1. Introduction
Artificial intelligence seeks to resemble the capabilities of human beings represented in machines, and it is involved in human fields such as learning, reasoning, adaptation, and self-correction [
1]. Within this field, neural networks and image processing are working together in order to generate an accurate classification model, improving learning from the extraction of characteristics and patterns. In addition, the combination of these two fields can be seen applied in regenerative medicine, microbiology, hematology, precision agriculture, and tumor identification, among others.
In Reference [
2], the authors obtained an improvement in the classification and learning speed of a model that adapts to the characteristics of the input data. The implementation of a convolutional neural network with image processing was performed, in which the results were produced similarly for different types of files and sizes, making use of semantic segmentation by the means of an FCN (fully convolutional network), in which the delimitation and separation of objects was carried out. In Reference [
2], the authors had difficulty in improving the resolution of high-quality images and videos from making use of convolutional neural networks; by analyzing each of the contained pixels, it was demonstrated that the implementation of a convolutional layer at the pixel level improved the quality of the images and videos without the need for a great expense of computational resources. This is why the combination of a convolutional neural network (CNN) with image segmentation methods offers the opportunity to optimize the results.
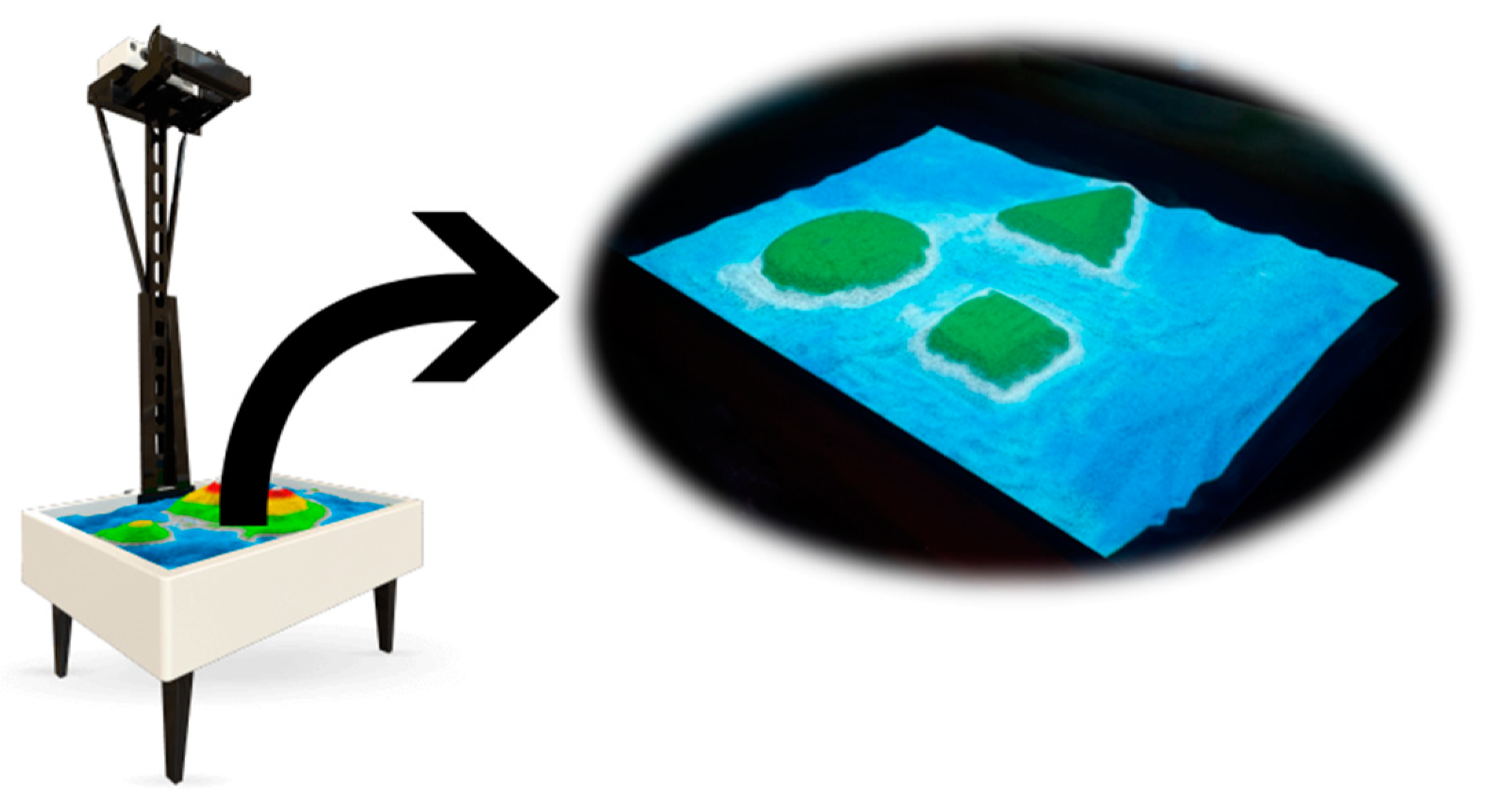
The use of an AR-Sandbox [
3,
4] is a technique that allows us to close the gap between two-dimensional (2D) and three-dimensional (3D) visualization when projecting a digital topographic map on a landscape of isolated space, improving spatial thinking and modeling skills of people, with the purpose of placing it within the use of early childhood education and rehabilitation through motor therapy, which will be expanded upon in the Image Acquisition Section. For these reasons, this study’s motivation is to create contributions towards image recognition based in other fields, such as immersive techniques, through deep learning by the means of convolutional neural networks, making use of hyperparameter optimization and image processing.
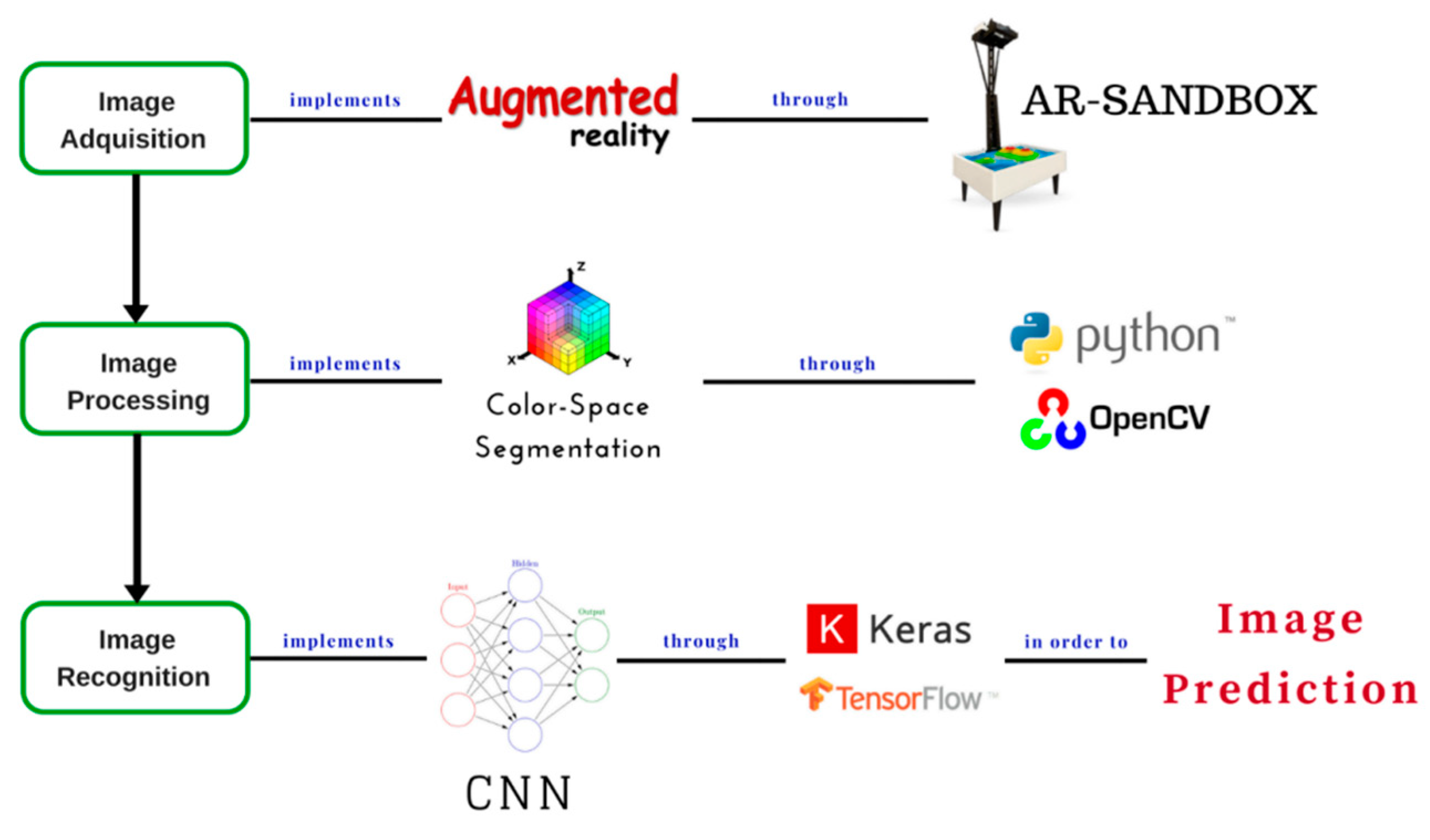
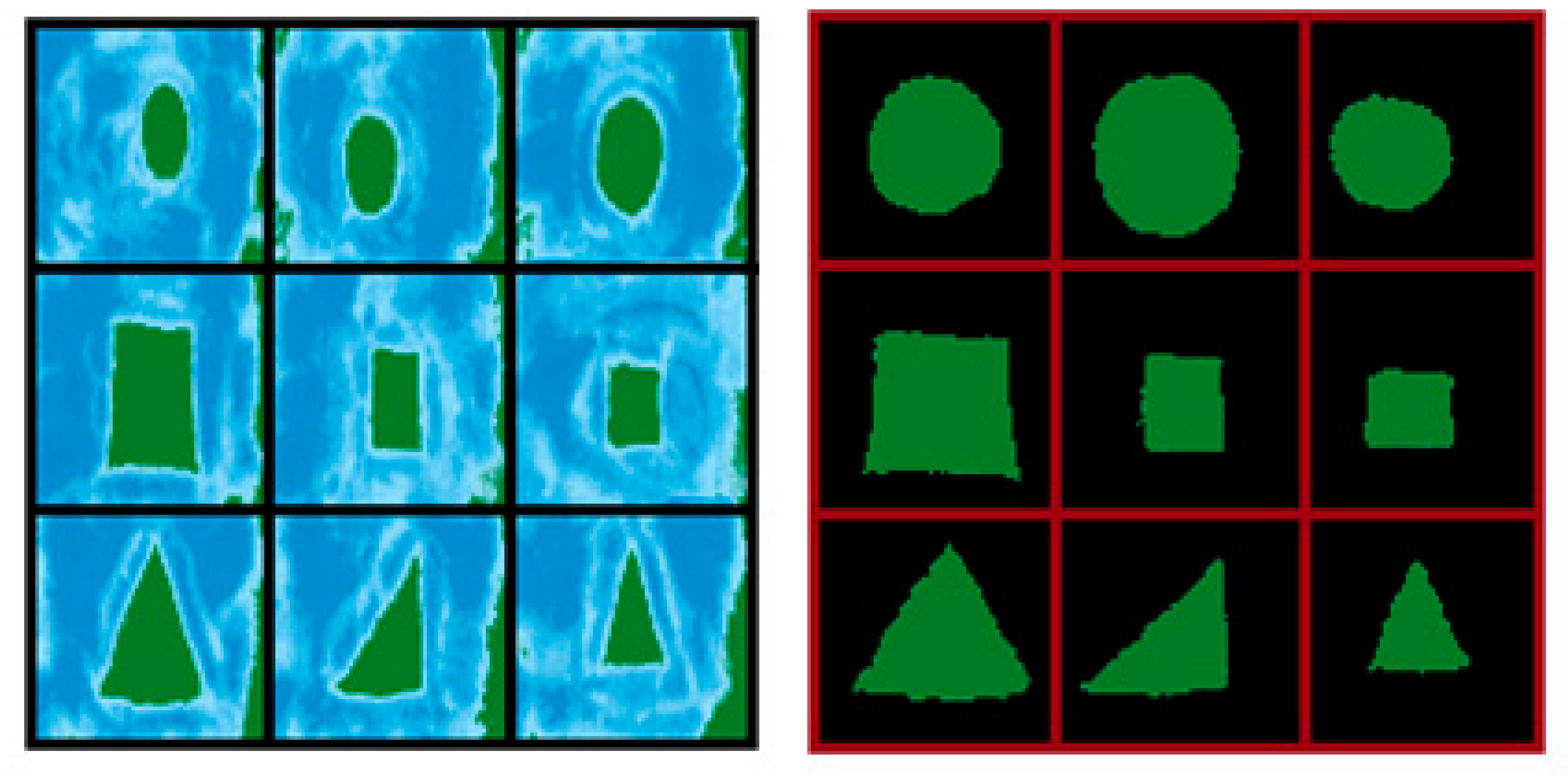
This article presents the preliminary results of a prediction model implementation that is based on convolutional neural networks for the classification of geometrical figures, contemplating previous steps such as the acquisition of images using the AR-Sandbox augmented reality device, and the processing of these images through segmentation by color–space. The purpose of applying this type of segmentation to the images acquired with the AR-Sandbox is to improve the model performance of the selected convolutional neural network, improving the extraction and identification of characteristics of the geometric figures in the prediction phase, and preserving a high percentage of similarity between the original image and the segmented image. To ensure this aspect, the efficiency of the segmentation method is evaluated by using the similarity coefficients of Jaccard and Sørensen-Dice [
5], which will be discussed later, in order to integrate the model with the AR-Sandbox for future applications in rehabilitation and education. The prediction model is made using the Keras neural network library under the TensorFlow framework and the Talos library in order to carry out the configuration, performance, and evaluation of hyperparameter optimization by implementing a Random Search algorithm, through an implementation made in Python. Additionally, the Talos library works with any Keras model [
5].
The rest of the article is organized as follows.
Section 2 consists of the background, where works related to the topics to be developed are discussed. Next,
Section 3 presents a macro-scenario that is subdivided into three components: approach, development, and testing, presenting the characteristics and the implementation of each of these components. In the scenario test component, two datasets with different characteristics are used in order to perform a comparative analysis and to establish the results. Finally, the conclusion and future works are presented.
2. Background
Image recognition through convolutional neural networks has had a large number of applications, together with the previous image processing, in order to optimize pattern recognition [
6]. Convolutional neural networks have had continuous improvements since their creation, through the innovation of new layers and making use of different computer vision techniques [
7]. In the regenerative medicine field, a study carried out by Reference [
8] has developed automatic cell culture systems, where a CNN was implemented as a deep learning method to automate the recognition of cellular differences by means of the contrast in the different images. On the other hand, in China, a study was carried out where white blood cell segmentation was implemented, which proposed a method for segmentation based on color–space, making a color adjustment before segmentation, where an accuracy of 95.7% and an overall accuracy of 91.3% were achieved for the segmentation of the nucleus and segmentation of the cytoplasm [
9].
On the other hand, based on studies carried out by Reference [
10], randomized trials are more efficient for the optimization of multiple parameters than tests in a grid; for this reason, the Random Search method was selected for the optimization of CNN hyperparameters. With this method, four tasks are performed: (1) the use of the same model with different initial parameters; (2) the best models discovered through cross-validation are taken; (3) different control points for each of the models are identified, and finally, (4) execution of the average of the parameters in the training stage.
For the implementation of Random Search in Python, the Talos library was used, which makes use of optimization algorithms such as Random Search, Grid Search, and correlation-based optimization. Talos has a POD strategy: Prepare, Optimize, and Deploy; this task is automated, and it produces results for prediction problems [
5].
In addition, in a study carried out by Reference [
11], they proposed a convolutional neuronal network for the improvement of thermal images, incorporating the domain of brightness with a residual learning technique, increasing the performance and speed of convergence. The fast development of precision agriculture has generated the need for agriculture production management and estimation through the classification of crops through satellite images, but due to the complexity and fragmentation of the characteristics, traditional methods have not been able to fulfill the standards of agricultural problems. For this reason, in Reference [
12], a classification method of agricultural remote sensing images was proposed based on convolutional neural networks, where the correct classification rate obtained was 99.55%. According to the above, neural networks, together with image processing, are a commonly used alternative for image classification [
13].
Additionally, statistical methods have been identified to verify the efficiency of the segmentation method. For this case, the Jaccard coefficient and the Sørensen-Dice coefficient were used, which make comparisons of images (original and with image segmentation) through the bitmaps of each one:
with
A and
B being the bitmaps of the selected images. The coefficient takes the intersection between
A and
B, which will be the points in common, and divides the union between
A and
B, this being the totality of the two images without repetition of data. This throws a value between 0 and 1 that is known as the Jaccard coefficient [
14], as shown in Equation (1).
with
A and
B being the bitmaps of the selected images, we take the rule of the intersection between
A and
B, which will be the points in common, multiplied by two; this is divided between the sum of the standard of
A and the norm of
B, which yields a value between 0 and 1, which is known as the Sørensen-Dice coefficient [
15], as shown in Equation (2). These coefficients are based on the Kappa coefficient, which is another statistical method that provides a probability of success [
16]. A good result of these coefficients is a number that is greater than 0.70, and this means that the segmentation method used supports image processing for prediction making.
In other methods, Reference [
17] uses image analysis and fractal dimensions to detect tumors in computed tomography (CT) scan images with high contrast, where the image preprocessing contrast of the cut images was improved by converting the values in the image intensity, using the histogram equalization to increase the accuracy of the tumor diagnosis. The image noise was reduced by the use of median filtering; finally, border detection was carried out by using the border function developed in MATLAB. This method has better performance and provides more acceptable responses than statistical algorithms. In Reference [
18], the authors describe the development and implementation of feature selection for content-based image retrieval (CBIR) through a system that automatically extracts features from images using color, texture, and shape in order to use feature selection by means of a genetic algorithm that searches for the best feature-use feature selection. The results of this study conclude that the CBIR system is more efficient, and that it performs better when using feature selection based on a genetic algorithm, because it reduces the time for retrieval and also increases the retrieval precision.
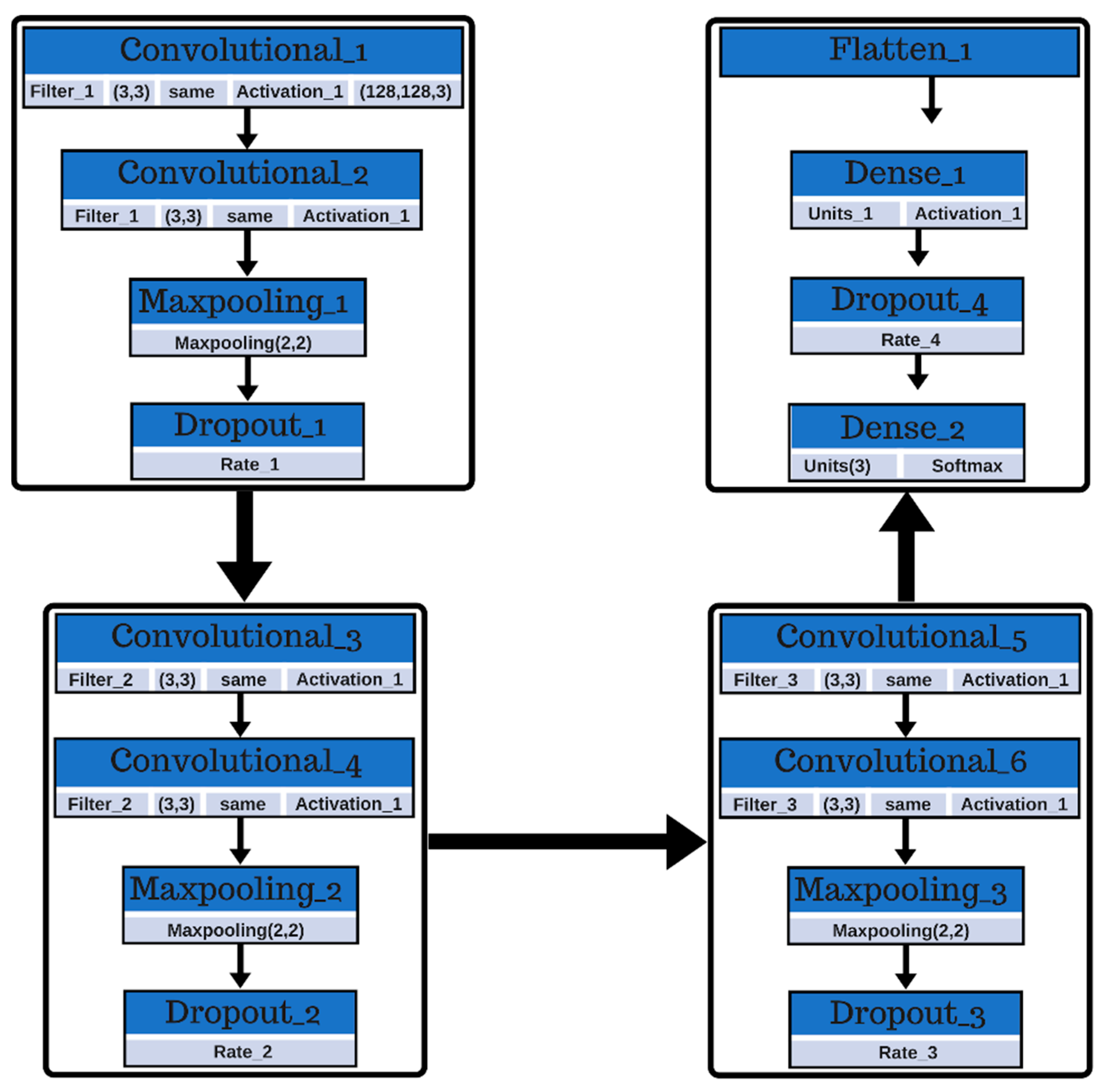
On the other hand, the authors in Reference [
19] presented a study that proposed a 14-layer convolutional neural network, combined with three advanced techniques: batch normalization, dropout, and stochastic pooling in order to carry out multiple sclerosis identification. The results of this study concluded that with this model they obtained an accuracy of 98.77 ± 0.39%. Results were compared with CNN when using maximum pooling and average pooling; the comparison showed that stochastic pooling gave a better performance than the other two pooling methods.
Furthermore, in Reference [
20], the researchers proposed to reconstruct objects based on incomplete images and some information on the 3D object using active and passive methods to reconstruct high-resolution items. Additionally, mathematics is essential for image processing; in Reference [
21], the researchers used an arithmetic method to find the right and more trustworthy solution for quantization tables, which are tables for image compression. In a similar field, the use of statistical analysis for algorithm optimization is a good way to optimize models; in Reference [
22], this type of analysis was used to enhance a cognitive model for a particle swarm based on vorticity, or the tendency of something to rotate.
On basis of this panorama, the present study intends to carry out the recognition of images by the means of deep learning techniques, such as convolutional neural networks and image processing by color–space segmentation, with the purpose of determining the performance variation of a convolutional neural network, applying color–space segmentation to one of the test datasets. In the next section, the selected study scenario is presented.
5. Results, Analysis, and Discussions
According to the
Table 3, taking as a guide Reference [
27], and the results and performances based on individual, average, and overall classification model accuracies, the 10 best models were selected by Random Search, and variations in the coefficients of loss, accuracy, and mean squared error that were input to their validation through dataSetOriginal and dataSetFilter, are presented. An average variation of 0.3945 in the function of loss, and an average variation of 0.1483 in the function accuracy, and a variation of 0.1118 in the function mean squared error, were obtained.
From the evaluation made for the CNN model, with dataSetOriginal and dataSetFilter, the following aspects were analyzed: loss function, hit metric, confusion matrix, and ROC curve, and the following results were obtained:
When evaluating the CNN with dataSetOriginal, 72% was obtained as a function of loss, while with dataSetFilter, 36% was obtained; therefore, a 36% decrease was obtained. On the other hand, through dataSetOriginal, a metric of hits was obtained with a percentage of 70%; while using dataSetFilter, 87% correct answers were obtained, presenting an increase of 17% between the two test datasets.
Regarding the confusion matrix, the percentage of correct answers when using dataSetOriginal was 61%, while with dataSetFilter, a percentage of 87.7% was obtained, presenting an increase of 26.7%. In addition, in
Table 7, the specific analysis for each of the geometric figures is presented.
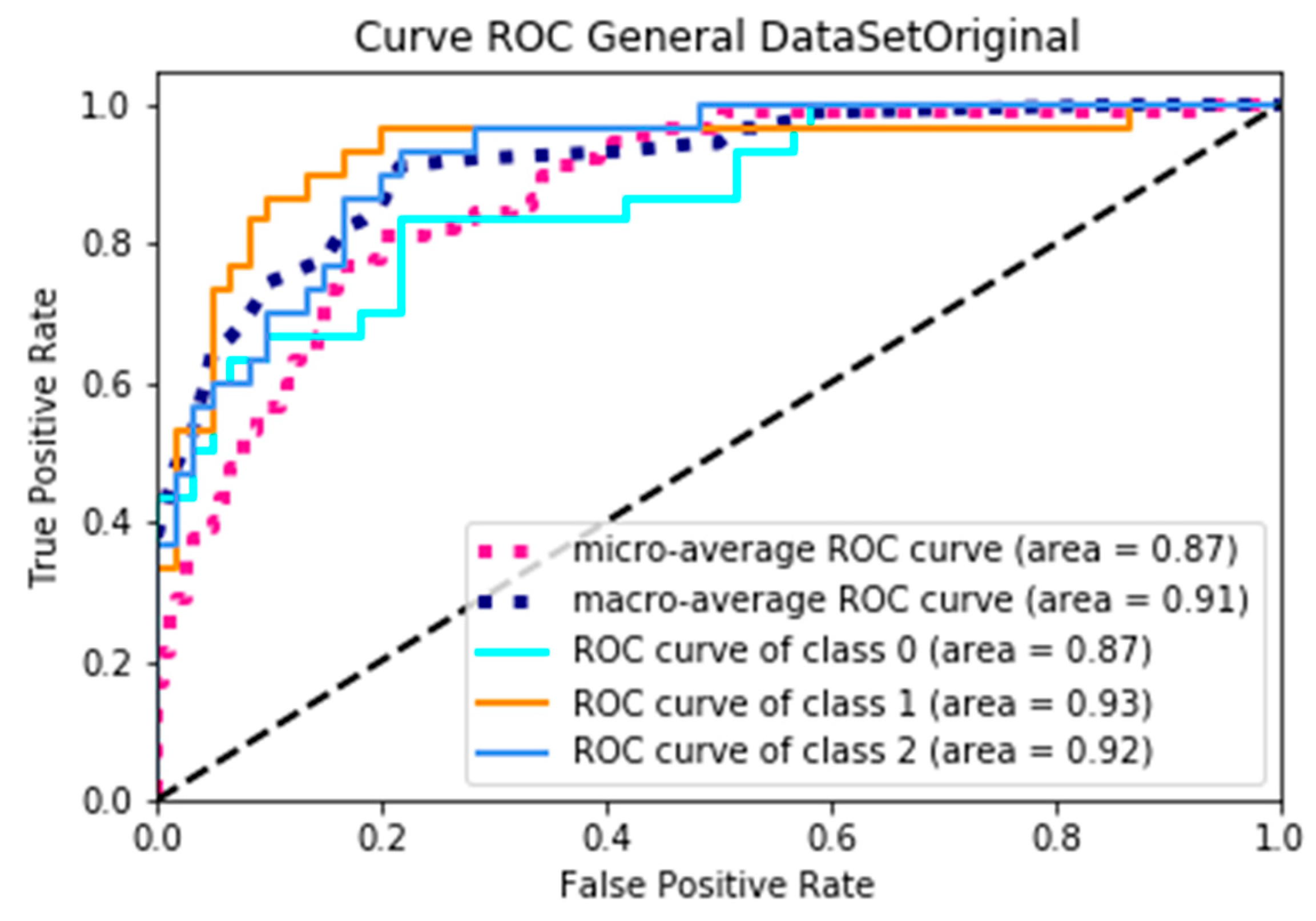
An ROC curve is a graph that shows the performance of a classification model in all of the classification thresholds [
28]. When analyzing an ROC curve, the determining parameter is the area under the curve (AUC); therefore, it is the factor that will be analyzed next.
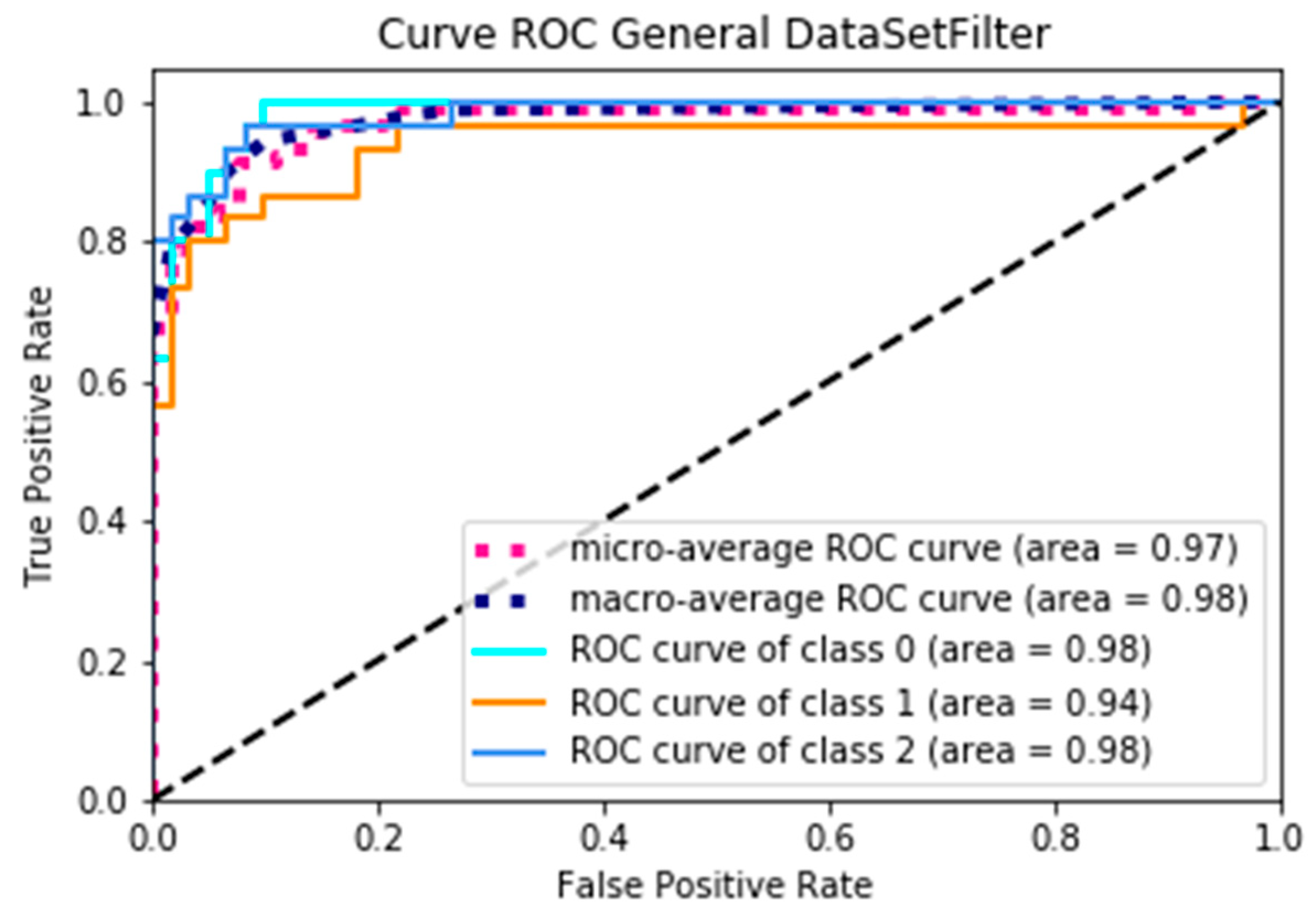
Figure 8 represents the ROC curve when using dataSetOriginal, where, according to the AUC, the minimum average is 0.87 and the maximum average is 0.91, obtaining an average yield of 0.89; on the other hand, Figure 9 presents the curve of the ROC when using dataSetFilter, where the minimum average of the AUC is 0.97 and the maximum average is 0.98, obtaining an average yield of 0.975, and presenting an increase of 0.085 yield. Through a specific analysis,
Table 8 presents the data and variations of the AUC of each of the classes by using the two test datasets.
From
Table 9, the performance in each of the classes presented variations with respect to the other test datasets, but the AUC value for image recognition, which previously had segmentation by color–space, was higher.
In addition, we agree with results shown in Reference [
29], since it states that for a prediction model to be considered optimal, the curve described must be convex, and in this case in Figure 9, taking the Macro curve as a reference, we can show that convexity occurs, since when making a region between this curve and the diagonal, putting any two points in the region, and passing a line through them, the curve is entirely within this region.
6. Conclusions and Future Works
From the analysis presented, it can be determined that: when evaluating the set of preselected CNN models with two data sets, one of them is previously processed by applying segmentation by color-space; an average decrease of 39.45% is obtained in the function of loss categorical_crossentropy, which is an increase of 14.83% on average in the accuracy coefficient, and a decrease of 11.18% on average in the regression coefficient.
On basis of the analysis presented, when evaluating the CNN with a previously processed dataset by applying color–space segmentation, a decrease of 36% in the loss value is obtained, increasing the value of hits generated by the accuracy metric by 17%, given that the distance between the value of the prediction and the expected value decreases. By implementing color–space segmentation to a set of test data, a positive contribution was made to the identification and extraction of patterns or characteristics that are necessary for the classification of images. Since the data has a coefficient of 0.8221 for the Jaccard method and a coefficient of 0.8767 for the Sørensen-Dice method, and since its probabilities are greater than 0.70, we can conclude that the segmentation of images by color–space contributes to image processing and subsequent prediction, given that through these values, it can be inferred that when applying segmentation by color space, the original and segmented images retain a large percentage of similarity, emphasizing the definition of the characteristics of the images.
When the area under the ROC curve is taken as a reference, an average of 0.975, corresponding to the CNN performance, is presented under the processed dataset, increasing by a value of 0.085 with respect to the average AUC generated from the raw dataset.
From the implementation of these tools, it is evident that the combination of areas, such as multimedia and artificial intelligence, provides a great field of action to continue researching and making proposals. The development of the technologies worked provides opportunities for making proposals in sectors such as health and education. In these sectors, proposals can be made for motor skill development, the development of basic knowledge in early childhood, reactions to certain situations, and psychology, among others.
In future work, it is proposed that an immersive environment is implemented by using augmented reality tools and devices to support motor therapy in children, continuously monitoring the child’s emotional behavior through brain–computer interfaces. On the other hand, an expansion of the scope of the developed CNN is planned, in order to achieve the identification of other geometric figures, forms, and symbols, in order to apply it to the abovementioned fields of action.