Using Bacteriophages to Treat Resilient Bacteria Found in Produced Water
Abstract
1. Introduction
2. Materials and Methods
2.1. Produced Water Collection and Processing
2.2. Model Strains and Culture Conditions
2.3. Bacteriophage Growth Conditions
2.4. Spot Assay and Bacteriophage Titer Determination
2.5. Strain Growth Curves in Produced Water
2.6. Individual Phage Treatment Assays
2.7. Phage Cocktail Assay Involving B. megaterium sp.
2.8. Statistical Analysis
3. Results and Discussion
3.1. Selection of Model Organisms
3.2. Produced Water Processing and Growth Assays
3.3. Phage Treatment Assays
3.3.1. Single Phage Treatment Involving P. aeruginosa
3.3.2. Single Phage Treatment Involving B. megaterium
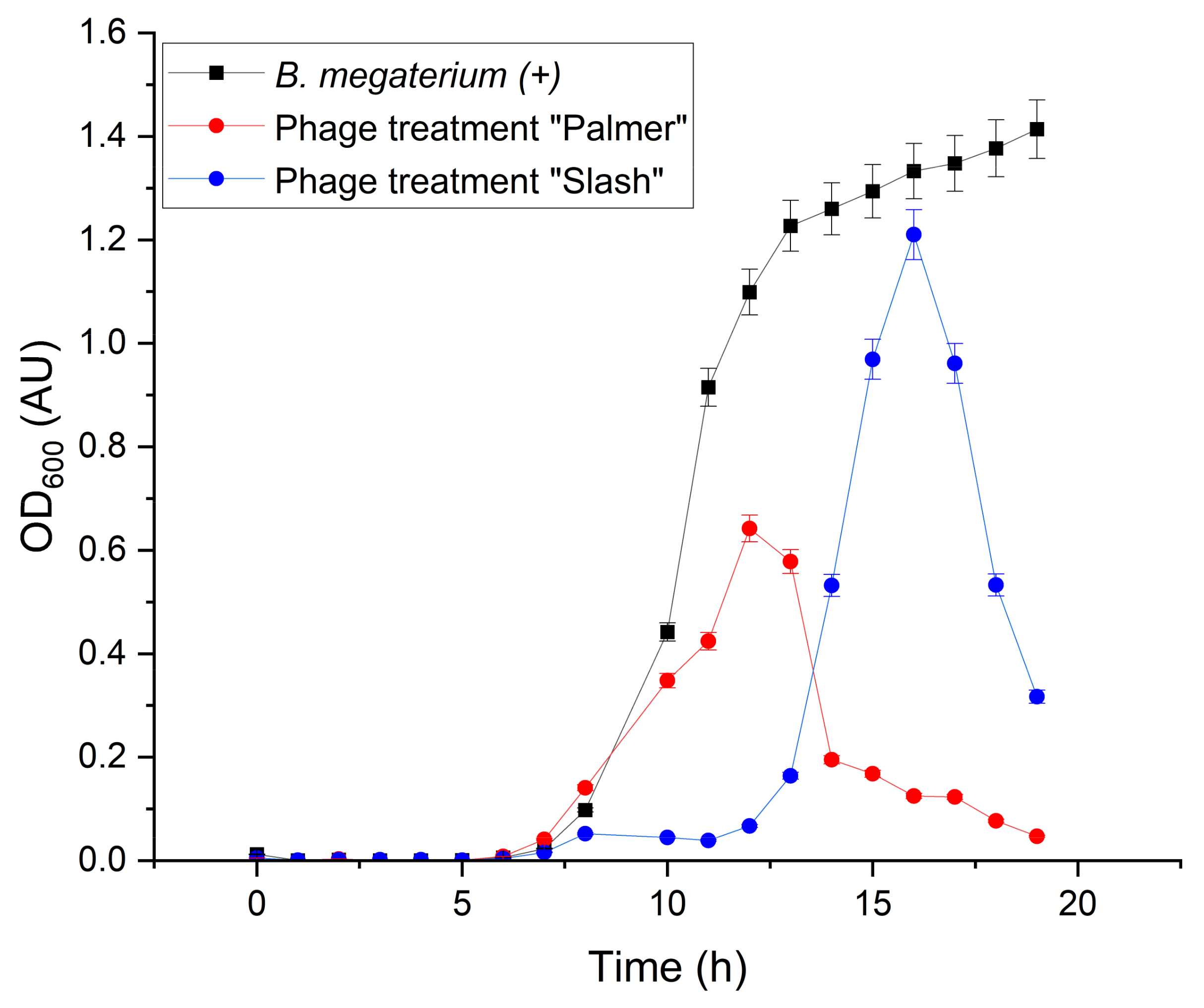
3.3.3. Phage Cocktail Treatment Involving B. megaterium
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Scanlon, B.R.; Reedy, R.C.; Xu, P.; Engle, M.; Nicot, J.P.; Yoxtheimer, D.; Yang, Q.; Ikonnikova, S. Can We Beneficially Reuse Produced Water from Oil and Gas Extraction in the U.S.? Sci. Total Environ. 2020, 717, 137085. [Google Scholar] [CrossRef]
- Al-Ghouti, M.A.; Al-Kaabi, M.A.; Ashfaq, M.Y.; Da’na, D.A. Produced Water Characteristics, Treatment and Reuse: A Review. J. Water Process. Eng. 2019, 28, 222–239. [Google Scholar] [CrossRef]
- Igunnu, E.T.; Chen, G.Z. Produced Water Treatment Technologies. Int. J. Low-Carbon Technol. 2014, 9, 157–177. [Google Scholar] [CrossRef]
- Rodriguez, A.Z.; Wang, H.; Hu, L.; Zhang, Y.; Xu, P. Treatment of produced water in the permian basin for hydraulic fracturing: Comparison of different coagulation processes and innovative filter media. Water 2020, 12, 770. [Google Scholar] [CrossRef]
- Xiao, F. Characterization and Treatment of Bakken Oilfield Produced Water as a Potential Source of Value-Added Elements. Sci. Total Environ. 2021, 770, 145283. [Google Scholar] [CrossRef] [PubMed]
- Barbot, E.; Vidic, N.S.; Gregory, K.B.; Vidic, R.D. Spatial and Temporal Correlation of Water Quality Parameters of Produced Waters from Devonian-Age Shale Following Hydraulic Fracturing. Environ. Sci. Technol. 2015, 47, 41–59. [Google Scholar] [CrossRef]
- Lewandowski, C.M.; Co-investigator, N.; Lewandowski, C.M. Naturally Occurring Radioactive Materials (NORM) in Produced Water and Scale from Texas Oil, Gas, and Geothermal Wells: Geographic, Geologic, and Geochemical Controls. Vr. Lanscapes Texas 2015, 1, 1689–1699. [Google Scholar]
- McMahon, P.B.; Galloway, J.M.; Hunt, A.G.; Belitz, K.; Jurgens, B.C.; Johnson, T.D. Geochemistry and Age of Groundwater in the Williston Basin, USA: Assessing Potential Effects of Shale-Oil Production on Groundwater Quality. Appl. Geochem. 2021, 125, 104833. [Google Scholar] [CrossRef]
- Varona-Torres, E.; Carlton, D.D.; Hildenbrand, Z.L.; Schug, K.A. Matrix-Effect-Free Determination of BTEX in Variable Soil Compositions Using Room Temperature Ionic Liquid Co-Solvents in Static Headspace Gas Chromatography Mass Spectrometry. Anal Chim Acta 2018, 1021, 41–50. [Google Scholar] [CrossRef] [PubMed]
- Hildenbrand, Z.L.; Santos, I.C.; Liden, T.; Carlton, D.D.; Varona-Torres, E.; Martin, M.S.; Reyes, M.L.; Mulla, S.R.; Schug, K.A. Characterizing Variable Biogeochemical Changes during the Treatment of Produced Oilfield Waste. Sci. Total Environ. 2018, 634, 1519–1529. [Google Scholar] [CrossRef] [PubMed]
- Abass, O.K.; Zhuo, M.; Zhang, K. Concomitant Degradation of Complex Organics and Metals Recovery from Fracking Wastewater: Roles of Nano Zerovalent Iron Initiated Oxidation and Adsorption. Chem. Eng. J. 2017, 328, 159–171. [Google Scholar] [CrossRef]
- De Oliveira, E.S.D.; Roseana, F.; Pereira, C.; Alice, M.; Lima, G.D.A. Study on Biofilm Forming Microorganisms Associated with the Biocorrosion of X80 Pipeline Steel in Produced Water from Oilfield. Mater. Res. 2021, 24. [Google Scholar] [CrossRef]
- Mohan, A.M.; Bibby, K.J.; Lipus, D.; Hammack, R.W.; Gregory, K.B. The Functional Potential of Microbial Communities in Hydraulic Fracturing Source Water and Produced Water from Natural Gas Extraction Characterized by Metagenomic Sequencing. PLoS ONE 2014, 9, e107682. [Google Scholar] [CrossRef] [PubMed]
- Kahrilas, G.A.; Blotevogel, J.; Stewart, P.S.; Borch, T. Biocides in Hydraulic Fracturing Fluids: A Critical Review of Their Usage, Mobility, Degradation, and Toxicity. Environ. Sci. Technol. 2015, 49, 16–32. [Google Scholar] [CrossRef] [PubMed]
- Akyon, B.; Lipus, D.; Bibby, K. Glutaraldehyde Inhibits Biological Treatment of Organic Additives in Hydraulic Fracturing Produced Water. Sci. Total Environ. 2019, 666, 1161–1168. [Google Scholar] [CrossRef]
- Santos, I.C.; Chaumette, A.; Smuts, J.; Hildenbrand, Z.L.; Schug, K.A. Analysis of Bacteria Stress Responses to Contaminants Derived from Shale Energy Extraction. Environ. Sci. Process Impacts 2019, 21, 269–278. [Google Scholar] [CrossRef]
- Sinha, S.; Grewal, R.K.; Roy, S. Modeling Bacteria–Phage Interactions and Its Implications for Phage Therapy; Elsevier Inc.: Amsterdam, The Netherlands, 2018; Volume 103. [Google Scholar]
- Burrowes, B.H.; Abedon, S.T.; Burrowes, B.H.; McConville, M.L.; Harper, D.R. Bacteriophages: Biology, Technology; Springer: Berlin/Heidelberg, Germany, 2018; ISBN 9783319419855. [Google Scholar]
- Sahota, J.S.; Smith, C.M.; Radhakrishnan, P.; Winstanley, C.; Goderdzishvili, M.; Chanishvili, N.; Kadioglu, A.; O’Callaghan, C.; Clokie, M.R.J. Bacteriophage Delivery by Nebulization and Efficacy Against Phenotypically Diverse Pseudomonas Aeruginosa from Cystic Fibrosis Patients. J. Aerosol Med. Pulm. Drug Deliv. 2015, 28, 353–360. [Google Scholar] [CrossRef]
- Klopatek, S.; Callaway, T.R.; Wickersham, T.; Sheridan, T.G.; Nisbet, D.J. Bacteriophage Utilization in Animal Hygiene. In Bacteriophages; Springer: Cham, Switzerland, 2018; pp. 1–28. [Google Scholar] [CrossRef]
- Fieseler, L.; Loessner, M.J.; Hagens, S. Bacteriophages and Food Safety. In Protective Cultures, Antimicrobial Metabolites and Bacteriophages for Food and Beverage Biopreservation; Woodhead Publishing: Sawston, UK, 2011; pp. 161–178. [Google Scholar]
- Breitbart, M.; Wegley, L.; Leeds, S.; Schoenfeld, T.; Rohwer, F. Phage Community Dynamics in Hot Springs. Appl. Environ. Microbiol. 2004, 70, 1633–1640. [Google Scholar] [CrossRef]
- Jepson, C.D.; March, J.B. Bacteriophage Lambda Is a Highly Stable DNA Vaccine Delivery Vehicle. Vaccine 2004, 22, 2413–2419. [Google Scholar] [CrossRef]
- Lu, Z.; Breidt, F.; Plengvidhya, V.; Fleming, H.P. Bacteriophage Ecology in Commercial Sauerkraut Fermentations. Appl. Environ. Microbiol. 2003, 69, 3192–3202. [Google Scholar] [CrossRef] [PubMed]
- Wilson, C.; Caton, T.M.; Buchheim, J.A.; Buchheim, M.A.; Schneegurt, M.A.; Miller, R.V. DNA-Repair Potential of Halomonas spp. from the Salt Plains Microbial Observatory of Oklahoma. Microb. Ecol. 2004, 48, 541–549. [Google Scholar] [CrossRef] [PubMed]
- Sanchez-Rosario, R.; Hildenbrand, Z.L. Produced Water Treatment and Valorization: A Techno-Economical Review. Energies 2022, 15, 4619. [Google Scholar] [CrossRef]
- Jiménez, S.; Micó, M.M.; Arnaldos, M.; Medina, F.; Contreras, S. State of the Art of Produced Water Treatment. Chemosphere 2018, 192, 186–208. [Google Scholar] [CrossRef] [PubMed]
- Chibani-Chennoufi, S.; Sidoti, J.; Bruttin, A.; Kutter, E.; Sarker, S.; Brüssow, H. In Vitro and in Vivo Bacteriolytic Activities of Escherichia Coli Phages: Implications for Phage Therapy. Antimicrob. Agents Chemother. 2004, 48, 2558–2569. [Google Scholar] [CrossRef] [PubMed]
- Jia, R.; Yang, D.; Xu, D.; Gu, T. Anaerobic Corrosion of 304 Stainless Steel Caused by the Pseudomonas Aeruginosa Biofilm. Front. Microbiol. 2017, 8, 2335. [Google Scholar] [CrossRef] [PubMed]
- Alnuaimi, M.T.; Taher, T.A.; Aljanabi, Z.Z.; Adel, M.M. High-Resolution Gc/Ms Study of Biodegradation of Crude Oil by Bacillus Megaterium. Res. Crops 2020, 21, 650–657. [Google Scholar] [CrossRef]
- Kokjohn, T.A.; Sayler, G. Attachment and Replication of Pseudomonas Aeruginosa Bacteriophages under Conditions Simulating Aquatic Environments. Microbiology 1991, 137, 661–666. [Google Scholar] [CrossRef]
- Zhang, Y.; Hunt, H.K.; Hu, Z. Application of Bacteriophages to Selectively Remove Pseudomonas Aeruginosa in Water and Wastewater Filtrationsystems. Water Res. 2013, 47, 4507–4518. [Google Scholar] [CrossRef]
- Zhang, Y.; Hu, Z. Combined Treatment of Pseudomonas Aeruginosa Biofilms with Bacteriophages and Chlorine. Biotechnol. Bioeng. 2013, 110, 286–295. [Google Scholar] [CrossRef]
- Magin, V.; Garrec, N.; Andrés, Y. Selection of Bacteriophages to Control in Vitro 24 h Old Biofilm of Pseudomonas Aeruginosa Isolated from Drinking and Thermal Water. Viruses 2019, 11, 749. [Google Scholar] [CrossRef]
- Kauppinen, A.; Siponen, S.; Pitkänen, T.; Holmfeldt, K.; Pursiainen, A.; Torvinen, E.; Miettinen, I.T. Phage Biocontrol of Pseudomonas Aeruginosa in Water. Viruses 2021, 13, 928. [Google Scholar] [CrossRef] [PubMed]
- Lenski, R.E.; Levin, B.R. Constraints on the coevolution of bacteria and virulent phage: A model, some experiments, and predictions for natural communities. Am. Nat. 1985, 125, 585–602. [Google Scholar] [CrossRef]
- Oechslin, F. Resistance Development to Bacteriophages Occurring during Bacteriophage Therapy. Viruses 2018, 10, 351. [Google Scholar] [CrossRef] [PubMed]
- DeCrescenzo, A.J.; Ritter, M.A.; Chamakura, K.R.; Kuty Everett, G.F. Complete Genome of Bacillus Megaterium Siphophage Slash. Genome Announc. 2013, 1, e00862-13. [Google Scholar] [CrossRef]
- Hargrove, E.C.; Lopez, M.S.; Hernandez, A.C.; Everett, G.F.K. Complete Genome Sequence of Bacillus Megaterium Podophage Palmer. Genome Announc. 2015, 3, e00358-15. [Google Scholar] [CrossRef]
- Huang, Y.; Sun, H.; Wei, S.; Cai, L.; Liu, L.; Jiang, Y.; Xin, J.; Chen, Z.; Que, Y.; Kong, Z.; et al. Structure and Proposed DNA Delivery Mechanism of a Marine Roseophage. Nat. Commun. 2023, 14, 3609. [Google Scholar] [CrossRef]
- Dennehy, J.J.; Abedon, S.T. Adsorption: Phage Acquisition of Bacteria. In Bacteriophages; Springer: Cham, Switzerland, 2021; pp. 93–117. [Google Scholar] [CrossRef]
- Nale, J.Y.; Spencer, J.; Hargreaves, K.R.; Trzepiński, P.; Douce, G.R.; Clokie, M.R.J. Bacteriophage Combinations Significantly Reduce Clostridium Difficile Growth in Vitro and Proliferation in Vivo. Antimicrob. Agents Chemother. 2016, 60, 968–981. [Google Scholar] [CrossRef] [PubMed]
- Sadeqi, S.; Shahraki, A.H.; Nikkhahi, F.; Javadi, A.; Mahmoud, S.; Marashi, A. Application of Bacteriophage Cocktails for Reducing the Bacterial Load of Nosocomial Pathogens in Hospital Wastewater. Iran. J. Microbiol. 2022, 14, 395–401. [Google Scholar] [CrossRef] [PubMed]
- Fiedler, A.W.; Gundersen, M.S.; Vo, T.P.; Almaas, E.; Vadstein, O.; Bakke, I. Phage Therapy Minimally Affects the Water Microbiota in an Atlantic Salmon (Salmo Salar) Rearing System While Still Preventing Infection. Sci. Rep. 2023, 13, 19145. [Google Scholar] [CrossRef]
- Abujayyab, M.A.; Hamouda, M.; Aly Hassan, A. Biological Treatment of Produced Water: A Comprehensive Review and Metadata Analysis. J. Pet. Sci. Eng. 2022, 209, 109914. [Google Scholar] [CrossRef]
- Beheshti Maal, K.; Delfan, A.S.; Salmanizadeh, S. Isolation and Identification of Two Novel Escherichia Coli Bacteriophages and Their Application in Wastewater Treatment and Coliform’s Phage Therapy. Jundishapur J. Microbiol. 2015, 8, e14945. [Google Scholar] [CrossRef]
- Mathieu, J.; Yu, P.; Zuo, P.; Da Silva, M.L.B.; Alvarez, P.J.J. Going Viral: Emerging Opportunities for Phage-Based Bacterial Control in Water Treatment and Reuse. Acc. Chem. Res. 2019, 52, 849–857. [Google Scholar] [CrossRef] [PubMed]
- Sachs, J.D.; Schmidt-Traub, G.; Mazzucato, M.; Messner, D.; Nakicenovic, N.; Rockström, J. Six Transformations to Achieve the Sustainable Development Goals. Nat. Sustain. 2019, 2, 805–814. [Google Scholar] [CrossRef]
- Amakiri, K.T.; Ogolo, N.A.; Angelis-Dimakis, A.; Albert, O. Physicochemical Assessment and Treatment of Produced Water: A Case Study in Niger Delta Nigeria. Pet. Res. 2023, 8, 87–95. [Google Scholar] [CrossRef]



Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sanchez-Rosario, R.; Garcia, J.; Rodriguez, V.; Schug, K.A.; Hildenbrand, Z.L.; Bernal, R.A. Using Bacteriophages to Treat Resilient Bacteria Found in Produced Water. Water 2024, 16, 797. https://doi.org/10.3390/w16060797
Sanchez-Rosario R, Garcia J, Rodriguez V, Schug KA, Hildenbrand ZL, Bernal RA. Using Bacteriophages to Treat Resilient Bacteria Found in Produced Water. Water. 2024; 16(6):797. https://doi.org/10.3390/w16060797
Chicago/Turabian StyleSanchez-Rosario, Ramon, Jesus Garcia, Vivian Rodriguez, Kevin A. Schug, Zacariah L. Hildenbrand, and Ricardo A. Bernal. 2024. "Using Bacteriophages to Treat Resilient Bacteria Found in Produced Water" Water 16, no. 6: 797. https://doi.org/10.3390/w16060797
APA StyleSanchez-Rosario, R., Garcia, J., Rodriguez, V., Schug, K. A., Hildenbrand, Z. L., & Bernal, R. A. (2024). Using Bacteriophages to Treat Resilient Bacteria Found in Produced Water. Water, 16(6), 797. https://doi.org/10.3390/w16060797






