The Influence of pH on Subsurface Denitrification Stimulated with Emulsified Vegetable Oil
Abstract
:1. Introduction
2. Materials and Methods
2.1. Materials
2.2. Batch Experiments
2.3. Analytical Methods
2.4. ASM3 Modeling Approach
3. Results and Discussions
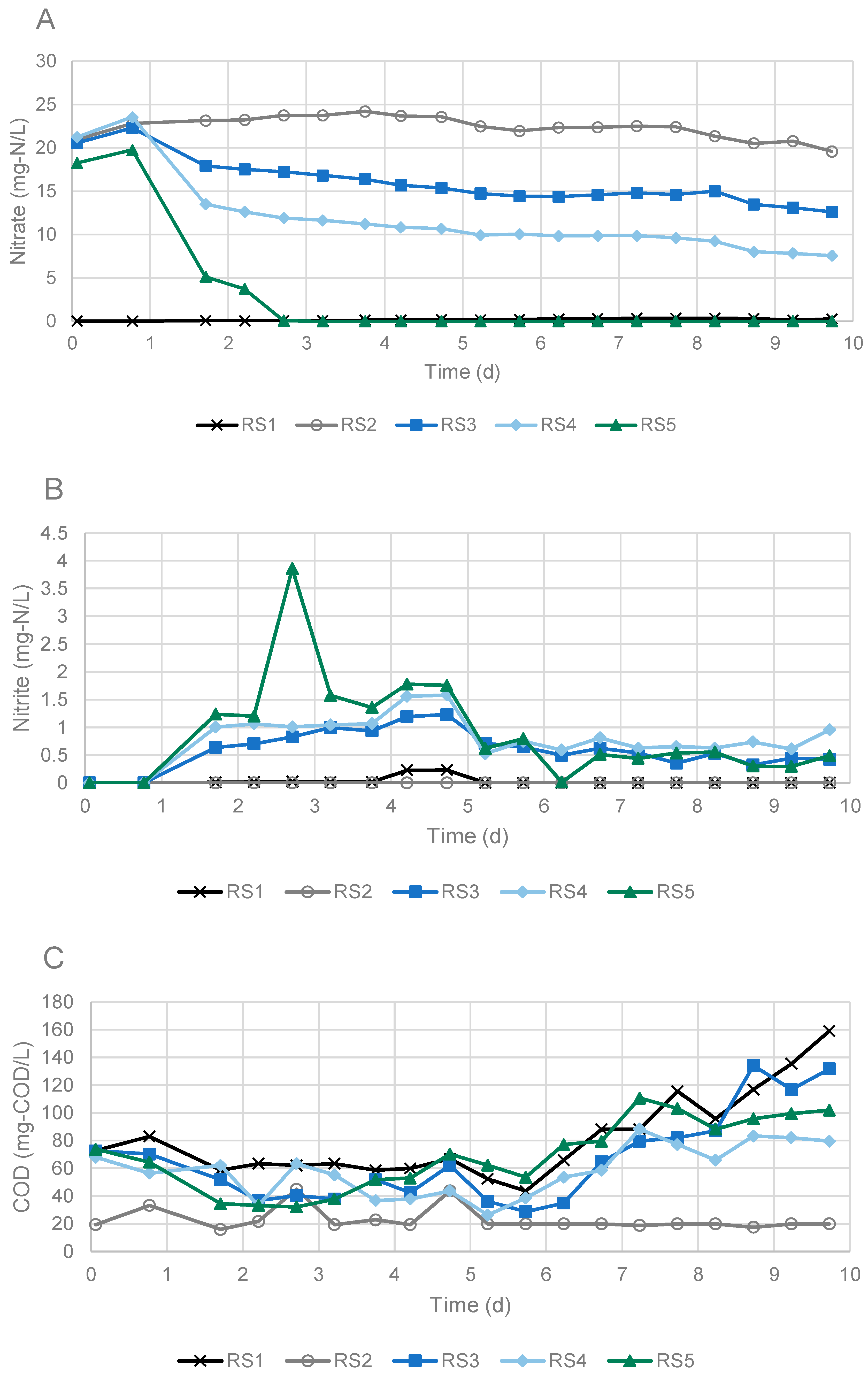
3.1. Denitrification in Experiments Using Sandwich and Falmouth Material at Neutral pH
3.2. Influence of pH on Denitrificaiton
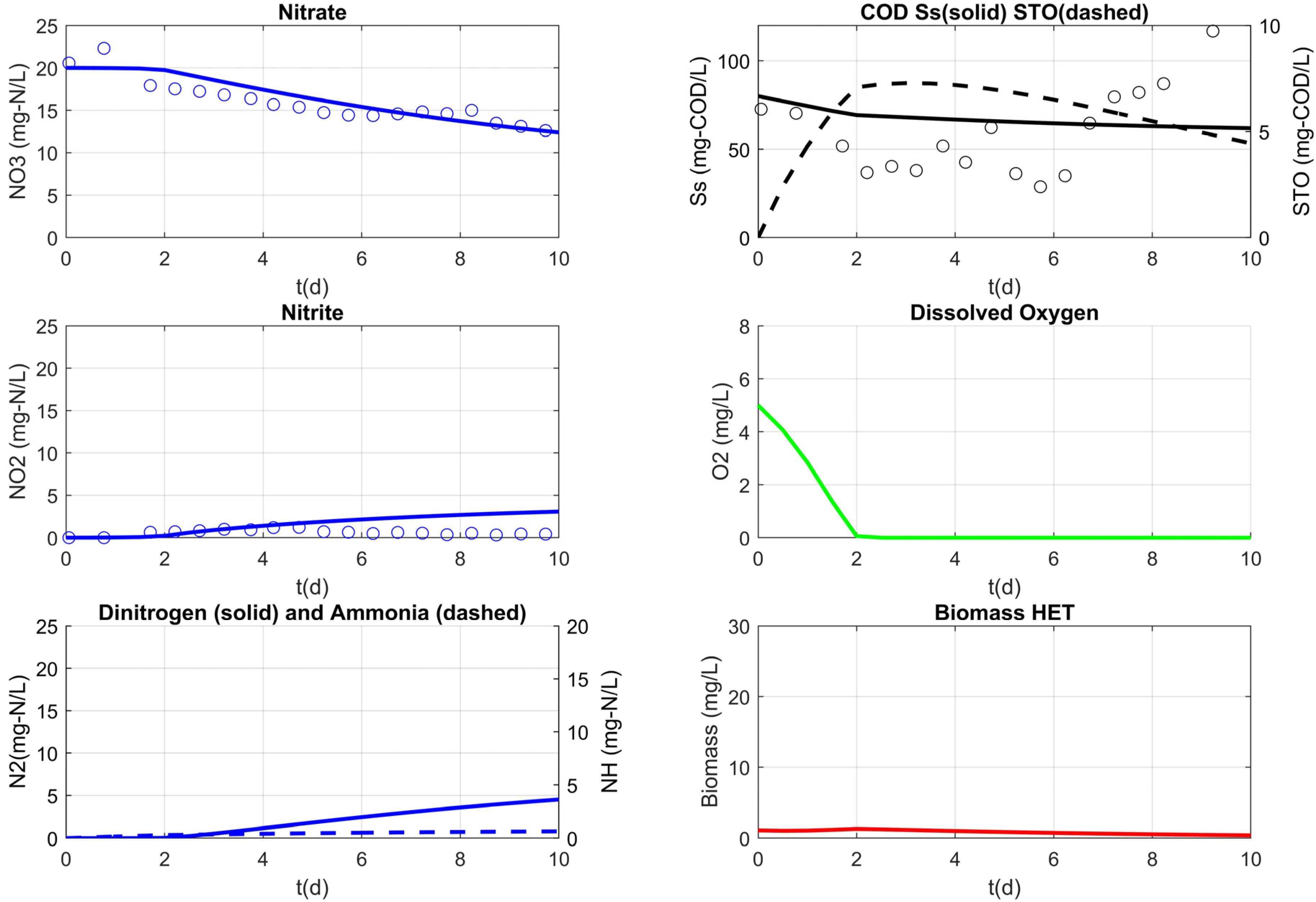
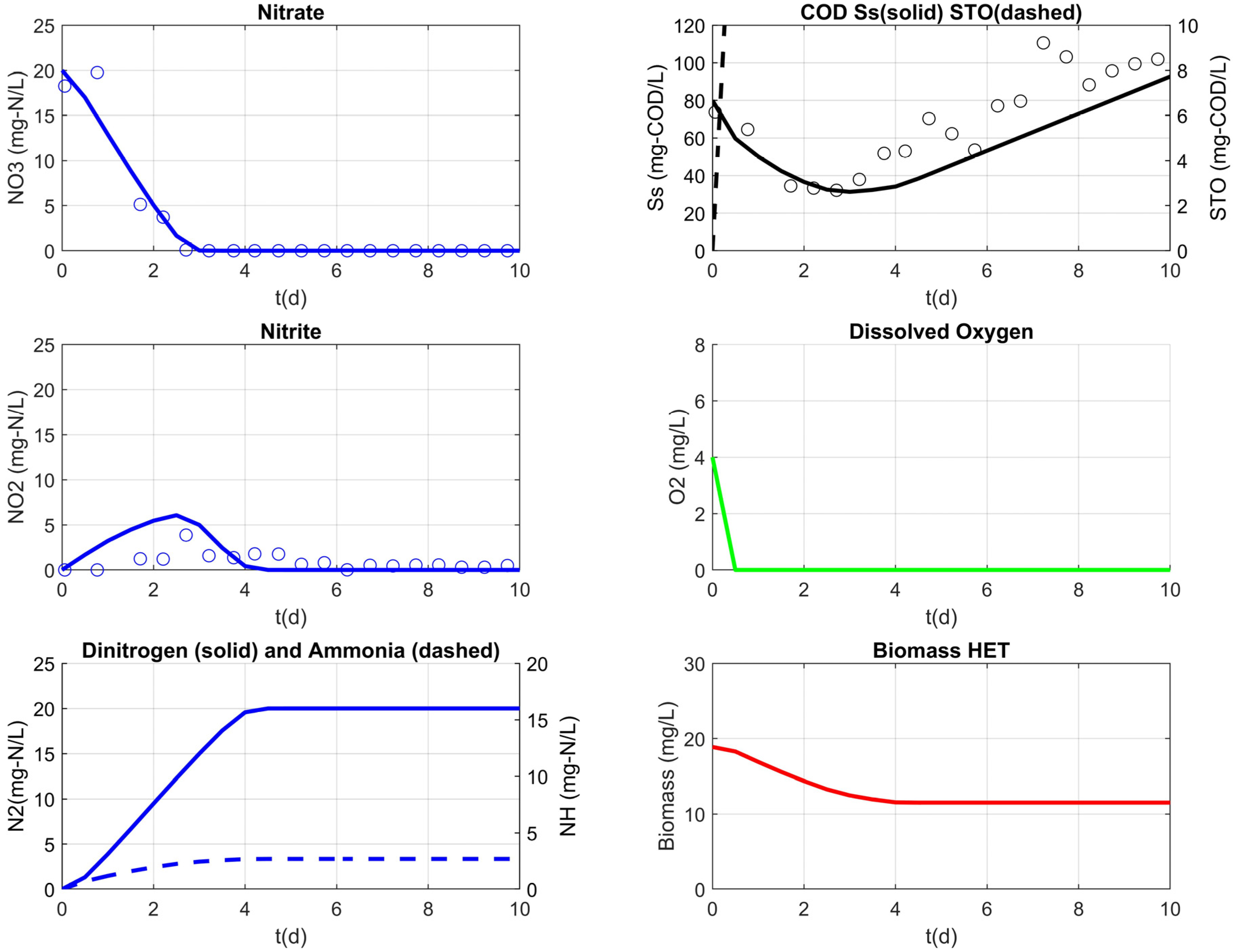
3.3. Model Results
4. Implications
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Robertson, W.D.; Blowes, D.W.; Ptacek, C.J.; Cherry, J.A. Long-Term Performance of In Situ Reactive Barriers for Nitrate Remediation. Ground Water 2000, 38, 689–695. [Google Scholar] [CrossRef]
- Korom, S.F. Natural Denitrification in the Saturated Zone: A Review. Water Resour. Res. 1992, 28, 1657–1668. [Google Scholar] [CrossRef] [Green Version]
- Lu, H.; Chandran, K. Diagnosis and Quantification of Glycerol Assimilating Denitrifying Bacteria in an Integrated Fixed-Film Activated Sludge Reactor via 13C DNA Stable-Isotope Probing. Environ. Sci. Technol. 2010, 44, 8943–8949. [Google Scholar] [CrossRef] [PubMed]
- Dahab, M.F. Comparison and Evaluation of In-Situ Bio-Denitrification Systems for Nitrate Reduction in Groundwater. Water Sci. Technol. 1993, 28, 359–368. [Google Scholar] [CrossRef]
- Hunter, W.J. Use of Vegetable Oil in a Pilot-Scale Denitrifying Barrier. J. Contam. Hydrol. 2001, 53, 119–131. [Google Scholar] [CrossRef]
- Friedman, L.; Mamane, H.; Avisar, D.; Chandran, K. The Role of Influent Organic Carbon-to-Nitrogen (COD/N) Ratio in Removal Rates and Shaping Microbial Ecology in Soil Aquifer Treatment (SAT). Water Res. 2018, 146, 197–205. [Google Scholar] [CrossRef]
- Slater, J.M.; Capone, D.G. Denitrification in Aquifer Soil and Nearshore Marine Sediments Influenced by Groundwater Nitrate. Appl. Environ. Microbiol. 1987, 53, 1292–1297. [Google Scholar] [CrossRef] [Green Version]
- Slater, L.; Binley, A. Evaluation of Permeable Reactive Barrier (PRB) Integrity Using Electrical Imaging Methods. Geophysics 2003, 68, 911–921. [Google Scholar] [CrossRef]
- Faisal, A.A.H.; Sulaymon, A.H.; Khaliefa, Q.M. A Review of Permeable Reactive Barrier as Passive Sustainable Technology for Groundwater Remediation. Int. J. Environ. Sci. Technol. 2018, 15, 1123–1138. [Google Scholar] [CrossRef]
- Robertson, W.D.; Cherry, J.A. In Situ Denitrification of Septic-System Nitrate Using Reactive Porous Media Barriers: Field Trials. Ground Water 1995, 33, 99–111. [Google Scholar] [CrossRef]
- Hiortdahl, K.M.; Borden, R.C. Enhanced Reductive Dechlorination of Tetrachloroethene Dense Nonaqueous Phase Liquid with EVO and Mg(OH)2. Environ. Sci. Technol. 2014, 48, 624–631. [Google Scholar] [CrossRef] [PubMed]
- Harkness, M.; Fisher, A. Use of Emulsified Vegetable Oil to Support Bioremediation of TCE DNAPL in Soil Columns. J. Contam. Hydrol. 2013, 151, 16–33. [Google Scholar] [CrossRef] [PubMed]
- Tang, G.; Wu, W.-M.; Watson, D.B.; Parker, J.C.; Schadt, C.W.; Shi, X.; Brooks, S.C. U(VI) Bioreduction with Emulsified Vegetable Oil as the Electron Donor–Microcosm Tests and Model Development. Environ. Sci. Technol. 2013, 47, 3209–3217. [Google Scholar] [CrossRef] [PubMed]
- Sleep, B.E.; Seepersad, D.J.; Mo, K.; Heidorn, C.M.; Hrapovic, L.; Morrill, P.L.; McMaster, M.L.; Hood, E.D.; LeBron, C.; Sherwood Lollar, B.; et al. Biological Enhancement of Tetrachloroethene Dissolution and Associated Microbial Community Changes. Environ. Sci. Technol. 2006, 40, 3623–3633. [Google Scholar] [CrossRef]
- Lee, M.D.; Hostrop, F.; Hauptmann, E.; Raymond, R., Jr.; Begley, J.F. Column Studies to Evaluate EVO for Nitrate Removal in Permeable Reactive Barriers on Cape Cod, MA. In Proceedings of the Remediation of Chlorinated and Recalcitrant Compounds—2016, Palm Springs, CA, USA, 22–26 May 2016; Battelle Memorial Institute: Columbus, OH, USA, 2016. [Google Scholar]
- Berge, N.D.; Ramsburg, C.A. Oil-in-Water Emulsions for Encapsulated Delivery of Reactive Iron Particles. Environ. Sci. Technol. 2009, 43, 5060–5066. [Google Scholar] [CrossRef]
- Muller, K.A.; Esfahani, S.G.; Chapra, S.C.; Ramsburg, C.A. Transport and Retention of Concentrated Oil-in-Water Emulsions in Porous Media. Environ. Sci. Technol. 2018, 52, 4256–4264. [Google Scholar] [CrossRef]
- Coulibaly, K.M.; Long, C.M.; Borden, R.C. Transport of Edible Oil Emulsions in Clayey Sands: One-Dimensional Column Results and Model Development. J. Hydrol. Eng. 2006, 11, 230–237. [Google Scholar] [CrossRef] [Green Version]
- Crocker, J.J.; Berge, N.D.; Ramsburg, C.A. Encapsulated Delivery of Reactive Iron Particles Using Oil-in-Water Emulsions. In Proceedings of the Securing Groundwater Quality in Urban and Industrial Environments; IAHS Press: Fremantle, Australia, 2008; pp. 242–249. [Google Scholar]
- Coulibaly, K.M.; Borden, R.C. Impact of Edible Oil Injection on the Permeability of Aquifer Sands. J. Contam. Hydrol. 2004, 71, 219–237. [Google Scholar] [CrossRef]
- Dong, J.; Yu, D.; Li, Y.; Li, B.; Bao, Q. Transport and Release of Electron Donors and Alkalinity during Reductive Dechlorination by Combined Emulsified Vegetable Oil and Colloidal Mg(OH)2: Laboratory Sand Column and Microcosm Tests. J. Contam. Hydrol. 2019, 225, 103501. [Google Scholar] [CrossRef]
- Smith, R.L.; Duff, J.H. Denitrification in a Sand and Gravel Aquifer. Appl. Environ. Microbiol. 1988, 54, 8. [Google Scholar] [CrossRef] [Green Version]
- Cape Cod Commission Section 208 Plan-Cape Cod Area Wide Water Quality Management Plan Update 2015. Available online: https://www.capecodcommission.org/resource-library/file/?url=/dept/commission/team/208/208%20Final/Cape_Cod_Area_Wide_Water_Quality_Management_Plan_Update_June_15_2015.pdf (accessed on 23 February 2023).
- Rakhimbekova, S.; O’Carroll, D.M.; Oldfield, L.E.; Ptacek, C.J.; Robinson, C.E. Spatiotemporal Controls on Septic System Derived Nutrients in a Nearshore Aquifer and Their Discharge to a Large Lake. Sci. Total Environ. 2021, 752, 141262. [Google Scholar] [CrossRef] [PubMed]
- Rust, C. Control of PH during Denitrification in Subsurface Sediment Microcosms Using Encapsulated Phosphate Buffer. Water Res. 2000, 34, 1447–1454. [Google Scholar] [CrossRef]
- Borden, R.C. Effective Distribution of Emulsified Edible Oil for Enhanced Anaerobic Bioremediation. J. Contam. Hydrol. 2007, 94, 1–12. [Google Scholar] [CrossRef]
- Borden, R.C.; Rodriguez, B.X. Evaluation of Slow Release Substrates for Anaerobic Bioremediation. Bioremediation J. 2006, 10, 59–69. [Google Scholar] [CrossRef]
- Duhamel, M.; Wehr, S.D.; Yu, L.; Rizvi, H.; Seepersad, D.; Dworatzek, S.; Cox, E.E.; Edwards, E.A. Comparison of Anaerobic Dechlorinating Enrichment Cultures Maintained on Tetrachloroethene, Trichloroethene, Cis-Dichloroethene and Vinyl Chloride. Water Res. 2002, 36, 4193–4202. [Google Scholar] [CrossRef]
- Löffler, F.E.; Cole, J.R.; Ritalahti, K.M.; Tiedje, J.M. Diversity of Dechlorinating Bacteria. In Dehalogenation: Microbial Processes and Environmental Applications; Häggblom, M.M., Bossert, I.D., Eds.; Springer: Boston, MA, USA, 2003; pp. 53–87. ISBN 978-0-306-48011-9. [Google Scholar]
- Gihring, T.M.; Zhang, G.; Brandt, C.C.; Brooks, S.C.; Campbell, J.H.; Carroll, S.; Criddle, C.S.; Green, S.J.; Jardine, P.; Kostka, J.E.; et al. A Limited Microbial Consortium Is Responsible for Extended Bioreduction of Uranium in a Contaminated Aquifer. Appl. Environ. Microbiol. 2011, 77, 5955–5965. [Google Scholar] [CrossRef] [Green Version]
- Calderer, M.; Jubany, I.; Pérez, R.; Martí, V.; de Pablo, J. Modelling Enhanced Groundwater Denitrification in Batch Micrococosm Tests. Chem. Eng. J. 2010, 165, 2–9. [Google Scholar] [CrossRef]
- Kornaros, M.; Lyberatos, G. Kinetic Modelling of Pseudomonas Denitrificans Growth and Denitrification under Aerobic, Anoxic and Transient Operating Conditions. Water Res. 1998, 32, 1912–1922. [Google Scholar] [CrossRef]
- Mekala, C.; Nambi, I.M. Transport of Ammonium and Nitrate in Saturated Porous Media Incorporating Physiobiotransformations and Bioclogging. Bioremediation J. 2016, 20, 117–132. [Google Scholar] [CrossRef]
- Clement, T.P.; Peyton, B.M.; Skeen, R.S.; Jennings, D.A.; Petersen, J.N. Microbial Growth and Transport in Porous Media under Denitrification Conditions: Experiments and Simulations. J. Contam. Hydrol. 1997, 24, 269–285. [Google Scholar] [CrossRef]
- Glass, C.; Silverstein, J. Denitrification Kinetics of High Nitrate Concentration Water: PH Effect on Inhibition and Nitrite Accumulation. Water Res. 1998, 32, 831–839. [Google Scholar] [CrossRef]
- Lu, B.; Liu, X.; Dong, P.; Tick, G.R.; Zheng, C.; Zhang, Y.; Mahmood-UI-Hassan, M.; Bai, H.; Lamy, E. Quantifying Fate and Transport of Nitrate in Saturated Soil Systems Using Fractional Derivative Model. Appl. Math. Model. 2020, 81, 279–295. [Google Scholar] [CrossRef]
- Rodríguez-Escales, P.; Folch, A.; van Breukelen, B.M.; Vidal-Gavilan, G.; Sanchez-Vila, X. Modeling Long Term Enhanced in Situ Biodenitrification and Induced Heterogeneity in Column Experiments under Different Feeding Strategies. J. Hydrol. 2016, 538, 127–137. [Google Scholar] [CrossRef] [Green Version]
- Sun, Y.; Petersen, J.N.; Clement, T.P.; Hooker, B.S. Effect of Reaction Kinetics on Predicted Concentration Profiles during Subsurface Bioremediation. J. Contam. Hydrol. 1998, 31, 359–372. [Google Scholar] [CrossRef]
- Wang, J.; Ma, R.; Guo, Z.; Qu, L.; Yin, M.; Zheng, C. Experiment and Multicomponent Model Based Analysis on the Effect of Flow Rate and Nitrate Concentration on Denitrification in Low-Permeability Media. J. Contam. Hydrol. 2020, 235, 103727. [Google Scholar] [CrossRef] [PubMed]
- Killingstad, M.W.; Widdowson, M.A.; Smith, R.L. Modeling Enhanced In Situ Denitrification in Groundwater. J. Environ. Eng. 2002, 128, 491–504. [Google Scholar] [CrossRef]
- Smith, R.L.; Miller, D.N.; Brooks, M.H.; Widdowson, M.A.; Killingstad, M.W. In Situ Stimulation of Groundwater Denitrification with Formate to Remediate Nitrate Contamination. Environ. Sci. Technol. 2001, 35, 196–203. [Google Scholar] [CrossRef]
- Kaelin, D.; Manser, R.; Rieger, L.; Eugster, J.; Rottermann, K.; Siegrist, H. Extension of ASM3 for Two-Step Nitrification and Denitrification and Its Calibration and Validation with Batch Tests and Pilot Scale Data. Water Res. 2009, 43, 1680–1692. [Google Scholar] [CrossRef]
- Koch, G.; Kühni, M.; Gujer, W.; Siegrist, H. Calibration and Validation of Activated Sludge Model No. 3 for Swiss Municipal Wastewater. Water Res. 2000, 34, 3580–3590. [Google Scholar] [CrossRef]
- Ni, B.-J.; Ruscalleda, M.; Pellicer-Nàcher, C.; Smets, B.F. Modeling Nitrous Oxide Production during Biological Nitrogen Removal via Nitrification and Denitrification: Extensions to the General ASM Models. Environ. Sci. Technol. 2011, 45, 7768–7776. [Google Scholar] [CrossRef]
- Ni, B.-J.; Yu, H.-Q. An Approach for Modeling Two-Step Denitrification in Activated Sludge Systems. Chem. Eng. Sci. 2008, 63, 1449–1459. [Google Scholar] [CrossRef]
- Lu, H.; Chandran, K.; Stensel, D. Microbial Ecology of Denitrification in Biological Wastewater Treatment. Water Res. 2014, 64, 237–254. [Google Scholar] [CrossRef] [PubMed]
- Perez-Garcia, O.; Mankelow, C.; Chandran, K.; Villas-Boas, S.G.; Singhal, N. Modulation of Nitrous Oxide (N2O) Accumulation by Primary Metabolites in Denitrifying Cultures Adapting to Changes in Environmental C and N. Environ. Sci. Technol. 2017, 51, 13678–13688. [Google Scholar] [CrossRef] [PubMed]
- Gujer, W.; Henze, M.; Mino, T.; van Loosdrecht, M. Activated Sludge Model No. 3. Water Sci. Technol. 1999, 39, 189–193. [Google Scholar] [CrossRef]
- Jenkins, C.; Ling, C.L.; Ciesielczuk, H.L.; Lockwood, J.; Hopkins, S.; McHugh, T.D.; Gillespie, S.H.; Kibbler, C.C. Detection and Identification of Bacteria in Clinical Samples by 16S RRNA Gene Sequencing: Comparison of Two Different Approaches in Clinical Practice. J. Med. Microbiol. 2012, 61, 483–488. [Google Scholar] [CrossRef] [PubMed]
- Graf, D.R.H.; Jones, C.M.; Hallin, S. Intergenomic Comparisons Highlight Modularity of the Denitrification Pathway and Underpin the Importance of Community Structure for N2O Emissions. PLoS ONE 2014, 9, e114118. [Google Scholar] [CrossRef] [Green Version]
- Baumann, B.; van der Meer, J.R.; Snozzi, M.; Zehnder, A.J.B. Inhibition of Denitrification Activity but Not of MRNA Induction in Paracoccus Denitrificans by Nitrite at a Suboptimal PH. Antonie Van Leeuwenhoek 1997, 72, 183–189. [Google Scholar] [CrossRef]
- Drtil, M.; Németh, P.; Kucman, K.; Bodík, I.; Kašperek, V. Acidobasic Balances in the Course of Heterotrophic Denitrification. Water Res. 1995, 29, 1353–1360. [Google Scholar] [CrossRef]
- Henry, S.; Baudoin, E.; López-Gutiérrez, J.C.; Martin-Laurent, F.; Brauman, A.; Philippot, L. Quantification of Denitrifying Bacteria in Soils by NirK Gene Targeted Real-Time PCR. J. Microbiol. Methods 2004, 59, 327–335. [Google Scholar] [CrossRef]
- Henze, M.; Grady, C.P.L.; Gujer, W.; Marais, G.V.R.; Matsuo, T. A General Model for Single-Sludge Wastewater Treatment Systems. Water Res. 1987, 21, 505–515. [Google Scholar] [CrossRef] [Green Version]
- Rivett, M.O.; Buss, S.R.; Morgan, P.; Smith, J.W.N.; Bemment, C.D. Nitrate Attenuation in Groundwater: A Review of Biogeochemical Controlling Processes. Water Res. 2008, 42, 4215–4232. [Google Scholar] [CrossRef] [PubMed]
- Parkin, T.B. Soil Microsites as a Source of Denitrification Variability 1. Soil Sci. Soc. Am. J. 1987, 51, 1194–1199. [Google Scholar] [CrossRef] [Green Version]
- Pan, Y.; Ni, B.-J.; Lu, H.; Chandran, K.; Richardson, D.; Yuan, Z. Evaluating Two Concepts for the Modelling of Intermediates Accumulation during Biological Denitrification in Wastewater Treatment. Water Res. 2015, 71, 21–31. [Google Scholar] [CrossRef] [PubMed] [Green Version]




| Reactor ID | Nitrate (mg-N/L) | Condition | EVO (mg/L) | Soil (g) | Water (mL) |
|---|---|---|---|---|---|
| R01 | 0.06 | control | 1216 | 50.9 | 502.1 |
| R02 | 17.97 | control | 0 | 51.0 | 501.6 |
| R03 | 17.79 | treatment | 1237 | 50.9 | 500.6 |
| R04 | 18.41 | treatment | 1096 | 50.9 | 502.2 |
| R05 | 17.55 | treatment | 1119 | 50.8 | 500.9 |
| RS1 | 0.03 | control | 817 | 51.1 | 501.5 |
| RS2 | 20.95 | control | 0 | 51.0 | 500.7 |
| RS3 | 20.55 | treatment | 1099 | 50.7 | 501.2 |
| RS4 | 21.24 | treatment | 1135 | 51.0 | 501.9 |
| RS5 | 18.25 | treatment | 1038 | 50.8 | 500.4 |
| Target pH | Reactor ID | Nitrate (mg-N/L) | EVO (mg/L) | Water (mL) | Soil (g) | Soil Type |
|---|---|---|---|---|---|---|
| 7.0 unadjusted | R10 | 6.8 | 3336 | 493.5 | 51.8 | OS |
| R11 | 5.3 | 3555 | 493.7 | 66.8 | SS | |
| 7.0 unadjusted | R70 | 22.2 | 3336 | 493.5 | 51.8 | OS |
| R71 | 22.2 | 3555 | 493.7 | 66.8 | SS | |
| 4.0 adjusted | R40 | 18.5 | 0 | 497.6 | 66.6 | SS |
| R41 | 15.0 | 422 | 497.4 | 66.9 | SS | |
| R41D | 15.6 | 221 | 497.6 | 69.2 | SS | |
| 4.0 unadjusted | R20 | 19.5 | 0 | 497.6 | 66.4 | SS |
| R21 | 18.7 | 1084 | 498.0 | 66.9 | SS | |
| R21D | 18.8 | 1065 | 497.8 | 67.2 | SS | |
| 8.0 adjusted | R80 | 17.1 | 0 | 248.5 | 32.2 | SS |
| R81 | 17.4 | 964 | 248.9 | 32 | SS | |
| R81D | 17.8 | 1247 | 248.7 | 32 | SS |
| Symbol | Characterization | Value | Units |
|---|---|---|---|
| N content of inert soluble COD SI | 0.01 | [g N/g COD] | |
| N content of readily biodegradable substrate | 0.03 | [g N/g COD] | |
| N content of inert particulate COD XI | 0.04 | [g N/g COD] | |
| N content of slowly biodegradable substrate | 0.03 | [g N/g COD] | |
| N content of biomass, XH, XA | 0.07 | [g N/g COD] | |
| Production of SI in hydrolysis | 0.00 | [g COD/g COD] | |
| Fraction of inert COD generated in biomass lysis | 0.2 | [g COD/g COD] | |
| YHO2 | Yield coeff. for heterotrophs in aerobic growth | 0.8 | [g COD/g COD] |
| YHNO3 | Yield coeff., heterotrophic anoxic growth using NO3 | 0.7 | [g COD/g COD] |
| YHNO2 | Yield coeff., heterotrophic anoxic growth using NO2 | 0.7 | [g COD/g COD] |
| YSTOO2 | Yield coeff. for XSTO in aerobic growth | 0.8 | [g COD/g COD] |
| YSTONO3 | Yield coeff. for XSTO in anoxic growth using NO3 | 0.7 | [g COD/g COD] |
| YSTONO2 | Yield coeff. for XSTO in anoxic growth using NO2 | 0.7 | [g COD/g COD] |
| Hydrolysis rate constant | 9 | [] | |
| Maximum storage rate | 12 | ] | |
| Maximum growth rate on substrate | 3 | ] | |
| Aerobic end. resp. rate for XH | 0.2 | ] [43] | |
| Aerobic end. resp. rate for XSTO | 0.3 | ] | |
| Reduction factor for NO3 reduction | 0.2 (0.15–0.25) | - | |
| Reduction factor for NO2 reduction | 0.2 (0.15–0.25) | - | |
| Reduction factor for bH using NO3 | 0.4 (0.25–0.50) | - | |
| Reduction factor for bH using NO2 | 0.5 (0.35–0.70) | - | |
| Reduction factor for bAOB and bNOB, anoxic | 0.1 | - | |
| Hydrolysis half saturation constant | 1.0 | ||
| Saturation/inhibition coeff. for oxygen, het. growth | 0.2 | ||
| Inhibition coefficient for oxygen, het. growth | 0.2 | ||
| Saturation coeff. for readily biodegradableSubstrates | 10 | ||
| Saturation/inhibition coefficient for ammonium | 0.01 | ||
| Saturation/inhibition coefficient for nitrate | 0.5 | ||
| Saturation/inhibition coefficient for nitrite | 0.5 | ||
| Inhibition coefficient for nitrite | 0.5 | ||
| Saturation coefficient for alkalinity | 0.1 | ||
| Saturation coefficient for storage products | 0.1 |
| Component | Symbol | RS1 | RS2 | RS3 | RS4 | RS5 |
|---|---|---|---|---|---|---|
| oxygen (mg/L) | SDO | 6 | 6 | 5 | 5 | 4 |
| nitrate (mg-N/L) | SNO3 | 0 | 20 | 20 | 20 | 20 |
| nitrite (mg-N/L) | SNO2 | 0 | 0 | 0 | 0 | 0 |
| nitrogen (mg-N/L) | SN2 | 0 | 0 | 0 | 0 | 0 |
| ammonia (mg-N/L) | SNH | 0 | 0 | 0 | 0 | 0 |
| EVO (mg-COD/L) | XS | 2860 | 0 | 2860 | 2860 | 2860 |
| soluble substrate (mg-COD/L) | SS | 80 | 0 | 80 | 80 | 80 |
| alkalinity (mole HCO3/m3) | Alk | 0 | 0 | 0 | 0 | 0 |
| particulate inert organic matter (mg-COD/L) | XI | 0 | 0 | 0 | 0 | 0 |
| storage (mg-COD/L) | XSTO | 0 | 0 | 0 | 0 | 0 |
| heterotrophic biomass (mg-COD/L) | XH | calibrated | ||||
| Parameter | Symbol | Fitted Value | Initial Guess | Literature Value |
|---|---|---|---|---|
| initial biomass (mg-COD/L) | not applicable | |||
| RS3 | 1.10 | 2 | ||
| RS4 | 2.56 | 2 | ||
| RS5 | 18.9 | 20 | ||
| nitrite inhibition (mg-N/L) | 0.0123 | 0.1 | 0.2 1 [42] | |
| half saturation for Ss (mg-COD/L) | 66.91 | 65 | 10 [43] | |
| 10–180 [54] | ||||
| 2 [48] | ||||
| hydrolysis rate constant (d−1) | 0.86 | 1 | 3 [43] | |
| 9 [42] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gonsalez, V.L.; Dombrowski, P.M.; Lee, M.D.; Ramsburg, C.A. The Influence of pH on Subsurface Denitrification Stimulated with Emulsified Vegetable Oil. Water 2023, 15, 883. https://doi.org/10.3390/w15050883
Gonsalez VL, Dombrowski PM, Lee MD, Ramsburg CA. The Influence of pH on Subsurface Denitrification Stimulated with Emulsified Vegetable Oil. Water. 2023; 15(5):883. https://doi.org/10.3390/w15050883
Chicago/Turabian StyleGonsalez, Veronica L., Paul M. Dombrowski, Michael D. Lee, and C. Andrew Ramsburg. 2023. "The Influence of pH on Subsurface Denitrification Stimulated with Emulsified Vegetable Oil" Water 15, no. 5: 883. https://doi.org/10.3390/w15050883
APA StyleGonsalez, V. L., Dombrowski, P. M., Lee, M. D., & Ramsburg, C. A. (2023). The Influence of pH on Subsurface Denitrification Stimulated with Emulsified Vegetable Oil. Water, 15(5), 883. https://doi.org/10.3390/w15050883






