CBRL and CBRC: Novel Algorithms for Improving Missing Value Imputation Accuracy Based on Bayesian Ridge Regression
Abstract
1. Introduction
1.1. Missingness Mechanisms
- Missing completely at random (MCAR) [12]: Assume that the missing value indicator matrix and the complete data . The missing data mechanism is described by the conditional distribution of given , say where represents the unknown parameter. If missingness does not depend on the values of the data , missing or observed, then
- Missing at random (MAR) [12]: Let and denote missing data and observed data, respectively. If the Missingness do not depend on the data that are missing, but depends only on of , then,
- Missing not at random (MNAR) [2]: When the missing data depends on both observed and missing data.
1.2. Dealing with Missing Data
- Simple linear regression: In which a linear relationship between the dependent and the independent variables holdswhere is the value of when is equal to zero, is estimated regression coefficient, and is the estimation error.
- Multiple linear regression: In which more independent variables work together to obtain better prediction. The linear relationship between the dependent and independent variables holds
1.3. Relevant Imputation Algorithms
2. Proposed Algorithms
- In the first step, each proposed algorithm takes a dataset as input that holds missing data, then splits it into two sets, the first set includes all complete features, and the second set includes all incomplete features. The authors assume that the target feature contains no missing data, so comprises all full features plus the target feature .
- In the second step, each proposed algorithm implements its feature selection condition to select the candidate feature to be imputed.
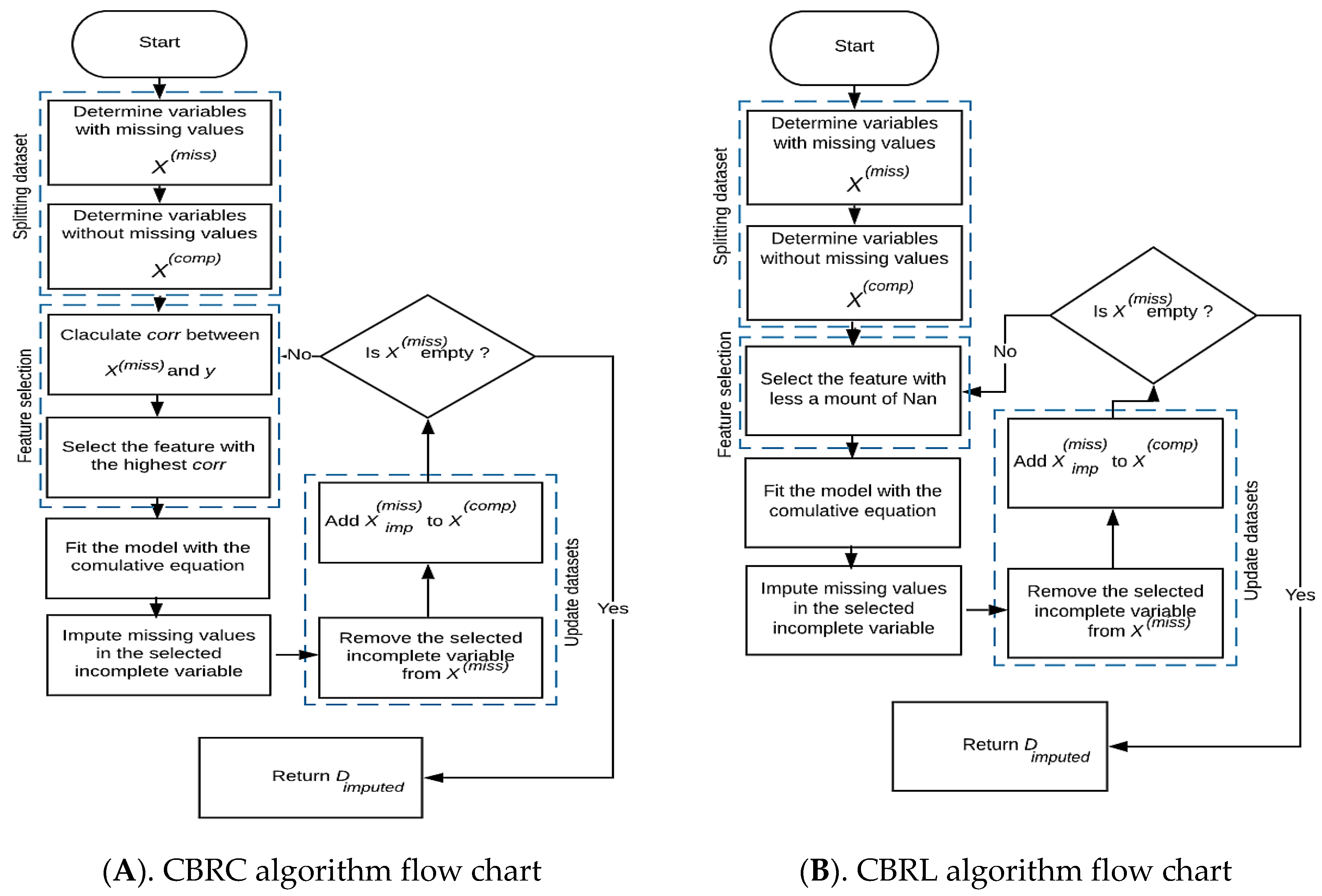
- The first algorithm, we called Cumulative Bayesian Ridge with Less NaN (CBRL), as its name indicates that this algorithm selects the feature that contains less missing data, which leads the model to be built on the most available information (Algorithm 1).
Algorithm 1 CBRL 1: Input:
2: D: A dataset with missing values containing instances.
3: Output:
4: Dimputed: A dataset with all missing features imputed.
5: Definitions:
6: Set of complete features.
7: Set of incomplete features.
8: Imputed feature from .
9: Number of features containing missing values.
10: Set of missing instances in the independent feature .
11: Number of missing values in the independent feature .
12: Begin
13: 1 Split D into and .
14: 2 From select that satisfies the condition:
15: Min (Card ()).
16: 3 While
17: i ← index of the candidate feature in .
18: ii Fit a Bayesian ridge regression model on as independent features and as dependent feature.
19: iii ← Impute the missing data in with the fitted model.
20: iv Delete from and add to .
21: End While
22: 4 return Dimputed ←
23: End - The second algorithm, we called Cumulative Bayesian Ridge with high correlation (CBRC), depends on the highest correlation between the candidate features that contain missing data and the target feature. CBRC chooses the feature that gives the highest correlation with the target feature. The correlation criterion (i.e., Pearson correlation coefficient) is given by Equation (4) [25]:where is the feature, is the output feature, is the variance, and is the covariance. Correlation ranking can only notice linear dependencies between the input feature and output feature (Algorithm 2).
Algorithm 2 CBRC 1: Input:
2: D: A dataset with missing values containing instances.
3: Output:
4: Dimputed: A dataset with all missing features imputed.
5: Definitions:
6: Set of complete features.
7: Set of incomplete features.
8: Imputed feature from .
9: Number of features containing missing values.
10: Correlation between and , .
11: Begin
11: 1 Split D into and .
12: 2 From select that satisfies the condition:
13: Max (Corr ()).
14: 3 While
15: i ← index of the candidate feature in .
16: ii Fit a Bayesian ridge regression model on as independent features and as dependent feature.
17: iii ← Impute the missing data in with the fitted model.
18: iv Delete from and add to .
19: End While
20: 4 return Dimputed ←
21: End
- After selecting the candidate feature , the model is fitted with the cumulative formula defined in Equation (5) using the candidate feature as dependent and the as the independent feature. The selected feature deleted from , and after imputation, the imputed feature is added to . Now consists of all complete features, and . Select another candidate feature from . Fit the model using the cumulative BRR formula with this candidate feature as the dependent feature and as an independent feature.where:where is the number of features containing missing values and is the number of complete independents.
- Repeat from step 2 of feature selection until is empty, then return the imputed dataset (), see Figure 1.
3. Experimental Implementation
3.1. Benchmark Datasets
3.2. Evaluation
3.2.1. RMSE and MAE
3.2.2. R2 Score
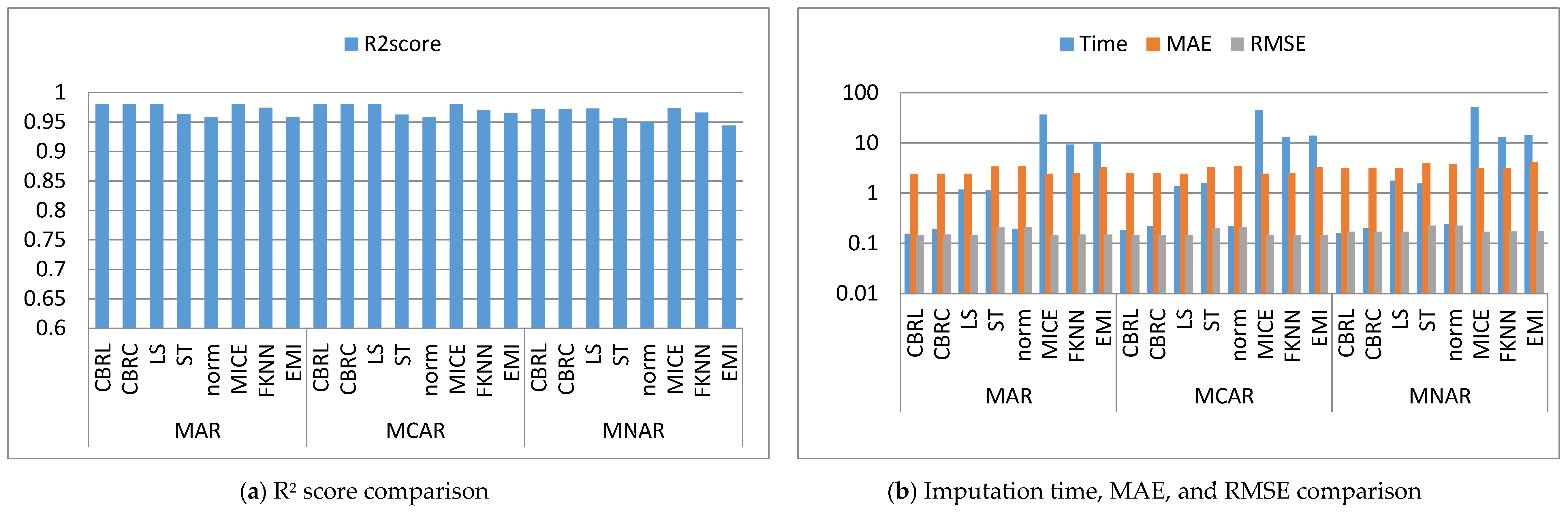
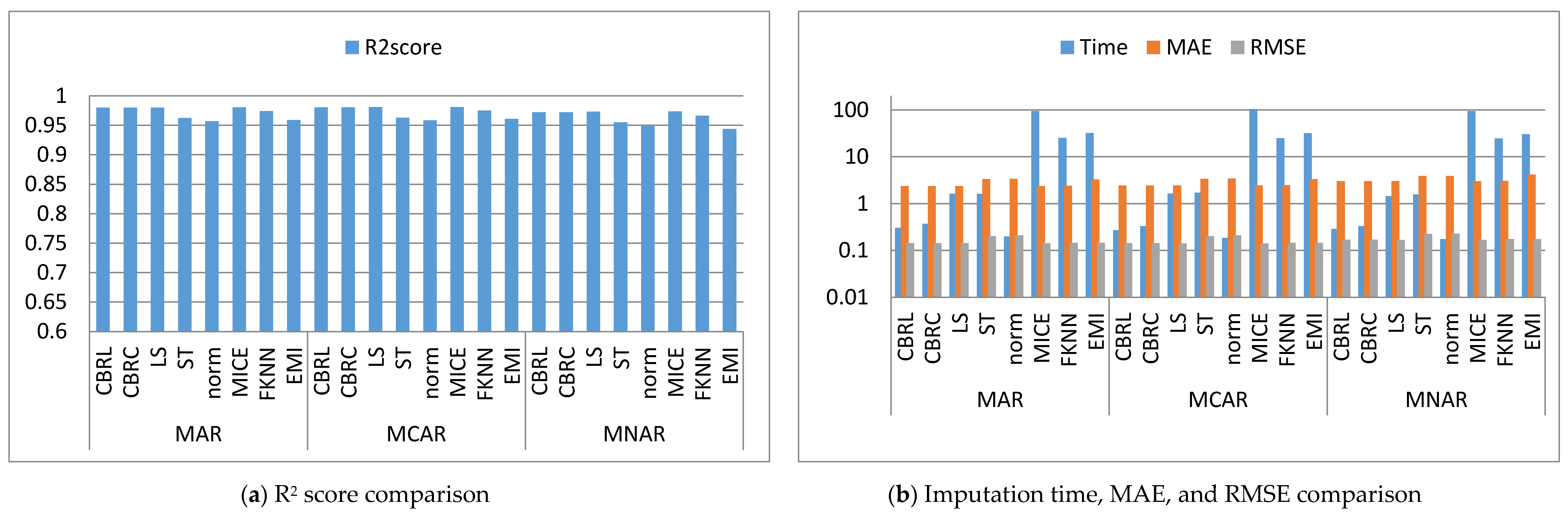
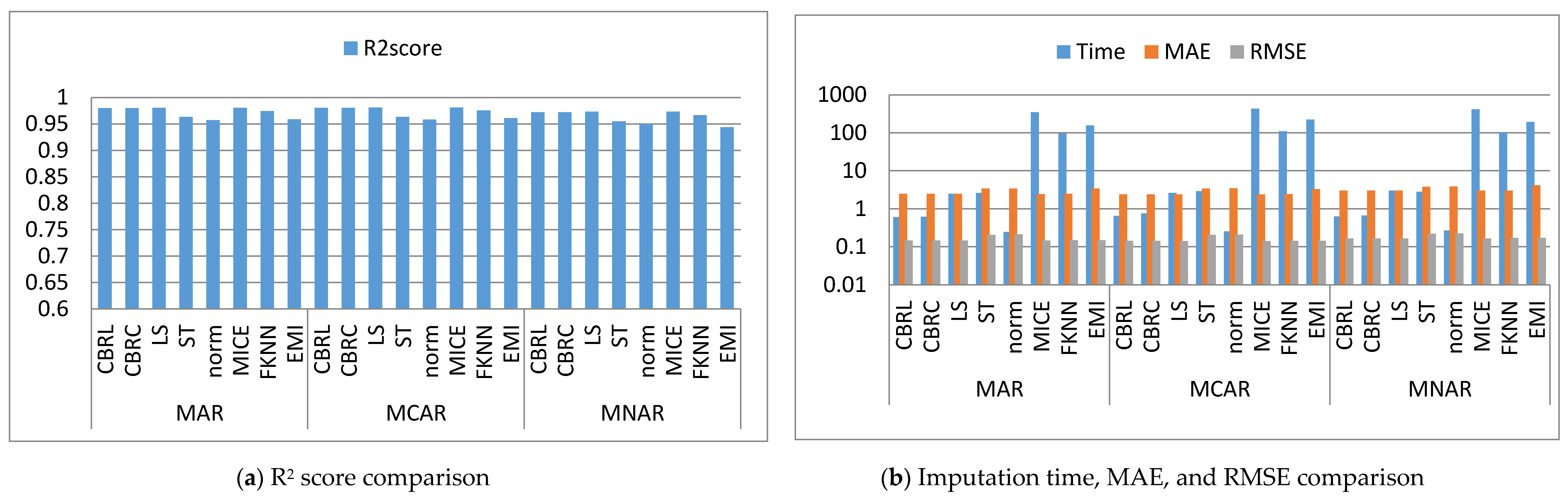
4. Results and Discussion
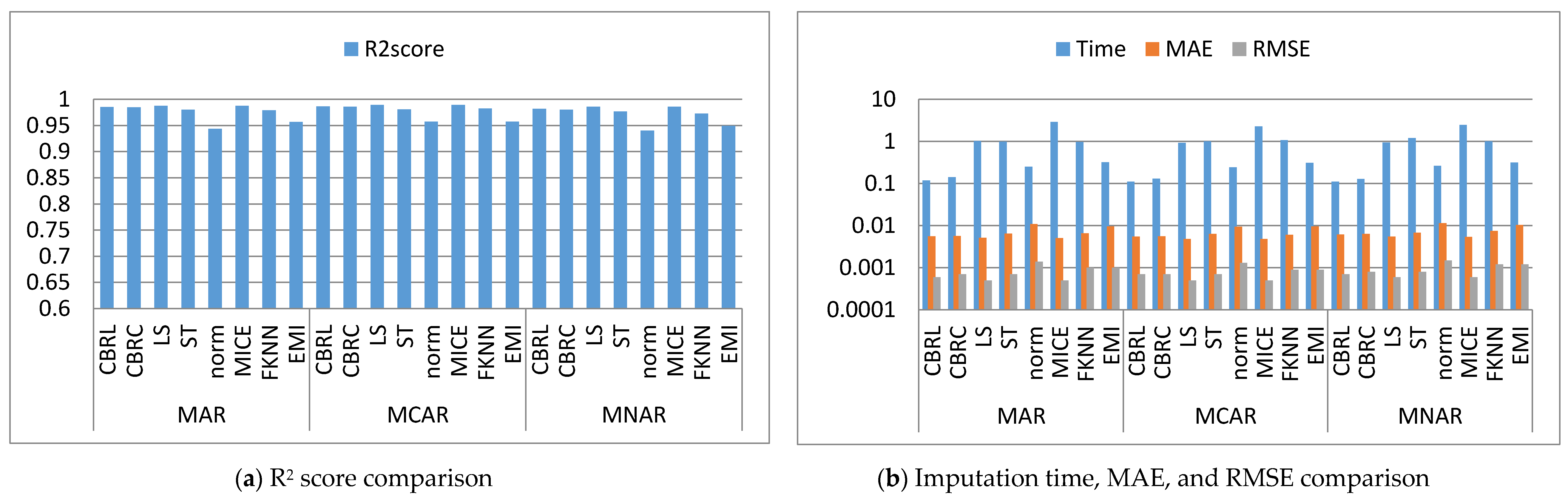
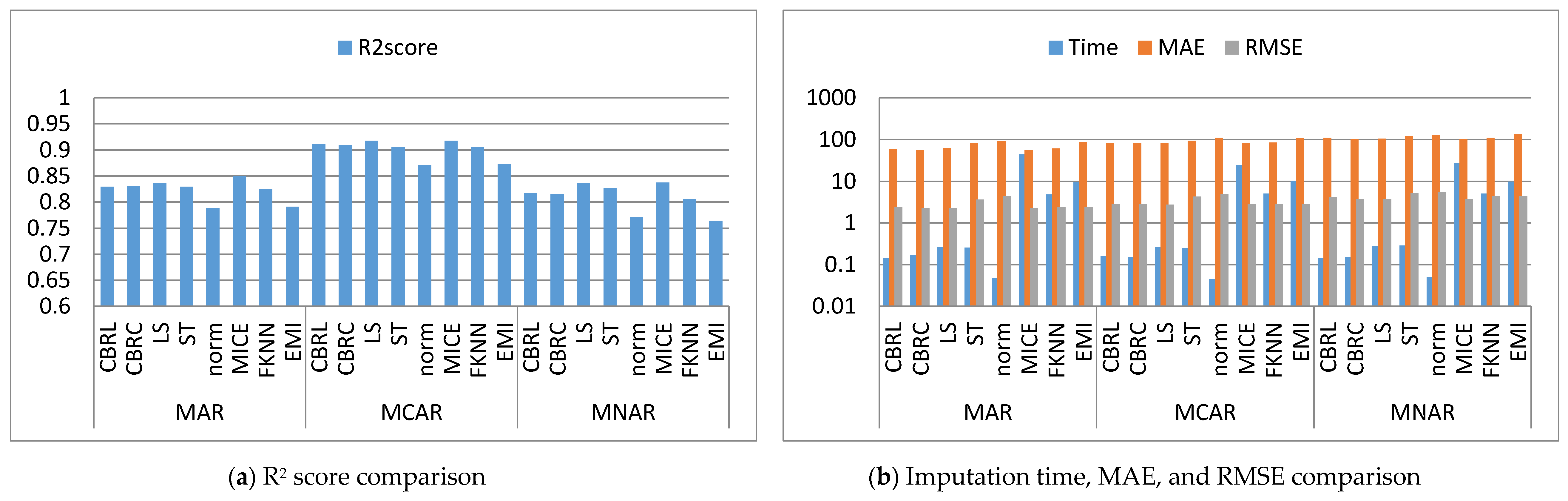
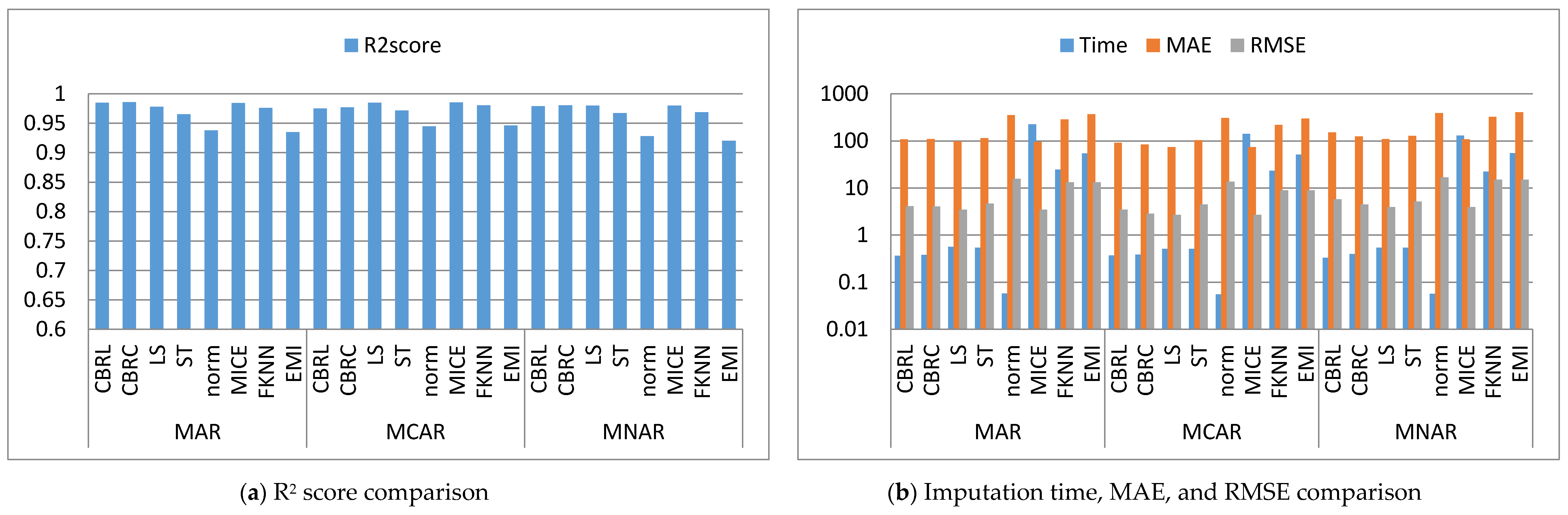
4.1. Error Analysis
4.2. Imputation Time
4.3. Accuracy Analysis
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Mostafa, S.M. Imputing missing values using cumulative linear regression. CAAI Trans. Intell. Technol. 2019, 4, 182–200. [Google Scholar] [CrossRef]
- Salgado, C.M.; Azevedo, C.; Manuel Proença, H.; Vieira, S.M. Missing data. Second. Anal. Electron. Health Rec. 2016, 143–162. [Google Scholar] [CrossRef]
- Hapfelmeier, A.; Hothorn, T.; Ulm, K.; Strobl, C. A new variable importance measure for random forests with missing data. Stat. Comput. 2014, 24, 21–34. [Google Scholar] [CrossRef]
- Batista, G.; Monard, M.-C. A study of k-nearest neighbour as an imputation method. Hybrid Intell. Syst. Ser. Front Artif. Intell. Appl. 2002, 87, 251–260. [Google Scholar]
- Aydilek, I.B.; Arslan, A. A hybrid method for imputation of missing values using optimized fuzzy c-means with support vector regression and a genetic algorithm. Inf. Sci. 2013, 233, 25–35. [Google Scholar] [CrossRef]
- Pampaka, M.; Hutcheson, G.; Williams, J. Handling missing data: Analysis of a challenging data set using multiple imputation. Int. J. Res. Method Educ. 2016, 39, 19–37. [Google Scholar] [CrossRef]
- Abdella, M.; Marwala, T. The use of genetic algorithms and neural networks to approximate missing data in database. Comput. Inform. 2005, 24, 577–589. [Google Scholar]
- Luengo, J.; García, S.; Herrera, F. On the choice of the best imputation methods for missing values considering three groups of classification methods. Knowl. Inf. Syst. 2012, 32, 77–108. [Google Scholar] [CrossRef]
- Donders, A.R.T.; van der Heijden, G.J.M.G.; Stijnen, T.; Moons, K.G.M. Review: A gentle introduction to imputation of missing values. J. Clin. Epidemiol. 2006, 59, 1087–1091. [Google Scholar] [CrossRef]
- Perkins, N.J.; Cole, S.R.; Harel, O.; Tchetgen Tchetgen, E.J.; Sun, B.; Mitchell, E.M.; Schisterman, E.F. Principled Approaches to Missing Data in Epidemiologic Studies. Am. J. Epidemiol. 2018, 187, 568–575. [Google Scholar] [CrossRef]
- Croiseau, P.; Génin, E.; Cordell, H.J. Dealing with missing data in family-based association studies: A multiple imputation approach. Hum. Hered. 2007, 63, 229–238. [Google Scholar] [CrossRef] [PubMed]
- Mostafa, S.M. Missing data imputation by the aid of features similarities. Int. J. Big Data Manag. 2020, 1, 81–103. [Google Scholar] [CrossRef]
- Iltache, S.; Comparot, C.; Mohammed, M.S.; Charrel, P.J. Using semantic perimeters with ontologies to evaluate the semantic similarity of scientific papers. Informatica 2018, 42, 375–399. [Google Scholar] [CrossRef]
- Yadav, M.L.; Roychoudhury, B. Handling missing values: A study of popular imputation packages in R. Knowl.-Based Syst. 2018, 160, 104–118. [Google Scholar] [CrossRef]
- Farhangfar, A.; Kurgan, L.A.; Pedrycz, W. A Novel Framework for Imputation of Missing Values in Databases. IEEE Trans. Syst. Man Cybern. Part A Syst. Hum. 2007, 37, 692–709. [Google Scholar] [CrossRef]
- Zahin, S.A.; Ahmed, C.F.; Alam, T. An effective method for classification with missing values. Appl. Intell. 2018, 48, 3209–3230. [Google Scholar] [CrossRef]
- Batista, G.E.A.P.A.; Monard, M.C. An analysis of four missing data treatment methods for supervised learning. Appl. Artif. Intell. 2003, 17, 519–533. [Google Scholar] [CrossRef]
- Acuña, E.; Rodriguez, C. The Treatment of Missing Values and its Effect on Classifier Accuracy. In Classification, Clustering, and Data Mining Applications; Springer: Berlin/Heidelberg, Germany, 2004; pp. 639–647. [Google Scholar]
- Li, D.; Deogun, J.; Spaulding, W.; Shuart, B. Towards Missing Data Imputation: A Study of Fuzzy K-means Clustering Method. In Proceedings of the International Conference on Rough Sets and Current Trends in Computing, Madrid, Spain, 9–13 July 2004; Springer: Berlin/Heidelberg, Germany, 2004; Volume 3066, pp. 573–579. [Google Scholar]
- Feng, H.; Chen, G.; Yin, C.; Yang, B.; Chen, Y. A SVM regression based approach to filling in missing values. In Proceedings of the International Conference on Knowledge-Based and Intelligent Information and Engineering Systems, Melbourne, Australia, 14–16 September 2005; Springer: Berlin/Heidelberg, Germany, 2005; Volume 3683, pp. 581–587. [Google Scholar] [CrossRef]
- Choudhury, S.J.; Pal, N.R. Imputation of missing data with neural networks for classification. Knowl.-Based Syst. 2019, 182. [Google Scholar] [CrossRef]
- Muñoz, J.F.; Rueda, M. New imputation methods for missing data using quantiles. J. Comput. Appl. Math. 2009, 232, 305–317. [Google Scholar] [CrossRef]
- Twala, B.; Jones, M.C.; Hand, D.J. Good methods for coping with missing data in decision trees. Pattern Recognit. Lett. 2008, 29, 950–956. [Google Scholar] [CrossRef]
- Varoquaux, G.; Buitinck, L.; Louppe, G.; Grisel, O.; Pedregosa, F.; Mueller, A. Scikit-learn. J. Mach. Learn. Res. 2011, 12, 2825–2830. [Google Scholar] [CrossRef]
- Chandrashekar, G.; Sahin, F. A survey on feature selection methods. Comput. Electr. Eng. 2014, 40, 16–28. [Google Scholar] [CrossRef]
- Van Buuren, S.; Groothuis-Oudshoorn, K.; Robitzsch, A.; Vink, G.; Doove, L.; Jolani, S.; Schouten, R.; Gaffert, P.; Meinfelder, F.; Gray, B. MICE: Multivariate Imputation by Chained Equations. 2019. Available online: https://cran.rproject.org/web/packages/mice/ (accessed on 15 March 2019).
- Efron, B.; Hastie, T.; Iain, J.; Robert, T. Diabetes Data. 2004. Available online: https://www4.stat.ncsu.edu/~boos/var.select/diabetes.html (accessed on 1 June 2019).
- Acharya, M.S. Graduate Admissions-1-6-2019. Available online: https://www.kaggle.com/mohansacharya/graduate-admissions (accessed on 1 June 2019).
- Stephen, B. Profit Estimation of Companies. Available online: https://github.com/boosuro/profit_estimation_of_companies (accessed on 8 August 2019).
- Kartik, P. Red & White Wine Dataset. Available online: https://www.kaggle.com/numberswithkartik/red-white-wine-dataset (accessed on 11 February 2019).
- Cam, N. California Housing Prices. Available online: https://www.kaggle.com/camnugent/california-housing-prices (accessed on 6 July 2019).
- Magrawal, S. Diamonds. Available online: https://www.kaggle.com/shivam2503/diamonds (accessed on 30 August 2019).
- Cattral, R.; Oppacher, F. Poker Hand Dataset. Available online: https://archive.ics.uci.edu/ml/datasets/Poker+Hand (accessed on 24 November 2019).
- Holmes, G.; Pfahringer, B.; van Rijn, J.; Vanschoren, J. BNG_heart_statlog. Available online: https://www.openml.org/d/267 (accessed on 11 September 2019).
- Kearney, J.; Barkat, S. Autoimpute. Available online: https://autoimpute.readthedocs.io/en/latest/ (accessed on 1 January 2020).
- Law, E. Impyute. Available online: https://impyute.readthedocs.io/en/latest/ (accessed on 8 August 2019).
- Chai, T.; Draxler, R.R. Root mean square error (RMSE) or mean absolute error (MAE)? -Arguments against avoiding RMSE in the literature. Geosci. Model Dev. 2014, 7, 1247–1250. [Google Scholar] [CrossRef]








| Dataset name | #Instances | #Features | Missingness Mechanism | ||
|---|---|---|---|---|---|
| MAR | MCAR | MNAR | |||
| Diabetes [27] | 442 | 11 | √ | √ | √ |
| graduate admissions [28] | 500 | 8 | √ | √ | √ |
| profit estimation of companies [29] | 1000 | 6 | √ | √ | √ |
| red & white wine [30] | 4898 | 12 | √ | √ | √ |
| California [31] | 20,640 | 9 | √ | √ | √ |
| Diamonds [32] | 53,940 | 10 | √ | √ | √ |
| Poker Hand [33] | 1,025,010 | 11 | √ | √ | √ |
| BNG_heart_statlog [34] | 1,000,000 | 14 | √ | √ | √ |
| Method Name | Function Name | Package | Description |
|---|---|---|---|
| MICE [9,26] | mice | impyute | implements the multivariate imputation by chained equations algorithm. |
| least squares (LS) [35] | SingleImputer | autoimpute | produces predictions using the least squares methodology. |
| norm [35] | SingleImputer | autoimpute | creates a normal distribution using the sample variance and mean of the detected data. |
| stochastic (ST) [35] | SingleImputer | autoimpute | samples from the regression’s error distribution and adds the random draw to the prediction. |
| Fast KNN (FKNN) [36] | fast_knn | impyute | uses K-Dimensional tree to find k nearest neighbor and imputes using the weighted average of them. |
| EMI [36] | em | impyute | imputes using Expectation-Maximization-Imputation. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
M. Mostafa, S.; S. Eladimy, A.; Hamad, S.; Amano, H. CBRL and CBRC: Novel Algorithms for Improving Missing Value Imputation Accuracy Based on Bayesian Ridge Regression. Symmetry 2020, 12, 1594. https://doi.org/10.3390/sym12101594
M. Mostafa S, S. Eladimy A, Hamad S, Amano H. CBRL and CBRC: Novel Algorithms for Improving Missing Value Imputation Accuracy Based on Bayesian Ridge Regression. Symmetry. 2020; 12(10):1594. https://doi.org/10.3390/sym12101594
Chicago/Turabian StyleM. Mostafa, Samih, Abdelrahman S. Eladimy, Safwat Hamad, and Hirofumi Amano. 2020. "CBRL and CBRC: Novel Algorithms for Improving Missing Value Imputation Accuracy Based on Bayesian Ridge Regression" Symmetry 12, no. 10: 1594. https://doi.org/10.3390/sym12101594
APA StyleM. Mostafa, S., S. Eladimy, A., Hamad, S., & Amano, H. (2020). CBRL and CBRC: Novel Algorithms for Improving Missing Value Imputation Accuracy Based on Bayesian Ridge Regression. Symmetry, 12(10), 1594. https://doi.org/10.3390/sym12101594






