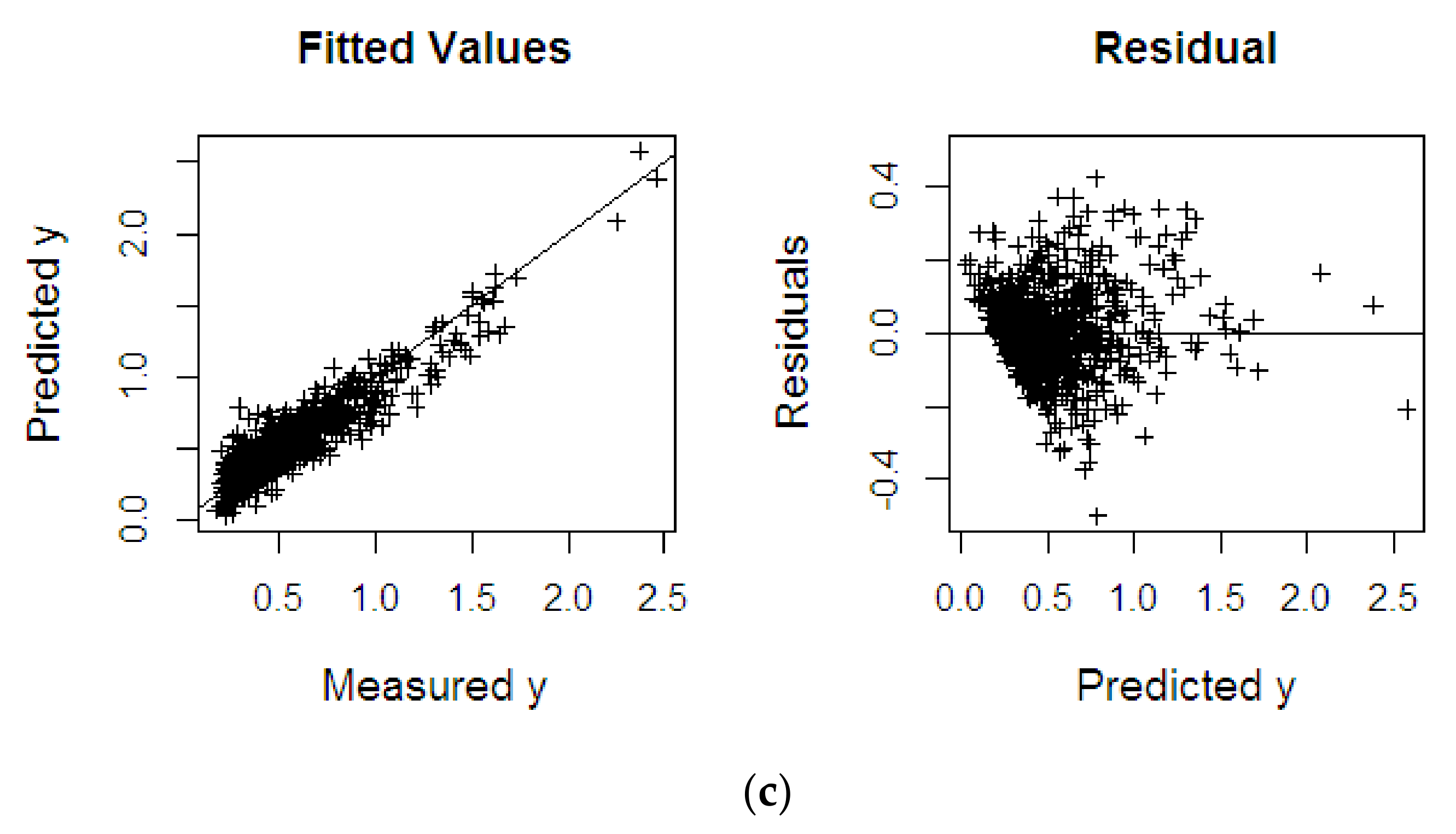
| Dataset | Methods | RMSEP | R2 | SE |
|---|---|---|---|---|
| %ODM | KPLS | 0.283 | 0.927 | 0.282 |
| KPRGM6 | 0.250 | 0.997 | 0.252 | |
| KPRMGM6 | 0.128 | 0.999 | 0.128 | |
| %OWM | KPLS | 0.404 | 0.967 | 0.402 |
| KPRGM6 | 0.364 | 0.998 | 0.365 | |
| KPRMGM6 | 0.301 | 0.999 | 0.301 | |
| %FFA | KPLS | 0.222 | 0.814 | 0.222 |
| KPRGM6 | 0.207 | 0.844 | 0.208 | |
| KPRMGM6 | 0.117 | 0.866 | 0.117 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Silalahi, D.D.; Midi, H.; Arasan, J.; Mustafa, M.S.; Caliman, J.-P. Kernel Partial Least Square Regression with High Resistance to Multiple Outliers and Bad Leverage Points on Near-Infrared Spectral Data Analysis. Symmetry 2021, 13, 547. https://doi.org/10.3390/sym13040547
Silalahi DD, Midi H, Arasan J, Mustafa MS, Caliman J-P. Kernel Partial Least Square Regression with High Resistance to Multiple Outliers and Bad Leverage Points on Near-Infrared Spectral Data Analysis. Symmetry. 2021; 13(4):547. https://doi.org/10.3390/sym13040547
Chicago/Turabian StyleSilalahi, Divo Dharma, Habshah Midi, Jayanthi Arasan, Mohd Shafie Mustafa, and Jean-Pierre Caliman. 2021. "Kernel Partial Least Square Regression with High Resistance to Multiple Outliers and Bad Leverage Points on Near-Infrared Spectral Data Analysis" Symmetry 13, no. 4: 547. https://doi.org/10.3390/sym13040547
APA StyleSilalahi, D. D., Midi, H., Arasan, J., Mustafa, M. S., & Caliman, J.-P. (2021). Kernel Partial Least Square Regression with High Resistance to Multiple Outliers and Bad Leverage Points on Near-Infrared Spectral Data Analysis. Symmetry, 13(4), 547. https://doi.org/10.3390/sym13040547






