Automatic Segmentation and Measurement of Infantile Hemangioma
Abstract
1. Introduction
The Novelty of This Paper
2. Materials and Methods
- Automatic computation of the image scale and the detection of the ruler direction line.
- Rotation of the image such that the ruler becomes vertical.
- Extraction of the centimeter area.
- Extraction of the digits from the centimeter area.
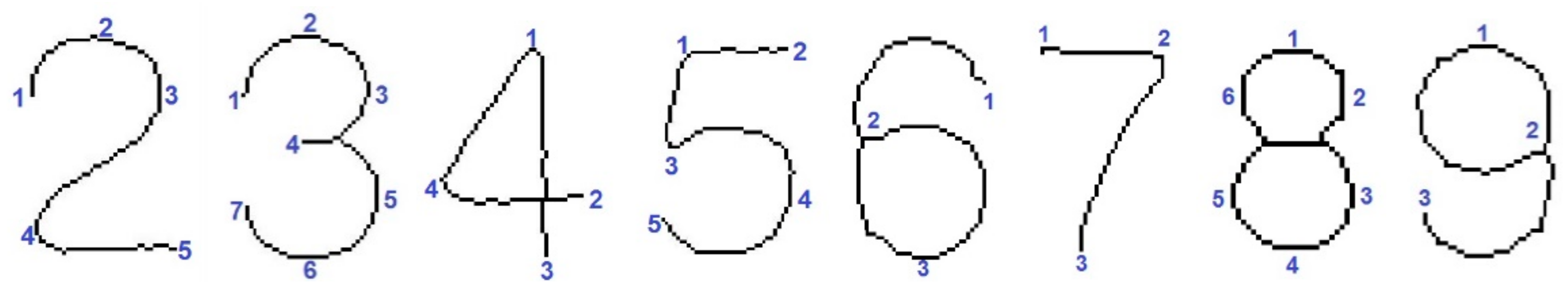
- Performance of optical character recognition (OCR) of each of the objects that could be a digit. We search for the digits: 2–9.
- Geometric transformation of the entire image such that the detected digit is geometrically aligned to its corresponding frontal template.
- Segmentation of the hemangioma.
- Computation of the lesion surface.
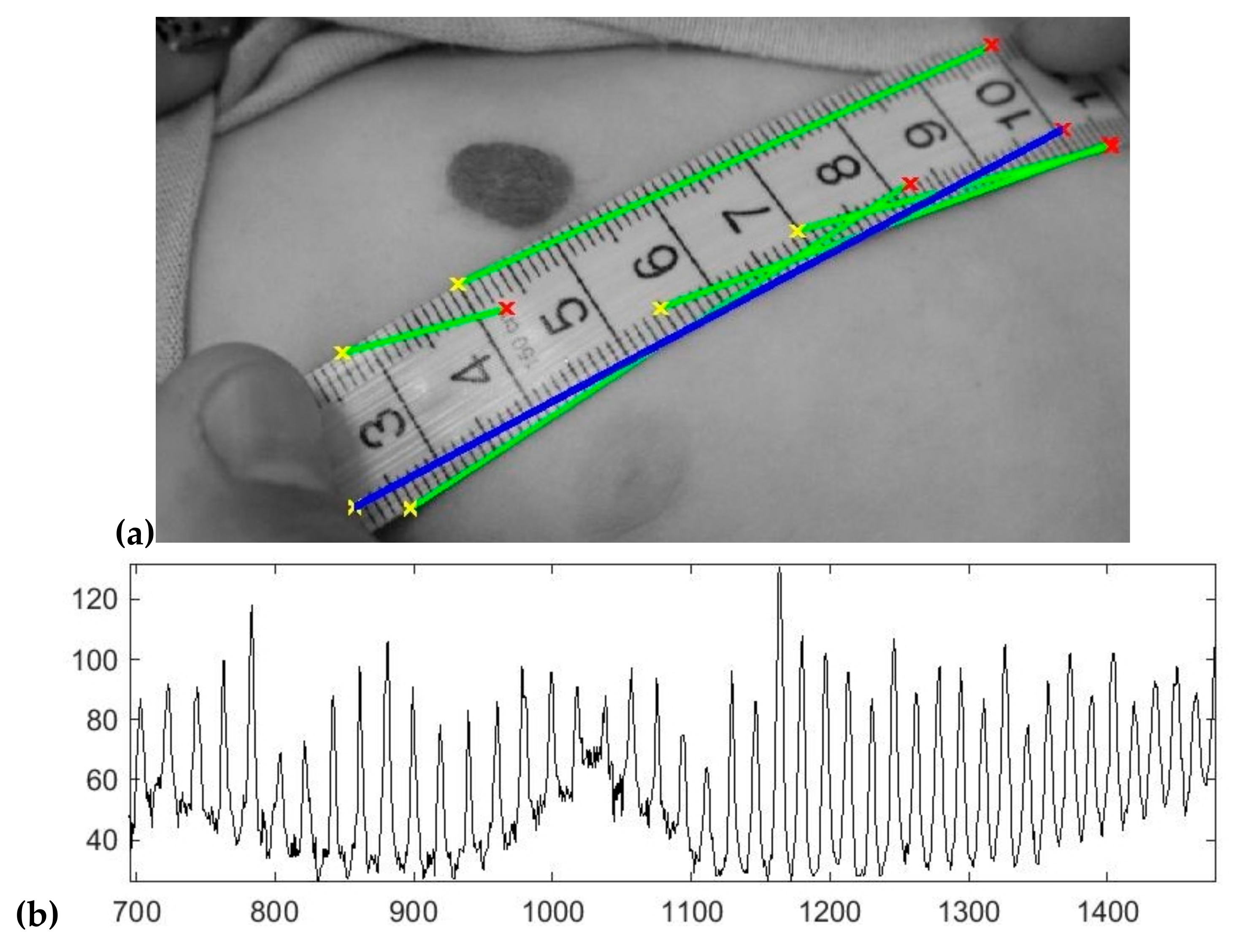
2.1. Detection of the Ruler and Computation of the Image Scale
- -
- Find the edges, using the Canny method;
- -
- Apply the Hough transform for lines on the edges of the image (obtained in the previous step), and keep only the lines that are at least 250 pixels long (the minimum length of 250 pixels was selected because it corresponds to around 1 cm length on the ruler at the resolution at which the images were acquired; if a smaller value is chosen, the algorithm still works, but the processing time increases due to the detection of more lines within the image);
- -
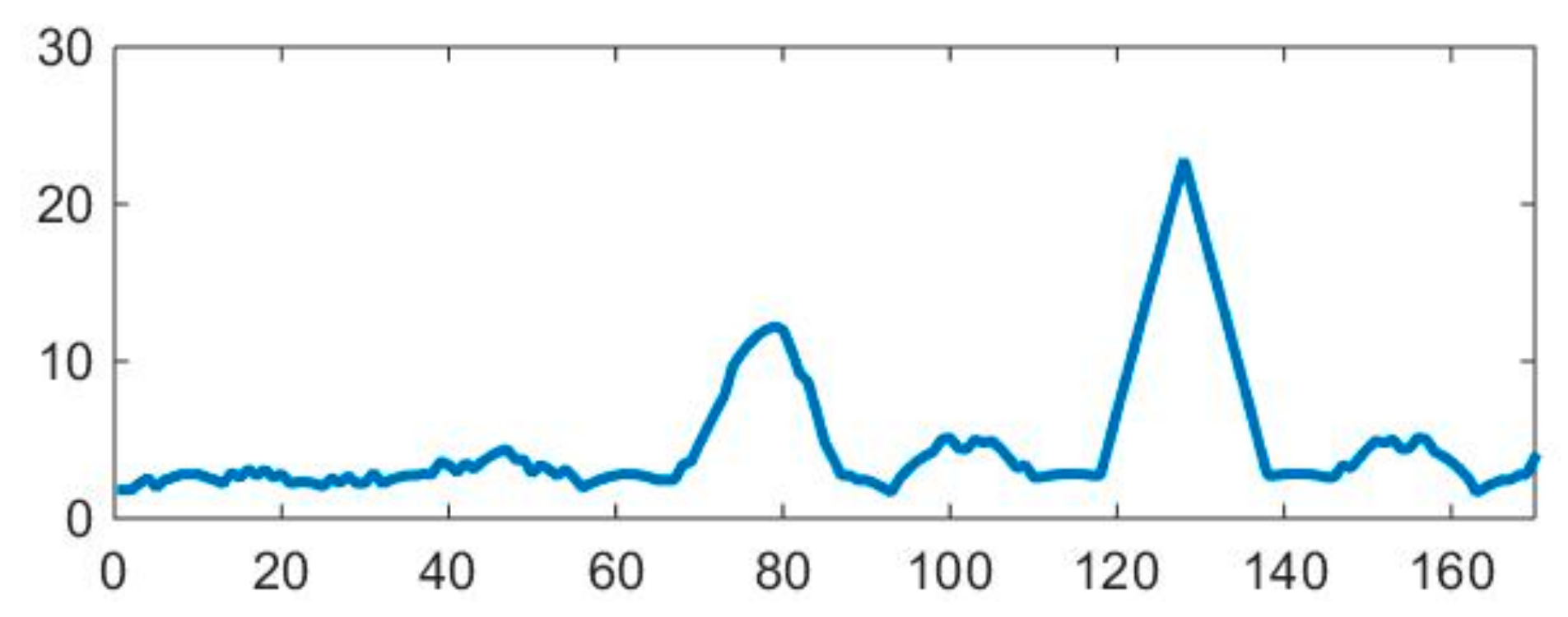
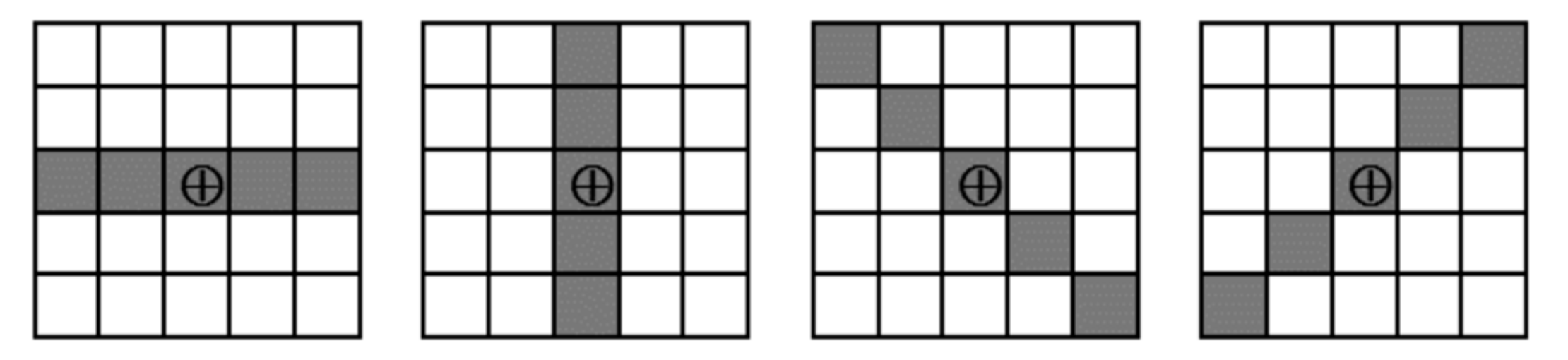
- For each extracted line, record the 1D intensity profile along the line and compute its total variation (1);
- -
- The line which has the maximum total variation is selected;
- -
- Detect the peaks from the intensity profile of the line and then compute the median distance between two consecutive peaks.
- Apply a Canny edge detection on the original image.
- Build an edge density image, D, by convolving the contour map with a blurring kernel.
- Construct a binary image B, which contains the higher edge density, by thresholding D with an adaptive threshold (set at 45% of the higher contour density).
- Keep the pixels from B that have a red component (from RGB) 55% higher than the green component and 55% higher than the blue one.
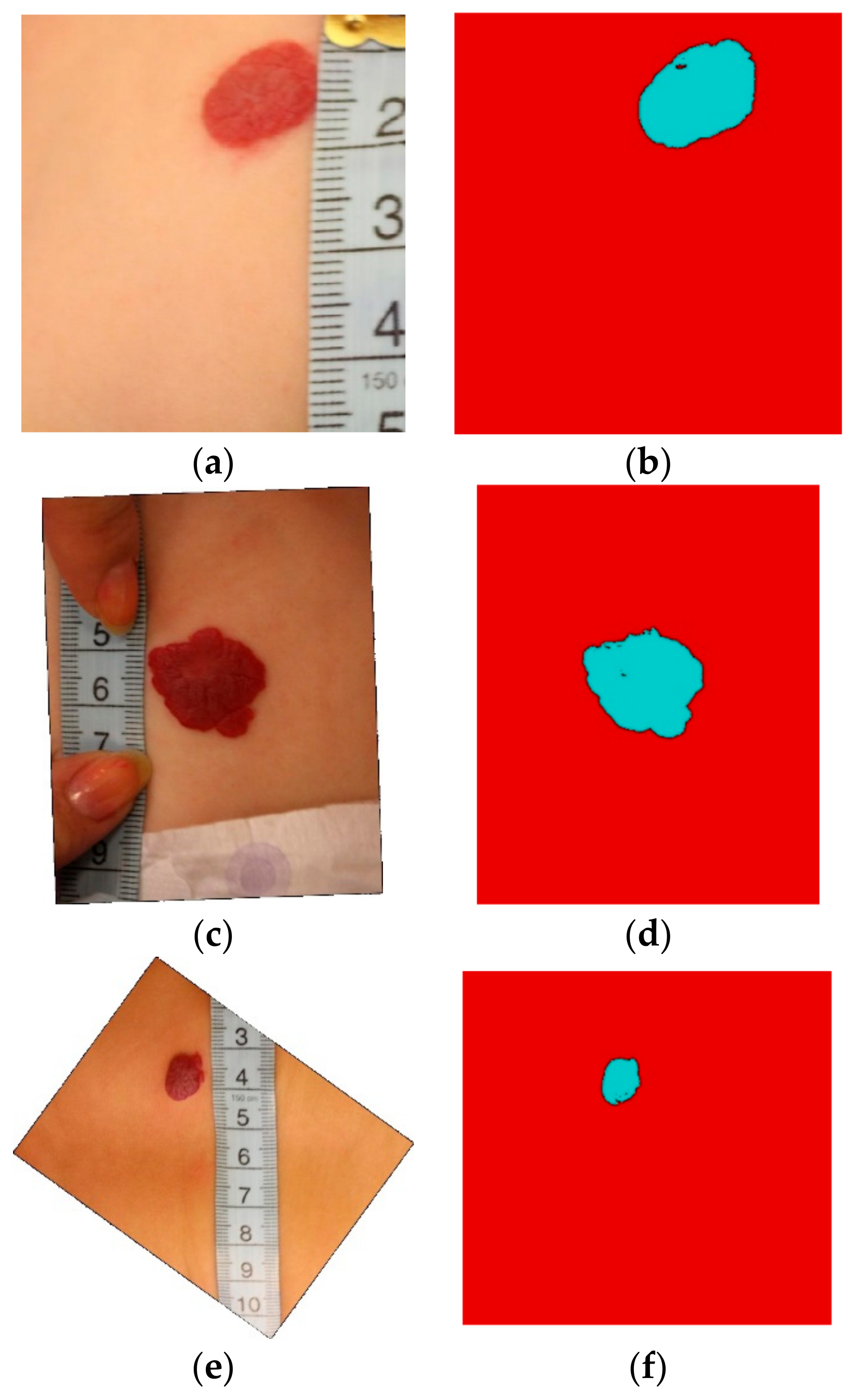
2.2. Hemangioma Lesion Segmentation
- construct a seven-plane image composed of:
- -
- the L* component of the image;
- -
- the a* plane;
- -
- the b* plane;
- -
- the TV image obtained by applying the first filter on the L* component of the image;
- -
- the TV image obtained by applying the second filter on the L* component of the image;
- -
- the H plane from HSV (shifted with 50° in order that the red color is not on the transition from 0° to 360°); and
- -
- the S plane from HSV;
- create a binary mask A, which keeps only the reddish pixels (that have the red component from RGB bigger than the green and blue components; an example of a pixel belonging to a hemangioma: R = 152, G = 61, B = 35), that can be hemangioma pixels, by setting a threshold of 85% from the maximum value of the a* plane from (L*a*b*); note that in L*a*b* there is a symmetrical distribution of colors in the a*b* planes;
- multiply each TV image with the mask A to keep only the hemangioma pixels and possibly also some reddish healthy skin pixels;
- apply the Karhunen-Loeve transform on the seven-plane image, and keep only the first five most important planes;
- run the K-means clustering on the five-plane image obtained in the previous step, with a number of 6 classes (the best number of classes was experimentally determined);
- from the segmented image, choose as the hemangioma class the one that has the maximum a* average value, or if there are two regions with the same a* average, choose the one that has the maximum TV (computed with the first filter) value;
- eliminate the obtained hemangioma regions that are smaller than a threshold, or the ones with weak borders.
| Algorithm 1 The proposed method for hemangioma segmentation |
| Read the color input image I G ← graylevel(I) BW ← Canny_edge_detection(G) Apply the Hough transform for lines on BW Compute the total variation for each line with Equation (1) Keep the line with the maximum TV value d ← median distance between two consecutive peaks on the line Rotate(I), such as the extracted line becomes vertical M ← extracted digits from the ruler using K-means clustering with 2 classes M ← eliminate objects from M that are too small or too big to represent digits D ← edge density image, obtained by convolving BW with a blurring kernel B ← binary image, where value 1 = if pixel is 45% of the highest contour density from D B ← pixels from B that have R (from RGB) 55% higher than G and 55% higher than B [Bx, By] ← mean location of the pixels with value 1 from B (lesion location estimation) Apply OCR on each region from M; choose the digit from 2 to 9 that is the closest to [Bx, By] Find the key points on the digit I ← geometrical transformation of I, using Equation (2) A1 ← L* from L*a*b* version of I A2 ← a* from L*a*b* version of I A3 ← b* from L*a*b* version of I A4 ← image obtained by applying the first TV filter on L* A5 ← image obtained by applying the second TV filter on L* A6 ← H from HSV (shifted with 50°) A7 ← S from HSV N ← 85% * max(a*), a binary mask where 1 = reddish pixels, 0 = other pixels A4 ← A4 pixelwise multiplied by N A5 ← A5 pixelwise multiplied by N A ← [A1 A2 A3 A4 A5 A6 A7] (7 plane image) X ← Karhunen-Loeve(A), keep only the 5 most important planes Y ← K-means(X), 6 classes S ← binary image, where 1= hemangioma class from Y S ← S without regions smaller than a threshold S ← only regions from S with strong borders R ← compute the lesion surface (number of pixels of value 1 from S * 1/d2) |
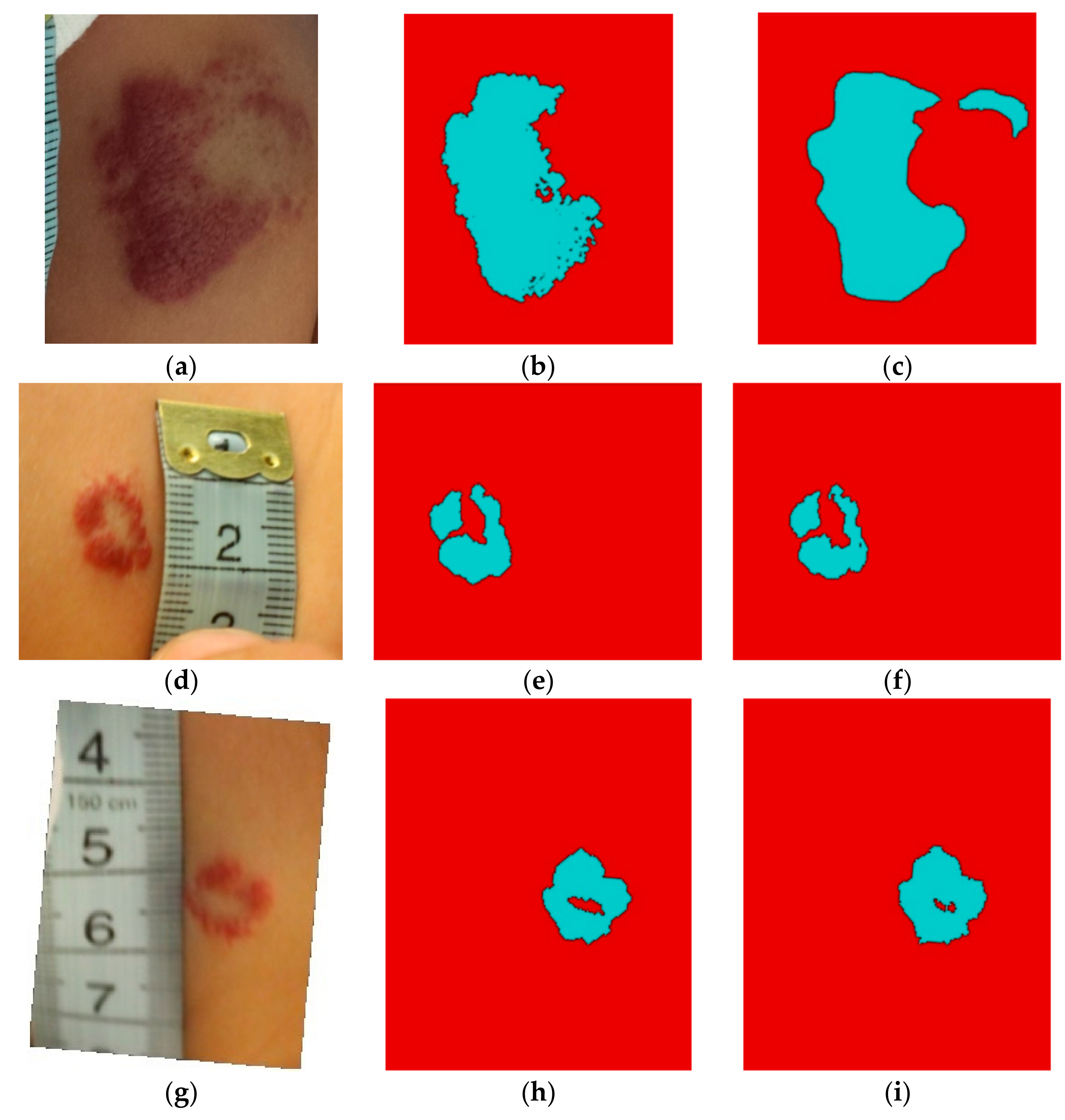
3. Results
Comparison of the Obtained Results with Other Methods
4. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Shwayder, T.; Schneider, S.L.; Icecreamwala, D.; Jahnke, M.N. Infantile Hemangiomas. In Longitudinal Observation of Pediatric Dermatology Patients; Springer: Cham, Switzerland, 2019. [Google Scholar] [CrossRef]
- Group, T.H.I.; Haggstrom, A.N.; Drolet, B.A.; Baselga, E.; Chamlin, S.L.; Garzon, M.C.; Horii, K.A.; Lucky, A.W.; Mancini, A.J.; Metry, D.W.; et al. Prospective Study of Infantile Hemangiomas: Demographic, Prenatal, and Perinatal Characteristics. J. Pediatrics 2007, 150, 291–294. [Google Scholar] [CrossRef]
- Chang, L.C.; Haggstrom, A.N.; Drolet, B.A.; Baselga, E.; Chamlin, S.L.; Garzon, M.C.; Horii, K.A.; Lucky, A.W.; Mancini, A.J.; Metry, D.W.; et al. Growth Characteristics of Infantile Hemangiomas: Implications for Management. Pediatrics 2008, 122, 360–367. [Google Scholar] [CrossRef] [PubMed]
- Rogowska, J. Chap. 5—Overview and Fundamentals of Medical Image Segmentation. In Handbook of Medical Image Processing and Analysis, 2nd ed.; Bankman, I.N., Ed.; Academic Press: Cambridge, MA, USA, 2009; pp. 73–90. [Google Scholar] [CrossRef]
- Gillmann, C.; Post, T.; Wischgoll, T.; Hagen, H.; Maciejewski, R. Hierarchical image semantics using probabilistic path propagations for biomedical research. IEEE Comput. Graph. Appl. 2019, 39, 86–101. [Google Scholar] [CrossRef] [PubMed]
- Zambanini, S.; Sablatnig, R.; Maier, H.; Langs, G. Automatic image-based assessment of lesion development during hemangioma follow-up examinations. Artif. Intell. Med. 2010, 50, 83–94. [Google Scholar] [CrossRef] [PubMed]
- Sultana, A.; Zamfir, M.; Ciuc, M.; Oprisescu, S.; Popescu, M. Automatic segmentation of infantile hemangiomas. In Proceedings of the Signals, Circuits and Systems (ISSCS), International Symposium on, Iasi, Romania, 9–10 July 2015; IEEE: New York, NY, USA, 2015; pp. 1–4. [Google Scholar] [CrossRef]
- Sultana, A.; Oprisescu, S.; Ciuc, M. Automatic evaluation of hemangiomas for follow-up monitoring. In Proceedings of the E-Health and Bioengineering Conference (EHB), Iasi, Romania, 19–21 November 2015; IEEE: New York, NY, USA, 2015; pp. 1–4. [Google Scholar] [CrossRef]
- Oprisescu, S.; Ciuc, M.; Sultana, A.; Vasile, I. Automatic segmentation of infantile hemangiomas within an optimally chosen color space. In Proceedings of the E-Health and Bioengineering Conference, Iasi, Romania, 19–21 November 2015; IEEE: New York, NY, USA, 2015; pp. 1–4. [Google Scholar] [CrossRef]
- Neghina, C.; Zamfir, M.; Sultana, A.; Ovreiu, E.; Ciuc, M. Automatic detection of hemangiomas using unsupervised segmentation of regions of interest. In Proceedings of the Communications (COMM), 2016 International Conference on, Bucharest, Romania, 9–10 June 2016; IEEE: New York, NY, USA, 2016; pp. 69–72. [Google Scholar] [CrossRef]
- Neghina, C.; Zamfir, M.; Ciuc, M.; Sultana, A. Automatic detection of hemangioma through a cascade of self-organizing map clustering and morphological operators. Procedia Comput. Sci. 2016, 90, 145–150. [Google Scholar] [CrossRef][Green Version]
- Alves, P.G.; Cardoso, J.S.; Bom-Sucesso, M.D. The Challenges of Applying Deep Learning for Hemangioma Lesion Segmentation. In Proceedings of the 2018 7th European Workshop on Visual Information Processing (EUVIP), Tampere, Finland, 26–28 November 2018; pp. 1–6. [Google Scholar] [CrossRef]
- Sultana, A.; Balazs, H.; Ovreiu, S.; Oprisescu, S.; Neghina, C. Infantile Hemangioma Detection using Deep Learning. In Proceedings of the 2020 13th International Conference on Communications (COMM), Bucharest, Romania, 18–20 June 2020; pp. 313–316. [Google Scholar] [CrossRef]
- Oprisescu, S.; Ciuc, M.; Sultana, A. Automatic measurement of infantile hemangiomas. In Proceedings of the 2017 E-Health and Bioengineering Conference (EHB), Sinaia, Romania, 22–24 June 2017; pp. 503–506. [Google Scholar] [CrossRef]
- Smith, R. An Overview of the Tesseract OCR Engine. In Proceedings of the Ninth International Conference on Document Analysis and Recognition (ICDAR 2007), Parana, Brazil, 23–26 September 2007; Volume 2, pp. 629–633. [Google Scholar]
- Sosa, N.L.D.; Noguera, J.L.V.; Silva, J.J.C.; Torres, M.G.; Ayala, H.L. RGB Inter-Channel Measures for Morphological Color Texture Characterization. Symmetry 2019, 11, 1190. [Google Scholar] [CrossRef]
- Ndajah, P.; Kikuchi, H. Total variation image edge detection. In Proceedings of the 10th WSEAS International Conference on Electronics, Hardware, Wireless and Optical Communications; Springer: New York, NY, USA, 2011; pp. 246–251. [Google Scholar]
- Gillmann, C.; Wischgoll, T.; Hamann, B.; Ahrens, J. Modeling and Visualization of Uncertainty-Aware Geometry Using Multi-variate Normal Distributions. In Proceedings of the 2018 IEEE Pacific Visualization Symposium (PacificVis), Kobe, Japan, 10–13 April 2018; pp. 106–110. [Google Scholar] [CrossRef]


Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Oprisescu, S.; Ciuc, M.; Sultana, A. Automatic Segmentation and Measurement of Infantile Hemangioma. Symmetry 2021, 13, 138. https://doi.org/10.3390/sym13010138
Oprisescu S, Ciuc M, Sultana A. Automatic Segmentation and Measurement of Infantile Hemangioma. Symmetry. 2021; 13(1):138. https://doi.org/10.3390/sym13010138
Chicago/Turabian StyleOprisescu, Serban, Mihai Ciuc, and Alina Sultana. 2021. "Automatic Segmentation and Measurement of Infantile Hemangioma" Symmetry 13, no. 1: 138. https://doi.org/10.3390/sym13010138
APA StyleOprisescu, S., Ciuc, M., & Sultana, A. (2021). Automatic Segmentation and Measurement of Infantile Hemangioma. Symmetry, 13(1), 138. https://doi.org/10.3390/sym13010138






