Exploring the Important Attributes of Human Immunodeficiency Virus and Generating Decision Rules
Abstract
1. Introduction
2. Related Works
2.1. HIV/AIDS Disease
2.2. RFM Model
2.3. Attribute Selection
2.4. Rough Set Theory
2.5. Related Classifier
2.5.1. Decision Trees
2.5.2. Support Vector Machine
2.5.3. Random Forest
2.5.4. Radial Basis Function Networks
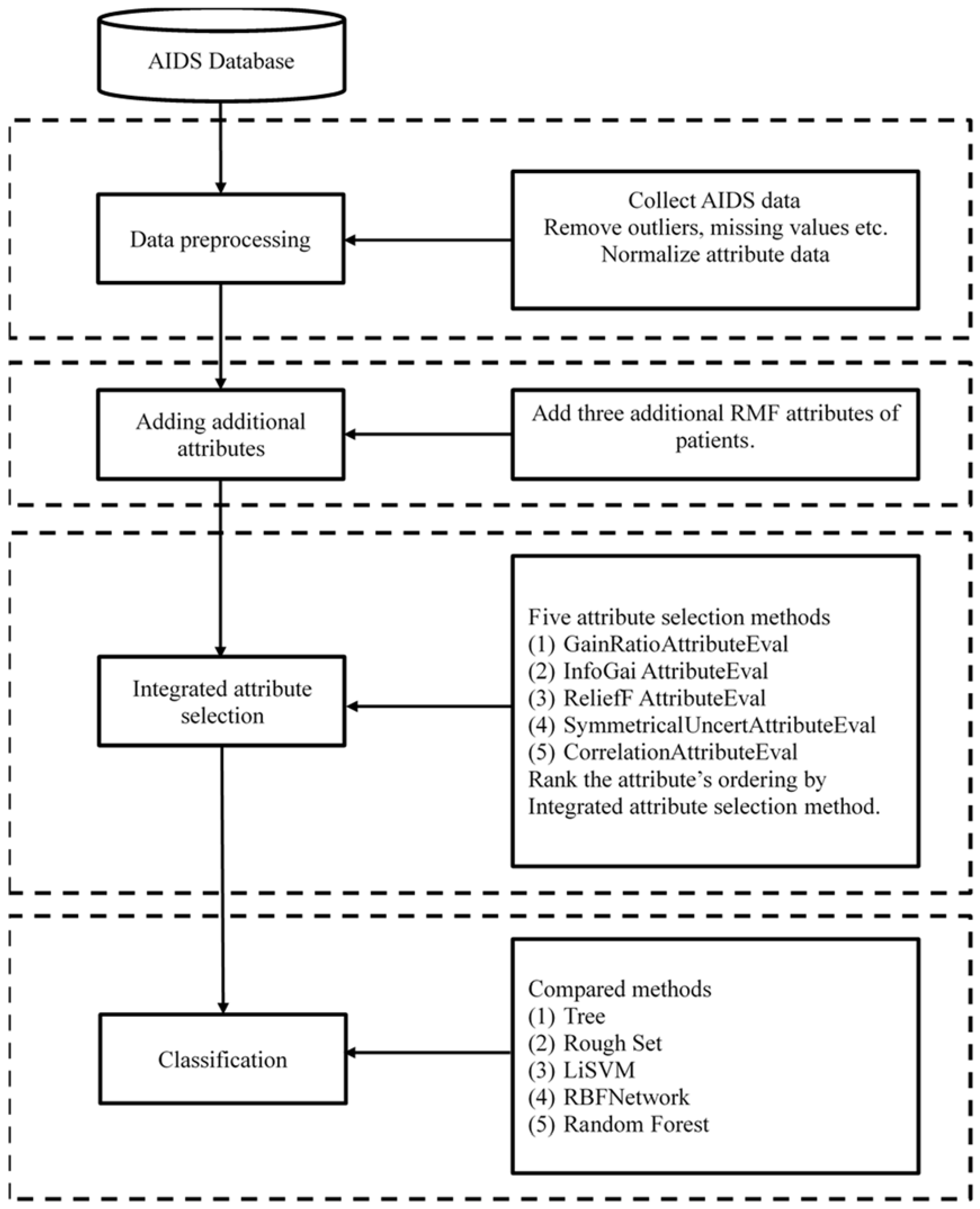
3. Proposed Method
- (1)
- R (the last physician visit): The first medical record in the AIDS data set was recorded on 1 January 2005, and the last on 19 January 2012. This study divided the time interval (seven years) into five groups that were represented as code numbers 1 to 5, respectively. For example, the first time-group was ~2005/01/01–2006/05/30. If the patient’s medical treatment time fell within this time, it was identified as code 1. The second time-group was ~2006/05/31–2007/10/27, which was designated as code 2, and so on.
- (2)
- F (the frequency of medical visits): The definition of F was the number of patients’ physician visits from 1 January 2005 to 19 January 2012. In the AIDS data set, the highest frequency of patients’ physician visit was 10 and the least was 1. This study divided the visit frequency into five groups. If the number of medical visits was one or two, it was assigned code 1. Code 2 was designated for three or four visits, code 3 was for five or six visits, code 4 was for seven or eight visits, and code 5 was for nine or 10 visits.
- (3)
- M (whether or not the patient continued to take their medicine): This attribute was divided into two groups. Code 1 described a patient stopping their HAART medication. If the patient interrupted or stopped their HAART treatment, the record (patient) was assigned to this group. Code 2 was assigned to patients who continuously took HAART.
4. Results
4.1. Experimental Data and Attribute Selection
4.2. Classification and Key Attributes
4.3. Decision Rule
5. Findings and Discussion
5.1. CD4+ Levels and Continued HAART Treatment
5.2. Lower Willingness to Continue Taking Antiretroviral Medications
5.3. The Relationship between Drug Abuse and AIDS
5.4. RFM Attributes and CD4+
5.5. Key Attributes
6. Conclusions
- (1)
- The addition of other attributes or the use of other models in HIV/AIDS;
- (2)
- The development of feasible mobile information systems, such as the automated evaluation of HIV+ patients to aid clinical settings in the future;
- (3)
- The discretization of HIV infection stages to more classes.
Author Contributions
Funding
Conflicts of Interest
References
- De, V.G.; Wulfsohn, M.; Fischl, M.A.; Tsiatis, A. Modeling the relationship between survival and CD4 lymphocytes in patients with AIDS and AIDS-related complex. J. Acquir. Immune Defic. Syndr. 1993, 6, 359–365. [Google Scholar]
- Montarroyos, U.R.; Miranda-Filho, D.B.; César, C.C.; Souza, W.V.; Lacerda, H.R.; Albuquerque Mde, F.; Aguiar, M.F.; Ximenes, R.A. Factors related to changes in CD4+ T-cell counts over time in patients living with HIV/AIDS: A multilevel analysis. PLoS ONE 2014, 9, e84276. [Google Scholar] [CrossRef] [PubMed]
- Langevin, S.; Pichon, M.; Smith, E.; Morrison, J.; Bent, Z.; Green, R.; Barker, K.; Solberg, O.; Gillet, Y.; Javouhey, E.; et al. Early nasopharyngeal microbial signature associated with severe influenza in children: A retrospective pilot study. J. Gen. Virol. 2017, 98, 2425–2437. [Google Scholar] [CrossRef] [PubMed]
- Oliveira, A.; Faria, B.M.; Gaio, R.A.; Reis, L. Data mining in HIV-AIDS surveillance system: Application to portuguese data. J. Med Syst. 2017, 41, 51. [Google Scholar] [CrossRef] [PubMed]
- Ko, G.M.; Reddy, A.S.; Kumar, S.; Bailey, B.A.; Garg, R. Computational analysis of HIV-1 protease protein binding pockets. J. Chem. Inf. Model. 2010, 50, 1759–1771. [Google Scholar] [CrossRef]
- Prabhu, P.; Duraiswamy, K. Feature selection for HIV database using rough system. In Proceedings of the 2010 Second International conference on Computing, Communication and Networking Technologies, Karur, India, 29–31 July 2010; pp. 1–6. [Google Scholar]
- Haile Mariam, T. Application of Data Mining Techniques for Predicting CD4 Status of Patients on ART in Jimma and Bonga Hospitals, Ethiopia. J. Health Med. Inform. 2015, 6, 208. [Google Scholar] [CrossRef]
- UNAIDS (The Joint United Nations Programme on HIV/AIDS). 2018. Available online: http://www.unaids.org/sites/default/files/media_asset/unaidsdata-2018_en.pdf#page=4&zoom=auto,-62,843 (accessed on 20 December 2018).
- Gallo, R.C.; Montagnier, L. The discovery of HIV as the cause of AIDS. N. Engl. J. Med. 2003, 349, 2283–2285. [Google Scholar] [CrossRef]
- Ford, N.; Meintjes, G.; Vitoria, M.; Greene, G.; Chiller, T. The evolving role of CD4 cell counts in HIV care. Curr. Opin. HIV AIDS 2017, 12, 123–128. [Google Scholar] [CrossRef]
- Pezzotti, P.; Napoli, P.A.; Acciai, S.; Boros, S.; Urciuoli, R.; Lazzeri, V.; Rezza, G.; Group, T.A.S. Increasing survival time after AIDS in Italy: The role of new combination antiretroviral therapies. AIDS 1999, 13, 249–255. [Google Scholar] [CrossRef]
- Hogg, R.S.; O’Shaughnessy, M.V.; Gataric, N.; Yip, B.; Craib, K.; Schechter, M.T.; Montaner, J.S. Decline in deaths from AIDS due to new antiretrovirals. Lancet 1997, 349, 1294. [Google Scholar] [CrossRef]
- WHO. HIV/AIDS. 15 November 2019. Available online: https://www.who.int/news-room/fact-sheets/detail/hiv-aids?fbclid=IwAR1y8FK_whe2s7K7SGGraydiwoMNjl6WGvnN4c9JYYGpRi2-50wPp7JLkew (accessed on 20 December 2019).
- Bangsberg, D.R.; Perry, S.; Charlebois, E.D.; Clark, R.A.; Roberston, M.; Zolopa, A.R.; Moss, A. Non-adherence to highly active antiretroviral therapy predicts progression to AIDS. AIDS 2001, 15, 1181–1183. [Google Scholar] [CrossRef] [PubMed]
- Tsiptsis, K.; Chorianopoulos, A. Data Mining Techniques in CRM: Inside Customer Segmentation; Wiley: Hoboken, NJ, USA, 2009. [Google Scholar]
- Cai, J.; Luo, J.; Wang, S.; Yang, S. Feature selection in machine learning: A new perspective. Neurocomputing 2018, 300, 70–79. [Google Scholar] [CrossRef]
- Tang, J.; Alelyani, S.; Liu, H. Feature selection for classification: A review. Data Classif. Algorithms Appl. 2014, 37, 37–64. [Google Scholar]
- Liu, H.; Yu, L. Toward integrating feature selection algorithms for classification and clustering. IEEE Trans. Knowl. Data Eng. 2005, 17, 491–502. [Google Scholar]
- Tuo, Q.; Zhao, H.; Hu, Q. Hierarchical feature selection with subtree based graph regularization. Knowl. Based Syst. 2019, 163, 996–1008. [Google Scholar] [CrossRef]
- Guyon, I.; Elisseeff, A. An introduction to variable and feature selection. J. Mach. Learn. Res. 2003, 3, 1157–1182. [Google Scholar]
- Chandrashekar, G.; Sahin, F. A survey on feature selection methods. Comput. Electr. Eng. 2014, 40, 16–28. [Google Scholar] [CrossRef]
- Pawlak, Z. Rough sets. Int. J. Comput. Inf. Sci. 1982, 11, 341–356. [Google Scholar] [CrossRef]
- Chen, H.; Li, T.; Fan, X.; Luo, C. Feature selection for imbalanced data based on neighborhood rough sets. Inf. Sci. 2019, 483, 1–20. [Google Scholar] [CrossRef]
- Sheeja, T.; Kuriakose, A.S. A novel feature selection method using fuzzy rough sets. Comput. Ind. 2018, 97, 111–121. [Google Scholar] [CrossRef]
- Kumar, S.S.; Inbarani, H.H. Optimistic multi-granulation rough set based classification for medical diagnosis. Procedia Comput. Sci. 2015, 47, 374–382. [Google Scholar] [CrossRef]
- Yıldırım, E.G.; Karahoca, A.; Uçar, T. Dosage planning for diabetes patients using data mining methods. Procedia Comput. Sci. 2011, 3, 1374–1380. [Google Scholar] [CrossRef][Green Version]
- Mahapatra, S.; Mahapatra, S. Attribute selection in marketing: A rough set approach. IIMB Manag. Rev. 2010, 22, 16–24. [Google Scholar] [CrossRef]
- Suraj, Z. An introduction to rough set theory and its applications. In Proceedings of the ICENCO’2004, Cairo, Egypt, 27–30 December 2004. [Google Scholar]
- Quinlan, J.R. Induction of decision trees. Mach. Learn. 1986, 1, 81–106. [Google Scholar] [CrossRef]
- Tayefi, M.; Esmaeili, H.; Karimian, M.S.; Zadeh, A.A.; Ebrahimi, M.; Safarian, M.; Nematy, M.; Parizadeh, S.M.R.; Ferns, G.A.; Ghayour-Mobarhan, M. The application of a decision tree to establish the parameters associated with hypertension. Comput. Methods Programs Biomed. 2017, 139, 83–91. [Google Scholar] [CrossRef]
- Ramezankhani, A.; Pournik, O.; Shahrabi, J.; Khalili, D.; Azizi, F.; Hadaegh, F. Applying decision tree for identification of a low risk population for type 2 diabetes. Tehran Lipid and Glucose Study. Diabetes Res. Clin. Pract. 2014, 105, 391–398. [Google Scholar] [CrossRef]
- Cortes, C.; Vapnik, V. Support-vector networks. Mach. Learn. 1995, 20, 273–297. [Google Scholar] [CrossRef]
- Chang, C.-C.; Lin, C.-J. LIBSVM: A library for support vector machines. ACM Trans. Intell. Syst. Technol. 2011, 2, 27. [Google Scholar] [CrossRef]
- Ho, T.K. Random decision forests. In Proceedings of the Third International Conference on Document Analysis and Recognition, Montreal, QC, Canada, 14–16 August 1995; pp. 278–282. [Google Scholar]
- Hsieh, C.-H.; Lu, R.-H.; Lee, N.-H.; Chiu, W.-T.; Hsu, M.-H.; Li, Y.-C.J. Novel solutions for an old disease: Diagnosis of acute appendicitis with random forest, support vector machines, and artificial neural networks. Surgery 2011, 149, 87–93. [Google Scholar] [CrossRef]
- Masetic, Z.; Subasi, A. Congestive heart failure detection using random forest classifier. Comput. Methods Programs Biomed. 2016, 130, 54–64. [Google Scholar] [CrossRef]
- Broomhead, D.S.; Lowe, D. Multivariable functional interpolation and adaptive networks. Complex Syst. 1988, 2, 321–355. [Google Scholar]
- Ganapathy, K.; Vaidehi, V.; Chandrasekar, J.B. Optimum steepest descent higher level learning radial basis function network. Expert Syst. Appl. 2015, 42, 8064–8077. [Google Scholar] [CrossRef]
- Brandstetter, P.; Kuchar, M. Sensorless control of variable speed induction motor drive using RBF neural network. J. Appl. Logic 2017, 24, 97–108. [Google Scholar] [CrossRef]
- Anish, C.M.; Majhi, B. Hybrid nonlinear adaptive scheme for stock market prediction using feedback FLANN and factor analysis. J. Korean Stat. Soc. 2016, 45, 64–76. [Google Scholar] [CrossRef]
- Pottmann, M.; Jörgl, H.P. Radial basis function networks for internal model control. Appl. Math. Comput. 1995, 70, 283–298. [Google Scholar] [CrossRef]
- Morlini, I. Radial basis function networks with partially classified data. Ecol. Model. 1999, 120, 109–118. [Google Scholar] [CrossRef]
- Project, T.W. The Body. Available online: http://www.thebody.com/content/58838/understanding-cd4-cells-and-cd4-cell-tests.html?getPage=1#hiv (accessed on 24 February 2019).
- Chen, Y.-S. A comprehensive identification-evidence based alternative for HIV/AIDS treatment with HAART in the healthcare industries. Comput. Methods Programs Biomed. 2016, 131, 111–126. [Google Scholar] [CrossRef]
- Department of Laboratory Medicine, National Taiwan University Hospital. Blood Biological Reference Interval. Available online: https://wwwsp.ntuh.gov.tw/labmed/%E6%AA%A2%E9%A9%97%E7%9B%AE%E9%8C%84/Lists/service/DispForm.aspx?ID=381&Source=https%3A%2F%2Fwwwsp.ntuh.gov.tw%2Flabmed%2F%25e6%25aa%25a2%25e9%25a9%2597%25e7%259b%25ae%25e9%258c%2584%2FPages%2FA.aspx%3FTXT%3DAST%25EF%25BC%2588GOT%25EF%25BC%2589%26button%3D%25E9%2580%2581%25E5%2587%25BA&ContentTypeId=0x01003E4C6DCCAFBEF64BACFA9B88DBDEF416&fbclid=IwAR2GLgtSjEWmkz-YvxxOkawv-yOUUZhVZyKeEUz5vZyPIgSfHoJ6GMVEOJ0 (accessed on 26 December 2019).
- Department of Laboratory Medicine, National Taiwan University Hospital. Clinical Laboratory Manual. Available online: https://wwwsp.ntuh.gov.tw/labmed/%E6%AA%A2%E9%A9%97%E7%9B%AE%E9%8C%84/Lists/service/DispForm.aspx?ID=377&Source=https%3A%2F%2Fwwwsp.ntuh.gov.tw%2Flabmed%2F%25e6%25aa%25a2%25e9%25a9%2597%25e7%259b%25ae%25e9%258c%2584%2FPages%2FA.aspx%3FTXT%3DALT%26button%3D%25E9%2580%2581%25E5%2587%25BA&ContentTypeId=0x01003E4C6DCCAFBEF64BACFA9B88DBDEF416&fbclid=IwAR1rOXhAWFODgGwxCAtWNdN2cKl4JlhNTcHfXR1mLHLeCH91x11O365PhDc (accessed on 26 December 2019).
- Department of Laboratory Medicine, National Taiwan University Hospital. Clinical Laboratory Manual. Available online: https://wwwsp.ntuh.gov.tw/labmed/%E6%AA%A2%E9%A9%97%E7%9B%AE%E9%8C%84/Lists/service/DispForm.aspx?ID=383&Source=https%3A%2F%2Fwwwsp.ntuh.gov.tw%2Flabmed%2F%25e6%25aa%25a2%25e9%25a9%2597%25e7%259b%25ae%25e9%258c%2584%2FPages%2FA.aspx%3FTXT%3DBilirubin-T%26button%3D%25E9%2580%2581%25E5%2587%25BA&ContentTypeId=0x01003E4C6DCCAFBEF64BACFA9B88DBDEF416&fbclid=IwAR3IQubED7Y7etirAdxwCS5WCgxfBtGPKSfu1X6k61i0LxhDv7po6w-opTY (accessed on 26 December 2019).
- Department of Laboratory Medicine, National Taiwan University Hospital. Clinical Laboratory Manual. Available online: https://health.ntuh.gov.tw/health/hrc_v3/DataFiles/kensa.htm (accessed on 26 December 2019).
- Han, J.; Pei, J.; Kamber, M. Data Mining: Concepts and Techniques; Elsevier: Amsterdam, The Netherlands, 2011. [Google Scholar]
- Hall, M.A. Correlation-Based Feature Selection for Machine Learning. Ph.D. Thesis, The University of Waikato, Hamilton, New Zealand, 1999. [Google Scholar]
- Kononenko, I. Estimating attributes: Analysis and extensions of RELIEF. In Proceedings of the 7th European Conference on Machine Learning (ECML’94), Catania, Italy, 6–8 April 1994; pp. 171–182. [Google Scholar]
- Kira, K.; Rendell, L.A. A practical approach to feature selection. In Proceedings of the Ninth International Workshop on Machine Learning, Aberdeen, Scotland, 1–3 July 1992; pp. 249–256. [Google Scholar]
- Forgy, E.W. Cluster analysis of multivariate data: Efficiency versus interpretability of classifications. Biometrics 1965, 21, 768–769. [Google Scholar]
| Attribute | Data Type | Description | #Sample (%) | Test p Value |
|---|---|---|---|---|
| Age | Numeric | Patient age, Min: 17~Max: 77 | 1308 (100%) | |
| PVL | Numeric | Plasma Viral Load, Min: 43~Max: 6641662 | 1308 (100%) | |
| Gender | M | Patient gender, Male | 1207 (92%) | 0.487 |
| F | Patient gender, Female | 101 (8%) | ||
| Cause | A | Cause of HIV infection (high-risk sexuality relationship) | 127 (10%) | 0.001 |
| B | Cause of HIV infection (Injection drug or sharing syringes) | 1181 (90%) | ||
| PLT_C | Y | Platelets, PLT ≥ 150 or PLT < 350 | 897 (69%) | 0.081 |
| N | Platelets, PLT ≥ 350 or PLT < 150 | 411 (31%) | ||
| T-bil_C | Y | Total bilirubin, T-bil ≥ 0.2 or T-bil ≤ 1.5 | 966 (74%) | 0.355 |
| N | Total bilirubin, T-bil > 1.5 or T-bil < 0.2 | 342 (26%) | ||
| AST_C | Y | Aspartate transaminase, AST ≤ 35 | 581 (44%) | 0.149 |
| N | Aspartate transaminase, AST > 35 | 727 (56%) | ||
| ALT_C | Y | Alanine transaminase, ALT ≤ 40 | 693 (53%) | 0.120 |
| N | Alanine transaminase, ALT > 40 | 615 (47%) | ||
| R | 1 | Last physician visit, 2005/01/01~2006/05/30 | 6 (0.5%) | 0.002 |
| 2 | Last physician visit, 2006/05/31~2007/10/27 | 177 (13.5%) | ||
| 3 | Last physician visit, 2007/10/28~2009/03/25 | 359 (27.4%) | ||
| 4 | Last physician visit, 2009/03/26~2010/08/22 | 434 (33.2%) | ||
| 5 | Last physician visit, 2010/08/23~2012/01/19 | 332 (25.4%) | ||
| F | 1 | The frequency of medical visits | 270(21%) | 0.000 |
| 2 | The frequency of medical visits | 378(29%) | ||
| 3 | The frequency of medical visits | 409 (31%) | ||
| 4 | The frequency of medical visits | 232 (18%) | ||
| 5 | The frequency of medical visits | 19 (1%) | ||
| M | 1 | Stop taking the medication | (71%) | 0.000 |
| 2 | Continue taking the medication | (29%) | ||
| CD4_C | A | the CD4+ cell counts < 500 (serious) | (73%) | |
| B | the CD4+ cell counts ≥ 500 (well control) | (27%) |
| Rank | Score | GainRatio | InfoGain | ReliefF | Symmetrical | Correlation |
|---|---|---|---|---|---|---|
| 1 | 11 | M | M | M | M | M |
| 2 | 10 | PVL | R | PLT_C | PVL | R |
| 3 | 9 | Age | PVL | R | R | F |
| 4 | 8 | R | Age | cause | Age | cause |
| 5 | 7 | cause | F | F | cause | Age |
| 6 | 6 | F | cause | T-bil_C | F | PVL |
| 7 | 5 | PLT_C | PLT_C | Age | PLT_C | PLT_C |
| 8 | 4 | ALT_C | ALT_C | Gender | ALT_C | ALT_C |
| 9 | 3 | AST_C | AST_C | AST_C | AST_C | AST_C |
| 10 | 2 | Gender | T-bil_C | ALT_C | T-bil_C | T-bil_C |
| 11 | 1 | T-bil_C | Gender | PVL | Gender | Gender |
| Rank | Attribute Name | Five Selection Methods’ Scores | Four Selection Methods’ Scores |
|---|---|---|---|
| 1 | M | 55 | 44 |
| 2 | R | 46 | 37 |
| 3 | PVL | 36 | 35 |
| 4 | Age | 37 | 32 |
| 5 | Cause | 36 | 28 |
| 6 | F | 35 | 28 |
| 7 | PLT_C | 30 | 20 |
| 8 | ALT_C | 18 | 16 |
| 9 | AST_C | 15 | 12 |
| 10 | T-bil_C | 13 | 7 |
| 11 | Gender | 9 | 5 |
| Classifier | Parameter |
|---|---|
| TREE | Confidence Factor = 0.25 |
| Rough set | Method: LEM2 algorithm Cover parameter = 0.9 |
| LibSVM | Kernel: radial basis function Cost = 1.0 Epsilon = 0.001 |
| RBF Network | clustering seed = 1 MinStdDev = 0.1 |
| Random Forest | Debug: false Maximum depth = 0 |
| Attributes | TREE | Rough Set | LibSVM | RBF Network | Random Forest |
|---|---|---|---|---|---|
| Original attribute set (no RFM) | 72.14 (1.86) | 77.42 (2.16) | 73.39 (0.31) | 73.18 (0.32) | 74.28 (1.60) |
| New attribute set (adding RFM) | 72.42 (1.73) | 79.14 (0.33) | 73.39 (0.32) | 73.13 (0.67) | 76.44 (1.58) |
| Delete Gender & T-bil_C & ALT_C | 73.08 (1.76) | 77.84 (1.06) | 73.38 (0.33) | 72.97 (0.76) | 76.12 (1.62) |
| Delete Gender & T-bil_C & AST_C & ALT_C | 73.20 (1.59) | 80.48 (1.81) | 73.40 (0.32) | 72.80 (1.03) | 76.12 (1.61) |
| Rank | Match | Decision Rules |
|---|---|---|
| Class A Rules | ||
| 1 | 52 | (cause=B)&(PLT_C=N)&(M=2)&(R=4)=>(CD4_C={A[52]}) |
| 2 | 36 | (cause=B)&(PLT_C=Y)&(M=2)&(R=5)=>(CD4_C={A[36]}) |
| 3 | 25 | (cause=B)&(PLT_C=N)&(M=2)&(R=5)=>(CD4_C={A[25]}) |
| Class B Rules | ||
| 1 | 6 | (cause=B)&(M=1)&(PLT_C=Y)&(R=3)&(F=4)&(Age="(44.5,45.5)")=>(CD4_C={B[6]}) |
| 2 | 6 | (cause=B)&(M=1)&(PLT_C=Y)&(R=4)&(F=3)&(Age="(24.5,25.5)")=>(CD4_C={B[6]}) |
| 3 | 4 | (cause=B)&(M=1)&(PLT_C=N)&(F=2)&(Age="(42.5,43.5)")=>(CD4_C={B[4]}) |
| Cluster 1 | Cluster 2 | Cluster 3 | Cluster 4 | Cluster 5 | |
|---|---|---|---|---|---|
| R | 2 | 4 | 2 | 4 | 5 |
| F | 4 | 1 | 2 | 3 | 4 |
| M | 1 | 1 | 1 | 1 | 1 |
| Total records | 132 | 412 | 86 | 613 | 65 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cheng, C.-H.; Wang, Y.-C. Exploring the Important Attributes of Human Immunodeficiency Virus and Generating Decision Rules. Symmetry 2020, 12, 67. https://doi.org/10.3390/sym12010067
Cheng C-H, Wang Y-C. Exploring the Important Attributes of Human Immunodeficiency Virus and Generating Decision Rules. Symmetry. 2020; 12(1):67. https://doi.org/10.3390/sym12010067
Chicago/Turabian StyleCheng, Ching-Hsue, and Yun-Chun Wang. 2020. "Exploring the Important Attributes of Human Immunodeficiency Virus and Generating Decision Rules" Symmetry 12, no. 1: 67. https://doi.org/10.3390/sym12010067
APA StyleCheng, C.-H., & Wang, Y.-C. (2020). Exploring the Important Attributes of Human Immunodeficiency Virus and Generating Decision Rules. Symmetry, 12(1), 67. https://doi.org/10.3390/sym12010067





