A Modified HSIFT Descriptor for Medical Image Classification of Anatomy Objects
Abstract
:1. Introduction
- We discuss the problem of intra-class and inter-class variability in medical image classification.
- We developed a robust classification method for classifying medical images based on modality and anatomy addressing the challenges of intra-class and inter-class variability.
- We provided a detailed comparative analysis of the conventional method and deep learning methods for medical image classification.
- We evaluate the efficacy of the developed method. The experiments demonstrate the effectiveness of the developed method for medical image classification based on modality and anatomy.
2. Related Work
3. Formulation of the Proposed HSIFT Model with Bag of Visual Words Representation
3.1. Data Collection
3.2. Feature Extraction
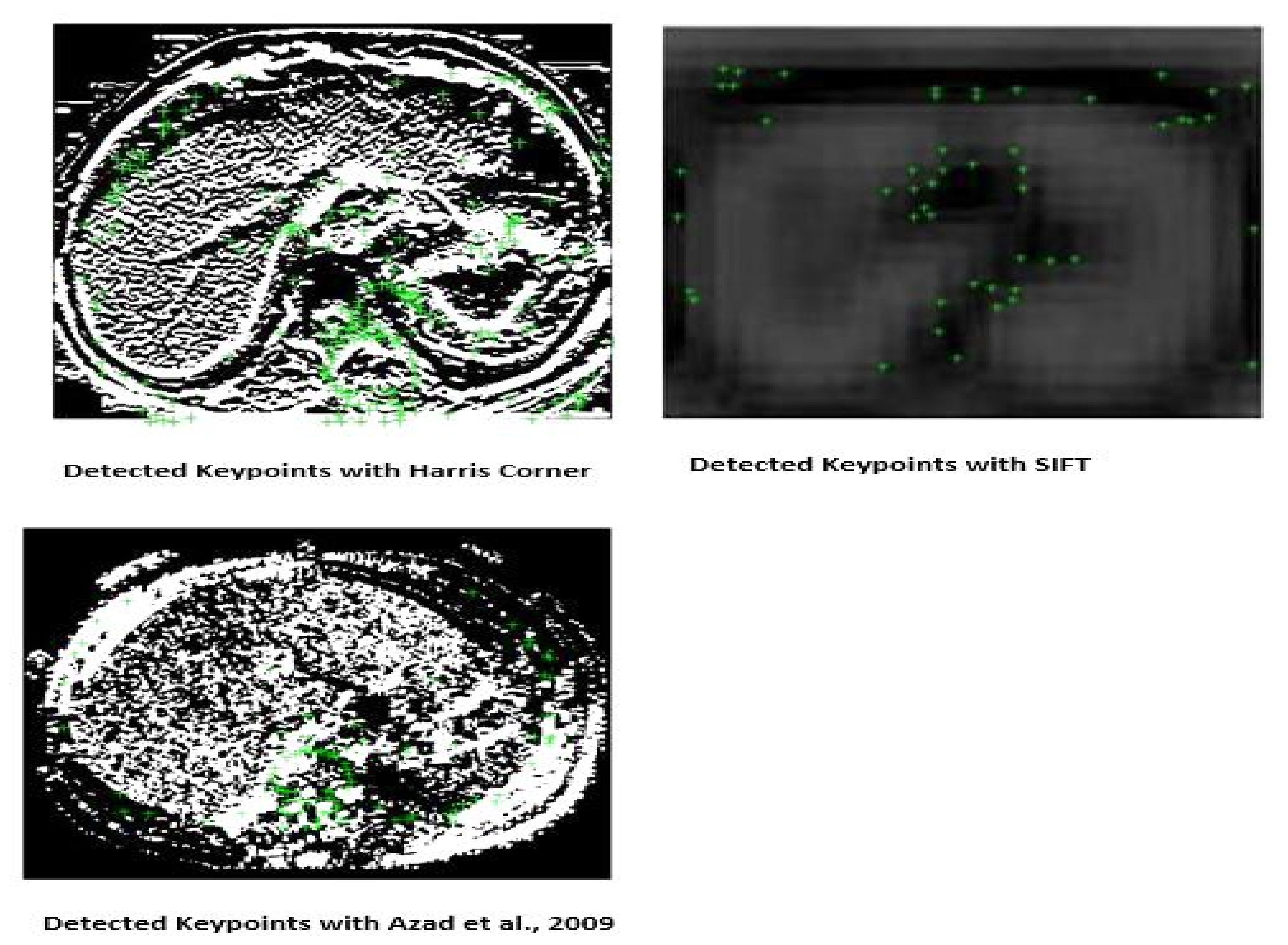
3.2.1. Detection of Harris Corner
- The difference between the original and transferred window is represented by E,
- the x-axis shift of the window is represented by u,
- The y-axis shift of the windows is represented by v,
- The window at (x, y) is represented by w(x, y). This appears to be a mask. Which ensures that only the appropriate window is used,
- The window function is represented by w(x,y),
- The shifted intensity is represented by I(x+u,y+v) and the intensity at (x,y) is represented by I(x,y).
- computing image derivatives: and are obtained by computing the x and y derivatives as:
- Calculate derivative products at each pixel as:
- Finally, the response of the detector at each pixel is computed as:
3.2.2. Key Points Orientation Assignment
3.3. Construction of the Codebook
3.4. Ensemble Classifier with Surrogate Splits
Bagging
| Algorithm 1: Bagging Algorithm |
| Input:training set S, Decision Tree I, integer T (number of bootstrap samples). |
| 1: for i : = 1 to T do |
| 2: S’ = bootstrap sample from S |
| 3: Ci = I’(S’) |
| 4 : end for |
| 5: (the most often predicted label y) |
| Output: classifier C* |
| Algorithm 2: Bagging with surrogate splits |
| 1: procedure BAGALGO () |
| 2: Initialize training set T |
| 3: for n = 1,.....,N do |
| 4: Create a surrogate split X,X ≤ sX, bootstrap replica by randomly sampling with replacement on training set T. |
| 5: Learn m individual classifiers Cm |
| 6: Create an ensemble classifier by aggregating individual classifiers Cm: m = 1,.....,M |
| 7: Classify sample ti to class sj according to the number of votes obtained from classifiers. |
| 8: end for |
| 9: end procedure |
4. Comparative Results for HSIFT and CNN and Experiment
4.1. Experimental Setup for HSIFT
4.2. Experimental Setup for Convolutional Neural Network (CNN)
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Zhou, X.; Depeursinge, A.; Müller, H. Hierarchical classification using a frequency-based weighting and simple visual features. Pattern Recognit. Lett. 2008, 29, 2011–2017. [Google Scholar] [CrossRef]
- Tommasi, T.; Orabona, F.; Caputo, B. Discriminative cue integration for medical image annotation. Pattern Recognit. Lett. 2008, 29, 1996–2002. [Google Scholar] [CrossRef] [Green Version]
- Kalpathy-Cramer, J.; Hersh, W. Effectiveness of global features for automatic medical image classification and retrieval–The experiences of OHSU at ImageCLEFmed. Pattern Recognit. Lett. 2008, 29, 2032–2038. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Avni, U.; Greenspan, H.; Konen, E.; Sharon, M.; Goldberger, J. X-ray categorization and retrieval on the organ and pathology level, using patch-based visual words. Med. Imaging IEEE Trans. 2011, 30, 733–746. [Google Scholar] [CrossRef] [PubMed]
- Depeursinge, A.; Vargas, A.; Platon, A.; Geissbuhler, A.; Poletti, P.; Müller, H. 3D case–based retrieval for interstitial lung diseases. Med-Content-Based Retr. Clin. Decis. Support 2010, 5853, 39–48. [Google Scholar]
- Rahman, M.; Davis, D. Addressing the class imbalance problem in medical datasets. Int. J. Mach. Learn. Comput. 2013, 3, 224–228. [Google Scholar] [CrossRef]
- Song, Y.; Cai, W.; Huang, H.; Zhou, Y.; Wang, Y.; Feng, D. Locality-constrained Subcluster Representation Ensemble for lung image classification. Med. Image Anal. 2015, 22, 102–111. [Google Scholar] [CrossRef] [Green Version]
- Srinivas, M.; Naidu, R.; Sastry, C.; Mohan, C. Content based medical image retrieval using dictionary learning. Neurocomputing 2015, 168, 880–895. [Google Scholar] [CrossRef]
- Magdy, E.; Zayed, N.; Fakhr, M. Automatic classification of normal and cancer lung CT images using multiscale AM-FM features. J. Biomed. Imaging 2015, 2015, 11. [Google Scholar] [CrossRef] [Green Version]
- Chen, D.; Chang, R.; Chen, C.; Ho, M.; Kuo, S.; Chen, S.; Hung, S.; Moon, W. Classification of breast ultrasound images using fractal feature. Clin. Imaging 2005, 29, 235–245. [Google Scholar] [CrossRef]
- Roth, H.; Lee, C.; Shin, H.; Seff, A.; Kim, L.; Yao, J.; Lu, L.; Summers, R. Anatomy-specific classification of medical images using deep convolutional nets. In Proceedings of the 2015 IEEE 12th International Symposium On Biomedical Imaging (ISBI), New York, NY, USA, 16–19 April 2015; pp. 101–104. [Google Scholar]
- Lyndon, D.; Kumar, A.; Kim, J.; Leong, P.; Feng, D. Convolutional Neural Networks for Medical Clustering. Ceur Workshop Proc. 2015. Available online: http://ceur-ws.org/Vol-1391/52-CR.pdf (accessed on 21 September 2021).
- Harris, C.; Stephens, M. A combined corner and edge detector. Alvey Vis. Conf. 1988, 15, 50. [Google Scholar]
- Lowe, D. Object recognition from local scale-invariant features. In Proceedings of the Seventh IEEE International Conference On Computer Vision, Kerkyra, Greece, 20–27 September 1999; pp. 1150–1157. [Google Scholar]
- Feelders, A. Handling missing data in trees: Surrogate splits or statistical imputation? In Proceedings of the European Conference on Principles of Data Mining and Knowledge Discovery, Prague, Czech Republic, 15–18 September 1999; pp. 329–334. [Google Scholar]
- Tartar, A.; Akan, A. Ensemble learning approaches to classification of pulmonary nodules. In Proceedings of the 2016 International Conference On Control, Decision Furthermore, Information Technologies (CoDIT), Saint Julian, Malta, 6–8 April 2016; pp. 472–477. [Google Scholar]
- Xia, J.; Zhang, S.; Cai, G.; Li, L.; Pan, Q.; Yan, J.; Ning, G. Adjusted weight voting algorithm for random forests in handling missing values. Pattern Recognit. 2017, 69, 52–60. [Google Scholar] [CrossRef]
- Zare, M.; Mueen, A.; Seng, W. Automatic medical X-ray image classification using annotation. J. Digit. Imaging 2014, 27, 77–89. [Google Scholar] [CrossRef] [Green Version]
- Kumar, A.; Dyer, S.; Li, C.; Leong, P.; Kim, J. Automatic Annotation of Liver CT Images: The Submission of the BMET Group to ImageCLEFmed 2014. In Proceedings of the CLEF (Work. Notes), Sheffield, UK, 15–18 September 2014; pp. 428–437. [Google Scholar]
- Yang, W.; Lu, Z.; Yu, M.; Huang, M.; Feng, Q.; Chen, W. Content-based retrieval of focal liver lesions using bag-of-visual-words representations of single-and multiphase contrast-enhanced CT images. J. Digit. Imaging 2012, 25, 708–719. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Désir, C.; Petitjean, C.; Heutte, L.; Thiberville, L.; Salaün, M. An SVM-based distal lung image classification using texture descriptors. Comput. Med. Imaging Graph. 2012, 36, 264–270. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lecron, F.; Benjelloun, M.; Mahmoudi, S. Descriptive image feature for object detection in medical images. In Proceedings of the International Conference Image Analysis and Recognition, Aveiro, Portugal, 25–27 June 2012; pp. 331–338. [Google Scholar]
- Sargent, D.; Chen, C.; Tsai, C.; Wang, Y.; Koppel, D. Feature detector and descriptor for medical images. SPIE Med. Imaging 2009, 7259, 72592Z. [Google Scholar]
- Cui, J.; Xie, J.; Liu, T.; Guo, X.; Chen, Z. Corners detection on finger vein images using the improved Harris algorithm. Opt.-Int. J. Light Electron Opt. 2014, 125, 4668–4671. [Google Scholar] [CrossRef]
- Kim, H.; Shin, S.; Wang, W.; Jeon, S. SVM-based Harris corner detection for breast mammogram image normal/abnormal classification. In Proceedings of the 2013 Research in Adaptive and Convergent Systems, Montreal, QC, Canada, 1–4 October 2013; pp. 187–191. [Google Scholar]
- Shim, J.; Park, K.; Ko, B.; Nam, J. X-Ray image classification and retrieval using ensemble combination of visual descriptors. In Proceedings of the Pacific-Rim Symposium on Image and Video Technology, Tokyo, Japan, 13–16 January 2009; pp. 738–747. [Google Scholar]
- Taheri, M.; Hamer, G.; Son, S.; Shin, S. Enhanced Breast Cancer Classification with Automatic Thresholding Using SVM and Harris Corner Detection. In Proceedings of the International Conference on Research in Adaptive and Convergent Systems, Odense, Denmark, 11–14 October 2016; pp. 56–60. [Google Scholar]
- Lee, C.; Wang, H.; Chen, C.; Chuang, C.; Chang, Y.; Chou, N. A modified Harris corner detection for breast IR image. Math. Probl. Eng. 2014, 2014, 902659. [Google Scholar] [CrossRef] [Green Version]
- Gao, L.; Pan, H.; Han, J.; Xie, X.; Zhang, Z.; Zhai, X. Corner detection and matching methods for brain medical image classification. In Proceedings of the 2016 IEEE International Conference On Bioinformatics Biomedicine (BIBM), Shenzhen, China, 15–18 December 2016; pp. 475–478. [Google Scholar]
- Zhou, D.; Gao, Y.; Lu, L.; Wang, H.; Li, Y.; Wang, P. Hybrid corner detection algorithm for brain magnetic resonance image registration. In Proceedings of the 2011 4th International Conference On Biomedical Engineering Furthermore, Informatics (BMEI), Shanghai, China, 15–17 October 2011; pp. 308–313. [Google Scholar]
- Biswas, B.; Dey, K.; Chakrabarti, A. Medical image registration based on grid matching using Hausdorff Distance and Near set. In Proceedings of the 2015 Eighth International Conference On Advances In Pattern Recognition (ICAPR), Kolkata, India, 4–7 January 2015; pp. 1–5. [Google Scholar]
- Zhang, R.; Zhou, W.; Li, Y.; Yu, S.; Xie, Y. Nonrigid registration of lung CT images based on tissue features. Comput. Math. Methods Med. 2013, 2013, 834192. [Google Scholar] [CrossRef]
- Chen, J.; Tian, J.; Lee, N.; Zheng, J.; Smith, R.; Laine, A. A partial intensity invariant feature descriptor for multimodal retinal image registration. IEEE Trans. Biomed. Eng. 2010, 57, 1707–1718. [Google Scholar] [CrossRef] [Green Version]
- Gharabaghi, S.; Daneshvar, S.; Sedaaghi, M. Retinal image registration using geometrical features. J. Digit. Imaging 2013, 26, 248–258. [Google Scholar] [CrossRef] [Green Version]
- Jin, D.; Zhu, S.; Cheng, Y. Salient object detection via harris corner. In Proceedings of the 2017 29th Chinese Control And Decision Conference (CCDC), Chongqing, China, 28–30 May 2017; pp. 1108–1112. [Google Scholar]
- Khan, S.; Yong, S.; Deng, J. Ensemble classification with modified SIFT descriptor for medical image modality. In Proceedings of the 2015 International Conference On Image Furthermore, Vision Computing New Zealand (IVCNZ), Auckland, New Zealand, 23–24 November 2015; pp. 1–6. [Google Scholar]
- Benjelloun, M.; Mahmoudi, S.; Lecron, F. A framework of vertebra segmentation using the active shape model-based approach. J. Biomed. Imaging 2011, 2011, 9. [Google Scholar] [CrossRef]
- Yan, Z.; Zhang, J.; Zhang, S.; Metaxas, D. Automatic Rapid Segmentation of Human Lung from 2D Chest X-Ray Images. In Proceedings of the Miccai Workshop Sparsity Tech. Med Imaging, Nice, France, 12–16 October 2012; pp. 1–8. [Google Scholar]
- Azad, P.; Asfour, T.; Dillmann, R. Combining Harris interest points and the SIFT descriptor for fast scale-invariant object recognition. In Proceedings of the IROS 2009. IEEE/RSJ International Conference On Intelligent Robots Furthermore Systems, St. Louis, MO, USA, 10–15 October 2009; pp. 4275–4280. [Google Scholar]
- Yang, M.; Yuan, Y.; Li, X.; Yan, P. Medical Image Segmentation Using Descriptive Image Features. BMVC. 2011, pp. 1–11. Available online: https://citeseerx.ist.psu.edu/viewdoc/download?doi=10.1.1.297.9559&rep=rep1&type=pdf (accessed on 21 September 2021).
- Moradi, M.; Abolmaesoumi, P.; Mousavi, P. Deformable registration using scale space keypoints. Med. Imaging 2006, 6144, 61442G. [Google Scholar]
- Cireşan, D.; Giusti, A.; Gambardella, L.; Schmidhuber, J. Mitosis detection in breast cancer histology images with deep neural networks. In Proceedings of the International Conference on Medical Image Computing and Computer-Assisted Intervention, Nagoya, Japan, 22–26 September 2013; pp. 411–418. [Google Scholar]
- Prasoon, A.; Petersen, K.; Igel, C.; Lauze, F.; Dam, E.; Nielsen, M. Deep feature learning for knee cartilage segmentation using a triplanar convolutional neural network. In Proceedings of the International Conference on Medical Image Computing and Computer-Assisted Intervention, Nagoya, Japan, 22–26 September 2013; pp. 246–253. [Google Scholar]
- Roth, H.; Lu, L.; Seff, A.; Cherry, K.; Hoffman, J.; Wang, S.; Liu, J.; Turkbey, E.; Summers, R. A new 2.5 D representation for lymph node detection using random sets of deep convolutional neural network observations. In Proceedings of the International Conference on Medical Image Computing and Computer-Assisted Intervention, Boston, MA, USA, 14–18 September 2014; pp. 520–527. [Google Scholar]
- Li, Q.; Cai, W.; Wang, X.; Zhou, Y.; Feng, D.; Chen, M. Medical image classification with convolutional neural network. In Proceedings of the 2014 13th International Conference On Control Automation Robotics & Vision (ICARCV), Marina Bay Sands, Singapore, 10–12 December 2014; pp. 844–848. [Google Scholar]
- Cho, J.; Lee, K.; Shin, E.; Choy, G.; Do, S. Medical Image Deep Learning with Hospital PACS Dataset. arXiv 2015, arXiv:1511.06348. [Google Scholar]
- Csurka, G.; Dance, C.; Fan, L.; Willamowski, J.; Bray, C. Visual categorization with bags of keypoints. Workshop Stat. Learn. Comput. Vis. 2004, 1, 1–2. [Google Scholar]
- Wang, J.; Yang, J.; Yu, K.; Lv, F.; Huang, T.; Gong, Y. Locality-constrained linear coding for image classification. In Proceedings of the 2010 IEEE Conference On Computer Vision Furthermore, Pattern Recognition (CVPR), San Francisco, CA, USA, 13–18 June 2010; pp. 3360–3367. [Google Scholar]
- Claesen, M.; De Smet, F.; Suykens, J.; De Moor, B. EnsembleSVM: A library for ensemble learning using support vector machines. J. Mach. Learn. Res. 2014, 15, 141–145. [Google Scholar]
- Valdiviezo, H.; Van Aelst, S. Tree-based prediction on incomplete data using imputation or surrogate decisions. Inf. Sci. 2015, 311, 163–181. [Google Scholar] [CrossRef]
- Breiman, L. Bagging predictors. Mach. Learn. 1996, 24, 123–140. [Google Scholar] [CrossRef] [Green Version]
- Zare, M.; Mueen, A.; Seng, W. Automatic classification of medical X-ray images using a bag of visual words. Comput. Vis. IET 2013, 7, 105–114. [Google Scholar] [CrossRef]
- Gál, V.; Kerre, E.; Nachtegael, M. Multiple kernel learning based modality classification for medical images. In Proceedings of the 2012 IEEE Computer Society Conference On Computer Vision Furthermore, Pattern Recognition Workshops (CVPRW), Providence, RI, USA, 16–21 June 2012; pp. 76–83. [Google Scholar]
- Krizhevsky, A.; Sutskever, I.; Hinton, G. Imagenet classification with deep convolutional neural networks. Adv. Neural Inf. Process. Syst. 2012, 25, 1097–1105. [Google Scholar] [CrossRef]
- Chen, W.; Song, Y.; Bai, H.; Lin, C.; Chang, E. Parallel spectral clustering in distributed systems. IEEE Trans. Pattern Anal. Mach. Intell. 2011, 33, 568–586. [Google Scholar] [CrossRef] [PubMed]
- LeCun, Y.; Bottou, L.; Bengio, Y.; Haffner, P. Gradient-based learning applied to document recognition. Proc. IEEE 1998, 86, 2278–2324. [Google Scholar] [CrossRef] [Green Version]
- Szegedy, C.; Liu, W.; Jia, Y.; Sermanet, P.; Reed, S.; Anguelov, D.; Erhan, D.; Vanhoucke, V.; Rabinovich, A. Going deeper with convolutions. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Boston, MA, USA, 7–12 June 2015; pp. 1–9. [Google Scholar]













| Feature Representation | SVM Error Rate % | Regular Bagging Error Rate % | Bagging with Surrogate Splits Error Rate % |
|---|---|---|---|
| SIFT [52] | 29.0 | 26.6 | 23.0 |
| SIFT + Harris corner [39] | 30.0 | 28.6 | 26.4 |
| BOVW(SIFT) [53] | 20.5 | 33.2 | 33.4 |
| BOVW(HSIFT) | 9.7 | 3.0 | 2.0 |
| Layers | LeNet | AlexNet | GoogLeNet |
|---|---|---|---|
| conv-1 | 0.22 | 0.66 | 0.56 |
| conv-2 | 0.28 | 0.70 | 0.62 |
| conv-3 | - | 0.74 | 0.51 |
| conv-4 | - | 0.75 | 0.53 |
| ip2/fc8/pool5/ | 0.30 | 0.79 | 0.63 |
| HSIFT | 0.93 | 0.93 | 0.93 |
| Model | Runtime in Seconds | Validation Loss | Validation Accuracy | Test Accuracy |
|---|---|---|---|---|
| LeNet [56] | 1655 | 1.3 | 58.0 | 59.0 |
| AlexNet [54] | 33,466 | 1.39 | 65.0 | 74.0 |
| GoogLeNet [57] | 52,470 | 1.2 | 55.0 | 45.0 |
| SIFT + Harris corner [39] | 648 | 26.4 | 73.6 | 73.8 |
| HSIFT-LBP [36] | 328 | 0.21 | 79.0 | 64.0 |
| Optimized CNN | 16,728 | 0.67 | 76.6 | 81.0 |
| Proposed BOVW(HSIFT) | 235 | 0.02 | 98.0 | 98.1 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Khan, S.A.; Gulzar, Y.; Turaev, S.; Peng, Y.S. A Modified HSIFT Descriptor for Medical Image Classification of Anatomy Objects. Symmetry 2021, 13, 1987. https://doi.org/10.3390/sym13111987
Khan SA, Gulzar Y, Turaev S, Peng YS. A Modified HSIFT Descriptor for Medical Image Classification of Anatomy Objects. Symmetry. 2021; 13(11):1987. https://doi.org/10.3390/sym13111987
Chicago/Turabian StyleKhan, Sumeer Ahmad, Yonis Gulzar, Sherzod Turaev, and Young Suet Peng. 2021. "A Modified HSIFT Descriptor for Medical Image Classification of Anatomy Objects" Symmetry 13, no. 11: 1987. https://doi.org/10.3390/sym13111987
APA StyleKhan, S. A., Gulzar, Y., Turaev, S., & Peng, Y. S. (2021). A Modified HSIFT Descriptor for Medical Image Classification of Anatomy Objects. Symmetry, 13(11), 1987. https://doi.org/10.3390/sym13111987








