Coprostanol as a Population Biomarker for SARS-CoV-2 Wastewater Surveillance Studies
Abstract
:1. Introduction
2. Methods
2.1. Sample Sites and Sample Collection
2.2. Composite Wastewater Influent Concentration and Nucleic Acid Extraction
2.3. qPCR Assays to Quantify SARS-CoV-2 RNA and Human Faecal Markers
2.4. Quantification of Coprostanol
2.5. Data Analysis
3. Results and Discussion
3.1. SARS-CoV-2 N1 RNA Can Be Quantified in Large and Medium WWTP Influent
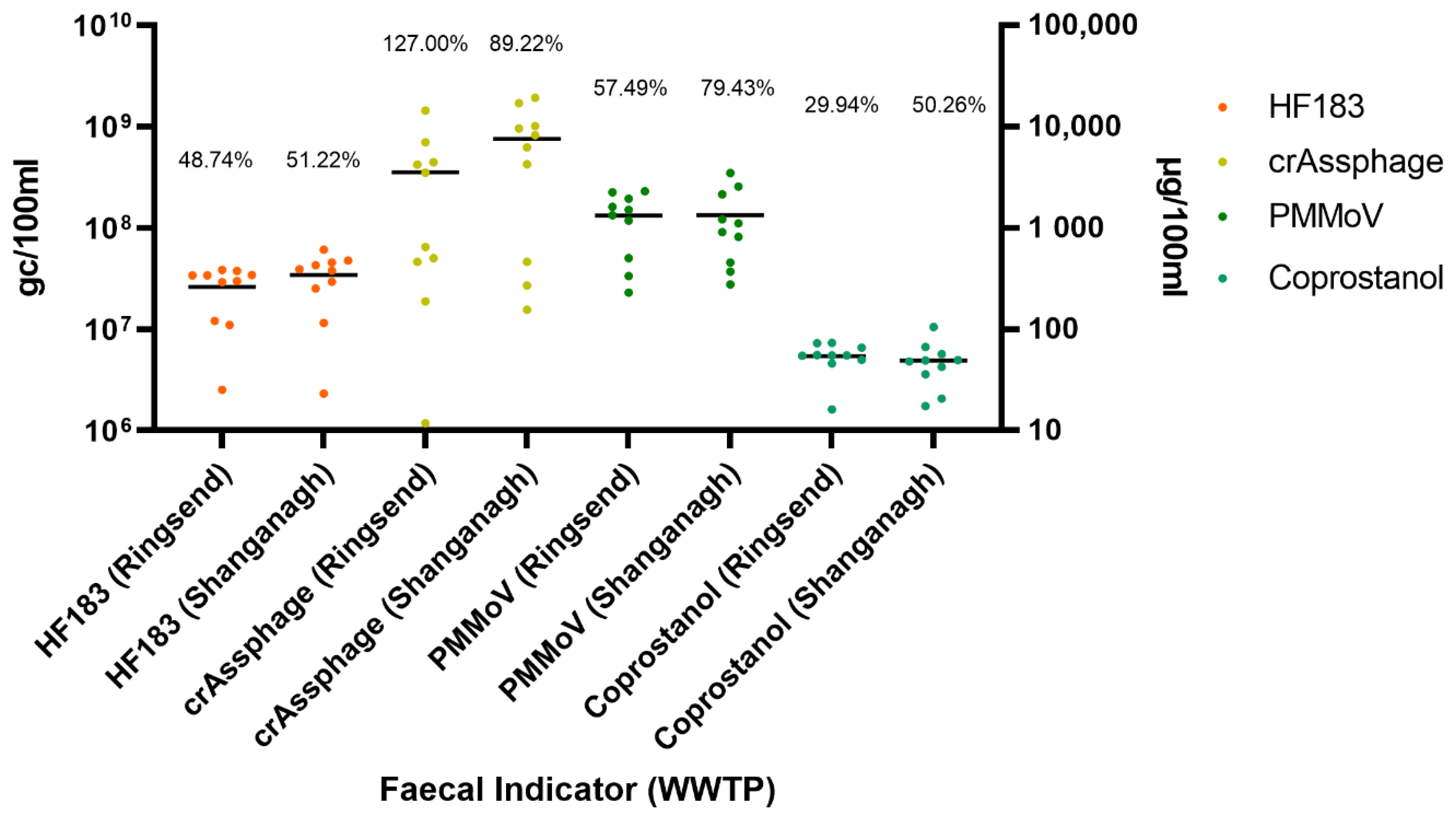
3.2. Faecal Indicators Are Abundant in Composite Influent
3.3. Coprostanol Concentrations Exhibit Less Variability than Viral and Bacterial Faecal Indicators
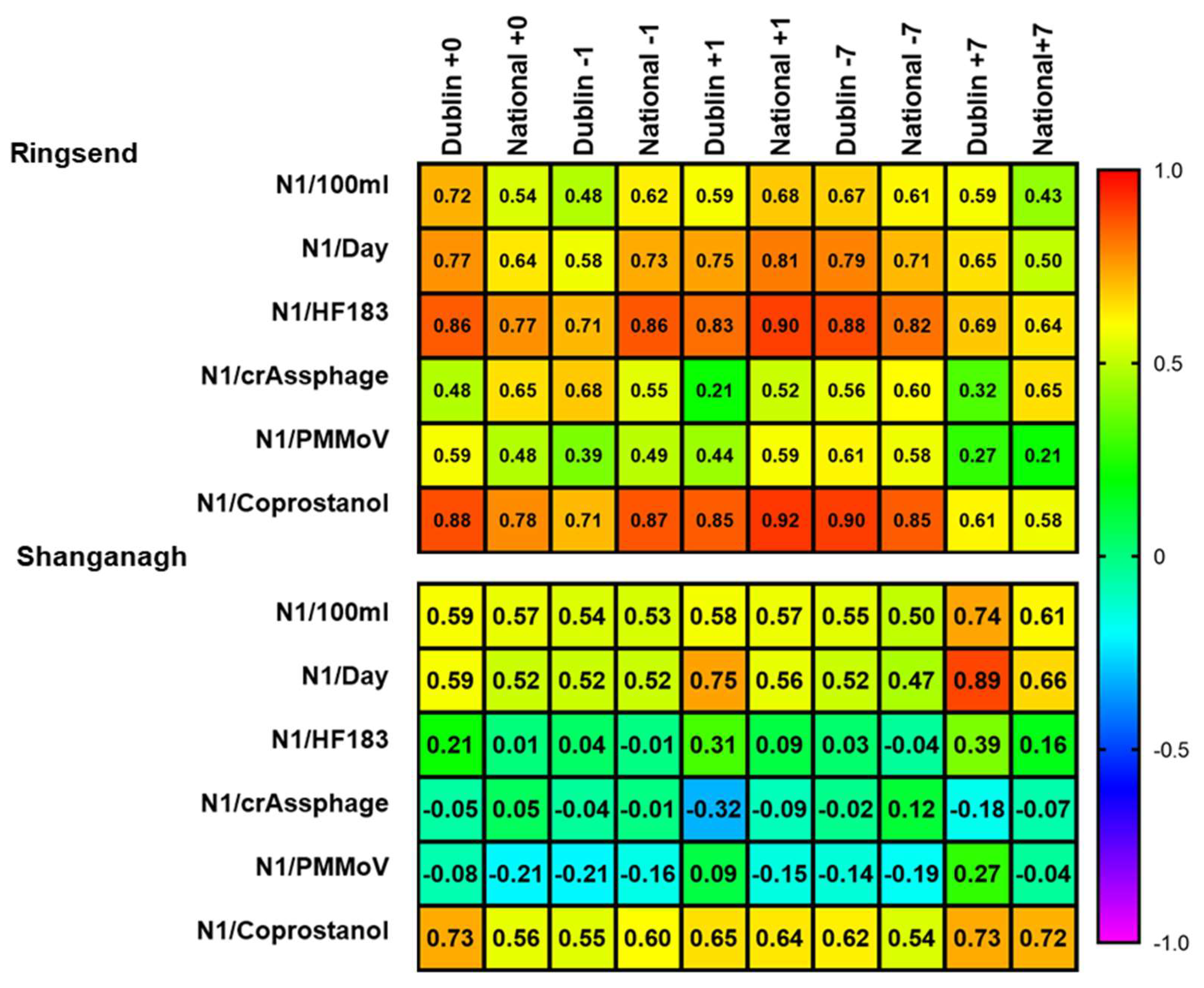
3.4. Coprostanol-Normalised SARS-CoV-2 N1 Levels Correlate with National and Dublin Case Numbers
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- WHO. Coronavirus Disease (COVID-19) Pandemic Dashboard; WHO: Geneva, Switzerland; Available online: https://covid19.who.int/ (accessed on 1 December 2021).
- Grant, M.C.; Geoghegan, L.; Arbyn, M.; Mohammed, Z.; McGuinness, L.; Clarke, E.L.; Wade, R.G. The prevalence of symptoms in 24,410 adults infected by the novel coronavirus (SARS-CoV-2; COVID-19): A systematic review and meta-analysis of 148 studies from 9 countries. PLoS ONE 2020, 15, e0234765. [Google Scholar] [CrossRef]
- Wölfel, R.; Corman, V.M.; Guggemos, W.; Seilmaier, M.; Zange, S.; Muller, M.A.; Niemeyer, D.; Jones, T.C.; Vollmar, P.; Rothe, C.; et al. Virological assessment of hospitalized patients with COVID-2019. Nature 2020, 581, 465–469. [Google Scholar] [CrossRef] [Green Version]
- Chen, Y.; Chen, L.; Deng, Q.; Zhang, G.; Wu, K.; Ni, L.; Yang, Y.; Liu, B.; Wang, W.; Wei, C.; et al. The presence of SARS-CoV-2 RNA in the feces of COVID-19 patients. J. Med. Virol. 2020, 92, 833–840. [Google Scholar] [CrossRef] [Green Version]
- Bivins, A.; Greaves, J.; Fischer, R.; Yinda, K.C.; Ahmed, W.; Kitajima, M.; Munster, V.J.; Bibby, K. Persistence of SARS-CoV-2 in Water and Wastewater. Environ. Sci. Technol. Lett. 2020, 7, 937–942. [Google Scholar] [CrossRef]
- Sala-Comorera, L.; Reynolds, L.J.; Martin, N.A.; O’Sullivan, J.J.; Meijer, W.G.; Fletcher, N.F. Decay of infectious SARS-CoV-2 and surrogates in aquatic environments. Water Res. 2021, 201, 117090. [Google Scholar] [CrossRef] [PubMed]
- Medema, G.; Heijnen, L.; Elsinga, G.; Italiaander, R.; Brouwer, A. Presence of SARS-Coronavirus-2 RNA in Sewage and Correlation with Reported COVID-19 Prevalence in the Early Stage of the Epidemic in The Netherlands. Environ. Sci. Technol. Lett. 2020, 7, 511–516. [Google Scholar] [CrossRef]
- Ahmed, W.; Angel, N.; Edson, J.; Bibby, K.; Bivins, A.; O’Brien, J.W.; Choi, P.M.; Kitajima, M.; Simpson, S.L.; Li, J.; et al. First confirmed detection of SARS-CoV-2 in untreated wastewater in Australia: A proof of concept for the wastewater surveillance of COVID-19 in the community. Sci. Total Environ. 2020, 728, 138764. [Google Scholar] [CrossRef]
- Hata, A.; Hara-Yamamura, H.; Meuchi, Y.; Imai, S.; Honda, R. Detection of SARS-CoV-2 in wastewater in Japan during a COVID-19 outbreak. Sci. Total Environ. 2021, 758, 143578. [Google Scholar] [CrossRef]
- Wurtzer, S.; Waldman, P.; Levert, M.; Mouchel, J.M.; Gorgé, O.; Boni, M.; Maday, Y.; Marechal, V.; Moulin, L. Monitoring the propagation of SARS CoV2 variants by tracking identified mutation in wastewater using specific RT-qPCR. medRxiv 2021. [Google Scholar] [CrossRef]
- Crits-Christoph, A.; Kantor, R.S.; Olm, M.R.; Whitney, O.N.; Al-Shayeb, B.; Lou, Y.C.; Flamholz, A.; Kennedy, L.C.; Greenwald, H.; Hinkle, A.; et al. Genome Sequencing of Sewage Detects Regionally Prevalent SARS-CoV-2 Variants. mBio 2021, 12, e02703–e02720. [Google Scholar] [CrossRef] [PubMed]
- Gunther, T.; Czech-Sioli, M.; Indenbirken, D.; Robitaille, A.; Tenhaken, P.; Exner, M.; Ottinger, M.; Fischer, N.; Grundhoff, A.; Brinkmann, M.M. SARS-CoV-2 outbreak investigation in a German meat processing plant. EMBO Mol. Med. 2020, 12, e13296. [Google Scholar] [CrossRef]
- Rennert, L.; McMahan, C.; Kalbaugh, C.A.; Yang, Y.; Lumsden, B.; Dean, D.; Pekarek, L.; Colenda, C.C. Surveillance-based informative testing for detection and containment of SARS-CoV-2 outbreaks on a public university campus: An observational and modelling study. Lancet Child Adolesc. Health 2021, 5, 428–436. [Google Scholar] [CrossRef]
- Daughton, C. The international imperative to rapidly and inexpensively monitor community-wide Covid-19 infection status and trends. Sci. Total Environ. 2020, 726, 138149. [Google Scholar] [CrossRef]
- Hamouda, M.; Mustafa, F.; Maraqa, M.; Rizvi, T.; Aly Hassan, A. Wastewater surveillance for SARS-CoV-2: Lessons learnt from recent studies to define future applications. Sci. Total Environ. 2021, 759, 143493. [Google Scholar] [CrossRef] [PubMed]
- Holcomb, D.A.; Stewart, J.R. Microbial Indicators of Fecal Pollution: Recent Progress and Challenges in Assessing Water Quality. Curr. Environ. Health Rep. 2020, 7, 311–324. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Aljaro, C.; Blanch, A.R.; Campos, C.; Jofre, J.; Lucena, F. Pathogens, faecal indicators and human-specific microbial source-tracking markers in sewage. J. Appl. Microbiol. 2019, 126, 701–717. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Haramoto, E.; Kitajima, M.; Kishida, N.; Konno, Y.; Katayama, H.; Asami, M.; Akiba, M. Occurrence of pepper mild mottle virus in drinking water sources in Japan. Appl. Environ. Microbiol. 2013, 79, 7413–7418. [Google Scholar] [CrossRef] [Green Version]
- Stachler, E.; Kelty, C.; Sivaganesan, M.; Li, X.; Bibby, K.; Shanks, O.C. Quantitative CrAssphage PCR Assays for Human Fecal Pollution Measurement. Environ. Sci. Technol. 2017, 51, 9146–9154. [Google Scholar] [CrossRef] [PubMed]
- Zhang, T.; Breitbart, M.; Lee, W.H.; Run, J.-Q.; Wei, C.L.; Soh, S.W.L.; Hibberd, M.L.; Liu, E.T.; Rohwer, F.; Ruan, Y. RNA Viral Community in Human Feces: Prevalence of Plant Pathogenic Viruses. PLoS Biol. 2005, 4, e3. [Google Scholar] [CrossRef] [Green Version]
- Ahmed, W.; Bivins, A.; Bertsch, P.M.; Bibby, K.; Gyawali, P.; Sherchan, S.P.; Simpson, S.L.; Thomas, K.V.; Verhagen, R.; Kitajima, M.; et al. Intraday variability of indicator and pathogenic viruses in 1-h and 24-h composite wastewater samples: Implications for wastewater-based epidemiology. Environ. Res. 2021, 193, 110531. [Google Scholar] [CrossRef]
- D’Aoust, P.M.; Mercier, É.; Montpetit, D.; Jia, J.-J.; Alexandrov, I.; Neault, N.; Baig, A.T.; Mayne, J.; Zhang, X.; Alain, T.; et al. Quantitative analysis of SARS-CoV-2 RNA from wastewater solids in communities with low COVID-19 incidence and prevalence. medRxiv 2020. [Google Scholar] [CrossRef] [PubMed]
- Reynolds, L.J.; Sala-Comorera, L.; Martin, N.A.; Nolan, T.M.; Stephens, J.H.; Gitto, A.; O’Hare, G.M.P.; O’Sullivan, J.J.; Meijer, W.G. Correlation between antimicrobial resistance and faecal contamination in small urban streams and bathing waters. Sci. Total Environ. 2020, 739, 140242. [Google Scholar] [CrossRef]
- Sala-Comorera, L.; Reynolds, L.J.; Martin, N.A.; Pascual-Benito, M.; Stephens, J.H.; Nolan, T.M.; Gitto, A.; O’Hare, G.M.P.; O’Sullivan, J.J.; García-Aljaro, C.; et al. crAssphage as a human molecular marker to evaluate temporal and spatial variability in faecal contamination of urban marine bathing waters. Sci. Total Environ. 2021, 789, 147828. [Google Scholar] [CrossRef]
- Leeming, R.; Ball, A.; Ashbolt, N.; Nichols, P. Using faecal sterols from humans and animals to distinguish faecal pollution in receiving waters. Water Res. 1996, 30, 2893–2900. [Google Scholar] [CrossRef]
- Daughton, C.G. Real-time estimation of small-area populations with human biomarkers in sewage. Sci. Total Environ. 2012, 414, 6–21. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rutledge, R.G.; Côté, C. Mathematics of quantitative kinetic PCR and the application of standard curves. Nucleic Acids Res. 2003, 31, e93. [Google Scholar] [CrossRef] [PubMed]
- Rutledge, R.G.; Stewart, D. Critical evaluation of methods used to determine amplification efficiency refutes the exponential character of real-time PCR. BMC Mol. Biol. 2008, 9, 96. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- CDC. 2019-Novel Coronavirus (2019-nCoV) Real-Time rRT-PCR Panel Primers and Probes. Available online: https://www.cdc.gov/coronavirus/2019-ncov/downloads/rt-pcr-panel-primer-probes.pdf (accessed on 1 July 2020).
- Seurinck, S.; Defoirdt, T.; Verstraete, W.; Siciliano, S.D. Detection and quantification of the human-specific HF183 Bacteroides 16S rRNA genetic marker with real-time PCR for assessment of human faecal pollution in freshwater. Environ. Microbiol. 2005, 7, 249–259. [Google Scholar] [CrossRef] [PubMed]
- Baz-Lomba, J.A.; Di Ruscio, F.; Amador, A.; Reid, M.; Thomas, K.V. Assessing Alternative Population Size Proxies in a Wastewater Catchment Area Using Mobile Device Data. Environ. Sci. Technol. 2019, 53, 1994–2001. [Google Scholar] [CrossRef]
- Korajkic, A.; McMinn, B.; Herrmann, M.P.; Sivaganesan, M.; Kelty, C.A.; Clinton, P.; Nash, M.S.; Shanks, O.C. Viral and Bacterial Fecal Indicators in Untreated Wastewater across the Contiguous United States Exhibit Geospatial Trends. Appl. Environ. Microbiol. 2020, 86, e02967-19. [Google Scholar] [CrossRef]
- Tandukar, S.; Sherchan, S.P.; Haramoto, E. Applicability of crAssphage, pepper mild mottle virus, and tobacco mosaic virus as indicators of reduction of enteric viruses during wastewater treatment. Sci. Rep. 2020, 10, 3616. [Google Scholar] [CrossRef] [Green Version]
- Daughton, C.G. Using biomarkers in sewage to monitor community-wide human health: Isoprostanes as conceptual prototype. Sci. Total Environ. 2012, 424, 16–38. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wu, F.; Zhang, J.; Xiao, A.; Gu, X.; Lee, W.L.; Armas, F.; Kauffman, K.; Hanage, W.; Matus, M.; Ghaeli, N.; et al. SARS-CoV-2 Titers in Wastewater Are Higher than Expected from Clinically Confirmed Cases. mSystems 2020, 5, e00614–e00620. [Google Scholar] [CrossRef]
- Graham, K.E.; Loeb, S.K.; Wolfe, M.K.; Catoe, D.; Sinnott-Armstrong, N.; Kim, S.; Yamahara, K.M.; Sassoubre, L.M.; Mendoza Grijalva, L.M.; Roldan-Hernandez, L.; et al. SARS-CoV-2 RNA in Wastewater Settled Solids Is Associated with COVID-19 Cases in a Large Urban Sewershed. Environ. Sci. Technol. 2021, 55, 488–498. [Google Scholar] [CrossRef] [PubMed]
- Haramoto, E.; Malla, B.; Thakali, O.; Kitajima, M. First environmental surveillance for the presence of SARS-CoV-2 RNA in wastewater and river water in Japan. Sci. Total Environ. 2020, 737, 140405. [Google Scholar] [CrossRef]
- Green, H.; Wilder, M.; Collins, M.; Fenty, A.; Gentile, K.; Kmush, B.L.; Zeng, T.; Middleton, F.A.; Larsen, D.A. Quantification of SARS-CoV-2 and cross-assembly phage (crAssphage) from wastewater to monitor coronavirus transmission within communities. medRxiv 2020. [Google Scholar] [CrossRef]
- Bivins, A.; Crank, K.; Greaves, J.; North, D.; Wu, Z.; Bibby, K. Cross-assembly phage and pepper mild mottle virus as viral water quality monitoring tools—potential, research gaps, and way forward. Curr. Opin. Environ. Sci. Health 2020, 16, 54–61. [Google Scholar] [CrossRef]
- Lyons, B.P.; Devlin, M.J.; Abdul Hamid, S.A.; Al-Otiabi, A.F.; Al-Enezi, M.; Massoud, M.S.; Al-Zaidan, A.S.; Smith, A.J.; Morris, S.; Bersuder, P.; et al. Microbial water quality and sedimentary faecal sterols as markers of sewage contamination in Kuwait. Mar. Pollut. Bull. 2015, 100, 689–698. [Google Scholar] [CrossRef]
- Carreon-Palau, L.; Parrish, C.C.; Perez-Espana, H. Urban sewage lipids in the suspended particulate matter of a coral reef under river influence in the South West Gulf of Mexico. Water Res. 2017, 123, 192–205. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Kostakis, C.; Gerber, J.P.; Tscharke, B.J.; Irvine, R.J.; White, J.M. Towards finding a population biomarker for wastewater epidemiology studies. Sci. Total Environ. 2014, 487, 621–628. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z.; Xu, G.; Reboud, J.; Kasprzyk-Hordern, B.; Cooper, J.M. Monitoring Genetic Population Biomarkers for Wastewater-Based Epidemiology. Anal. Chem. 2017, 89, 9941–9945. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Daughton, C.G. Monitoring wastewater for assessing community health: Sewage Chemical-Information Mining (SCIM). Sci. Total Environ. 2018, 619–620, 748–764. [Google Scholar] [CrossRef] [PubMed]
- Maldonado, C.; Venkatesan, M.I.; Phillips, C.R.; Bayona, J.M. Distribution of Trialkylamines and Coprostanol in San Pedro Shelf Sediments Adjacent to a Sewage Outfall. Mar. Pollut. Bull. 2000, 40, 680–687. [Google Scholar] [CrossRef]
- Kitajima, M.; Sassi, H.P.; Torrey, J.R. Pepper mild mottle virus as a water quality indicator. NPJ Clean Water 2018, 1, 19. [Google Scholar] [CrossRef] [Green Version]
- Ballesté, E.; Demeter, K.; Masterson, B.; Timoneda, N.; Sala-Comorera, L.; Meijer, W.G. Implementation and integration of microbial source tracking in a river watershed monitoring plan. Sci. Total Environ. 2020, 736, 139573. [Google Scholar] [CrossRef]
- Weidhaas, J.; Aanderud, Z.T.; Roper, D.K.; VanDerslice, J.; Gaddis, E.B.; Ostermiller, J.; Hoffman, K.; Jamal, R.; Heck, P.; Zhang, Y.; et al. Correlation of SARS-CoV-2 RNA in wastewater with COVID-19 disease burden in sewersheds. Sci. Total Environ. 2021, 775, 145790. [Google Scholar] [CrossRef]
- Peccia, J.; Zulli, A.; Brackney, D.E.; Grubaugh, N.D.; Kaplan, E.H.; Casanovas-Massana, A.; Ko, A.I.; Malik, A.A.; Wang, D.; Wang, M.; et al. Measurement of SARS-CoV-2 RNA in wastewater tracks community infection dynamics. Nat. Biotechnol. 2020, 38, 1164–1167. [Google Scholar] [CrossRef] [PubMed]
- Nemudryi, A.; Nemudraia, A.; Wiegand, T.; Surya, K.; Buyukyoruk, M.; Cicha, C.; Vanderwood, K.K.; Wilkinson, R.; Wiedenheft, B. Temporal Detection and Phylogenetic Assessment of SARS-CoV-2 in Municipal Wastewater. Cell Rep. Med. 2020, 1, 100098. [Google Scholar] [CrossRef]
- Wolfe, M.K.; Archana, A.; Catoe, D.; Coffman, M.M.; Dorevich, S.; Graham, K.E.; Kim, S.; Grijalva, L.M.; Roldan-Hernandez, L.; Silverman, A.I.; et al. Scaling of SARS-CoV-2 RNA in Settled Solids from Multiple Wastewater Treatment Plants to Compare Incidence Rates of Laboratory-Confirmed COVID-19 in Their Sewersheds. Environ. Sci. Technol. Lett. 2021, 8, 398–404. [Google Scholar] [CrossRef]
| Target Gene | Primer Sequence | Cycling Conditions | Amplicon Size (Standard) | Slope/y-Intercept/R2 | Reaction LoD (LoQ) | Reference |
|---|---|---|---|---|---|---|
| SARS-CoV-2 N1 | F: GACCCCAAAATCAGCGAAAT R: TCTGGTTACTGCCAGTTGAATCTG Probe: FAM-ACCCCGCATTACGTTTGGTGGACC-BHQ1 | RT (50 °C-600 s), 95 °C-30 s, 45 cycles (95 °C-5 s, 60 °C-30 s) | 73 bp (2019-nCoV_N_Positive Control Plasmid, Integrated DNA Technologies, Coralville, IA, USA) | −3.2177/ 37.82/0.99 | 5 gc/µL (5 gc/µL) | [29] |
| HF183 | F: ATCATGAGTTCACATGTCCG R: TACCCCGCCTACTATCTAATG | 45 cycles (95 °C-5 s, 60 °C-15 s, 72 °C-20 s) | 82 bp (HF183 Amplicon cloned into pBLUE Plasmid, linear, Addgene, Watertown, MA, USA) | −3.4902/ 40.2/1.00 | 1.1 gc/µL (2.2 gc/µL) | [30] |
| crAssphage | F: CAGAAGTACAAACTCCTAAAAAACGTAGAG R: GATGACCAATAAACAAGCCATTAGC Probe: FAM-AATAACGATTTACGTGATGTAAC-MGB | 95 °C-600 s, 40 cycles (95 °C-15 s, 60 °C-60 s) | 126 bp (gBlock, Integrated DNA Technologies, Coralville, IA, USA) | −3.398/ 37.01/1.00 | 4.7 gc/µL (4.7 gc/µL) | [19] |
| PMMoV | F: GAGTGGTTTGACCTTAACGTTTGA R: TTGTCGGTTGCAATGCAAGT Probe:FAM-CCTACCGAAGCAAATG-BHQ1 | RT (50 °C-600 s), 95 °C-30 s, 45 cycles (95 °C-15 s, 60 °C-60 s) | 68 bp (gBlock, Integrated DNA Technologies, Coralville, IA, USA) | −3.2312/ 46.76/0.99 | 160 gc/µL (160 gc/µL) | [18,20] |
| Wastewater Treatment Plant | SARS-CoV-2 RNA Concentration (gc/100 mL) | SARS-CoV-2 RNA Daily Loading (gc/day) | SARS-CoV-2 RNA Daily Loading Per Person (gc/day/person) | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Median | Min. | Max. | CV | Median | Min. | Max. | CV | Median | Min. | Max. | CV | |
| Ringsend | 4355 | 1450 | 12,100 | 65.93% | 2.32 × 1013 | 9.24 × 1012 | 7.95 × 1013 | 82.18% | 1.17 × 107 | 4.67 × 106 | 4.01 × 107 | 82.24% |
| Shanganagh | 1890 | 552 | 8540 | 99.56% | 8.25 × 1011 | 2.94 × 1011 | 4.48 × 1012 | 107.7% | 3.32 × 106 | 1.19 × 106 | 1.81 × 107 | 107.9% |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Reynolds, L.J.; Sala-Comorera, L.; Khan, M.F.; Martin, N.A.; Whitty, M.; Stephens, J.H.; Nolan, T.M.; Joyce, E.; Fletcher, N.F.; Murphy, C.D.; et al. Coprostanol as a Population Biomarker for SARS-CoV-2 Wastewater Surveillance Studies. Water 2022, 14, 225. https://doi.org/10.3390/w14020225
Reynolds LJ, Sala-Comorera L, Khan MF, Martin NA, Whitty M, Stephens JH, Nolan TM, Joyce E, Fletcher NF, Murphy CD, et al. Coprostanol as a Population Biomarker for SARS-CoV-2 Wastewater Surveillance Studies. Water. 2022; 14(2):225. https://doi.org/10.3390/w14020225
Chicago/Turabian StyleReynolds, Liam J., Laura Sala-Comorera, Mohd Faheem Khan, Niamh A. Martin, Megan Whitty, Jayne H. Stephens, Tristan M. Nolan, Eadaoin Joyce, Nicola F. Fletcher, Cormac D. Murphy, and et al. 2022. "Coprostanol as a Population Biomarker for SARS-CoV-2 Wastewater Surveillance Studies" Water 14, no. 2: 225. https://doi.org/10.3390/w14020225








