Abstract
With the complexity of Near Infrared (NIR) spectral data, the selection of the optimal number of Partial Least Squares (PLS) components in the fitted Partial Least Squares Regression (PLSR) model is very important. Selecting a small number of PLS components leads to under fitting, whereas selecting a large number of PLS components results in over fitting. Several methods exist in the selection procedure, and each yields a different result. However, so far no one has been able to determine the more superior method. In addition, the current methods are susceptible to the presence of outliers and High Leverage Points (HLP) in a dataset. In this study, a new automated fitting process method on PLSR model is introduced. The method is called the Robust Reliable Weighted Average—PLS (RRWA-PLS), and it is less sensitive to the optimum number of PLS components. The RRWA-PLS uses the weighted average strategy from multiple PLSR models generated by the different complexities of the PLS components. The method assigns robust procedures in the weighing schemes as an improvement to the existing Weighted Average—PLS (WA-PLS) method. The weighing schemes in the proposed method are resistant to outliers and HLP and thus, preserve the contribution of the most relevant variables in the fitted model. The evaluation was done by utilizing artificial data with the Monte Carlo simulation and NIR spectral data of oil palm (Elaeis guineensis Jacq.) fruit mesocarp. Based on the results, the method claims to have shown its superiority in the improvement of the weight and variable selection procedures in the WA-PLS. It is also resistant to the influence of outliers and HLP in the dataset. The RRWA-PLS method provides a promising robust solution for the automated fitting process in the PLSR model as unlike the classical PLS, it does not require the selection of an optimal number of PLS components.
1. Introduction
The Near Infrared Spectroscopy (NIRS) has recently been attracting a lot of attention as a secondary analytical tool for quality control of agricultural products. In some applications (see [1,2,3,4,5]), it has been proven that the NIRS offers a non-destructive, reliable, accurate, and rapid tool, particularly for quantitative and qualitative assessments. Theoretically, NIRS is a type of vibrational spectroscopic that produces rich information in a spectral dataset as a result of the interaction between optical light and the physical matter of the sample. This spectral is commonly presented in terms of spectral absorbance using wide wavelengths that range from 350 nm to 2500 nm, primarily attributed to the overtone or combination bands of C-H (fats, oil, hydrocarbons), O-H (water), and N-H (protein) [6]. The NIR spectral data are classified as high dimension due to the large sample size and wide wavelength collected as a dataset. In spectral processing, chemometric methods have been utilized as the standard processing method (see [7,8,9]). The methods combine the mathematical and multivariate statistical methods in order to pre-process, examine, and understand as much relevant information as possible from the spectral data. Comparing some of the existing chemometric methods, the Partial Least Squares Regression (PLSR) seems to be the most preferred one [10,11,12].
PLSR decomposes both the spectral and reference information (from wet chemistry analysis), simultaneously. It has the ability to screen unwanted samples in a dataset as a result of experimental error and instrumentation problem [13], distribution-free assumption [14,15], and handling the multicollinearity in dataset [16]. However, despite having these benefits, several studies have reported its weakness due to its robustness. The fitted model performs poorly when outliers and leverage points are present in a dataset [17,18], as it fails to fit the nonlinear behavior in the input space [19,20]. In addition, the contamination of irrelevant variables involves during the fitting process [21,22,23] is a popular topic in most discussions. However, so far, less attention has been paid to the basic principles of PLSR in selecting the optimal number of Partial Least Squares (PLS) components which is crucial [24]. Applying fewer number of components produced under fitting, while applying a large number of components results in over fitting. Some methods available in the selection procedure are the cross-validation with one-sigma heuristic [25], permutation approach [26], bootstrap [27], smoothed PLS–PoLiSh [28], weight randomization test [29], and Monte Carlo resampling [30]. These different methods suggest different optimal numbers of the PLS components and to date, there has been no claim made as to which method is superior to the other. These methods suffer from the presence outliers and High Leverage Points (HLP) in the dataset. Consequently, recalculation of the number of PLS components used in the model is required each time the dataset is updated. This would result in different accuracy achievements and sometimes, misleading interpretations. It has been observed that there are only a few studies that have highlighted the robust process. As such, a robust PLSR with less sensitivity to the selection of optimal number of PLS components is needed. This study provides another perspective of applying a robust procedure in the PLSR model with regard to the selection number of the factors used in the fitted model.
The automated fitting process on PLSR model using weighted average strategy has been introduced in several papers (see [31,32,33]). The method is known as the Locally Weighted Average PLS [31,32] or simply called the Local-WA-PLS. The Local-WA-PLS is an extension of the Locally Weighted Regression [33] which is used to fit a local linear regression based on the classification of similarity between the calibration and testing (or unknown) sample. This similarity is classified using the famous Euclidean distance and Mahalanobis distance method. Although the Local-WA-PLS has been widely used, it has been reported that the method works adequately only with a large spectral dataset (see [34,35]). As an improvement, the modified method by Zhang et al. [35], the Weighted Average PLS (WA-PLS), is suggested as it uses a different weighting scheme that is computationally simpler and comparable to the Local-WA-PLS. However, both methods are not able to handle the problems of outliers and HLP that may exist in the dataset from affecting their performances. In addition, the Local-WA-PLS and WA-PLS do not take into consideration the influence of some irrelevant variables in the model that might decrease their estimation accuracy. This has motivated the current study to propose another improvement to robustify the existing WA-PLS procedure. Our strategies were to employ the weighting schemes that are resistant to outliers and HLP and preserve the contribution of the most relevant variables in the fitted model. The utilization of the robust PLSR [36] is incorporated in the establishment of the proposed procedures.
The main objectives of this study are: (1) to establish an improved procedure for the automated fitting process in the PLSR model known as the Robust Reliable Weighted Average PLS (RRWA-PLS). This proposed method is expected to be less sensitive to the selection of the optimal number of PLS components; (2) to evaluate the performance of the proposed RRWA-PLS method with the classical PLSR using optimal number of PLS components, WA-PLS, and a slight modification method in WA-PLS using a robust weight procedure called MWA-PLS; (3) to apply the proposed method on the artificial data and NIR spectra of oil palm (Elaeis guineensis Jacq.) fruit mesocarp (fresh and dried ground). This study provides a significant contribution to the development of process control, particularly for research methodology in the vibrational spectroscopy area.
2. Materials and Methods
2.1. Partial Least Squares Regression
The PLSR model [14] is an iterative procedure of the multivariate statistical method. The method is used to derive original predictor variables that may have a multicollinearity problem into smaller uncorrelated new variables called components. The PLSR constructs a regression model using the new components against its response variable through covariance structure of these two spaces. In chemometric analysis, the PLSR has been widely used for dimensional reduction of high dimensionality problem in the NIR spectral dataset (see [37,38]). In this study, we limited the study only in the case of , where refers to the number of observations, and represents the number of predictor variables.
Let us define a multiple regression model which consists of two different sets of multiple predictor and a single response ,
where , are vector; is matrix; and is vector. Since the dataset contains high dimension of predictors, there will be an infinite number of solution for estimator . Considering is singular, it does not meet the usual trivial theorem on rank in the classical regression. To overcome this, new latent variables need to be produced by summarizing the covariance between predictor and response variable associating to the center values of these two sets [39].
Initializing a starting score vector of from the single ; there exists an outer relation for predictor in Equation (1) as
where is a (for ) matrix of the vector ; and is the column vector of scores in . The is a matrix consisting column vector of loading . The is a vector of weight for and is a matrix of residual in outer relation for . In addition, there is a linear inner relation between the and block scores, calculated as or written as
with is a vector of regression coefficient as the solution using Ordinary Least Square (OLS) on the decomposition of vector , and is vector of residual in the inner relation. Following the Nonlinear Iterative Partial Least Squares (NIPALS) algorithm (see [14]), the mixed relation in the PLSR model can be defined as
where is vector coefficient; represents vector coefficient which is ; and denotes vector of residual in mixed relation that has to be minimized. The estimator for parameter is given as
with denotes dimensional vector of regression coefficient in PLSR.
2.2. Partial Robust M-Regression
An alternative robust version of PLSR introduced by Serneel et al. [36] is the partial robust M-regression or simply known as PRM-Regression. The method assigns a generalized weight function using a modified robust M-estimate [40]. This weight is obtained from the iterative reweighting scheme (see [41]) to identify the outliers and HLP, both in each observation and score vector . Let us consider the regression in Equation (1), for , the least square estimator of is defined as
The least square is optimal if and or where ; otherwise, it fails to satisfy the normal assumption. When it does not satisfy this assumption, the least square losses its optimality; hence, a robust estimator such as M-estimates results in a better solution.
In Serneels et al. [36], the robust M-estimates reestablish the squares term into giving
where , as is defined to be loss function which is symmetric and nondecreasing. Recall the as residual column vector related to Equation (7), then . Using partial derivative and following the iterative reweighting scheme, there exists a weight in each observation as , taking , the Equation (7) can be rewritten as
It is considered that the weight in Equation (8) is only sensitive to the vertical outlier as improvement of another weight is added to identify the leverage points. The criteria would be identified as the leverage points. The modified final estimator in Equation (8) is given as
where is the generalized weight. Replacing the residual in Equation (9) with vector of residual in Equation (4), then giving the solution of the partial robust M-regression as
with the weights and are given as
where uses the robust , is the weight function of iterative reweighting.
is Euclidean norm; is a robust estimator of the center of the dimensional score vectors; and is the vector of component score matrix that needs to be estimated. The fair weight function in is preferred instead of other weights.
2.3. Weighted Average PLS
The WA-PLS method was introduced by Zhang [35] to encounter the sensitivity of PLSR toward the specific number of PLS components used. The method applies the averaging strategy to accommodate the whole possible complexity of the model. This complexity means that some models were initiated based on the increase from the th to the th number of PLS components used in the fitting model. Instead of applying the same weight in each PLSR model, the WA-PLS proposes different weights using variance weighting to each coefficient in the PLSR model with the complexity of .
where the Root Mean Square Error Cross Validation (RMSECV) in each different number of th PLS components is calculated as
where is the predicted value of the actual value of using the fitted model which is built without sample and is under the complexity of . The WA-PLS formula using weight and average from th to the th number of PLS components can then be written as
3. Robust Reliable Weighted Average
Following Zhang’s et al. [35] weighted average calculation on each coefficient of different numbers of PLS components, a robust version of the modified weighted average is developed. The method is called the Robust Reliable Weighted Average (RRWA) which accommodates two weights in the calculation of the PLSR model. It is expected that by assigning the weighted average method in the PLSR model, the model becomes less sensitive to the number of PLS components used.
In the first weight , the calculation uses the Standard Error Prediction (SEP) which is done iteratively based on the re-sampling procedure of -fold cross validation by splitting a dataset into -subsets [42]. This procedure is the most used approach to retrieve a good estimate of error rate in the model selection. Nonetheless, it is anticipated that 20% of the highest absolute values of residuals may still be included in the calculation of . In order to remove those residuals, the trimmed version (20%) SEP from the cross validation () is applied. The assigned weight to each coefficient of different numbers of PLS components is calculated as
where the values are calculated using the collection of the from -subsets starting from th to the th number of PLS components. The calculation for is given as
where is the residual from predicted value of and actual value of with the complexity of , and is the arithmetic mean of the residuals. It corresponds to the where the bias is identically equal to 0, then the is equals to . While the bias is identically (almost) zero, the squared root of which is
is (almost) equal to the . This alternative weight could be called as a modified weight in WA-PLS, and is simply denoted as the MWA-PLS method which is also included as an alternative proposed method in this study.
In the classical WA-PLS, the number of possible irrelevant variables is still involved in the model. Eliminating these irrelevant variables would result in under or over fitting. Here, a downgrading procedure by assigning the second weight to each variable in terms of reliability values [21] is proposed. The procedure is based on the PLSR coefficient that is applicable to increase the contribution of most relevant variables in the model, and downgrade the irrelevant variables. The reliability of each variable is obtained by
where the calculation is based on the robust measure of central tendency and the robust measure of variability on each th WA-PLS coefficient from th to the th numbers of PLS components. The robust weight in Equation (16) is preferred instead of the weight in Equation (13). In relation to the PLSR model, this reliability value is converted into a diagonal matrix with size . This diagonal matrix where is then used to transform the original input variables into the scaled input variables for the RRWA-PLS model.
To prevent the influence of outliers and HLP that may exist in the NIR spectral dataset, the calculation of and reliability values are based on the PRM regression coefficient through a cross-validation procedure. The proposed modification of the WA-PLS known as the RRWA-PLS can be rewritten as
where is the RRWA-PLS coefficient using the scaled input variables .
4. Monte Carlo Simulation Study
To examine the performance of the proposed RRWA-PLS and to compare its performance with the classical WA-PLS and MWA-PLS, a study using the Monte Carlo simulation was carried out. Following a simulation study by Kim [43], an artificial dataset which contained added noise that follows the Normal distribution was randomly generated using a Uniform distribution and included. This dataset was then applied in the linear combination equation with different scenarios. Three sample sizes ( 60, 200, 400), three levels of numbers of predictor variables ( 41, 101, 201), three levels of relevant variables ( 0.1, 0.3, 0.5), and three different levels of outliers and high leverage points ( 0.00, 0.05, 0.20) were considered. The of the predictor variables were randomly selected as relevant variables, and the remaining were considered as less relevant. The formulation of this simulation can be defined as follows:
where is the total number of predictors used; is the number of observable variables; and the is the number of artificial noise variable. These artificial variables are applied to evaluate the stability of the methods. The follows the Uniform distribution (1,10) with size . The artificial noise variables are added to the predictor and follow the Uniform distribution (5,20) with size . This is classified as an irrelevant variable. The follows the standard normal distribution with size , and represents a vector coefficient for selected relevant variables which follows the Uniform distribution (0,7) with size . The , and are independent of each other. The is the set of selected relevant variables in , and is the added error in the linear combination of . and are illustrated as observable variables. The high leverage points in the dimensions are created by generating following the Uniform distribution (1,10) with size . Corresponding to the vertical outlier, if the observation is considered as an outlier, follows the Uniform distribution (0,2) with size ; otherwise, it is considered as high leverage points and follows the Uniform distribution (1,7) with size . The different ranges applied in the uniform distribution are used to fit the different scenarios according to the added artificial noise, vertical outliers, and high leverage points in the dataset. By default, the predictor and response variable should be centered and scaled before the analysis. In the PLSR model, the selection on optimal number of PLSR components used in the model fitting is very important to prevent the model from becoming over- or under-prediction.
To assess the performance of the methods, several statistical measures such as desirability indices are used: Root Mean Square Error (RMSE), Coefficient of Determination (R2), and Standard Error (SE). The RMSE measures the absolute error of the predicted model; R2 is the proportion of variation in the data summarized by the model and indicates the reliability of the goodness of fit for model; and SE measures the uncertainty in the prediction. Here, the RPD parameter has no more used because it is not different than R2 to classify the model is poor or not [44]. Using the classical PLSR, the RMSECV which is the RMSE obtained through cross-validation, is calculated, along with the increasing number of PLS components. The RMSEP value is the RMSE obtained using the fitted model. In the simulation study, the maximum number of PLS components used was limited up to 20. Some different scenarios were applied to see the stability of classical PLSR model based on sample size, number of predictors, number of important variables, and the contamination of outlier and high leverage points in the dataset. In Figure 1, with no contamination in the data it can be seen that using small sample size ( = 60), small number of predictors ( = 41), and 10% relevant variable ( = 10%) the discrepancy between RMSECV and RMSEP is about two to five times. While using higher number of predictors ( = 101) the discrepancy then become larger. Another scenario using bigger sample size ( = 200), small number of predictors ( = 41), and 30% relevant variable ( = 30%) the discrepancy between RMSECV and RMSEP relatively smaller. While using higher number of predictors ( = 101) the discrepancy increases about two times. This shows that the classical PLS become instable and loss it accuracy when the number of sample size is small and number of predictor higher than sample size. In addition, with less number of relevant variable in the predictor variable also impacts to decrease the model accuracy. Using bigger sample size (for example = 200) as the number of PLS components increases the discrepancy between RMSECV and RMSEP become smaller hence improve the model accuracy and reliability. The rule is the gap between RMSEC and RMSEP values should very small and close to 0. This condition guarantees the reliability of the calibrated model and prevent the model becomes over-under fitting.
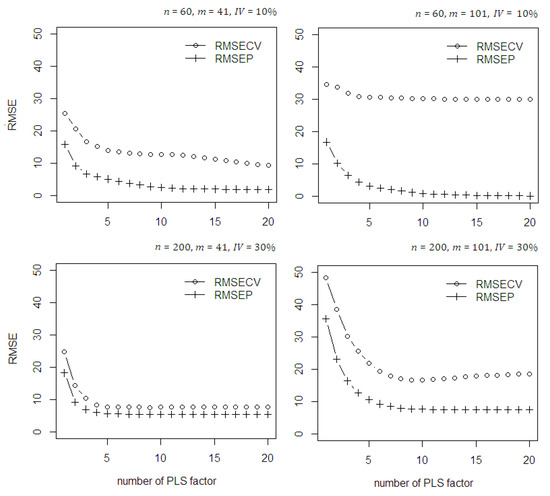
Figure 1.
The RMSECV and RMSEP of the classical PLSR on the simulated data with no contamination of outlier and high leverage points.
The stability of the classical PLSR model then is evaluated by introducing the presence of outlier and leverage points in the dataset (see Figure 2). According to the scenarios given, the classical PLSR model failed to converge even using higher number of PLS components. This can be investigated through RMSECV values which become large and fail to be minimum. In addition, the discrepancy between both RMSECV and RMSEP values also large. This gives evidence that the presence of outlier and HLP in the dataset will destroy the convergence and results to the poor model fitting.
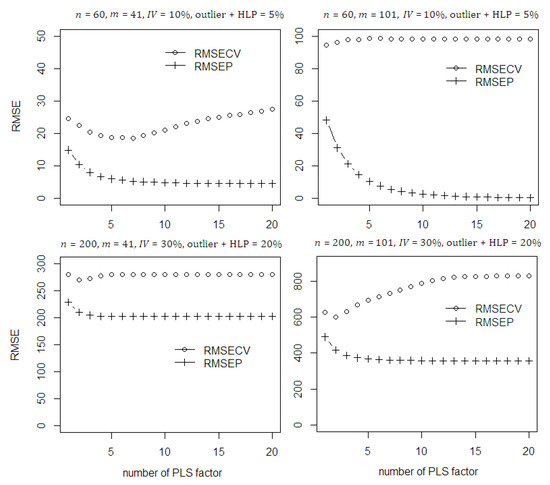
Figure 2.
The RMSECV and RMSEP of the classical PLSR on the simulated data with contamination of outlier and high leverage points.
In the proposed RRWA-PLS, the 20% trimmed SEP was used to calculate the weight by removing the 20% highest absolute residual. This procedure is suggested to produce the robust weight instead of using the whole residual. In the calculation of trimmed SEP in each PLS component, using the cross validation procedure the median is preferred. In general, using different dataset scenarios with contamination of outlier and HLP (see Figure 3) the proposed robust trimmed SEP median succeed to remove the influence by removing 20% highest absolute residual. The SEP mean is suffered both with small ( = 60) and bigger ( = 200) sample size due to the contamination. This results in the SEP values of SEP mean becomes two to four times greater than trimmed SEP median. The SEP median lost its advantage when bigger sample size ( = 200) is used. This results in the SEP values of SEP median becomes four times greater than trimmed SEP median. The SEP values using trimmed SEP median is lower than trimmed SEP mean thus improves model accuracy. This proves the robustness of the trimmed SEP median in weight calculation which irrespective of sample size, number of important variables, and percentage of contamination of outlier and HLP in the dataset.
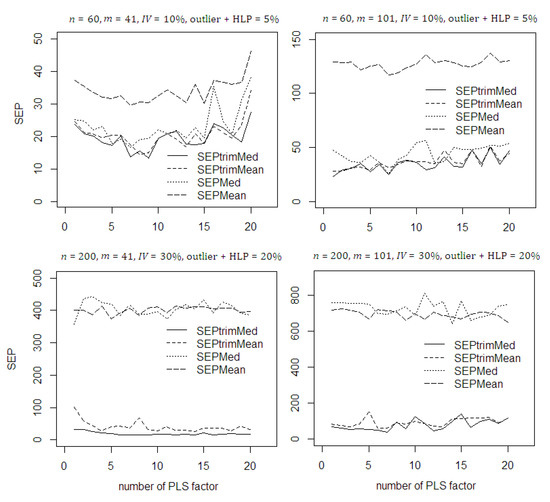
Figure 3.
SEP values in the RRWA-PLS using different approach on the simulated data with contamination of outlier and HLP.
It is very important to compare the weighting schemes between the WA-PLS and RRWA-PLS. This weight provides the contribution of predictors based on the aggregation of the PLS components used in the model. In Figure 4, the mean weights of the methods are also shown to illustrate two conditions: no contamination and with contamination of outlier and high leverage points. For no contamination, the weights in both WA-PLS and RRWA-PLS methods increase as the number of PLS component increases. The weight of RRWA-PLS is relatively smaller than that of the WA-PLS. In cases where the number of PLS components are greater than 10, the weight in both methods are not so much affected by the increasing number of PLS used in the model. On the other hand, when contaminated with outlier and HLP, the weights in both WA-PLS and RRWA-PLS methods decrease as the number of PLS component increases. Based on these scenarios, the WA-PLS still produces higher RMSE than RRWA-PLS. In general, according to the less weight value used in the model, the RRWA-PLS method is still superior and more efficient than the WA-PLS.
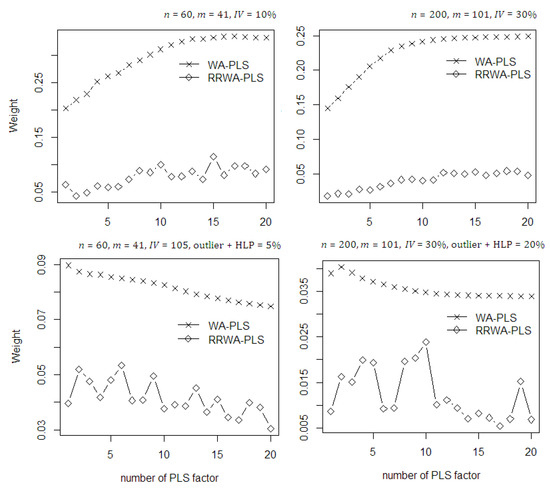
Figure 4.
The mean weights of the WA-PLS and RRWA-PLS on the simulated data with and without contamination of outlier and HLP.
In Figure 5, the prediction accuracy of the methods is evaluated through their RMSEP values. To get a better illustration, the maximum number of PLS components was limited to 15 components. With no contamination of outlier and HLP in the dataset, in the first 6 number of PLS components, the RRWA-PLS is less efficient than the classical PLS and WA-PLS. However, as the number of PLS component increases up to 15, the RRWA-PLS is comparable to the classical PLS and WA-PLS. The proposed RRWA-PLS shows its robustness when the contaminations of outlier and HLP exist in the dataset. It has succeeded to prevent the influence of the outlier and HLP during model fitting. On the other hand, the classical PLS and WA-PLS suffer from the influence of outlier and HLP both in low and high level percentage of contamination, resulting in poor accuracy.
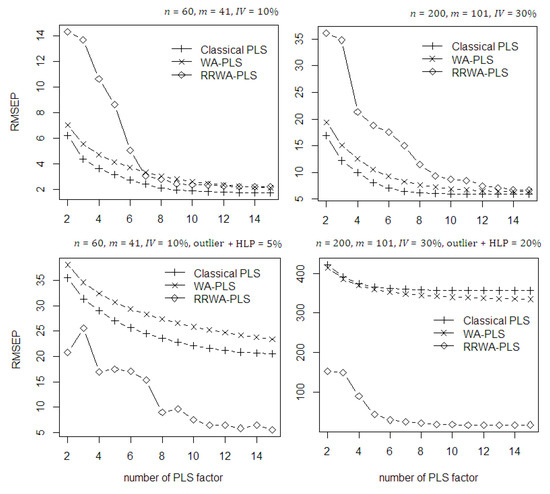
Figure 5.
The RMSEP values of the classical PLS, WA-PLS, and RRWA-PLS on the simulated data with and without contamination of outlier and HLP.
To further evaluate the methods, the Monte Carlo simulation was run 10,000 times on different dataset scenarios. The results, based on the average of statistical measures, are shown in Table 1. As mentioned earlier, in the fitting process, the number of PLS components used in the proposed methods was limited to 15. We use the term “PLS with opt.” to refer to the classical PLS with optimal number of PLS component selected through the “onesigma” approach and cross-validation. We also include a weight improvement procedure in the WA-PLS known as MWA-PLS. The MWA-PLS uses the robust weight version in the RRWA-PLS to replace the non-robust weight in WA-PLS. Based on the results, with no outliers and HLP in the dataset, the non-robust PLSR coupled with optimal components and WA-PLS are comparable to the MWA-PLS and RRWA-PLS. On the other hand, in the presence of outliers and HLP, the proposed RRWA-PLS method is superior to the classical PLS, WA-PLS, and MWA-PLS. Replacing the weight in the WA-PLS with the weight of the robust version improves the model accuracy with lower SE and better R2 values. The classical PLS fails to find the optimal number of PLS components due to the influence of 5–10% contamination of outliers and HLP during the fitting process. The WA-PLS also fails to fit the predicted model due to the impact of the contamination. The proposed RRWA-PLS consistently has the lowest RMSE, SE, and better R2 compared to the other methods, irrespective of the sample sizes, number of important variables, and percentages of contamination of outliers and HLP in the dataset.

Table 1.
The RMSE, R2, and SE in the weighted methods using the Monte Carlo Simulation with different dataset scenarios.
The prediction ability of the methods using the contamination data was evaluated by plotting the predicted values against the actual values (see Figure 6). The classical PLS and WA-PLS suffered from the contamination of outliers and HLP in the dataset, which resulted in a poor prediction. This is because the PLSR estimator is not resistant to the contamination hence, biasing the estimated model. The MWA-PLS and proposed RRWA-PLS are completely free from the impact of outliers and HLP in the dataset. The influential observations are removed far from the fitting line, while good observations are closed to the fitted regression line. The prediction ability in RRWA-PLS is better than the MWA-PLS; the method ensures the best prediction capabilities with better accuracy than the other methods. The RRWA-PLS shows its robustness which is not affected by the inclusion of model with the number of PLS component used and is resistant to the influence of outliers and HLP.
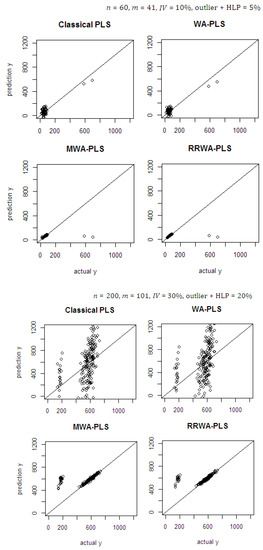
Figure 6.
Predicted against actual values on the simulated data using PLS with opt., WA-PLS, MWA-PLS, and RRWA-PLS.
5. NIR Spectral Dataset
NIR spectral data from oil palm fruit mesocarp were collected to evaluate the methods. The spectral data use light absorbance in each wavelength bands adopted from Beer-Lambert Law [6], and the data are presented in column vector using the log base 10. The spectral measurement was performed by scanning (in contact) the fruit mesocarp using a Portable Handheld NIR spectrometer, QualitySpec Trek, from Analytical Spectral Devices (ASD Inc., Boulder, Colorado (CO), USA). A total of 80 fruit bunches were harvested from the site of breeding trial in Palapa Estate, PT. Ivomas Tunggal, Riau Province, Indonesia. There were 12 fruit mesocarp samples in a bunch collected from different sampling positions. The sampling positions comprised the vertical and horizontal lines in a bunch (see [23]): bottom-front, bottom-left, bottom-back, bottom-right, equator-front, equator-left, equator-back, equator-right, top-front, top-left, top-back, and top-right. Right after collection, the fruit mesocarp samples were sent immediately to the laboratory for spectral measurement and wet chemistry analysis. The source of variability such as planting materials (Dami Mas, Clone, Benin, Cameroon, Angola, Colombia), planting year (2010, 2011, 2012) and ripeness level (unripe, under ripe, ripe, over ripe) were also considered to cover the different sources of variation in the palm population as much as possible.
Two sets of NIR spectral data with different sample properties, the fresh fruit mesocarp and dried ground mesocarp, were used in the study. The average of three spectra measurement on each fruit sample mesocarp was used in the computation. The fresh fruit mesocarp was used to estimate the percentage of Oil to Dry Mesocarp (%ODM) and percentage of Oil to Wet Mesocarp (%OWM), while the dried ground mesocarp was used to estimate the percentage of Fat Fatty Acids (%FFA). These parameters were analyzed through conventional analytical chemistry that adopts standard test methods from the Palm Oil Research Institute of Malaysia (PORIM) [45,46]. The %ODM was calculated in dry matter basis, which removes the weight of water content, while the %OWM used wet matter basis. Statistically, the distribution range of %ODM used as dataset is 56.38–86.9%; the %OWM is 19.75–64.81%, and the %FFA is 0.17–6.3%. The NIR spectra on oil palm fruit mesocarp (both in fresh and dried ground mesocarp) and its frequency distribution on response variables, the %ODM, %OWM, and %FFA, can be seen in the previous study (see [23]). It is important to note that no prior knowledge on whether or not outliers and high leverage points are present in this dataset. The discussions were therefore, addressed to evaluate the methods based on their accuracy improvement through its desirability index.
5.1. Oil to Dry Mesocarp
A total of 960 observations which comprised 488 wavelengths (range 550–2500 nm: 4 nm interval) of NIR spectral of fresh fruit mesocarp were used in this study. Following a prior procedure, the cross validation scheme was employed to obtain the RMSECV value in parallel to the increasing number of PLS components. To evaluate the RMSE values both in fitting and prediction ability performance, the scree plot is presented in Figure 7. This plot is essential to observe when the slope starts leveling off and illustrate the gap difference between the RMSECV and RMSEP values. The maximum number of PLS components was limited to 30 for computation efficiency purpose.
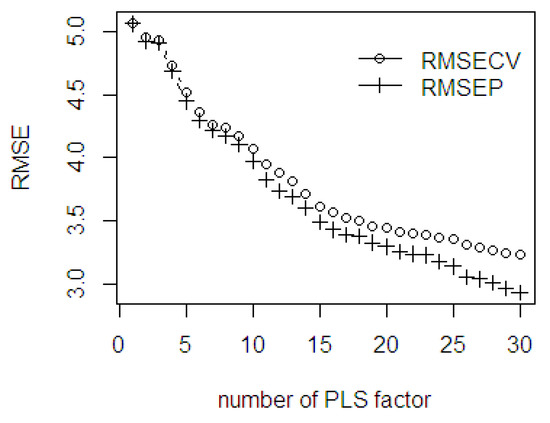
Figure 7.
The RMSE of the fitted PLSR through cross validation and the prediction ability using %ODM dataset.
As seen in Figure 7, the stages of where the slope starts leveling off are at 7, 16, and 26 PLS components. The gap difference between the RMSECV and RMSEP values is wider after 26 PLS components, but both errors gradually become smaller. A larger discrepancy between the values indicates an over fitted which decreases the model accuracy. This indicates that despite using higher components, improvement in the accuracy is not guaranteed.
The mean weights of the fitted PLS both using WA-PLS and RRWA-PLS are plotted in Figure 8. It can be seen that the weight of the WA-PLS rapidly increases as the number of PLS components increases. This shows that using a higher number of PLS components improves accuracy. In RRWA-PLS, the weights are relatively comparable as the PLS components increases. It is interesting to observe that by using the weight strategy in RRWA-PLS, some components show lower mean weights compared to the others even though they have less and higher PLS components. For instance, applying 2 and 5 PLS components results in the signal for under fitting while applying 35 and 45 PLS components results in over fitting. The weighting scheme in the WA-PLS and RRWA-PLS depends on the number of PLS components used in the PLSR model. In fact, using a higher number of PLS components may risk in the inclusion of more noise, yielding a larger variation in the predicted model. The WA-PLS is known to be only suitable in preventing a large regression coefficient which indicates an over fitting. Through its corrected weights using reliability values, the RRWA-PLS does not only prevent over fitting but also under fitting.
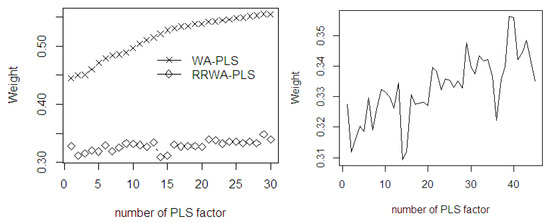
Figure 8.
The mean weights of the fitted PLSR in WA-PLS and RRWA-PLS methods using %ODM dataset.
As seen in Figure 9, the prediction ability error between the classical PLS, WA-PLS, and RRWA-PLS are comparable to each other. The first minimal RMSEP was obtained with 7 PLS components. After the 7 PLS components, the WA-PLS produced a higher RMSEP than the classical PLS and RRWA-PLS. The second minimal RMSEP was obtained with 16 PLS components, and the third minimal RMSEP was obtained with 26 PLS components. The classical PLS and RRWA-PLS have similar prediction ability error with 4, 9, 27, 28, 29, and 30 PLS components used in the PLSR model. In general, the RMSEP values using RRWA-PLS method are always within the range and fall around the classical PLS. The RMSEP curve decreases slightly up to 30 PLS components. In an industrial application, the number of factor greater than 30 PLS components is not recommended. Although this would yield better prediction ability, it is computationally intensive.
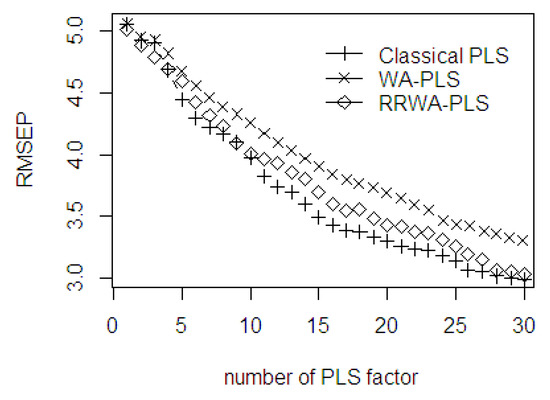
Figure 9.
The RMSEP values of classical PLS, WA-PLS, RRWA-PLS method using %ODM dataset.
The performance of WA-PLS is not better than its modified weight in MWA-PLS (see Table 2). However, the accuracy of MWA-PLS is still lower than that of the RRWA-PLS. This is due to the weight in the WA-PLS which is not able to capture the reliability of the predictor variables. Comparing the prediction ability in the classical PLS with optimum PLS component (27), the RRWA-PLS is still superior. To prevent the influence of noise to the final model, we eliminated the first PLS component from the RRWA-PLS model. This is due to the fact that the first PLS component is usually less accurate if it is still included in the procedure.

Table 2.
The RMSE, R2, and SE in the weighted methods using %ODM data.
5.2. Oil to Wet Mesocarp
In this section, the %OWM is considered as the response variable of the NIR spectral fresh fruit mesocarp dataset. The evaluation on the RMSE values, both in fitting and prediction ability, is presented in the scree plot. The maximum number of PLS components was limited to 30 for computation efficiency purposes. As seen in Figure 10, the slope of scree plot starts leveling off at 7, 16, and 22 PLS components. The gap difference between the RMSECV and RMSEP values are wider after the 22 PLS components. Contrarily, even though both errors become slightly smaller, a large difference between the RMSECV and RMSEP would lead to over fitting and make the predicted model unstable.
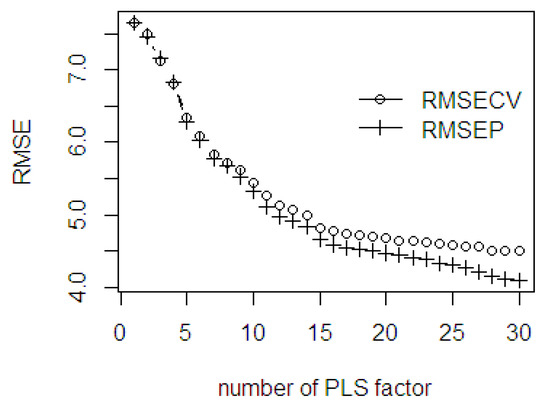
Figure 10.
The RMSE of the fitted PLSR through cross validation and the prediction ability using %OWM dataset.
With the increasing number of PLS components, the mean weights of both WA-PLS and RRWA-PLS also increase (see Figure 11). The mean weights of WA-PLS method are comparably higher to the weight in the RRWA-PLS where accuracy is improved by employing a higher number of PLS components. There are some components in the RRWA-PLS with mean weights lower than those of other PLS components although they have a higher number of components. Applying 2 and 5 PLS components results under fitting, while applying 26 and 29 PLS components results in over fitting. The RRWA-PLS shows its robustness which is not dependent on the increasing number of PLS components used. Its weighing scheme is based on the selection of the relevant aggregate number of PLS components used as factors in the PLSR model. The most relevant PLS components will get a higher weight, while the less relevant will obtain a lower weight.
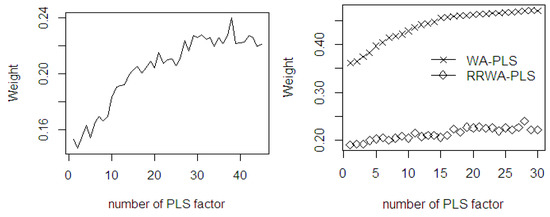
Figure 11.
The mean weights of the fitted PLSR in WA-PLS and RRWA-PLS methods using %OWM dataset.
Figure 12 indicates that the prediction ability of the three methods using the first 5 components is fairly close to each other, but afterwards their performances seem to be different in terms of accuracy. The first minimal RMSEP is obtained with 8 PLS components. After the 8 PLS components, the WA-PLS produces higher RMSEP than the classical PLS and RRWA-PLS. The second minimal RMSEP is obtained with 14 PLS components, and the third minimal RMSEP is obtained with 23 PLS components. The classical PLS with 15 to 22 PLS components produces lower RMSEP values; however, after 24 PLS components, the accuracy between RRWA-PLS and classical PLS becomes closer. In general, the RMSEP values using RRWA-PLS method are always within the range, and the values are reasonably close to the classical PLS. The RMSEP curves slightly decrease which begin from 17 to 30 PLS components. The WA-PLS relatively has low accuracy compared to the RRWA-PLS and classical PLS. The WA-PLS suffers from over-under fitting due to several irrelevant variables, but it may still possibly be included in the fitting process.
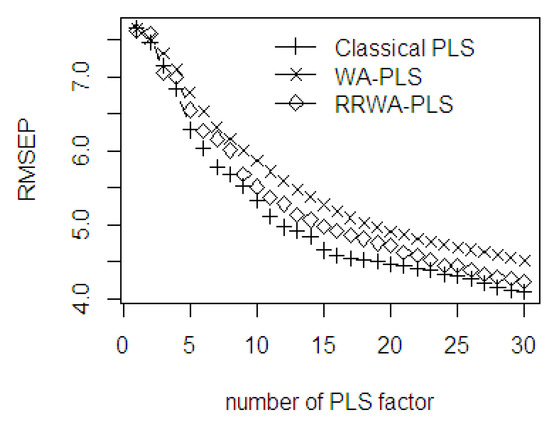
Figure 12.
The RMSEP values of classical PLS, WA-PLS, RRWA-PLS method using %OWM dataset.
Using its optimum at 22 PLS components, the classical PLS with the optimum number of PLS components is indeed inconsistent and sensitive to the number of PLS components used. By comparing the RMSE, R2, and SE values in Table 3, it can be concluded that the proposed RRWA-PLS produces better accuracy than the other methods. The modified weight in MWA-PLS has improved the accuracy of the predicted model; however, it cannot outperform the RRWA-PLS. The robust weighted-average strategy prevents the PLSR model from depending on the specific number of PLS components used in the fitting process.

Table 3.
The RMSE, R2, and SE in the weighted methods using %OWM data.
5.3. Fat Fatty Acids
The NIR spectral of dried ground mesocarp with a total of 839 observations and 500 wavelengths (range 500–2500 nm: 4 nm interval) were utilized as predictor variables. Here, the %FFA was used as the response variable. In the scree plot (Figure 13), the RMSECV and RMSEP curves gradually decrease when the number of PLS components increases. Within the first 10 PLS components, the gap difference between RMSECV and RMSEP is small, but after 10 PLS components, the gap difference starts to increase continuously. The slope of the scree plot starts leveling off at the 6, 16, 22, and 27 PLS components. The gap difference between the RMSECV and RMSEP values becomes wider after 16 PLS components. Therefore, the use of specific number of PLS components affects the accuracy of the fitted model.
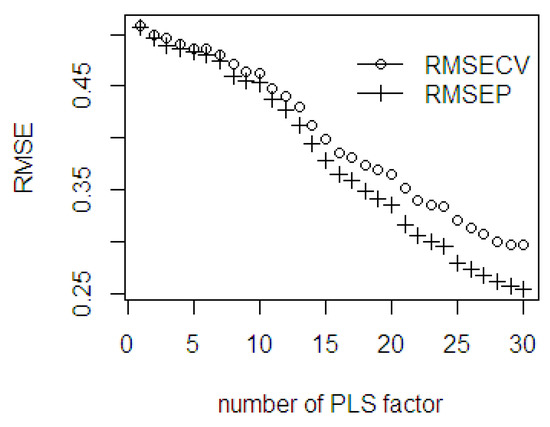
Figure 13.
The RMSE of the fitted PLSR through cross validation and the prediction ability using %FFA dataset.
The mean weights of both WA-PLS and RRWA-PLS increase as the number of PLS component (see Figure 14) increases. Using %FFA dataset, the weight of RRWA-PLS is higher than that of the WA-PLS. The mean weights of the WA-PLS method increase more steeply as the number of PLS components increases. This indicates that the predicted model tends to be over fitting. The weight of the RRWA-PLS is robust since its weight does not depend on the aggregation number of PLS components used, irrespective of the number of sample size and the number of important variables. Moreover, the weight is resistant to the influence of outliers and HLP that may exist in the dataset.
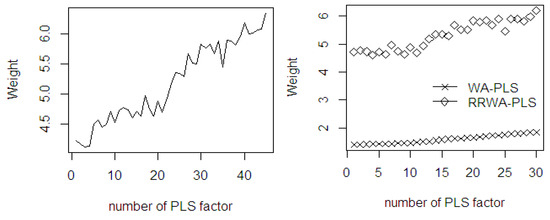
Figure 14.
The mean weights of the fitted PLSR in WA-PLS and RRWA-PLS methods using %FFA dataset.
In the first 6 components, the prediction ability of the three methods is comparable to each other (see Figure 15). After 10 components, the WA-PLS has less accuracy than the classical PLS and the RRWA-PLS. The first minimal RMSEP is obtained at 8 PLS components; after 8 PLS components, the WA-PLS produces larger RMSEP than the classical PLS and the RRWA-PLS. The WA-PLS shows the worst performance using this %FFA dataset. The second minimal RMSEP is obtained at 17 PLS components, and the third minimal RMSEP is obtained at 27 PLS components. The RMSEP values using RRWA-PLS method are always within the range and close to the classical PLS. The RMSEP in the classical PLS is not robust when it comes to the number of PLS components used as using any selection methods to find the optimal number of PLS components to be used in the PLSR model will result in unstable results. The application of an improper method in the selection will produce a less accurate result. The solution in using the robust weighted average is then suggested as it is unnecessary to find the optimal components. This is the automated fitting process in the PLSR model.
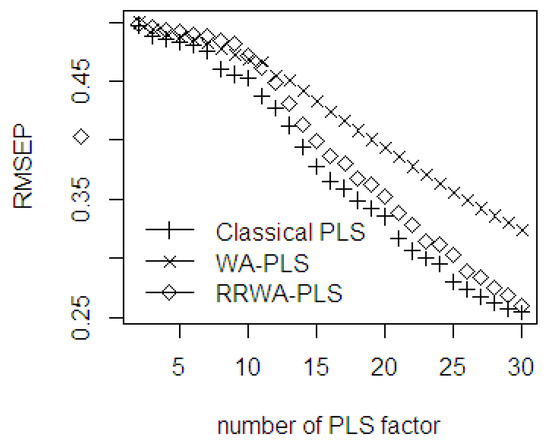
Figure 15.
The RMSEP values of classical PLS, WA-PLS, RRWA-PLS method using %FFA dataset.
In Table 4, the classical PLS really suffers from the model complexity used in the fitting process. Using the one-sigma heuristic method in component selection, the accuracy of the selected optimal number of PLS components is not better than the PLS with a higher number of components. This shows the weakness of using a specific number of PLS components in the PLSR model. The robust RRWA-PLS is free from the complexity of the aggregation number of PLS components used. As seen in Table 3, the WA-PLS has the worst performance compared to the MWA-PLS, classical PLS, and RRWA-PLS. The use of RRWA-PLS method is preferred to the classical PLS because it does not require the selection of an optimal number of PLS components to be used in the final PLSR model. In addition, the method offers better reliability of the goodness-of-fit for the model.

Table 4.
The RMSE, R2, and SE in the weighted methods using %FFA data.
6. Reliability Values
A number of irrelevant variables most probably still exist in the dataset. If the PLSR method fails to screen and downgrade the contribution of these irrelevant variables, it might decrease the accuracy of the final fitted model. The use of RRWA-PLS on the artificial dataset (see Figure 16a) helps the method to screen the most relevant variables and downgrade the irrelevant variables in the dataset successfully. The use of NIR spectral data with different response variables (%ODM, %OWM, and %FFA) has allowed the method to show its potential in the wavelength selection process. The method highlights the most relevant wavelengths and downgrades the influence of irrelevant wavelengths based on spectra absorption (see Figure 16b–d). The reliability values are important in order to increase the computational speed in the fitting process, improve the accuracy, and provide better interpretation of the NIR spectral dataset.
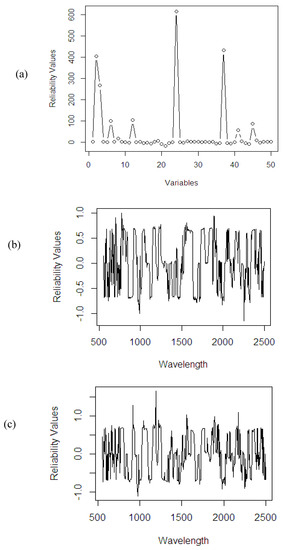
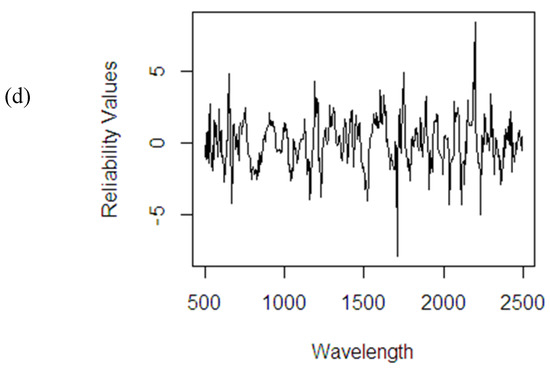
Figure 16.
Reliability values using RRWA-PLS method on different datasets: (a) artificial data; NIR spectral dataset: (b) %ODM; (c) %OWM; (d) %FFA.
7. Conclusions
This study has shown the robustness in the chemometric analysis of NIR spectral data related to the aggregate number of PLS components and the resistance against outliers and HLP. The rich and abundant information in the NIR spectral requires advanced chemometric analysis to classify the most and least relevant wavelengths used in computation. Based on the results, the proposed RRWA-PLS method is the most preferred method compared to other methods due to its robustness. The weight improvement in MWA-PLS gives a better solution in improving the accuracy and reliability of WA-PLS. In the selection of the optimal number of PLS components, the classical PLS still needs the re-computational process to determine a specific complexity each time the model is updated. The proposed RRWA-PLS shows its superiority in the improvement of weight and variable selection process. It is also resistant to the contamination of outliers and HLP in the dataset. In addition, the RRWA-PLS method offers a solution for automated fitting process in the PLSR model as it does not require the selection of the optimal number of PLS components unlike in the classical PLS.
Author Contributions
Conceptualization and methodology: D.D.S., H.M., J.A., M.S.M., J.-P.C.; Data Collection: D.D.S., H.M., J.-P.C.; Computational and Validation: H.M., J.A., M.S.M.; First draft preparation: D.D.S., H.M.; Writing up to review and editing: D.D.S., H.M., J.A., M.S.M., J.-P.C. All authors have read and agreed to the published version of the manuscript.
Funding
The present research was partially supported by the Universiti Putra Malaysia Grant under Putra Grant (GPB) with project number GPB/2018/9629700.
Acknowledgments
This work was supported by a research grant and scholarship from the Southeast Asian Regional Center for Graduate Study and Research in Agriculture (SEARCA). We are also grateful to SMARTRI, PT. SMART TBK for providing the portable handheld NIRS instrument, research site, and analytical laboratory services. We would like to thank Universiti Putra Malaysia for the journal publication fund support. Special thanks are also extended to all research staff and operator of SMARTRI for their cooperation and outstanding help with data collection.
Conflicts of Interest
The authors declare no conflict of interest.
Declaration
The results of this study were presented at the NIR 2019—the 19th biennial meeting of the International Council for NIR Spectroscopy (ICNIRS) held in Gold Coast, Queensland, Australia, from 15–20 September 2019. Some inputs and comments from the audiences and reviewers were included in this paper.
References
- Rodriguez-Saona, L.E.; Fry, F.S.; McLaughlin, M.A.; Calvey, E.M. Rapid analysis of sugars in fruit juices by FT-NIR spectroscopy. Carbohydr. Res. 2001, 336, 63–74. [Google Scholar] [CrossRef]
- Blanco, M.; Villarroya, I.N.I.R. NIR spectroscopy: A rapid-response analytical tool. Trends Anal. Chem. 2002, 21, 240–250. [Google Scholar] [CrossRef]
- Alander, J.T.; Bochko, V.; Martinkauppi, B.; Saranwong, S.; Mantere, T. A review of optical nondestructive visual and near-infrared methods for food quality and safety. Int. J. Spectrosc. 2013, 2013, 341402. [Google Scholar] [CrossRef]
- Lee, C.; Polari, J.J.; Kramer, K.E.; Wang, S.C. Near-Infrared (NIR) Spectrometry as a Fast and Reliable Tool for Fat and Moisture Analyses in Olives. ACS Omega. 2018, 3, 16081–16088. [Google Scholar] [CrossRef] [PubMed]
- Levasseur-Garcia, C. Updated overview of infrared spectroscopy methods for detecting mycotoxins on cereals (corn, wheat, and barley). Toxins 2018, 10, 38. [Google Scholar] [CrossRef]
- Stuart, B. Infrared Spectroscopy: Fundamentals and Applications; Wiley: Toronto, ON, Canada, 2004; pp. 167–185. [Google Scholar]
- Mark, H. Chemometrics in near-infrared spectroscopy. Anal. Chim. Acta 1989, 223, 75–93. [Google Scholar] [CrossRef]
- Cozzolino, D.; Morón, A. Potential of near-infrared reflectance spectroscopy and chemometrics to predict soil organic carbon fractions. Soil Tillage Res. 2006, 85, 78–85. [Google Scholar] [CrossRef]
- Roggo, Y.; Chalus, P.; Maurer, L.; Lema-Martinez, C.; Edmond, A.; Jent, N. A review of near infrared spectroscopy and chemometrics in pharmaceutical technologies. J. Pharm. Biomed. Anal. 2007, 44, 683–700. [Google Scholar] [CrossRef]
- Garthwaite, P.H. An interpretation of partial least squares. J. Am. Stat. Assoc. 1994, 89, 122–127. [Google Scholar] [CrossRef]
- Cozzolino, D.; Kwiatkowski, M.J.; Dambergs, R.G.; Cynkar, W.U.; Janik, L.J.; Skouroumounis, G.; Gishen, M. Analysis of elements in wine using near infrared spectroscopy and partial least squares regression. Talanta 2008, 74, 711–716. [Google Scholar] [CrossRef]
- McLeod, G.; Clelland, K.; Tapp, H.; Kemsley, E.K.; Wilson, R.H.; Poulter, G.; Coombs, D.; Hewitt, C.J. A comparison of variate pre-selection methods for use in partial least squares regression: A case study on NIR spectroscopy applied to monitoring beer fermentation. J. Food Eng. 2009, 90, 300–307. [Google Scholar] [CrossRef]
- Xu, L.; Cai, C.B.; Deng, D.H. Multivariate quality control solved by one-class partial least squares regression: Identification of adulterated peanut oils by mid-infrared spectroscopy. J. Chemom. 2011, 25, 568–574. [Google Scholar] [CrossRef]
- Wold, H. Model construction and evaluation when theoretical knowledge is scarce: Theory and application of partial least squares. In Evaluation of Econometric Models; Elsevier: Amsterdam, The Netherlands, 1980; pp. 47–74. [Google Scholar]
- Manne, R. Analysis of two partial-least-squares algorithms for multivariate calibration. Chemom. Intell. Lab. Syst. 1987, 2, 187–197. [Google Scholar] [CrossRef]
- Haenlein, M.; Kaplan, A.M. A beginner’s guide to partial least squares analysis. Understt. Satistics 2004, 3, 283–297. [Google Scholar] [CrossRef]
- Hubert, M.; Branden, K.V. Robust methods for partial least squares regression. J. Chemom. A J. Chemom. Soc. 2003, 17, 537–549. [Google Scholar] [CrossRef]
- Silalahi, D.D.; Midi, H.; Arasan, J.; Mustafa, M.S.; Caliman, J.P. Robust generalized multiplicative scatter correction algorithm on pretreatment of near infrared spectral data. Vib. Spectrosc. 2018, 97, 55–65. [Google Scholar] [CrossRef]
- Rosipal, R.; Trejo, L.J. Kernel partial least squares regression in reproducing kernel hilbert space. J. Mach. Learn. Res. 2001, 2, 97–123. [Google Scholar]
- Silalahi, D.D.; Midi, H.; Arasan, J.; Mustafa, M.S.; Caliman, J.P. Kernel partial diagnostic robust potential to handle high-dimensional and irregular data space on near infrared spectral data. Heliyon 2020, 6, e03176. [Google Scholar] [CrossRef]
- Centner, V.; Massart, D.L.; De Noord, O.E.; De Jong, S.; Vandeginste, B.M.; Sterna, C. Elimination of uninformative variables for multivariate calibration. Anal. Chem. 1996, 68, 3851–3858. [Google Scholar] [CrossRef]
- Mehmood, T.; Liland, K.H.; Snipen, L.; Sæbø, S. A review of variable selection methods in partial least squares regression. Chemom. Intell. Lab. Syst. 2012, 118, 62–69. [Google Scholar] [CrossRef]
- Silalahi, D.D.; Midi, H.; Arasan, J.; Mustafa, M.S.; Caliman, J.P. Robust Wavelength Selection Using Filter-Wrapper Method and Input Scaling on Near Infrared Spectral Data. Sensors 2020, 20, 5001. [Google Scholar] [CrossRef] [PubMed]
- Wiklund, S.; Nilsson, D.; Eriksson, L.; Sjöström, M.; Wold, S.; Faber, K. A randomization test for PLS component selection. J. Chemom. A J. Chemom. Soc. 2007, 21, 427–439. [Google Scholar] [CrossRef]
- Hastie, T.; Tibshirani, R.; Friedman, J. The elements of statistical learning: Data mining, inference, and prediction. In Springer Series in Statistics; Springer: Berlin/Heidelberg, Germany, 2009. [Google Scholar]
- Van Der Voet, H. Comparing the predictive accuracy of models using a simple randomization test. Chemom. Intell. Lab. Syst. 1994, 25, 313–323. [Google Scholar] [CrossRef]
- Efron, B. Bootstrap Methods: Another Look at the Jackknife. Annal. Stat. 1979, 7, 1–26. [Google Scholar] [CrossRef]
- Gómez-Carracedo, M.P.; Andrade, J.M.; Rutledge, D.N.; Faber, N.M. Selecting the optimum number of partial least squares components for the calibration of attenuated total reflectance-mid-infrared spectra of undesigned kerosene samples. Anal. Chim. Acta 2007, 585, 253–265. [Google Scholar] [CrossRef] [PubMed]
- Tran, T.; Szymańska, E.; Gerretzen, J.; Buydens, L.; Afanador, N.L.; Blanchet, L. Weight randomization test for the selection of the number of components in PLS models. J. Chemom. 2017, 31, e2887. [Google Scholar] [CrossRef]
- Kvalheim, O.M.; Arneberg, R.; Grung, B.; Rajalahti, T. Determination of optimum number of components in partial least squares regression from distributions of the root-mean-squared error obtained by Monte Carlo resampling. J. Chemom. 2018, 32, e2993. [Google Scholar] [CrossRef]
- Shenk, J.S.; Westerhaus, M.O.; Berzaghi, P. Investigation of a LOCAL calibration procedure for near infrared instruments. J. Near Infrared Spectrosc. 1997, 5, 223–232. [Google Scholar] [CrossRef]
- Barton, F.E.; Shenk, J.S.; Westerhaus, M.O.; Funk, D.B. The development of near infrared wheat quality models by locally weighted regressions. J. Near Infrared Spectrosc. 2000, 8, 201–208. [Google Scholar] [CrossRef]
- Naes, T.; Isaksson, T.; Kowalski, B. Locally weighted regression and scatter correction for near-infrared reflectance data. Anal. Chem. 1990, 62, 664–673. [Google Scholar] [CrossRef]
- Dardenne, P.; Sinnaeve, G.; Baeten, V. Multivariate calibration and chemometrics for near infrared spectroscopy: Which method? J. Near Infrared Spectrosc. 2000, 8, 229–237. [Google Scholar] [CrossRef]
- Zhang, M.H.; Xu, Q.S.; Massart, D.L. Averaged and weighted average partial least squares. Anal. Chim. Acta 2004, 504, 279–289. [Google Scholar] [CrossRef]
- Serneels, S.; Croux, C.; Filzmoser, P.; Van Espen, P.J. Partial robust M-regression. Chemom. Intell. Lab. Syst. 2005, 79, 55–64. [Google Scholar] [CrossRef]
- Cui, C.; Fearn, T. Comparison of partial least squares regression, least squares support vector machines, and Gaussian process regression for a near infrared calibration. J. Near Infrared Spectrosc. 2017, 25, 5–14. [Google Scholar] [CrossRef]
- Song, W.; Wang, H.; Maguire, P.; Nibouche, O. Local Partial Least Square classifier in high dimensionality classification. Neurocomputing 2017, 234, 126–136. [Google Scholar] [CrossRef]
- Martens, H.; Naes, T. Multivariate Calibration; John Wiley & Sons: Hoboken, NJ, USA, 1992. [Google Scholar]
- Huber, P.J. Robust regression: Asymptotics, conjectures and Monte Carlo. Ann. Stat. 1973, 1, 799–821. [Google Scholar] [CrossRef]
- Cummins, D.J.; Andrews, C.W. Iteratively reweighted partial least squares: A performance analysis by Monte Carlo simulation. J. Chemom. 1995, 9, 489–507. [Google Scholar] [CrossRef]
- Fushiki, T. Estimation of prediction error by using K-fold cross-validation. Stat. Comput. 2011, 21, 137–146. [Google Scholar] [CrossRef]
- Kim, S.; Okajima, R.; Kano, M.; Hasebe, S. Development of soft-sensor using locally weighted PLS with adaptive similarity measure. Chemom. Intell. Lab. Syst. 2013, 124, 43–49. [Google Scholar] [CrossRef]
- Minasny, B.; McBratney, A. Why you don’t need to use RPD. Pedometron 2013, 33, 14–15. [Google Scholar]
- Siew, W.L.; Tan, Y.A.; Tang, T.S. Methods of Test for Palm Oil and Palm Oil Products: Compiled; Lin, S.W., Sue, T.T., Ai, T.Y., Eds.; Palm Oil Research Institute of Malaysia: Selangor, Malaysia, 1995.
- Rao, V.; Soh, A.C.; Corley, R.H.V.; Lee, C.H.; Rajanaidu, N. Critical Reexamination of the Method of Bunch Quality Analysis in Oil Palm Breeding; PORIM Occasional Paper; FAO: Rome, Italy, 1983; Available online: https://agris.fao.org/agris-search/search.do?recordID=US201302543052 (accessed on 13 October 2020).
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).