3. Discussion
By virtue of the rapid advances in molecular biology, more and more information hidden in the genes of various organisms have been deciphered. The completion of the Human Genomic Project showed that the blueprint of a person can be determined. It has greatly impacted the fields of medicine, biotechnology, and life sciences. Important information closely related to public social life have come from the correct reading of the primary structure of DNA. For example, DNA profiling became a key piece of evidence in courts, the production and dispute over genetically modified organisms (GMO), the development of gene therapies, etc.
Dichotomy tells us that the double helix has its side effects or defects. Many students and experts were unintentionally or imperceptibly misguided by the double helix. Hence, the side effect of the double helix on the scientists’ psychology is probably indescribable and impalpable. Their implicit bias is hard to cure. Accompanying the great success of DNA sequencing and many amazing advances in molecular biology, overpraise and dazzling glory pushed the double helix model to irrational heights. This caused the public, as well as many scientists and journal editors, to lose the ability to question and doubt the double helix. This deterred Robert Chambers from further investigating his unexpected finding and prevented T. T. Wu, among other scientists to get grant support [
32,
33]. Perhaps, it also hindered many other individual scholars to produce or publish unorthodox ideas or findings.
In contrast to the primary structure of DNA, the attention on the secondary structure of DNA is pitiful. Most people assumed that in nature, the DNA structure is in the B-DNA form that had been convincingly proven using X-ray crystallography. It became a doctrine written in all textbooks. As a rare supplement (see
Supplementary Materials), Z-DNA was also proven by X-ray crystallography. This dote sign had better to be changed as an exclamation mark.
Nobody doubts the result obtained from X–ray crystallography! The problem is that all X-ray crystallography evidence was obtained from short fragments of DNA. It is plausible that extrapolating the correct result from short fragments to long DNA is incorrect. Just as someone focusing on a leaf may not see the tree and even less the forest. As a microscopic method, X-ray crystallography is not suitable to determine the molecular structure of very long DNA.
Our investigation uses the mathematical rule, linking number Lk is always equivalent to the sum of total twist of the double helix Tw and the writhing number Wr, i.e., Lk = Tw + Wr, to analyze and evaluate the experimental phenomenon observed from covalently closed circular DNA. It is a macroscopic method. The conclusion derived from experiment reflects the property of the whole plasmid molecule. As shown above, all specially designed experimental results and many facts obtained from various laboratories are consistent with each other. The evidence is self-consistent. This constitutes a chain of evidence sufficient to deny one of the main claims of the double helix, i.e., the two DNA strands are always winding right-handedly or plectonemically.
Similar to a court debate, all true evidence has to be put on the table for careful analysis. While judging the suspected secondary structure of DNA, all evidence collected from both microscopic and macroscopic methods should be fairly considered with no bias.
Different from the idea of God or ghosts, the structure of DNA is supposed to be an objective reality and a recognizable entity. No matter from when, where, or whom a correct DNA structure originates, it should be supported by all evidence that is reasonable and self-consistent.
What really is DNA? In the eyes of human beings, pure DNA is just some kind of white stuff or transparent fiber. Whereas, in the minds of different generations of people, guided by the most knowledgeable scholars at that time, DNA differs greatly. The Watson-Crick double helix model proposed on insufficient evidence in 1953 was reasonable and admirable. The history of DNA repeatedly tells us that our knowledge has to be changed according to the available evidence. The advancement of molecular biology is so fast that it pushes us to update incorrect scientific knowledge or change inappropriate conceptions.
Scientific knowledge is non-rivalrous and non-excludable, whereas different claims or hypotheses may be rivalrous and exclusive. All human knowledge is partial; nobody can reach the final truth. There is probably no standard answer to the question of “what is DNA”. Based on available evidence, a provisional answer closer to the truth is appropriate for the moment. We can see something further than before just because we are standing on the shoulders of many great scientists. Nobody complains that there are defects or incorrect information in the canonical double helix model since knowledge comes from evidence and collecting evidence needs time, chance, and vision.
Presently, all evidence shows that native DNA under physiological conditions is dynamic, polymorphous, and ambidextrous, rather than a monotonous plectonemic right-handed double helix.
The plectonemic DNA model seems to go against Occam’s razor, i.e., the simplest solution tends to be the correct one. However, as Crick pointed out, implementing Occam’s razor in biology can be very dangerous [
5]. Unlike cellulose or tendons, DNA is the critical macromolecule of all living cells responsible for various important biological functions; its structure should be very stable to preserve the fidelity of its genetic code and flexible for the convenience of code recognition. It is probably the reasonable choice of nature after billions years of evolution, although the exact reason is unclear.
In the eyes of future historians, ambidextrous DNA is just a slight modification of the double helix, i.e., from plectonemic winding to ambidextrous winding. The rational core of the double helix is unchanged, only the incorrect part about the winding of the two strands was amended. However, it is a big leap forward in the conception of the secondary structure of DNA and consequently has a big impact on DNA topology.
The most easily noted difference between two DNA models is on the understanding of supercoiling.
More than 50 years ago, while an extrachromosomal DNA, the polyoma virus DNA was studied in the laboratories of Dulbecco and Vinograd. They found this DNA is circular and resistant to heat or formamide denaturation. The purified DNA contains three components that were named DNA I (supercoiled DNA), DNA II (nicked DNA), and DNA III (linear DNA) afterwards. Vinograd and his colleagues realized that DNA I, normally found under experimental conditions, is a negatively supercoiled DNA; they also assumed it is due to the number of winding turns between two strands less than that of its relaxed counterparts in B-DNA form [
34].
A reasonable deduction is that in positively supercoiled DNA, the two strands should be over-wound. The question is how many water molecules can be squeezed out in a positively supercoiled plasmid under physiological conditions. Although B-DNA can be dehydrated in air and turned into more compact A-DNA, it does not mean the two strands can freely tighten with no limits in aqueous solution because the atoms are incompressible and many water molecules are tightly hydrogen-bonded with DNA [
35]. The highly positively supercoiled plasmid found by D. Lockshon and D.R. Morris is unexpected [
36]. It triggered a question: can a hyper-positively supercoiled plasmid be made as that of a hyper-negatively supercoiled plasmid [
37,
38,
39]? Nobody knows how the two strands are over-wound in the positively supercoiled DNA if the DNA is in the double helix model.
However, according to the ambidextrous DNA model, the linking number of a relaxed plasmid is zero. The linking numbers of positively or negatively supercoiled plasmids must be much higher than that of their relaxed counterparts. As shown in
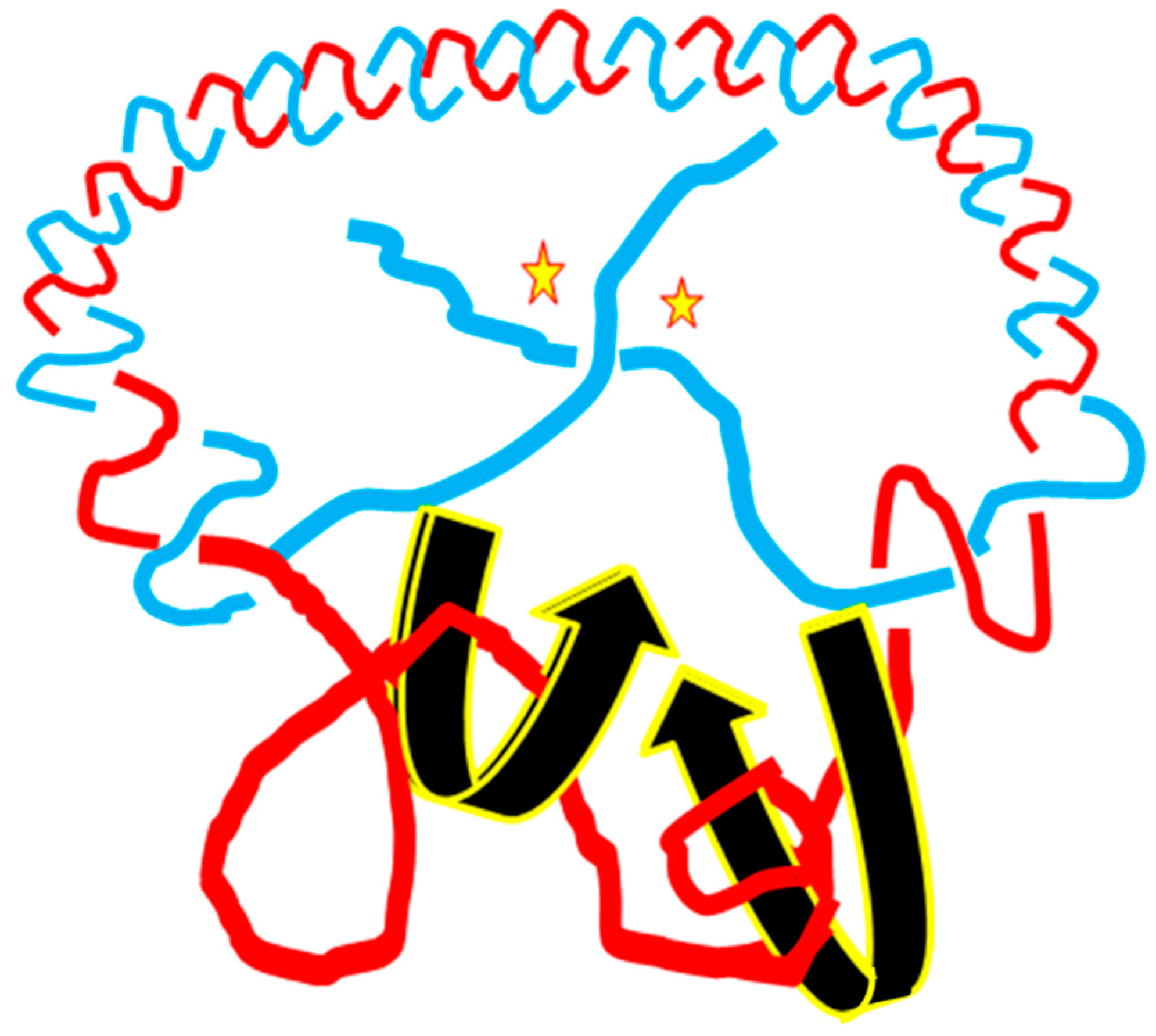
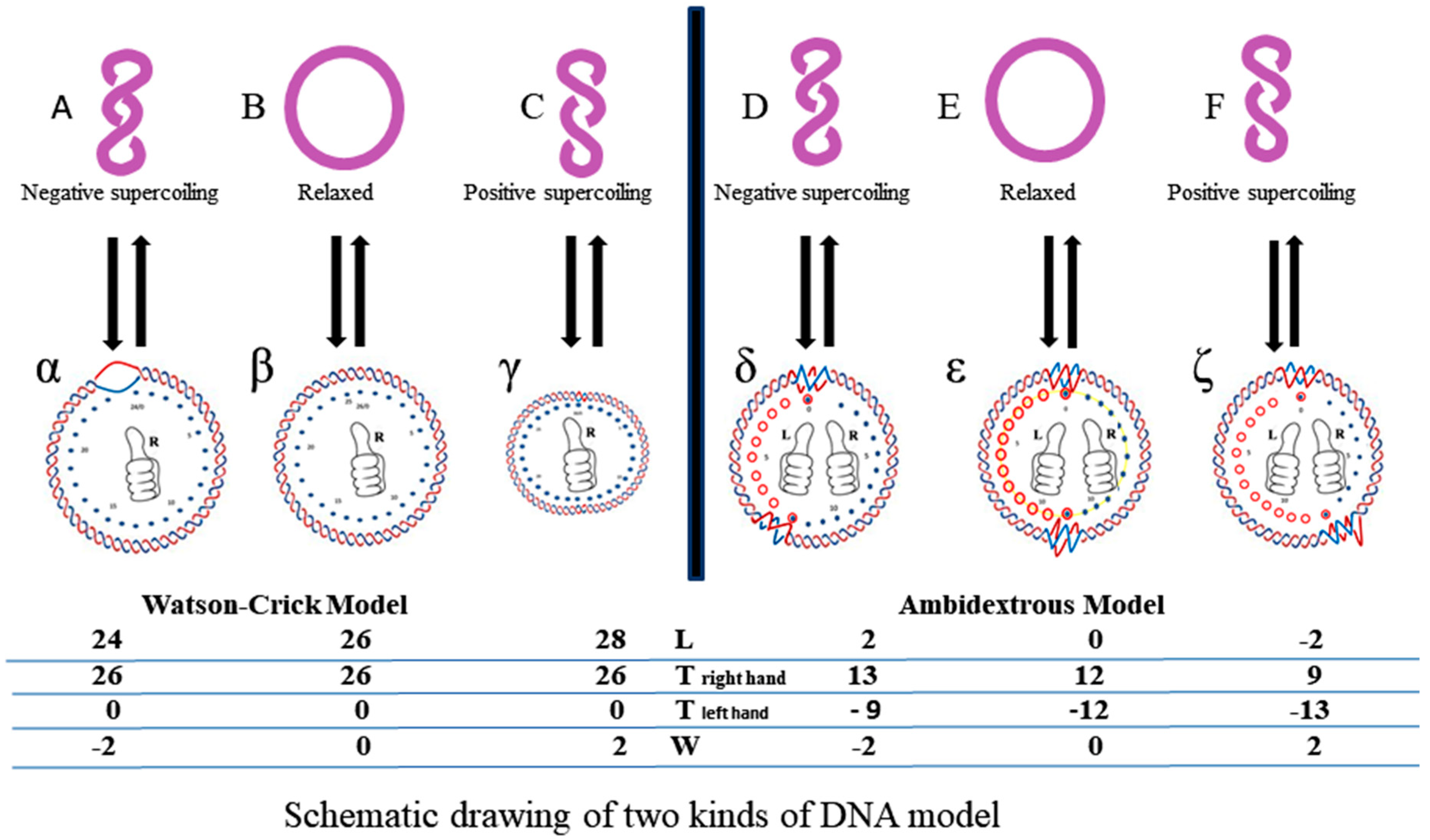
Figure 10, schematic drawings of the plasmid in two DNA models are put on a two-dimensional plane to represent the big difference between them.
In this simplified drawing of a 260-bp circular DNA, all forms of DNA are designed with 10 bp per turn. In the pictures, A, B, C, D, E, F are the general profiles of the plasmid; α, β, γ, δ, ε, ζ are the detailed images of six pairs of fully expanded circular DNAs projected onto a two-dimensional plane. The two complementary circular strands are always hydrogen bonded except there is no hydrogen bond in the loop region in
Figure 10A.
The finding of a zero-linking-number topoisomer indicated that the number of the two opposite-winding turns of that molecule should be the same, i.e., Tright-handed = Tleft-handed. The cancellation of the equal amount of opposite winding strands lead to the non-linkage of the two ssc DNAs. i.e., Lk = 0. A reasonable deduction is that in a negatively supercoiled plasmid, the turns of the right-handed DNA should be more than that of the left-handed DNA, i.e., Tright-handed > Tleft-handed. Likewise, in a positively supercoiled plasmid, the case is reversed, i.e., Tright-handed < Tleft-handed.
A second suggestion from the finding of a zero-linking-number topoisomer is that the left-handed DNA is a routine DNA, as common as the right-handed DNA. Left-handed DNA is no more limited to Z-DNA, which has a relatively strict sequence requirement of alternative purine and pyrimidine. An independent investigation on pBR322 pointed out that the helical repeats of left-handed DNA is 12 base pairs per turn, which matches the data of Z-DNA obtained from X-ray crystallography [
40]. However, the two strands of left-handed DNA are not more tightly tangled with each other, which is not consistent with the expectation of the canonical double helix model. Further investigation may show whether it is coincidental. In short, it is safe to say that Z-DNA is just a member of the left-handed family.
These deductions violated an old concept in DNA topology. It seems a new definition has to be set up to substitute the old one. For example, according to the Watson–Crick model, in a plasmid with N base pairs, DNA is supposed to have 10 base pairs per turn and the linking number of a relaxed plasmid should be a big integer, i.e., L0 ≈ N/10 >> 0. The superhelical density σ is defined as the linking number difference between supercoiled DNA (L) and relaxed DNA (L0) divided by the linking number of relaxed DNA, i.e., σ = (L − L0)/L0. However, it would be unreasonable to put zero as a denominator to indicate the superhelical property of the plasmid if the linking number of the relaxed DNA is zero.
A superhelical index, Si, was proposed as a convenient compromise to indicate the superhelical property of a circular DNA [
24]. Si is related to only two experimentally measurable parameters: the writhing number W
r and the base pair number N of the plasmid, i.e., Si = W
r/(N/10.4). Si can be used as an index to indicate and compare the supercoiling of different plasmid molecules in any DNA model. The reported σ value can still be used as the Si value.
Some readers may ask: if the plasmid DNA is actually following the ambidextrous model, what about linear DNA?
From the experiment of Depew and Wang [
41], it is understandable that before and after the ligation, the winding state of the two strands is unchanged. Hence the two strands of linear DNA should always be winding ambidextrously. This statement may cause great confusion or worry. A possible explanation is that the tertiary structure of DNA is dynamic and polymorphous. It can adopt its most probable shape under the provided conditions.
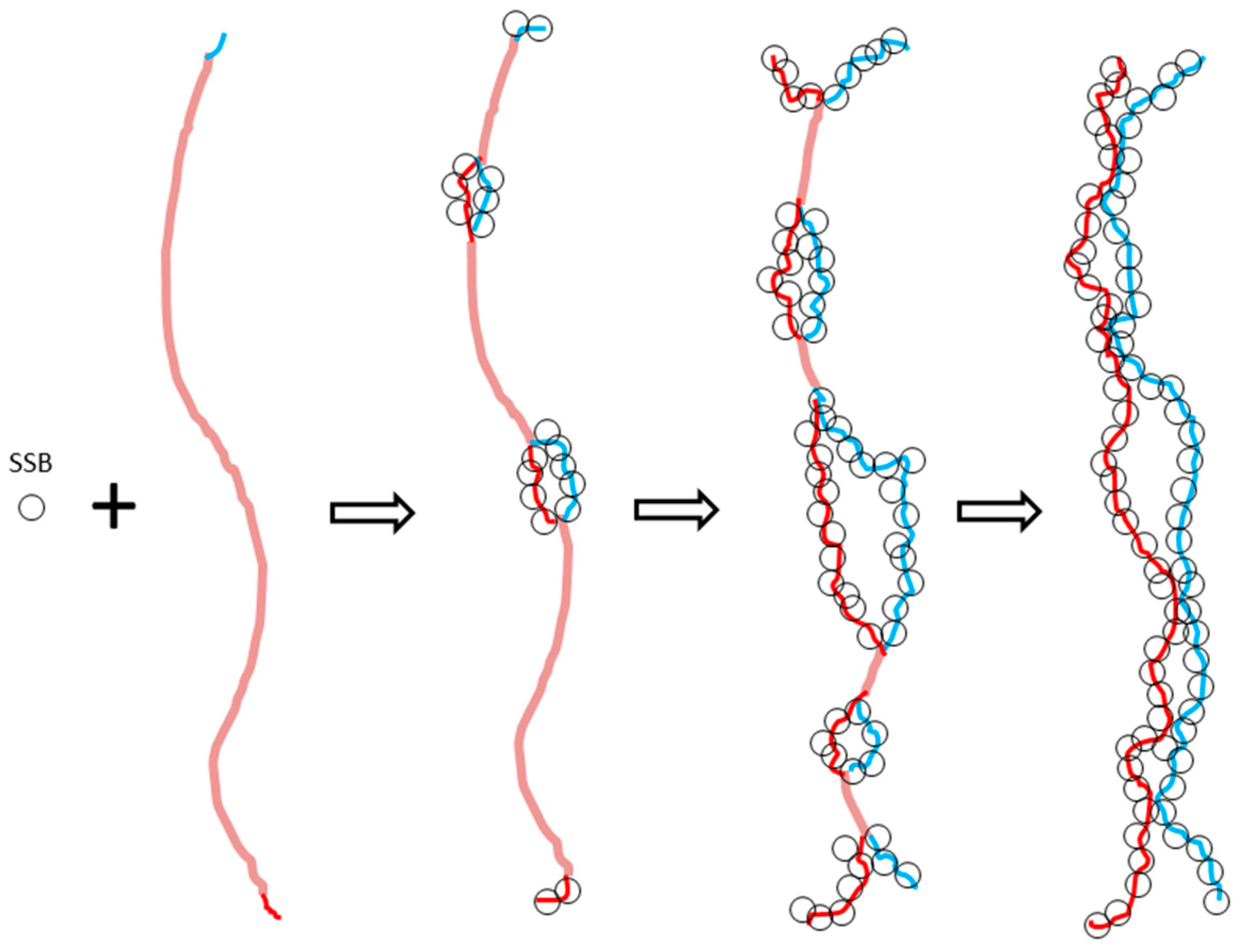
The aim of the proposed ambidextrous DNA model is to try to solve the untangling problem in DNA replication. Evidently, if the two strands of circular E. coli chromosomal DNA are not always tightly tangled with each other, their quick separation would have no topological difficulty. The puzzling problem regarding the quick unwinding of DNA disappears radically.
It is probable that the new DNA model can also help explain the mechanism of CRISPR (clustered regularly interspaced short palindromic repeats). During the long evolution process, bacteria developed a protection against invading viral DNA. CRISPR can be applied for precise and efficient genome editing [
42]. The mechanism of it is rather complicated. A critical step is that bacteria’s gRNA can easily and accurately find its target on the virus and tightly bind to it for its helper enzyme (Cas9) to snip the virus gene. The problem is how the gRNA, about 20 nucleotides long, can find the correct sequence hidden inside the two turns of viral DNA. Just like searching huge crowds, if everybody is tightly masked, a clever scout drone would be unable to identify or distinguish the terrorist. It would be much easier for the gRNA to search its matching DNA sequences if the two strands of the virus gene are not always tightly tangled with each other, as required in the Watson–Crick model.
As the main genetic message carrier, the secondary structure of DNA found in plasmids should not be limited to prokaryotes.
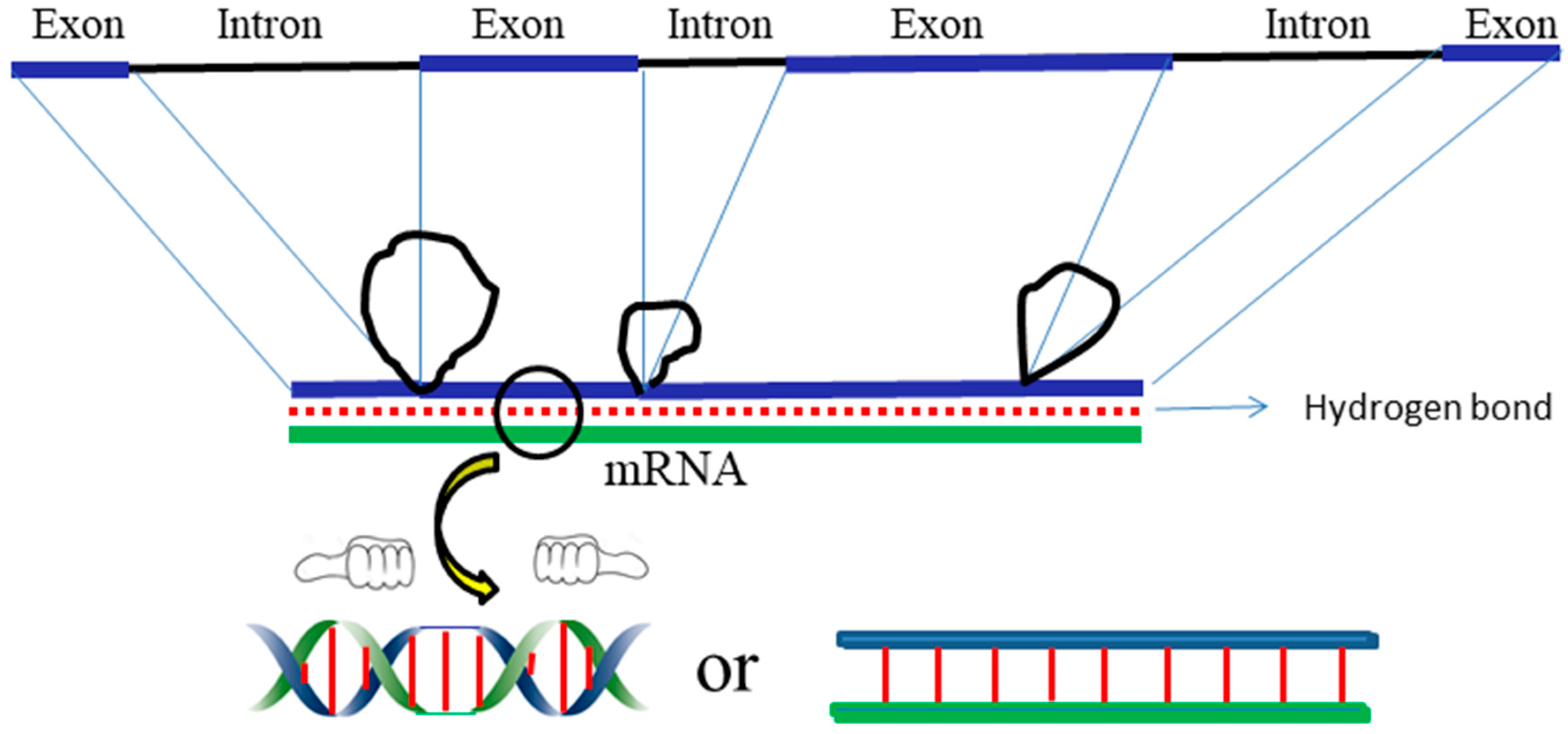
In eukaryotes, DNA is further wound on histones. The coding message becomes more difficult to decipher if the DNA is really in the canonical double helix model. While the genetic code is tightly hidden in the chromosome, the question regarding how it can be read by similar homeobox proteins at specific times and places remains un-answered [
43]. The ambidextrous DNA model enables easier recognition of the message hidden inside DNA.
The finding of a zero-linking-number topoisomer is a crucial fact that serves as a touchstone for the judgement of two different hypotheses. It acts as a fatal weapon on the Achilles heel of the double helix model, since its occurrence cannot be explained by the canonical double helix model.
The ambidextrous DNA model is not perfect or irrefutable. If the ambidextrous model is correct, a reasonable consequence is that, in any circular DNA, there is at least a pair of transitional sections between the oppositely wound helixes. According to the finding of Ha et al. [
44], there are two extruded bases at the junction between the B-DNA and Z-DNA of a 15mer oligonucleotides found using X-ray crystallography. This possibly implies that in any circular DNA, there are at least four pairs of bases that lost their hydrogen bonds at the two junctions.
This may be one of the possible refutations to the ambidextrous model. However, it should be noted that under physiological conditions, DNA structure is dynamic. Its tertiary structure is frequently affected by the surrounding conditions, including temperature, solvent, pH, cations, intercalating reagents, etc. It is unlikely that 8 or 12 deprived hydrogen bonds at the two junctions can be freely moving alone in the circular DNA. Furthermore, it is unfavorable for the stability of a circular DNA to always to keep some unpaired regions. Perhaps the extruded bases found in that short linear DNA can only occur under static conditions during crystallization.
It is probable that our hypothesis and related claims are not easy to be accepted by many readers. We welcome any rational criticism or accusation. It is the normal process of the self-adjustment of science based on testable explanations and predictions. The clash of different ideas or opinions may produce valuable products and is beneficial to the advancement of science.
The strength of a scientific theory is not only in that it can reasonably explain many facts, but also correctly predict what will happen or forecast some previously unknown phenomenon. Crick told us: “The job of theorists, especially in biology, is to suggest new experiments. A good theory makes not only predictions, but surprising predictions that then turn out to be true. (If its predictions appear obvious to experimentalists, why they need a theory?)” [
5].
We predicted that there is a zero-linking-number topoisomer in any kind of plasmid [
21]. One way of testing the ambidextrous double helix is to examine whether this prediction is correct.
To most scientists, this double helix conjecture is not obvious or even impossible. If proven, it would have surprising effects and would ease the understanding of the ambidextrous DNA model amongst scientists, as well as the public with no background knowledge of DNA topology. Now, we can provide a rather detailed and promising research plan for scientists to prove it or disprove it.
4. Promising and Feasible Research Proposal
A large amount of evidence leads to refuting one of the main claims of the double helix model, i.e., the two strands of native DNA are always wound plectonemically. Although the ambidextrous DNA model has got the support of much independent evidence, it would be better if one of its predictions can be verified.
Based on many years of trial and error tests, we learned that the two strands of a plasmid can be efficiently dissociated under very mild conditions [
24]. This paves the way for the demonstration of the double helix conjecture.
Actually, this is the test Crick et al. originally proposed in 1979. They assumed that if the two complementary DNAs of a plasmid are unlinked, simply heating can dissociate the two circular DNA strands. However, all native plasmid samples occur as a group with different linking numbers; therefore, it is a prerequisite to find out which one is a potential candidate and then get enough samples to do the test.
To find a non-linked plasmid or zero-linking-number topoisomer is not easy. The critical point is where can it be found and how can it be found. According to our experience, the zero-linking-number topoisomer is among the relaxed DNA. By following the procedures described here, the double helix conjecture can be verified. A dedicated scientist can accomplish the project with a low investment of time and money in most biochemical laboratories. A detailed description of the operation can be found from the references marked in parenthesis.
1. Choose a plasmid as the research object and make enough pure supercoiled DNA sample.
2. Make a set of relaxed topoisomers from the plasmid. This can be achieved via three different ways:
(a) Relaxing the supercoiled DNA with topoisomerase 1 under different conditions by varying temperature, ethidium bromide (EB)EB concentration, positive ions concentration, etc. [
41,
45].
(b) Turning the supercoiled plasmid into singly nicked DNA and then ligating them with DNA ligase under different conditions by varying the temperature, EB concentration, positive ions concentration, etc. [
41,
46].
(c) If a special strain is available, the whole set of topoisomer can be obtained using a novobiocin treatment [
36]. This would be the most convenient way to get whole set of relaxed plasmids.
3. The critical step is to completely denature the whole set of relaxed topoisomers slowly and carefully by incubating the plasmid in pure water at an appropriate temperature for a given time [
24].
4. The denatured plasmid can be renatured in the appropriate solvent, salt concentration, and temperature for a given time. This step may be the most time consuming. It needs to be tested several times to find the best reproducible condition [
24].
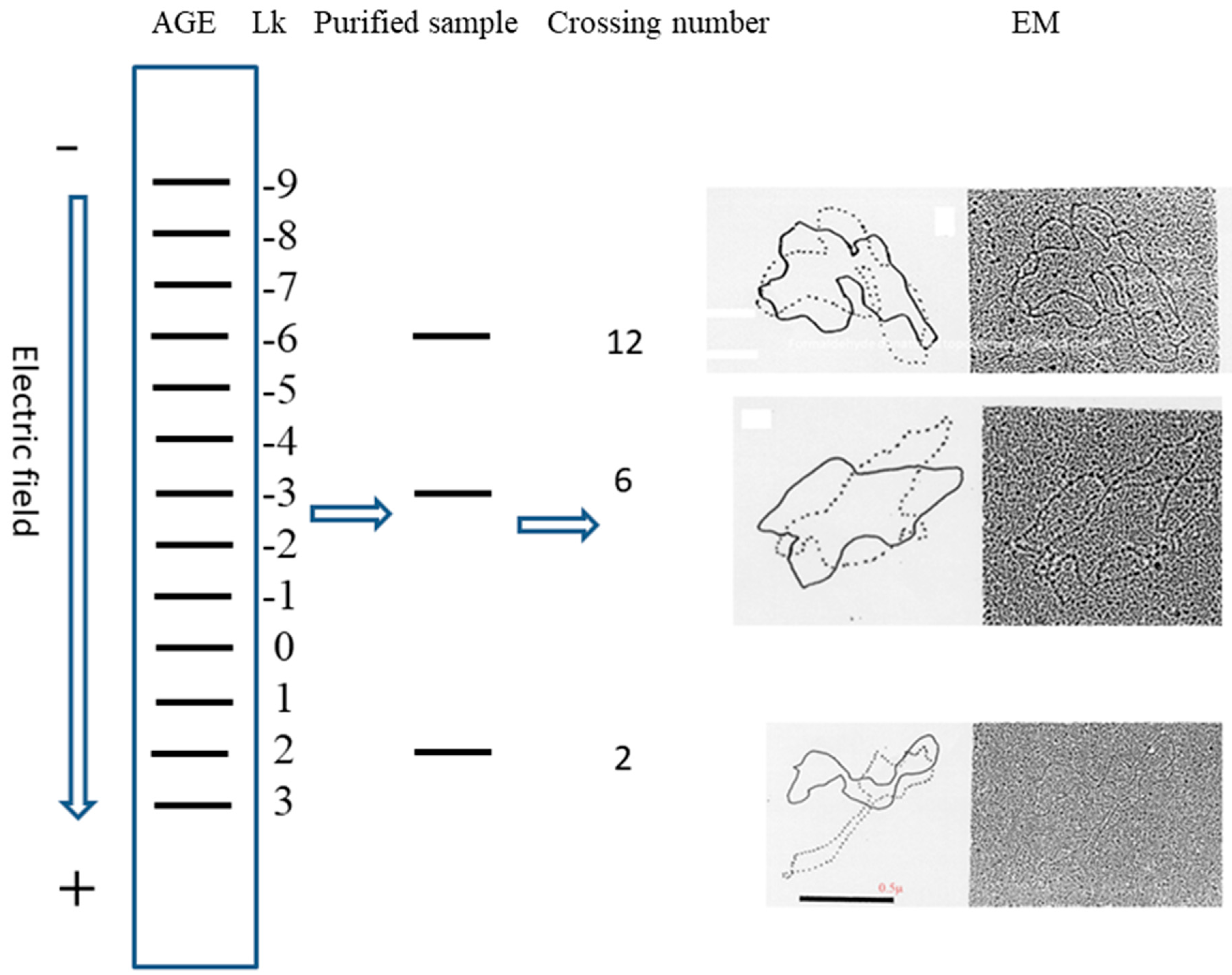
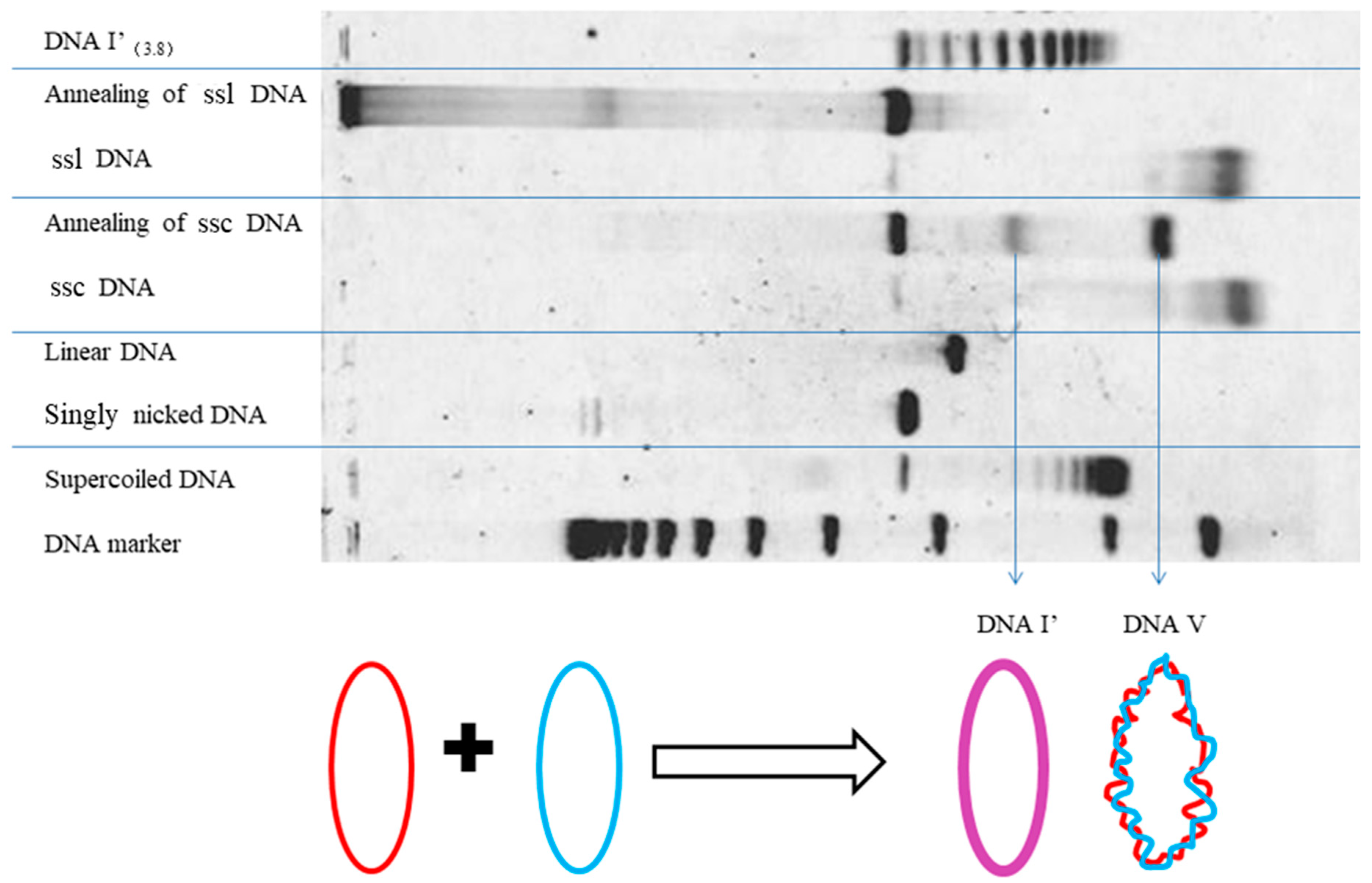
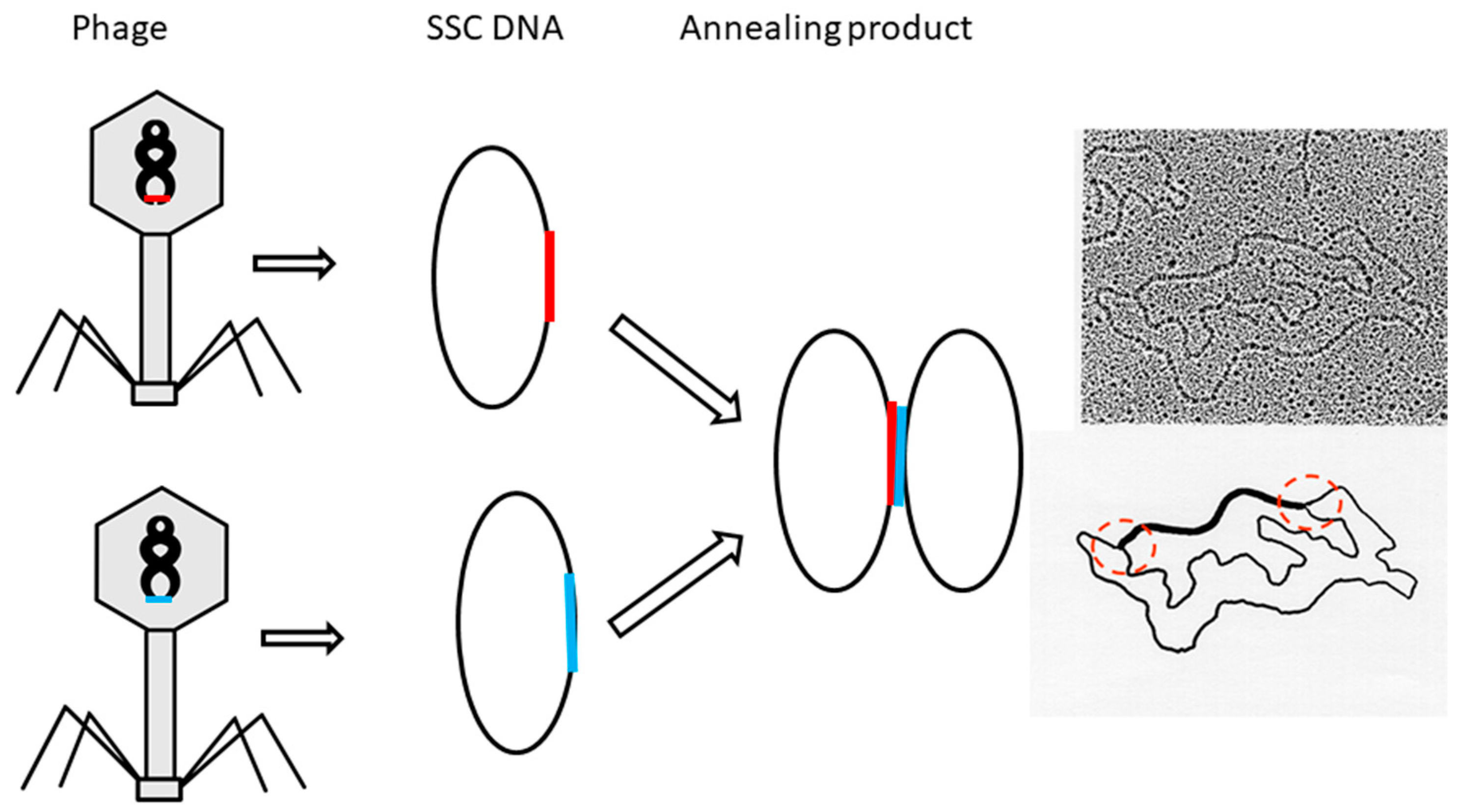
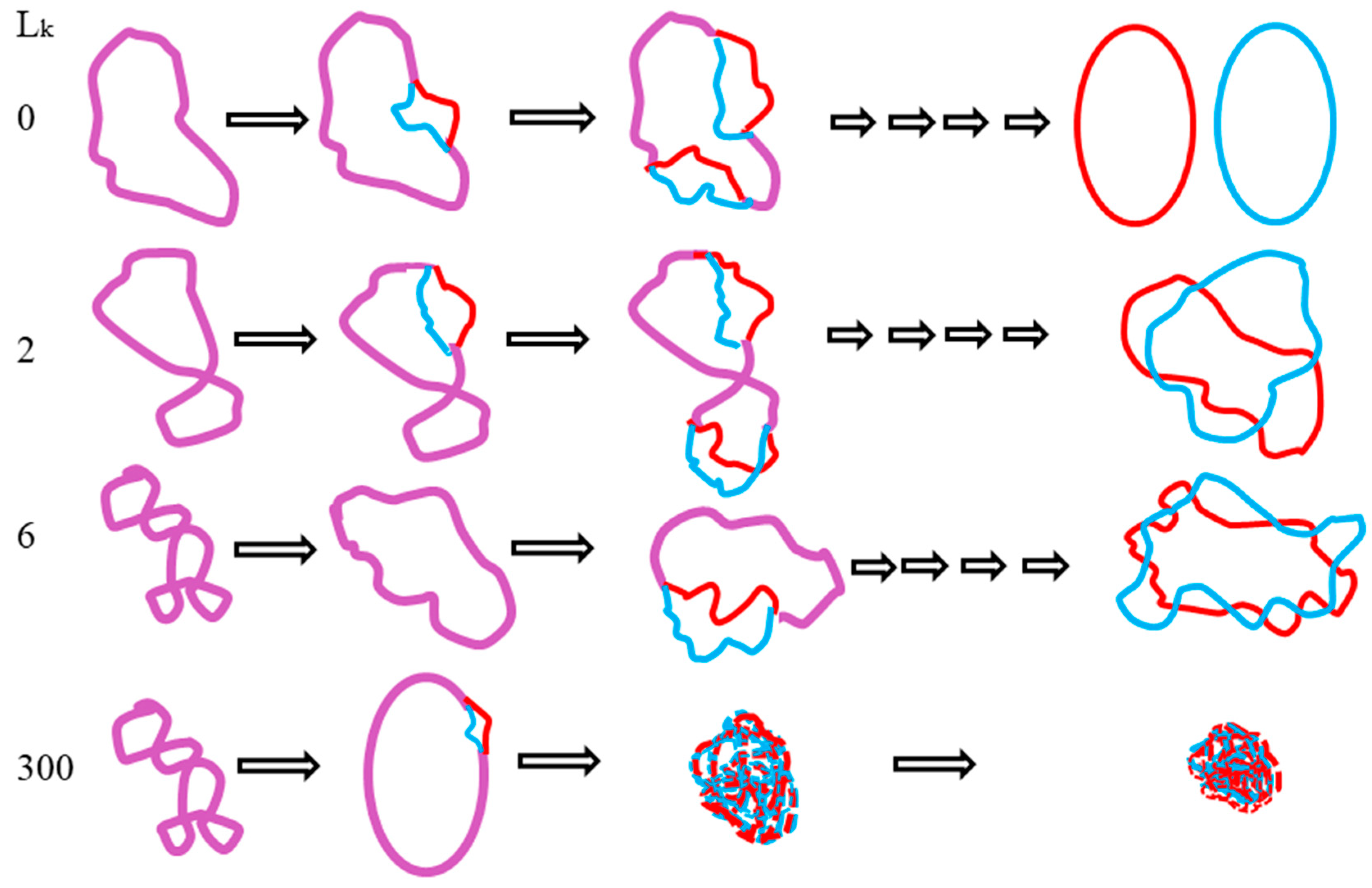
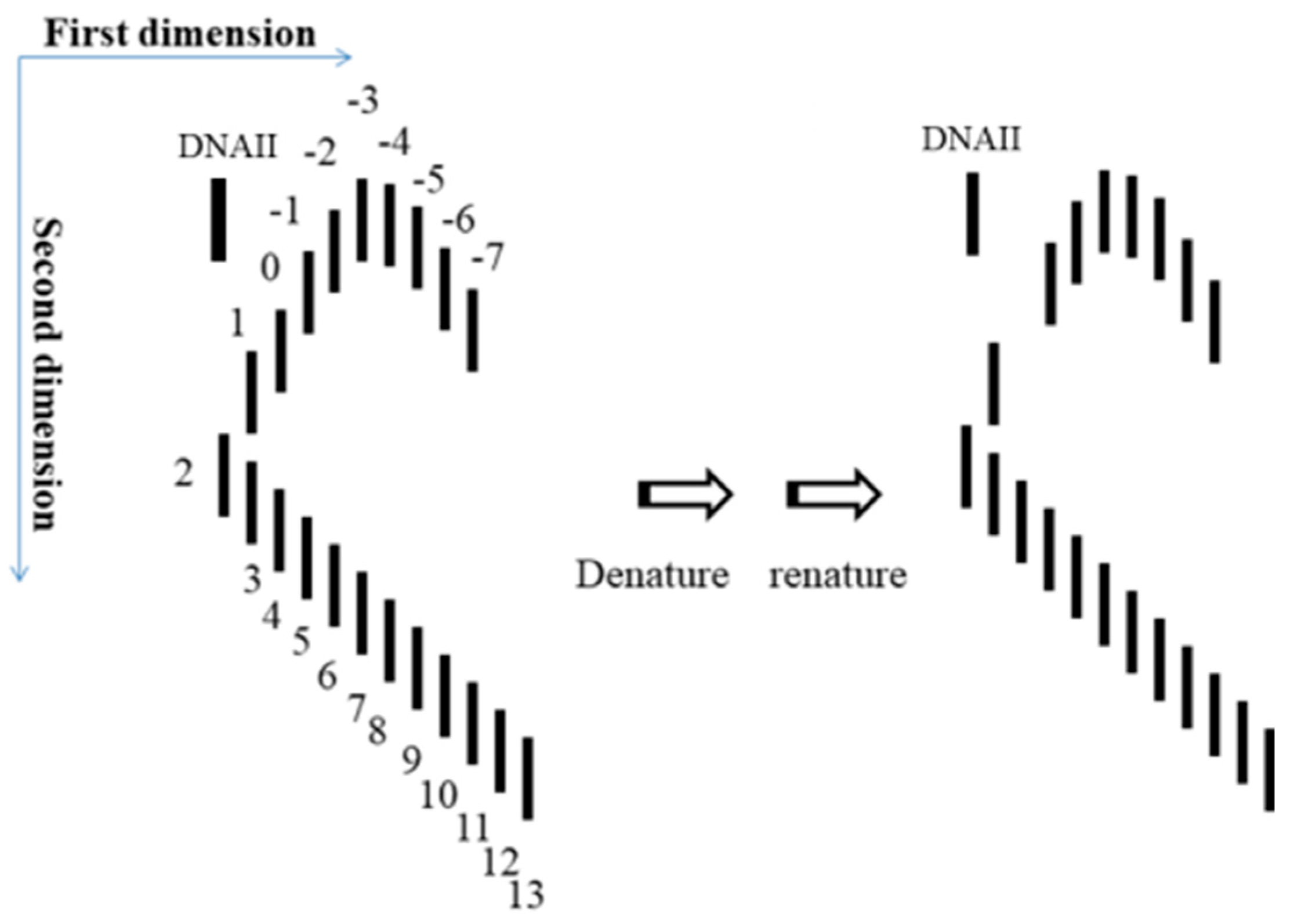
5. After the plasmid passes the manipulation of denaturing and renaturing, the two-dimensional AGE with control (same sample before manipulation) can be done. A zero-linking-number topoisomer can be located at the same location of control as that of the only missing band on the agarose gel. A schematic drawing is presented in
Figure 11 and the linking number of each band is marked beside.
Once the location of the zero-linking-number topoisomer is found, further purification of it may not be necessary. This is because this test already clearly indicated that the missing topoisomer must be a non-linked plasmid. However, if sufficient non-linked plasmids are sample is available, performing the test suggested by Crick et al. becomes feasible Additionally, the pure topoisomer may be of interest to some scholars for further investigation regarding its physical and chemical properties. The data of melting temperature (Tm), circular dichroism (CD) Tm, CD, etc., obtained from it or other individual topoisomers, would help us to get more information from DNA.
Two useful reminders are provided for those who may try this experiment: (a) During the denaturing process, DNA concentration plays an important role. The unlinked ssc DNA should separate better at a lower DNA concentration, whereas other denatured paired ssc DNAs are not affected by DNA concentration. (b) The binding force of hydrogen bonds between two complementary strands of DNA is closely related to the temperature, whereas the repelling force of the two strands in low salt solution or pure water is not.