The Genetic Component of Seagrass Restoration: What We Know and the Way Forwards
Abstract
1. Introduction
2. A Brief Glance at Factors Shaping Genetic Diversity and Population Structure in Seagrasses
2.1. Reproductive Strategies, Mutations
2.2. Level of Genetic Connectivity, Population Size, and Genetic Drift
2.3. Phenotypic Plasticity and Local Adaptation
2.4. Disturbances
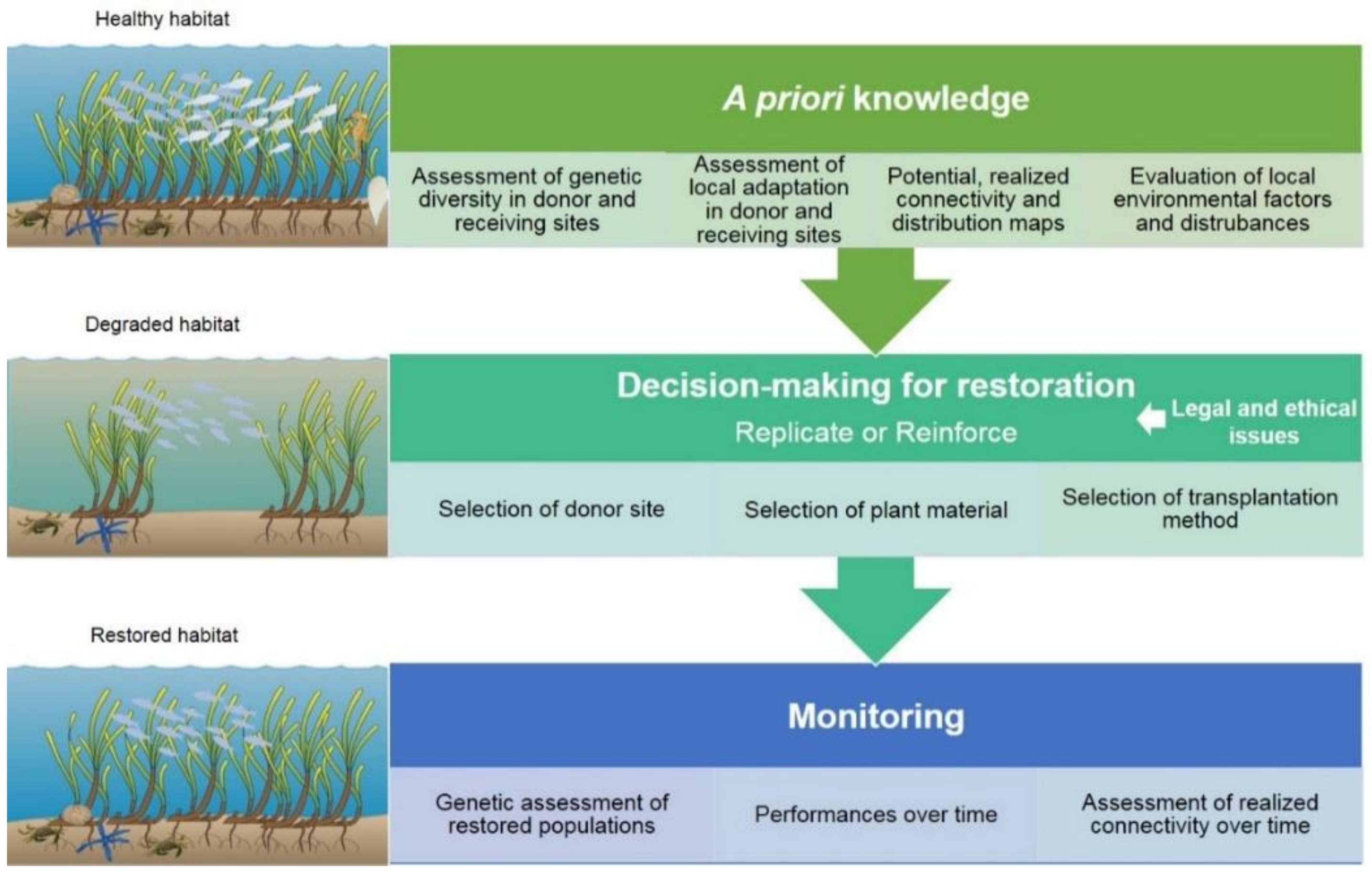
3. Integration of Genetic Research into Seagrass Restoration
3.1. Selection of Donor Sites
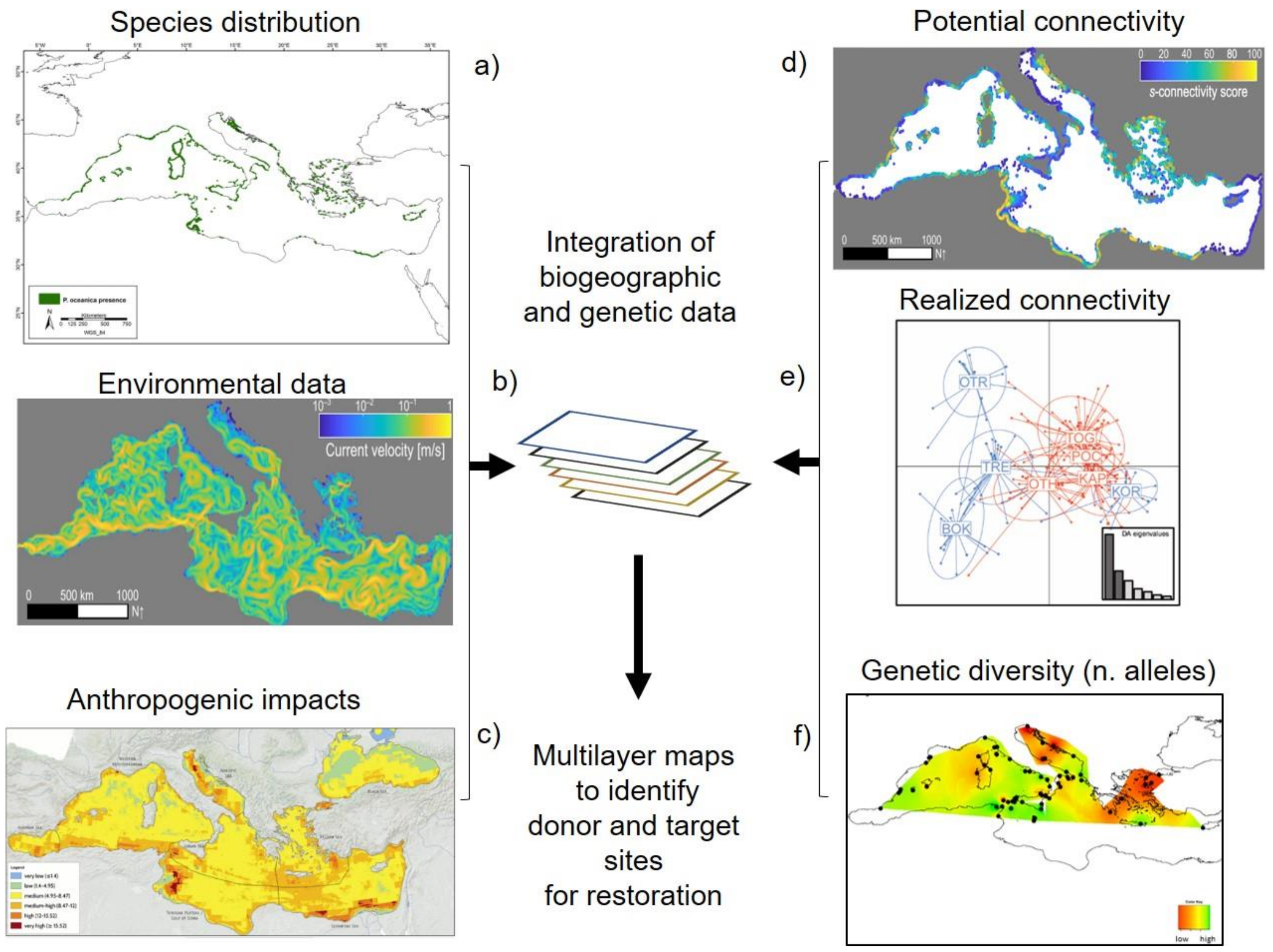
3.2. Integration of Biogeographic and Genetic Data
3.3. Selection of the Plant Material
3.4. Genetic Assessment of Transplantation Success
4. Future Directions in Seagrass Restoration
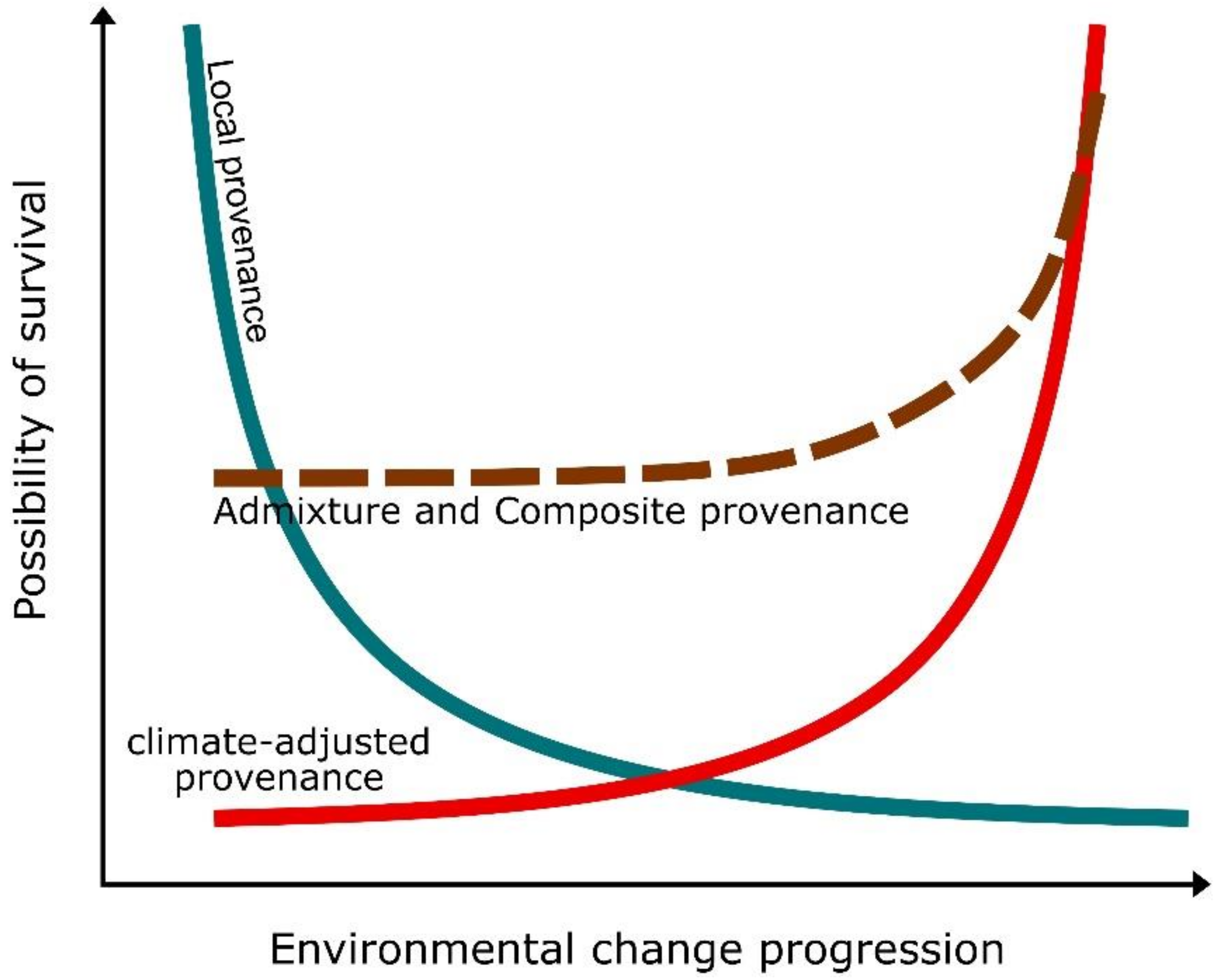
4.1. Improving Transplant Performances through Assisted Evolution
4.2. Potential of Epigenetics in Seagrass Restoration
5. Legal and Ethical Issues Related to Genetic Aspects of Seagrass Restoration
6. Recommendations and Conclusions
Author Contributions
Funding
Conflicts of Interest
Glossary
References
- Ceballos, G.; Ehrlich, P.R.; Barnosky, A.D.; García, A.; Pringle, R.M.; Palmer, T.M. Accelerated modern human-induced species losses: Entering the sixth mass extinction. Science 2015, 1, e1400253. [Google Scholar] [CrossRef]
- Hobbs, R.J.; Higgs, E.S.; Hall, C.M. Expanding the Portfolio: Conserving Nature’s Masterpieces in a Changing World. Bioscience 2017, 67, 568–575. [Google Scholar] [CrossRef]
- Perring, M.P.; Standish, R.J.; Price, J.N.; Craig, M.D.; Erickson, T.E.; Ruthrof, K.X.; Whiteley, A.S.; Valentine, L.E.; Hobbs, R.J. Advances in restoration ecology: Rising to the challenges of the coming decades. Ecosphere 2015, 6, 1–25. [Google Scholar] [CrossRef]
- Benayas, J.M.R.; Newton, A.C.; Diaz, A.; Bullock, J.M. Enhancement of biodiversity and ecosystem services by ecological restoration: A meta-analysis. Science 2009, 325, 1121–1124. [Google Scholar] [CrossRef] [PubMed]
- Reynolds, L.K.; Waycott, M.; McGlathery, K.J.; Orth, R.J. Ecosystem services returned through seagrass restoration. Restor. Ecol. 2016, 24, 583–588. [Google Scholar] [CrossRef]
- Wood, G.; Marzinelli, E.M.; Coleman, M.A.; Campbell, A.H.; Santini, N.S.; Kajlich, L.; Verdura, J.; Wodak, J.; Steinberg, P.D.; Vergés, A. Restoring subtidal marine macrophytes in the Anthropocene: Trajectories and future-proofing. Mar. Freshw. Res. 2019, 70, 936–951. [Google Scholar] [CrossRef]
- Boström-Einarsson, L.; Babcock, R.C.; Bayraktarov, E.; Ceccarelli, D.; Cook, N.; Ferse, S.C.A.; Hancock, B.; Harrison, P.; Hein, M.; Shaver, E. Coral restoration–A systematic review of current methods, successes, failures and future directions. PLoS ONE 2020, 15, e0226631. [Google Scholar] [CrossRef]
- Duarte, C.M.; Agusti, S.; Barbier, E.; Britten, G.L.; Castilla, J.C.; Gattuso, J.-P.; Fulweiler, R.W.; Hughes, T.P.; Knowlton, N.; Lovelock, C.E. Rebuilding marine life. Nature 2020, 580, 39–51. [Google Scholar] [CrossRef]
- Layton, C.; Coleman, M.A.; Marzinelli, E.M.; Steinberg, P.D.; Swearer, S.E.; Vergés, A.; Wernberg, T.; Johnson, C.R. Kelp forest restoration in Australia. Front. Mar. Sci. 2020, 7, 74. [Google Scholar] [CrossRef]
- Tan, Y.M.; Dalby, O.; Kendrick, G.A.; Statton, J.; Sinclair, E.A.; Fraser, M.W.; Macreadie, P.I.; Gillies, C.L.; Coleman, R.A.; Waycott, M. Seagrass restoration is possible: Insights and lessons from Australia and New Zealand. Front. Mar. Sci. 2020, 7, 617. [Google Scholar] [CrossRef]
- Evans, S.M.; Sinclair, E.A.; Poore, A.G.B.; Bain, K.F.; Vergés, A. Assessing the effect of genetic diversity on the early establishment of the threatened seagrass Posidonia australis using a reciprocal-transplant experiment. Restor. Ecol. 2018, 26, 570–580. [Google Scholar] [CrossRef]
- Short, F.T.; Carruthers, T.; Dennison, W.; Waycott, M. Global seagrass distribution and diversity: A bioregional model. J. Exp. Mar. Bio. Ecol. 2007, 350, 3–20. [Google Scholar] [CrossRef]
- Bertelli, C.M.; Unsworth, R.K.F. Protecting the hand that feeds us: Seagrass (Zostera marina) serves as commercial juvenile fish habitat. Mar. Pollut. Bull. 2014, 83, 425–429. [Google Scholar] [CrossRef] [PubMed]
- Nordlund, L.M.; Jackson, E.L.; Nakaoka, M.; Samper-Villarreal, J.; Beca-Carretero, P.; Creed, J.C. Seagrass ecosystem services—What’s next? Mar. Pollut. Bull. 2018, 134, 145–151. [Google Scholar] [CrossRef] [PubMed]
- Costanza, R.; De Groot, R.; Sutton, P.; Van der Ploeg, S.; Anderson, S.J.; Kubiszewski, I.; Farber, S.; Turner, R.K. Changes in the global value of ecosystem services. Glob. Environ. Chang. 2014, 26, 152–158. [Google Scholar] [CrossRef]
- Buia, M.C.; Mazzella, L. Reproductive phenology of the Mediterranean seagrasses Posidonia oceanica (L.) Delile, Cymodocea nodosa (Ucria) Aschers. and Zostera noltii Hornem. Aquat. Bot. 1991, 40, 343–362. [Google Scholar] [CrossRef]
- Jahnke, M.; Olsen, J.L.; Procaccini, G. A meta-analysis reveals a positive correlation between genetic diversity metrics and environmental status in the long-lived seagrass Posidonia oceanica. Mol. Ecol. 2015, 24, 2336–2348. [Google Scholar] [CrossRef]
- Paulo, D.; Diekmann, O.; Ramos, A.A.; Alberto, F.; Serrão, E.A. Sexual reproduction vs. Clonal propagation in the recovery of a seagrass meadow after an extreme weather event. Sci. Mar. 2019, 83, 357–363. [Google Scholar] [CrossRef]
- Marín-Guirao, L.; Entrambasaguas, L.; Ruiz, J.M.; Procaccini, G. Heat-stress induced flowering can be a potential adaptive response to ocean warming for the iconic seagrass Posidonia oceanica. Mol. Ecol. 2019, 28, 1–16. [Google Scholar] [CrossRef]
- Pazzaglia, J.; Reusch, T.B.H.; Terlizzi, A.; Marin Guirao, L.; Procaccini, G. Prompt phenotypic plasticity under rapid global changes: The intrinsic force for future seagrasses survival. Evol. Appl. 2021, in press. [Google Scholar] [CrossRef]
- Arnaud-Haond, S.; Duarte, C.M.; Diaz-Almela, E.; Marbà, N.; Sintes, T.; Serrão, E.A. Implications of extreme life span in clonal organisms: Millenary clones in meadows of the threatened seagrass posidonia oceanica. PLoS ONE 2012, 7. [Google Scholar] [CrossRef]
- Migliaccio, M.; De Martino, F.; Silvestre, F.; Procaccini, G. Meadow-scale genetic structure in Posidonia oceanica. Mar. Ecol. Prog. Ser. 2005, 304, 55–65. [Google Scholar] [CrossRef]
- Vallejo-Marín, M.; Dorken, M.E.; Barrett, S.C.H. The Ecological and Evolutionary Consequences of Clonality for Plant Mating. Annu. Rev. Ecol. Evol. Syst. 2010, 41, 193–213. [Google Scholar] [CrossRef]
- Ruocco, M.; Entrambasaguas, L.; Dattolo, E.; Milito, A.; Marín-Guirao, L.; Procaccini, G. A king and vassals’ tale: Molecular signatures of clonal integration in Posidonia oceanica under chronic light shortage. J. Ecol. 2020, 109, 294–312. [Google Scholar] [CrossRef]
- Procaccini, G.; Mazzella, L. Population genetic structure and gene flow in the seagrass Posidonia oceanica assessed using microsatellite analysis. Mar. Ecol. Prog. Ser. 1998, 169, 133–141. [Google Scholar] [CrossRef]
- Alberto, F.; Mata, L.; Santos, R. Genetic homogeneity in the seagrass Cymodocea nodosa at its northern Atlantic limit revealed through RAPD. Mar. Ecol. Prog. Ser. 2001, 221, 299–301. [Google Scholar] [CrossRef][Green Version]
- Procaccini, G.; Orsini, L.; Ruggiero, M.V.; Scardi, M. Spatial patterns of genetic diversity in Posidonia oceanica, an endemic Mediterranean seagrass. Mol. Ecol. 2001, 10, 1413–1421. [Google Scholar] [CrossRef] [PubMed]
- Marbà, N.; Duarte, C.M. Mediterranean warming triggers seagrass (Posidonia oceanica) shoot mortality. Glob. Chang. Biol. 2010, 16, 2366–2375. [Google Scholar] [CrossRef]
- Strydom, S.; Murray, K.; Wilson, S.; Huntley, B.; Rule, M.; Heithaus, M.; Bessey, C.; Kendrick, G.A.; Burkholder, D.; Holmes, T.; et al. Too hot to handle: Unprecedented seagrass death driven by marine heatwave in a World Heritage Area. Glob. Chang. Biol. 2020, 26. [Google Scholar] [CrossRef]
- Waycott, M.; Duarte, C.M.; Carruthers, T.J.B.; Orth, R.J.; Dennison, W.C.; Olyarnik, S.; Calladine, A.; Fourqurean, J.W.; Heck, K.L.; Hughes, A.R. Accelerating loss of seagrasses across the globe threatens coastal ecosystems. Proc. Natl. Acad. Sci. USA 2009, 106, 12377–12381. [Google Scholar] [CrossRef] [PubMed]
- Short, F.T.; Polidoro, B.; Livingstone, S.R.; Carpenter, K.E.; Bandeira, S.; Bujang, J.S.; Calumpong, H.P.; Carruthers, T.J.B.; Coles, R.G.; Dennison, W.C. Extinction risk assessment of the world’s seagrass species. Biol. Conserv. 2011, 144, 1961–1971. [Google Scholar] [CrossRef]
- Pazzaglia, J.; Santillán-Sarmiento, A.; Helber, S.B.; Ruocco, M.; Terlizzi, A.; Marín-Guirao, L.; Procaccini, G. Does Warming Enhance the Effects of Eutrophication in the Seagrass Posidonia oceanica? Front. Mar. Sci. 2020, 7, 1067. [Google Scholar] [CrossRef]
- Ruocco, M.; Marín-Guirao, L.; Ravaglioli, C.; Bulleri, F.; Procaccini, G. Molecular level responses to chronic versus pulse nutrient loading in the seagrass Posidonia oceanica undergoing herbivore pressure. Oecologia 2018, 188, 23–39. [Google Scholar] [CrossRef] [PubMed]
- Orth, R.J.; Carruthers, T.J.B.; Dennison, W.C.; Duarte, C.M.; Fourqurean, J.W.; Heck, K.L.; Hughes, A.R.; Kendrick, G.A.; Kenworthy, W.J.; Olyarnik, S. A global crisis for seagrass ecosystems. AIBS Bull. 2006, 56, 987–996. [Google Scholar] [CrossRef]
- Van Oppen, M.J.H.; Gates, R.D.; Blackall, L.L.; Cantin, N.; Chakravarti, L.J.; Chan, W.Y.; Cormick, C.; Crean, A.; Damjanovic, K.; Epstein, H. Shifting paradigms in restoration of the world’s coral reefs. Glob. Chang. Biol. 2017, 23, 3437–3448. [Google Scholar] [CrossRef]
- Coleman, M.A.; Wood, G.; Filbee-Dexter, K.; Minne, A.J.P.; Goold, H.D.; Vergés, A.; Marzinelli, E.M.; Steinberg, P.D.; Wernberg, T. Restore or redefine: Future trajectories for restoration. Front. Mar. Sci. 2020, 7, 237. [Google Scholar] [CrossRef]
- Abelson, A.; Reed, D.C.; Edgar, G.J.; Smith, C.S.; Kendrick, G.A.; Orth, R.J.; Airoldi, L.; Silliman, B.; Beck, M.W.; Krause, G. Challenges for restoration of coastal marine ecosystems in the Anthropocene. Front. Mar. Sci. 2020, 7, 892. [Google Scholar] [CrossRef]
- Fitzpatrick, M.C.; Keller, S.R. Ecological genomics meets community-level modelling of biodiversity: Mapping the genomic landscape of current and future environmental adaptation. Ecol. Lett. 2014, 18, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Eisele, F.; Seockhwan Hwang, B. New UN Decade on Ecosystem Restoration offers unparalleled opportunity for job creation, food security and addressing climate change. N. For. 2019, 50, 139–151. [Google Scholar]
- Hughes, A.R.; Inouye, B.D.; Johnson, M.T.J.; Underwood, N.; Vellend, M. Ecological consequences of genetic diversity. Ecol. Lett. 2008, 11, 609–623. [Google Scholar] [CrossRef]
- Ferber, S.; Stam, W.T.; Olsen, J.L. Genetic diversity and connectivity remain high in eelgrass Zostera marina populations in the Wadden Sea, despite major impacts. Mar. Ecol. Prog. Ser. 2008, 372, 87–96. [Google Scholar] [CrossRef]
- Bricker, E.; Waycott, M.; Calladine, A.; Zieman, J.C. High connectivity across environmental gradients and implications for phenotypic plasticity in a marine plant. Mar. Ecol. Prog. Ser. 2011, 423, 57–67. [Google Scholar] [CrossRef]
- Larkum, A.W.D.; Orth, R.J.; Duarte, C.M. Seagrasses: Biology, ecology and conservation. Seagrasses Biol. Ecol. Conserv. 2006, 1–691. [Google Scholar] [CrossRef]
- Den Hartog, C. Sea-Grasses of the World; North-Holland Publication Co.: Amsterdam, The Netherlands, 1970. [Google Scholar]
- Les, D.H.; Cleland, M.A.; Waycott, M. Phylogenetic studies in Alismatidae, II: Evolution of marine angiosperms (seagrasses) and hydrophily. Syst. Bot. 1997, 22, 443–463. [Google Scholar] [CrossRef]
- Reusch, T.B.H. Five microsatellite loci in eelgrass Zostera marina and a test of cross-species amplification in Z. noltii and Z. japonica. Mol. Ecol. 2000, 9, 371–373. [Google Scholar] [CrossRef] [PubMed]
- Ruggiero, M.V.; Capone, S.; Pirozzi, P.; Reusch, T.B.H.; Procaccini, G. Mating system and clonal architecture: A comparative study in two marine angiosperms. Evol. Ecol. 2005, 19, 487–499. [Google Scholar] [CrossRef]
- Zipperle, A.M.; Coyer, J.A.; Reise, K.; Stam, W.T.; Olsen, J.L. An evaluation of small-scale genetic diversity and the mating system in Zostera noltii on an intertidal sandflat in the Wadden Sea. Ann. Bot. 2011, 107, 127–134. [Google Scholar] [CrossRef] [PubMed]
- Evans, S.M.; Sinclair, E.A.; Poore, A.G.B.; Steinberg, P.D.; Kendrick, G.A.; Vergés, A. Genetic diversity in threatened Posidonia australis seagrass meadows. Conserv. Genet. 2014, 15, 717–728. [Google Scholar] [CrossRef]
- Arnaud-Haond, S.; Stoeckel, S.; Bailleul, D. New insights into the population genetics of partially clonal organisms: When seagrass data meet theoretical expectations. Mol. Ecol. 2020, 29, 3248–3260. [Google Scholar] [CrossRef]
- Reusch, T.B.H.; Boström, C. Widespread genetic mosaicism in the marine angiosperm Zostera marina is correlated with clonal reproduction. Evol. Ecol. 2011, 25, 899–913. [Google Scholar] [CrossRef]
- Harper, J.L. Population Biology of Plants; CABI: Wallingford, UK, 1977. [Google Scholar]
- Yu, L.; Boström, C.; Franzenburg, S.; Bayer, T.; Dagan, T.; Reusch, T.B.H. Somatic genetic drift and multi-level selection in modular species. Nat. Ecol. Evol. 2020, 4, 952–962. [Google Scholar] [CrossRef]
- Kendrick, G.A.; Orth, R.J.; Statton, J.; Hovey, R.; Montoya, L.R.; Lowe, R.J.; Krauss, S.L.; Sinclair, E.A. Demographic and genetic connectivity: The role and consequences of reproduction, dispersal and recruitment in seagrasses. Biol. Rev. 2017, 92, 921–938. [Google Scholar] [CrossRef]
- Tomasello, A.; Di Maida, G.; Calvo, S.; Pirrotta, M.; Borra, M.; Procaccini, G. Seagrass meadows at the extreme of environmental tolerance: The case of Posidonia oceanica in a semi-enclosed coastal lagoon. Mar. Ecol. 2009, 30, 288–300. [Google Scholar] [CrossRef]
- Serra, I.A.; Innocenti, A.M.; Di Maida, G.; Calvo, S.; Migliaccio, M.; Zambianchi, E.; Pizzigalli, C.; Arnaud-Haond, S.; Duarte, C.M.; Serrao, E.A. Genetic structure in the Mediterranean seagrass Posidonia oceanica: Disentangling past vicariance events from contemporary patterns of gene flow. Mol. Ecol. 2010, 19, 557–568. [Google Scholar] [CrossRef]
- Jahnke, M.; Casagrandi, R.; Melià, P.; Schiavina, M.; Schultz, S.T.; Zane, L.; Procaccini, G. Potential and realized connectivity of the seagrass Posidonia oceanica and their implication for conservation. Divers. Distrib. 2017, 23, 1423–1434. [Google Scholar] [CrossRef]
- Mari, L.; Melià, P.; Fraschetti, S.; Gatto, M.; Casagrandi, R. Spatial patterns and temporal variability of seagrass connectivity in the Mediterranean Sea. Divers. Distrib. 2020, 26, 169–182. [Google Scholar] [CrossRef]
- Kendrick, G.A.; Waycott, M.; Carruthers, T.J.B.; Cambridge, M.L.; Hovey, R.; Krauss, S.L.; Lavery, P.S.; Les, D.H.; Lowe, R.J.; Vidal, O.M.I. The central role of dispersal in the maintenance and persistence of seagrass populations. Bioscience 2012, 62, 56–65. [Google Scholar] [CrossRef]
- Alotaibi, N.M.; Kenyon, E.J.; Cook, K.J.; Börger, L.; Bull, J.C. Low genotypic diversity and long-term ecological decline in a spatially structured seagrass population. Sci. Rep. 2019, 9, 1–11. [Google Scholar] [CrossRef]
- Willi, Y.; Van Buskirk, J.; Schmid, B.; Fischer, M. Genetic isolation of fragmented populations is exacerbated by drift and selection. J. Evol. Biol. 2007, 20, 534–542. [Google Scholar] [CrossRef]
- Bijlsma, R.; Loeschcke, V. Genetic erosion impedes adaptive responses to stressful environments. Evol. Appl. 2012, 5, 117–129. [Google Scholar] [CrossRef]
- Kawecki, T.J.; Ebert, D. Conceptual issues in local adaptation. Ecol. Lett. 2004, 7, 1225–1241. [Google Scholar] [CrossRef]
- Fournier-Level, A.; Korte, A.; Cooper, M.D.; Nordborg, M.; Schmitt, J.; Wilczek, A.M. A map of local adaptation in Arabidopsis thaliana. Science 2011, 334, 86–89. [Google Scholar] [CrossRef] [PubMed]
- Savolainen, O.; Lascoux, M.; Merilä, J. Ecological genomics of local adaptation. Nat. Rev. Genet. 2013, 14, 807–820. [Google Scholar] [CrossRef] [PubMed]
- Tiffin, P.; Ross-Ibarra, J. Advances and limits of using population genetics to understand local adaptation. Trends Ecol. Evol. 2014, 29, 673–680. [Google Scholar] [CrossRef] [PubMed]
- Franssen, S.U.; Gu, J.; Bergmann, N.; Winters, G.; Klostermeier, U.C.; Rosenstiel, P.; Bornberg-Bauer, E.; Reusch, T.B.H. Transcriptomic resilience to global warming in the seagrass Zostera marina, a marine foundation species. Proc. Natl. Acad. Sci. USA 2011, 108, 19276–19281. [Google Scholar] [CrossRef] [PubMed]
- Franssen, S.U.; Gu, J.; Winters, G.; Huylmans, A.-K.; Wienpahl, I.; Sparwel, M.; Coyer, J.A.; Olsen, J.L.; Reusch, T.B.H.; Bornberg-Bauer, E. Genome-wide transcriptomic responses of the seagrasses Zostera marina and Nanozostera noltii under a simulated heatwave confirm functional types. Mar. Genom. 2014, 15, 65–73. [Google Scholar] [CrossRef]
- Marín-Guirao, L.; Bernardeau-Esteller, J.; García-Muñoz, R.; Ramos, A.; Ontoria, Y.; Romero, J.; Pérez, M.; Ruiz, J.M.; Procaccini, G. Carbon economy of Mediterranean seagrasses in response to thermal stress. Mar. Pollut. Bull. 2018, 135, 617–629. [Google Scholar] [CrossRef]
- Marín-Guirao, L.; Entrambasaguas, L.; Dattolo, E.; Ruiz, J.M.; Procaccini, G. Molecular mechanisms behind the physiological resistance to intense transient warming in an iconic marine plant. Front. Plant. Sci. 2017, 8, 1142. [Google Scholar] [CrossRef]
- Dattolo, E.; Marín-Guirao, L.; Ruiz, J.M.; Procaccini, G.; Marín-Guirao, L.; Ruiz, J.M.; Procaccini, G. Long-term acclimation to reciprocal light conditions suggests depth-related selection in the marine foundation species Posidonia oceanica. Ecol. Evol. 2017, 7, 1148–1164. [Google Scholar] [CrossRef]
- Bernard, A.; Marrano, A.; Donkpegan, A.; Brown, P.J.; Leslie, C.A.; Neale, D.B.; Lheureux, F.; Dirlewanger, E. Association and linkage mapping to unravel genetic architecture of phenological traits and lateral bearing in Persian walnut (Juglans regia L.). BMC Genom. 2020, 21, 1–25. [Google Scholar] [CrossRef]
- Jahnke, M.; D’Esposito, D.; Orrù, L.; Lamontanara, A.; Dattolo, E.; Badalamenti, F.; Mazzuca, S.; Procaccini, G.; Orsini, L. Adaptive responses along a depth and a latitudinal gradient in the endemic seagrass Posidonia oceanica. Heredity 2019, 122, 233–243. [Google Scholar] [CrossRef]
- Hughes, A.R.; Stachowicz, J.J. Genetic diversity enhances the resistance of a seagrass ecosystem to disturbance. Proc. Natl. Acad. Sci. USA 2004, 101, 8998–9002. [Google Scholar] [CrossRef] [PubMed]
- Reusch, T.B.H.; Ehlers, A.; Hämmerli, A.; Worm, B. Ecosystem recovery after climatic extremes enhanced by genotypic diversity. Proc. Natl. Acad. Sci. USA 2005, 102, 2826–2831. [Google Scholar] [CrossRef]
- Ehlers, A.; Worm, B.; Reusch, T.B.H. Importance of genetic diversity in eelgrass Zostera marina for its resilience to global warming. Mar. Ecol. Prog. Ser. 2008, 355, 1–7. [Google Scholar] [CrossRef]
- Evans, S.M.; Vergés, A.; Poore, A.G.B. Genotypic diversity and short-term response to shading stress in a threatened seagrass: Does low diversity mean low resilience? Front. Plant. Sci. 2017, 8, 1417. [Google Scholar] [CrossRef] [PubMed]
- Connolly, R.M.; Smith, T.M.; Maxwell, P.S.; Olds, A.D.; Macreadie, P.I.; Sherman, C.D.H. Highly disturbed populations of seagrass show increased resilience but lower genotypic diversity. Front. Plant. Sci. 2018, 9, 894. [Google Scholar] [CrossRef] [PubMed]
- Crutsinger, G.M.; Collins, M.D.; Fordyce, J.A.; Gompert, Z.; Nice, C.C.; Sanders, N.J. Plant genotypic diversity predicts community structure and governs an ecosystem process. Science 2006, 313, 966–968. [Google Scholar] [CrossRef] [PubMed]
- Whitlock, R. Relationships between adaptive and neutral genetic diversity and ecological structure and functioning: A meta-analysis. J. Ecol. 2014, 102, 857–872. [Google Scholar] [CrossRef] [PubMed]
- Williams, S.L. Reduced genetic diversity in eelgrass transplantations affects both population growth and individual fitness. Ecol. Appl. 2001, 11, 1472–1488. [Google Scholar] [CrossRef]
- Jahnke, M.; Serra, I.A.; Bernard, G.; Procaccini, G. The importance of genetic make-up in seagrass restoration: A case study of the seagrass Zostera noltei. Mar. Ecol. Prog. Ser. 2015, 532, 111–122. [Google Scholar] [CrossRef]
- Reynolds, L.K.; Waycott, M.; McGlathery, K.J.; Orth, R.J.; Zieman, J.C. Eelgrass restoration by seed maintains genetic diversity: Case study from a coastal bay system. Mar. Ecol. Prog. Ser. 2012, 448, 223–233. [Google Scholar] [CrossRef]
- Reynolds, L.K.; McGlathery, K.J.; Waycott, M. Genetic diversity enhances restoration success by augmenting ecosystem services. PLoS ONE 2012, 7, e38397. [Google Scholar] [CrossRef] [PubMed]
- Sinclair, E.A.; Verduin, J.; Krauss, S.L.; Hardinge, J.; Anthony, J.; Kendrick, G.A. A genetic assessment of a successful seagrass meadow (Posidonia australis) restoration trial. Ecol. Manag. Restor. 2013, 14, 68–71. [Google Scholar] [CrossRef]
- Campanella, J.J.; Bologna, P.A.X.; Smalley, J.V.; Avila, D.N.; Lee, K.N.; Areche, E.C.; Slavin, L.J. An analysis of the population genetics of restored Zostera marina plantings in Barnegat Bay, New Jersey. Popul. Ecol. 2013, 55, 121–133. [Google Scholar] [CrossRef]
- Hämmerli, A.; Reusch, T.B.H. Local adaptation and transplant dominance in genets of the marine clonal plant Zostera marina. Mar. Ecol. Prog. Ser. 2002, 242, 111–118. [Google Scholar] [CrossRef]
- Reynolds, L.K.; Waycott, M.; McGlathery, K.J. Restoration recovers population structure and landscape genetic connectivity in a dispersal-limited ecosystem. J. Ecol. 2013, 101, 1288–1297. [Google Scholar] [CrossRef]
- Procaccini, G.; Piazzi, L. Genetic polymorphism and transplantation success in the Mediterranean seagrass Posidonia oceanica. Restor. Ecol. 2001, 9, 332–338. [Google Scholar] [CrossRef]
- Travis, S.E.; Sheridan, P. Genetic structure of natural and restored shoalgrass Halodule wrightii populations in the NW Gulf of Mexico. Mar. Ecol. Prog. Ser. 2006, 322, 117–127. [Google Scholar] [CrossRef]
- Williams, S.L.; Davis, C.A. Population genetic analyses of transplanted eelgrass (Zostera marina) beds reveal reduced genetic diversity in southern California. Restor. Ecol. 1996, 4, 163–180. [Google Scholar] [CrossRef]
- Sgrò, C.M.; Lowe, A.J.; Hoffmann, A.A. Building evolutionary resilience for conserving biodiversity under climate change. Evol. Appl. 2011, 4, 326–337. [Google Scholar] [CrossRef] [PubMed]
- Breed, M.F.; Harrison, P.A.; Bischoff, A.; Durruty, P.; Gellie, N.J.C.; Gonzales, E.K.; Havens, K.; Karmann, M.; Kilkenny, F.F.; Krauss, S.L. Priority actions to improve provenance decision-making. Bioscience 2018, 68, 510–516. [Google Scholar] [CrossRef]
- Broadhurst, L.M.; Lowe, A.; Coates, D.J.; Cunningham, S.A.; McDonald, M.; Vesk, P.A.; Yates, C. Seed supply for broadscale restoration: Maximizing evolutionary potential. Evol. Appl. 2008, 1, 587–597. [Google Scholar] [CrossRef] [PubMed]
- Aitken, S.N.; Whitlock, M.C. Assisted gene flow to facilitate local adaptation to climate change. Annu. Rev. Ecol. Evol. Syst. 2013, 44, 367–388. [Google Scholar] [CrossRef]
- Nguyen, V.X.; Detcharoen, M.; Tuntiprapas, P.; Soe-Htun, U.; Sidik, J.B.; Harah, M.Z.; Prathep, A.; Papenbrock, J. Genetic species identification and population structure of Halophila (Hydrocharitaceae) from the Western Pacific to the Eastern Indian Ocean. BMC Evol. Biol. 2014, 14, 1–18. [Google Scholar] [CrossRef] [PubMed]
- Todesco, M.; Pascual, M.A.; Owens, G.L.; Ostevik, K.L.; Moyers, B.T.; Hübner, S.; Heredia, S.M.; Hahn, M.A.; Caseys, C.; Bock, D.G. Hybridization and extinction. Evol. Appl. 2016, 9, 892–908. [Google Scholar] [CrossRef]
- Liu, S.Y.V.; Kumara, T.P.; Hsu, C.-H. Genetic identification and hybridization in the seagrass genus Halophila (Hydrocharitaceae) in Sri Lankan waters. PeerJ 2020, 8, e10027. [Google Scholar] [CrossRef]
- Prober, S.M.; Byrne, M.; McLean, E.H.; Steane, D.A.; Potts, B.M.; Vaillancourt, R.E.; Stock, W.D. Climate-adjusted provenancing: A strategy for climate-resilient ecological restoration. Front. Ecol. Evol. 2015, 3, 65. [Google Scholar] [CrossRef]
- Green, E.P.; Short, F.T.; Frederick, T. The World Atlas of Seagrasses; University of California Press: Berkeley, CA, USA, 2003. [Google Scholar]
- Vander Mijnsbrugge, K.; Bischoff, A.; Smith, B. A question of origin: Where and how to collect seed for ecological restoration. Basic Appl. Ecol. 2010, 11, 300–311. [Google Scholar] [CrossRef]
- McMahon, K.; Sinclair, E.A.; Sherman, C.D.H.; Van Dijk, K.-J.; Hernawan, U.E.; Verduin, J.; Waycott, M. Genetic connectivity in tropical and temperate Australian seagrass species. In Seagrasses of Australia; Springer: Berlin/Heidelberg, Germany, 2018; pp. 155–194. [Google Scholar]
- Arnaud-Haond, S.; Migliaccio, M.; Diaz-Almela, E.; Teixeira, S.; Van De Vliet, M.S.; Alberto, F.; Procaccini, G.; Duarte, C.M.; Serrão, E.A. Vicariance patterns in the Mediterranean Sea: East-west cleavage and low dispersal in the endemic seagrass Posidonia Oceanica. J. Biogeogr. 2007, 34, 963–976. [Google Scholar] [CrossRef]
- Procaccini, G.; Olsen, J.L.; Reusch, T.B.H. Contribution of genetics and genomics to seagrass biology and conservation. J. Exp. Mar. Bio. Ecol. 2007, 350, 234–259. [Google Scholar] [CrossRef]
- Lanuru, M.; Mashoreng, S.; Amri, K. Using site-selection model to identify suitable sites for seagrass transplantation in the west coast of South Sulawesi. J. Phys. Conf. Ser. 2018, 979, 12007. [Google Scholar] [CrossRef]
- Ferdinando, B.; Federica, F.; Simona, F.; Paul, G.; Macpherson, E.; Serge, P.; Takvor, S. CoCoNet: Towards coast to coast networks of marine protected areas (from the shore to the high and deep sea), coupled with sea-based wind energy potential. Sci. Res. Inf. Technol. 2016, 6, 1–96. [Google Scholar]
- Valle, M.; Chust, G.; Del Campo, A.; Wisz, M.S.; Olsen, S.M.; Garmendia, J.M.; Borja, Á. Projecting future distribution of the seagrass Zostera noltii under global warming and sea level rise. Biol. Conserv. 2014, 170, 74–85. [Google Scholar] [CrossRef]
- Chefaoui, R.M.; Assis, J.; Duarte, C.M.; Serrão, E.A. Large-Scale Prediction of Seagrass Distribution Integrating Landscape Metrics and Environmental Factors: The Case of Cymodocea nodosa (Mediterranean-Atlantic). Estuaries Coasts 2016, 39, 123–137. [Google Scholar] [CrossRef]
- Oreska, M.P.J.; McGlathery, K.J.; Wiberg, P.L.; Orth, R.J.; Wicox, D.J. Defining the Zostera marina (Eelgrass) Niche from Long-Term Success of Restored and Naturally Colonized Meadows: Implications for Seagrass Restoration. Estuaries Coasts 2021, 44, 396–411. [Google Scholar] [CrossRef]
- McKay, J.K.; Christian, C.E.; Harrison, S.; Rice, K.J. How local is local? A review of practical and conceptual issues in the genetics of restoration. Restor. Ecol. 2005, 13, 432–440. [Google Scholar] [CrossRef]
- Sinclair, E.A.; Edgeloe, J.M.; Anthony, J.M.; Statton, J.; Breed, M.F.; Kendrick, G.A. Variation in reproductive effort, genetic diversity and mating systems across Posidonia australis seagrass meadows in Western Australia. AoB Plants 2020, 12, plaa038. [Google Scholar] [CrossRef] [PubMed]
- Sexton, J.P.; Strauss, S.Y.; Rice, K.J. Gene flow increases fitness at the warm edge of a specie’s range. Proc. Natl. Acad. Sci. USA 2011, 108, 11704–11709. [Google Scholar] [CrossRef]
- Sinclair, E.A.; Anthony, J.M.; Greer, D.; Ruiz-Montoya, L.; Evans, S.M.; Krauss, S.L.; Kendrick, G.A. Genetic signatures of Bassian glacial refugia and contemporary connectivity in a marine foundation species. J. Biogeogr. 2016, 43, 2209–2222. [Google Scholar] [CrossRef]
- Campanella, J.J.; Bologna, P.A.X.; Smith, S.M.; Rosenzweig, E.B.; Smalley, J.V. Zostera marina population genetics in Barnegat Bay, New Jersey, and implications for grass bed restoration. Popul. Ecol. 2010, 52, 181–190. [Google Scholar] [CrossRef]
- Kim, J.H.; Kang, J.H.; Jang, J.E.; Choi, S.K.; Kim, M.J.; Park, S.R.; Lee, H.J. Population genetic structure of eelgrass (Zostera marina) on the Korean coast: Current status and conservation implications for future management. PLoS ONE 2017, 12, e0174105. [Google Scholar] [CrossRef]
- Smith, T.M.; York, P.H.; Macreadie, P.I.; Keough, M.J.; Ross, D.J.; Sherman, C.D.H. Spatial variation in reproductive effort of a southern Australian seagrass. Mar. Environ. Res. 2016, 120, 214–224. [Google Scholar] [CrossRef]
- Procaccini, G.; Ruocco, M.; Marín-Guirao, L.; Dattolo, E.; Brunet, C.; D’Esposito, D.; Lauritano, C.; Mazzuca, S.; Serra, I.A.; Bernardo, L.; et al. Depth-specific fluctuations of gene expression and protein abundance modulate the photophysiology in the seagrass Posidonia oceanica. Sci. Rep. 2017, 7. [Google Scholar] [CrossRef]
- D’Esposito, D.; Dattolo, E.; Badalamenti, F.; Orsini, L.; Procaccini, G. Comparative analysis of genetic diversity of posidonia oceanica along a depth gradient using neutral and selective/non neutral microsatellites markers. Biol. Mar. Mediterr. 2012, 19, 45. [Google Scholar]
- Telesca, L.; Belluscio, A.; Criscoli, A.; Ardizzone, G.; Apostolaki, E.T.; Fraschetti, S.; Gristina, M.; Knittweis, L.; Martin, C.S.; Pergent, G. Seagrass meadows (Posidonia oceanica) distribution and trajectories of change. Sci. Rep. 2015, 5, 12505. [Google Scholar] [CrossRef] [PubMed]
- Micheli, F.; Halpern, B.S.; Walbridge, S.; Ciriaco, S.; Ferretti, F.; Fraschetti, S.; Lewison, R.; Nykjaer, L.; Rosenberg, A.A. Cumulative human impacts on Mediterranean and Black Sea marine ecosystems: Assessing current pressures and opportunities. PLoS ONE 2013, 8, e79889. [Google Scholar] [CrossRef] [PubMed]
- Jahnke, M.F. Population Connectivity and Genetic Diversity in Mediterranean Seagrasses in the Framework of Management and Conservation of the Coastline. Ph.D. Thesis, The Open University, Milton Keynes, UK, September 2015. [Google Scholar]
- Van Katwijk, M.M.; Thorhaug, A.; Marbà, N.; Orth, R.J.; Duarte, C.M.; Kendrick, G.A.; Althuizen, I.H.J.; Balestri, E.; Bernard, G.; Cambridge, M.L.; et al. Global analysis of seagrass restoration: The importance of large-scale planting. J. Appl. Ecol. 2016, 53, 567–578. [Google Scholar] [CrossRef]
- Kettenring, K.M.; Tarsa, E.E. Need to seed? Ecological, genetic, and evolutionary keys to seed-based wetland restoration. Front. Environ. Sci. 2020, 8, 109. [Google Scholar] [CrossRef]
- Orth, R.J.; Moore, K.A.; Marion, S.R.; Wilcox, D.J.; Parrish, D.B. Seed addition facilitates eelgrass recovery in a coastal bay system. Mar. Ecol. Prog. Ser. 2012, 448, 177–195. [Google Scholar] [CrossRef]
- Orth, R.J.; Lefcheck, J.S.; McGlathery, K.S.; Aoki, L.; Luckenbach, M.W.; Moore, K.A.; Oreska, M.P.J.; Snyder, R.; Wilcox, D.J.; Lusk, B. Restoration of seagrass habitat leads to rapid recovery of coastal ecosystem services. Sci. Adv. 2020, 6, 1–10. [Google Scholar] [CrossRef]
- Marion, S.R.; Orth, R.J. Innovative Techniques for large-scale seagrass restoration using Zostera marina (eelgrass) seeds. Restor. Ecol. 2010, 18, 514–526. [Google Scholar] [CrossRef]
- Statton, J.; Montoya, L.R.; Orth, R.J.; Dixon, K.W.; Kendrick, G.A. Identifying critical recruitment bottlenecks limiting seedling establishment in a degraded seagrass ecosystem. Sci. Rep. 2017, 7, 1–12. [Google Scholar] [CrossRef]
- Orth, R.J.; Luckenbach, M.L.; Marion, S.R.; Moore, K.A.; Wilcox, D.J. Seagrass recovery in the Delmarva coastal bays, USA. Aquat. Bot. 2006, 84, 26–36. [Google Scholar] [CrossRef]
- Van Rossum, F.; Hardy, O.J.; Le Pajolec, S.; Raspé, O. Genetic monitoring of translocated plant populations in practice. Mol. Ecol. 2020, 29, 4040–4058. [Google Scholar] [CrossRef]
- Lindenmayer, D. Improving restoration programs through greater connection with ecological theory and better monitoring. Front. Ecol. Evol. 2020, 8, 50. [Google Scholar] [CrossRef]
- Cook, C.N.; Sgrò, C.M. Understanding managers’ and scientists’ perspectives on opportunities to achieve more evolutionarily enlightened management in conservation. Evol. Appl. 2018, 11, 1371–1388. [Google Scholar] [CrossRef]
- Schwartz, M.K.; Luikart, G.; Waples, R.S. Genetic monitoring as a promising tool for conservation and management. Trends Ecol. Evol. 2007, 22, 25–33. [Google Scholar] [CrossRef] [PubMed]
- Mijangos, J.L.; Pacioni, C.; Spencer, P.B.S.; Craig, M.D. Contribution of genetics to ecological restoration. Mol. Ecol. 2015, 24, 22–37. [Google Scholar] [CrossRef] [PubMed]
- Jackson, E.L.; Smith, T.M.; York, P.H.; Nielsen, J.; Irving, A.D.; Sherman, C.D.H. An assessment of the seascape genetic structure and hydrodynamic connectivity for subtropical seagrass restoration. Restor. Ecol. 2020, 29, e13269. [Google Scholar] [CrossRef]
- Hori, M.; Sato, M. Genetic effects of eelgrass restoration efforts by fisher’s seeding to recover seagrass beds as an important natural capital for coastal ecosystem services. Popul. Ecol. 2021, 63, 92–101. [Google Scholar] [CrossRef]
- Birchler, J.A.; Yao, H.; Chudalayandi, S.; Vaiman, D.; Veitia, R.A. Heterosis. Plant. Cell 2010, 22, 2105–2112. [Google Scholar] [CrossRef]
- Statton, J.; Dixon, K.W.; Hovey, R.K.; Kendrick, G.A. A comparative assessment of approaches and outcomes for seagrass revegetation in Shark Bay and Florida Bay. Mar. Freshw. Res. 2012, 63, 984–993. [Google Scholar] [CrossRef]
- Procaccini, G.; Beer, S.; Björk, M.; Olsen, J.; Mazzuca, S.; Santos, R. Seagrass ecophysiology meets ecological genomics: Are we ready? Mar. Ecol. 2012, 33, 522–527. [Google Scholar] [CrossRef]
- Merilä, J.; Hendry, A.P. Climate change, adaptation, and phenotypic plasticity: The problem and the evidence. Evol. Appl. 2014, 7, 1–14. [Google Scholar] [CrossRef]
- Van Oppen, M.J.H.; Oliver, J.K.; Putnam, H.M.; Gates, R.D. Building coral reef resilience through assisted evolution. Proc. Natl. Acad. Sci. USA 2015, 112, 2307–2313. [Google Scholar] [CrossRef] [PubMed]
- Jones, T.A.; Monaco, T.A. A role for assisted evolution in designing native plant materials for domesticated landscapes. Front. Ecol. Environ. 2009, 7, 541–547. [Google Scholar] [CrossRef]
- Anderson, K.; Taylor, D.A.; Thompson, E.L.; Melwani, A.R.; Nair, S.V.; Raftos, D.A. Meta-analysis of studies using suppression subtractive hybridization and microarrays to investigate the effects of environmental stress on gene transcription in oysters. PLoS ONE 2015, 10, e0118839. [Google Scholar] [CrossRef]
- Fernandez i Marti, A.; Dodd, R.S. Using CRISPR as a gene editing tool for validating adaptive gene function in tree landscape genomics. Front. Ecol. Evol. 2018, 6, 76. [Google Scholar] [CrossRef]
- Breed, M.F.; Harrison, P.A.; Blyth, C.; Byrne, M.; Gaget, V.; Gellie, N.J.C.; Groom, S.V.C.; Hodgson, R.; Mills, J.G.; Prowse, T.A.A. The potential of genomics for restoring ecosystems and biodiversity. Nat. Rev. Genet. 2019, 20, 615–628. [Google Scholar] [CrossRef]
- Hartung, F.; Schiemann, J. Precise plant breeding using new genome editing techniques: Opportunities, safety and regulation in the EU. Plant. J. 2014, 78, 742–752. [Google Scholar] [CrossRef] [PubMed]
- Tanner, C.E.; Parham, T. Growing Zostera marina (eelgrass) from seeds in land-based culture systems for use in restoration projects. Restor. Ecol. 2010, 18, 527–537. [Google Scholar] [CrossRef]
- Jisha, K.C.; Vijayakumari, K.; Puthur, J.T. Seed priming for abiotic stress tolerance: An overview. Acta Physiol. Plant. 2013, 35, 1381–1396. [Google Scholar] [CrossRef]
- Vriet, C.; Hennig, L.; Laloi, C. Stress-induced chromatin changes in plants: Of memories, metabolites and crop improvement. Cell. Mol. Life Sci. 2015, 72, 1261–1273. [Google Scholar] [CrossRef]
- Nguyen, H.M.; Kim, M.; Ralph, P.J.; Marín-Guirao, L.; Pernice, M.; Procaccini, G. Stress memory in seagrasses: First insight into the effects of thermal priming and the role of epigenetic modifications. Front. Plant. Sci. 2020, 11, 494. [Google Scholar] [CrossRef]
- Jueterbock, A.; Boström, C.; Coyer, J.A.; Olsen, J.L.; Kopp, M.; Dhanasiri, A.K.S.; Smolina, I.; Arnaud-Haond, S.; Van de Peer, Y.; Hoarau, G. The seagrass methylome is associated with variation in photosynthetic performance among clonal shoots. Front. Plant. Sci. 2020, 11, 1387. [Google Scholar] [CrossRef] [PubMed]
- Bossdorf, O.; Richards, C.L.; Pigliucci, M. Epigenetics for ecologists. Ecol. Lett. 2008, 11, 106–115. [Google Scholar] [CrossRef] [PubMed]
- Richards, C.L.; Alonso, C.; Becker, C.; Bossdorf, O.; Bucher, E.; Colomé-Tatché, M.; Durka, W.; Engelhardt, J.; Gaspar, B.; Gogol-Döring, A.; et al. Ecological plant epigenetics: Evidence from model and non-model species, and the way forward. Ecol. Lett. 2017, 20, 1576–1590. [Google Scholar] [CrossRef] [PubMed]
- Holliday, R. Epigenetics: A historical overview. Epigenetics 2006, 1, 76–80. [Google Scholar] [CrossRef] [PubMed]
- Herman, J.J.; Spencer, H.G.; Donohue, K.; Sultan, S.E. How stable ‘should’epigenetic modifications be? Insights from adaptive plasticity and bet hedging. Evolution 2014, 68, 632–643. [Google Scholar] [CrossRef]
- Douhovnikoff, V.; Dodd, R.S. Epigenetics: A potential mechanism for clonal plant success. Plant. Ecol. 2015, 216, 227–233. [Google Scholar] [CrossRef]
- Dodd, R.S.; Douhovnikoff, V. Adjusting to global change through clonal growth and epigenetic variation. Front. Ecol. Evol. 2016, 4, 86. [Google Scholar] [CrossRef]
- Verhoeven, K.J.F.; Preite, V. Epigenetic variation in asexually reproducing organisms. Evolution 2014, 68, 644–655. [Google Scholar] [CrossRef]
- Greco, M.; Chiappetta, A.; Bruno, L.; Bitonti, M.B. In Posidonia oceanica cadmium induces changes in DNA methylation and chromatin patterning. J. Exp. Bot. 2012, 63, 695–709. [Google Scholar] [CrossRef]
- Greco, M.; Chiappetta, A.; Bruno, L.; Bitonti, M.B. Effects of light deficiency on genome methylation in Posidonia oceanica. Mar. Ecol. Prog. Ser. 2013, 473, 103–114. [Google Scholar] [CrossRef]
- Ruocco, M.; Marín-Guirao, L.; Procaccini, G. Within and among-leaf variations in photo-physiological functions, gene expression and DNA methylation patterns in the large-sized seagrass Posidonia oceanica. Mar. Biol. 2019, 166, 24. [Google Scholar] [CrossRef]
- Latzel, V.I.T.; Allan, E.; Silveira, A.B.; Colot, V.; Fischer, M.; Bossdorf, O. Epigenetic diversity increases the productivity and stability of plant populations. Nat. Commun. 2013, 4, 2875. [Google Scholar] [CrossRef]
- Gáspár, B.; Bossdorf, O.; Durka, W. Structure, stability and ecological significance of natural epigenetic variation: A large-scale survey in Plantago lanceolata. N. Phytol. 2019, 221, 1585–1596. [Google Scholar] [CrossRef]
- Rey, O.; Eizaguirre, C.; Angers, B.; Baltazar-Soares, M.; Sagonas, K.; Prunier, J.G.; Blanchet, S. Linking epigenetics and biological conservation: Towards a conservation epigenetics perspective. Funct. Ecol. 2020, 34, 414–427. [Google Scholar] [CrossRef]
- Farber, S.; Costanza, R.; Childers, D.L.; Erickson, J.; Gross, K.; Grove, M.; Hopkinson, C.S.; Kahn, J.; Pincetl, S.; Troy, A.; et al. Linking ecology and economics for ecosystem management. Bioscience 2006, 56, 121–133. [Google Scholar] [CrossRef]
- Prach, K.; Tolvanen, A. How can we restore biodiversity and ecosystem services in mining and industrial sites? Environ. Sci. Pollut. Res. 2016, 23, 13587–13590. [Google Scholar] [CrossRef] [PubMed]
- Laikre, L.; Allendorf, F.W.; Aroner, L.C.; Baker, C.S.; Gregovich, D.P.; Hansen, M.M.; Jackson, J.A.; Kendall, K.C.; Mckelvey, K.; Neel, M.C. Neglect of genetic diversity in implementation of the convention on biological diversity. Conserv. Biol. 2010, 24, 86–88. [Google Scholar] [CrossRef] [PubMed]
- De Los Santos, C.B.; Sigurðardóttir, R.; Cunha, A.; Cook, K.; Wiktor, J.M.; Tatarek, A.; Santos, R. A survey-based assessment of seagrass status, management and legislation in Europe. Front. Mar. Sci. Int. Meet. Mar. Res. 2014, 1. [Google Scholar] [CrossRef]
- FAO New United Nations Decade on Ecosystem Restoration Offers Unparalleled Opportunity for Job Creation, Food Security and Addressing Climate Change. Available online: http://www.fao.org/news/story/en/item/1182090/icode (accessed on 10 October 2020).
- Secretariat of the Convention on Biological Diversity. Bonn Guidelines on Access to Genetic Resources and Fair and Equitable Sharing of the Benefits Arising out of their Utilization; Secretariat of the Convention on Biological Diversity: Montreal, QC, Canada, 2002. [Google Scholar]
- Text And Annex of the Nagoya Protocol on Access to Genetic Resources and the Fair and Equitable Sharing of Benefits Arising from their Utilization to the Convention on Biological Diversity, 1st ed.; United Nations: Montreal, QC, Canada, 2015; Available online: https://www.cbd.int/abs/text/default.shtml (accessed on 10 November 2020).
- Zhu, Y.; Chen, H.; Fan, J.; Wang, Y.; Li, Y.; Chen, J.; Fan, J.X.; Yang, S.; Hu, L.; Leung, H.; et al. Genetic diversity and disease control in rice. Nature 2000, 406, 718–722. [Google Scholar] [CrossRef] [PubMed]
- Wood, D.; Lenné, J.M. Agrobiodiversity: Characterization, Utilization and Management; CABI: Wallingford, UK, 1999; ISBN 0851993370. [Google Scholar]
- Tilford, D.S. Saving the blueprints: The international legal regime for plant resources. J. Int. Law 1998, 30, 373. [Google Scholar]
- Guerrant, E.O.; Havens, K.; Maunder, M.; Havens, K. Ex Situ Plant. Conservation: Supporting Species Survival in the Wild; Island Press: Washington, DC, USA, 2004. [Google Scholar]
- Walck, J.L.; Baskin, J.M.; Baskin, C.C.; Hidayati, S.N. Defining transient and persistent seed banks in species with pronounced seasonal dormancy and germination patterns. Seed Sci. Res. 2005, 15, 189–196. [Google Scholar] [CrossRef]
- Falk, D.A.; Holsinger, K.E. Genetic sampling guidelines for conservation collections of endangered plants. In Genetics and Conservation of Rare Plants; Oxford University Press: New York, NY, USA, 1991. [Google Scholar]
- IUCN RSG. IUCN Guidelines for Reintroductions and other Conservation Translocations; IUCN: Gland, Switzerland, 2012. [Google Scholar]
- Filbee-Dexter, K.; Smajdor, A. Ethics of assisted evolution in marine conservation. Front. Mar. Sci. 2019, 6, 1–6. [Google Scholar] [CrossRef]
- Agarwal, M.; Shrivastava, N.; Padh, H. Advances in molecular marker techniques and their applications in plant sciences. Plant. Cell Rep. 2008, 27, 617–631. [Google Scholar] [CrossRef]
- Davey, J.W.; Hohenlohe, P.A.; Etter, P.D.; Boone, J.Q.; Catchen, J.M.; Blaxter, M.L. Genome-wide genetic marker discovery and genotyping using next-generation sequencing. Nat. Rev. Genet. 2011, 12, 499–510. [Google Scholar] [CrossRef]
- Ekblom, R.; Galindo, J. Applications of next generation sequencing in molecular ecology of non-model organisms. Heredity 2011, 107, 1–15. [Google Scholar] [CrossRef]
- McCormack, J.E.; Hird, S.M.; Zellmer, A.J.; Carstens, B.C.; Brumfield, R.T. Applications of next-generation sequencing to phylogeography and phylogenetics. Mol. Phylogenet. Evol. 2013, 66, 526–538. [Google Scholar] [CrossRef] [PubMed]
- Shafer, A.B.A.; Peart, C.R.; Tusso, S.; Maayan, I.; Brelsford, A.; Wheat, C.W.; Wolf, J.B.W. Bioinformatic processing of RAD-seq data dramatically impacts downstream population genetic inference. Methods Ecol. Evol. 2017, 8, 907–917. [Google Scholar] [CrossRef]
- Wang, S.; Meyer, E.; McKay, J.K.; Matz, M. V 2b-RAD: A simple and flexible method for genome-wide genotyping. Nat. Methods 2012, 9, 808–810. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Liu, P.; Lv, J.; Li, Y.; Cheng, T.; Zhang, L.; Xia, Y.; Sun, H.; Hu, X.; Bao, Z. Serial sequencing of isolength RAD tags for cost-efficient genome-wide profiling of genetic and epigenetic variations. Nat. Protoc. 2016, 11, 2189–2200. [Google Scholar] [CrossRef] [PubMed]
- Peterson, B.K.; Weber, J.N.; Kay, E.H.; Fisher, H.S.; Hoekstra, H.E. Double digest RADseq: An inexpensive method for de novo SNP discovery and genotyping in model and non-model species. PLoS ONE 2012, 7, e37135. [Google Scholar] [CrossRef]
- Andrews, K.R.; Good, J.M.; Miller, M.R.; Luikart, G.; Hohenlohe, P.A. Harnessing the power of RADseq for ecological and evolutionary genomics. Nat. Rev. Conserv. 2016, 17, 81. [Google Scholar] [CrossRef]
- Olsen, J.L.; Rouzé, P.; Verhelst, B.; Lin, Y.C.; Bayer, T.; Collen, J.; Dattolo, E.; De Paoli, E.; Dittami, S.; Maumus, F.; et al. The genome of the seagrass Zostera marina reveals angiosperm adaptation to the sea. Nature 2016, 530, 331–335. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.; Golicz, A.A.; Bayer, P.E.; Jiao, Y.; Tang, H.; Paterson, A.H.; Sablok, G.; Krishnaraj, R.R.; Chan, C.-K.K.; Batley, J.; et al. The Genome of a Southern Hemisphere Seagrass Species (Zostera muelleri). Plant. Physiol. 2016, 172, 272–283. [Google Scholar] [CrossRef] [PubMed]
- Phair, N.L.; Toonen, R.J.; Knapp, I.; Von der Heyden, S. Shared genomic outliers across two divergent population clusters of a highly threatened seagrass. PeerJ 2019, 7, e6806. [Google Scholar] [CrossRef]
- Phair, N.L.; Toonen, R.J.; Knapp, I.S.S.; Von der Heyden, S. Anthropogenic pressures negatively impact genomic diversity of the vulnerable seagrass Zostera capensis. J. Environ. Manag. 2020, 255, 109831. [Google Scholar] [CrossRef]
- Paun, O.; Verhoeven, K.J.F.; Richards, C.L. Opportunities and limitations of reduced representation bisulfite sequencing in plant ecological epigenomics. N. Phytol. 2019, 221, 738–742. [Google Scholar] [CrossRef]
- Olova, N.; Krueger, F.; Andrews, S.; Oxley, D.; Berrens, R.V.; Branco, M.R.; Reik, W. Comparison of whole-genome bisulfite sequencing library preparation strategies identifies sources of biases affecting DNA methylation data. Genom. Biol. 2018, 19, 1–19. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Lv, J.; Zhang, L.; Dou, J.; Sun, Y.; Li, X.; Fu, X.; Dou, H.; Mao, J.; Hu, X. MethylRAD: A simple and scalable method for genome-wide DNA methylation profiling using methylation-dependent restriction enzymes. Open Biol. 2015, 5, 150130. [Google Scholar] [CrossRef] [PubMed]
- Van Gurp, T.P.; Wagemaker, N.C.A.M.; Wouters, B.; Vergeer, P.; Ouborg, J.N.J.; Verhoeven, K.J.F. epiGBS: Reference-free reduced representation bisulfite sequencing. Nat. Methods 2016, 13, 322–324. [Google Scholar] [CrossRef] [PubMed]
- Trucchi, E.; Mazzarella, A.B.; Gilfillan, G.D.; Lorenzo, M.T.; Schönswetter, P.; Paun, O. Bs RAD seq: Screening DNA methylation in natural populations of non-model species. Mol. Ecol. 2016, 25, 1697–1713. [Google Scholar] [CrossRef]
- Williams, A.V.; Nevill, P.G.; Krauss, S.L. Next generation restoration genetics: Applications and opportunities. Trends Plant. Sci. 2014, 19, 529–537. [Google Scholar] [CrossRef] [PubMed]
- Corlett, R.T. Restoration, reintroduction, and rewilding in a changing world. Trends Ecol. Evol. 2016, 31, 453–462. [Google Scholar] [CrossRef]
- Montalvo, A.M.; Williams, S.L.; Rice, K.J.; Buchmann, S.L.; Cory, C.; Handel, S.N.; Nabhan, G.P.; Primack, R.; Robichaux, R.H. Restoration biology: A population biology perspective. Restor. Ecol. 1997, 5, 277–290. [Google Scholar] [CrossRef]
| Species | Year | Donor Location | Restored Location | Plant Material | Area | Duration | Genetic Diversity Assessment | Ref. |
|---|---|---|---|---|---|---|---|---|
| Posidonia australis | 2013 | Jervis Bay (Australia) reciprocal transplant study | St. Georges Basin (Australia) reciprocal transplant study | Adult plants | na | 6 months | Eight microsatellites | [11] |
| Zostera noltei | 2009 | Carteau in the Gulf of Fos (France) | Berre lagoon (France) | Adult plants | 450 m2 | 4 years | Nine microsatellites | [82] |
| Zostera marina | 2007 | Mobjack Bay, Chesapeake Bay, South Bay, USA | Hog Island Bay, USA | Seeds | 128 m2 | 20 months | Eight microsatellites | [83] |
| Zostera marina | 2006–2007 * | Chesapeake Bay (USA) | Virginia coastal bays (USA) | Seeds | na | 2–3 years | Eight microsatellites | [84] |
| Posidonia australis | 2004 | Parmelia Bank, Cockburn Sound (Australia) | Southern Flats, Cockburn Sound (Australia) | Adult plants | 3.2 ha | 4 years | Seven microsatellites | [85] |
| Zostera marina | 2001–2008 * | Related ref | related ref | Adult plants | related ref | 10 years | Seven microsatellites | [86] |
| Zostera marina | 2000 | Two sites along the German Baltic Coast | Two sites along the German Baltic Coast | Adult plants | 450 m2 | 11 weeks | Four microsatellites | [87] |
| Zostera marina | Late 1990s | Chesapeake Bay | Twenty-three meadows along the eastern coast of North America | Seeds | 1600 ha | 15 years | Seven microsatellites | [88] |
| Posidonia oceanica | 1994 | Gorgona Island, Pantelleria Island (Italy) | Vada (Italy) | Adult plants | na | 3 years | Six microsatellites | [89] |
| Halodule wrightii | 1993–2000 * | Related ref | Related ref | Adult plants | na | 2–7 years | 98 AFLPs | [90] |
| Zostera marina | 1993 | South San Diego Bay (USA) | North San Diego Bay (USA) | Adult plants | na | 2 years | Allozyme electrophoresis | [81] |
| Zostera marina | related ref * | Related ref | Related ref | Adult plants | Related ref | 3–16 years | Allozyme electrophoresis | [91] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pazzaglia, J.; Nguyen, H.M.; Santillán-Sarmiento, A.; Ruocco, M.; Dattolo, E.; Marín-Guirao, L.; Procaccini, G. The Genetic Component of Seagrass Restoration: What We Know and the Way Forwards. Water 2021, 13, 829. https://doi.org/10.3390/w13060829
Pazzaglia J, Nguyen HM, Santillán-Sarmiento A, Ruocco M, Dattolo E, Marín-Guirao L, Procaccini G. The Genetic Component of Seagrass Restoration: What We Know and the Way Forwards. Water. 2021; 13(6):829. https://doi.org/10.3390/w13060829
Chicago/Turabian StylePazzaglia, Jessica, Hung Manh Nguyen, Alex Santillán-Sarmiento, Miriam Ruocco, Emanuela Dattolo, Lázaro Marín-Guirao, and Gabriele Procaccini. 2021. "The Genetic Component of Seagrass Restoration: What We Know and the Way Forwards" Water 13, no. 6: 829. https://doi.org/10.3390/w13060829
APA StylePazzaglia, J., Nguyen, H. M., Santillán-Sarmiento, A., Ruocco, M., Dattolo, E., Marín-Guirao, L., & Procaccini, G. (2021). The Genetic Component of Seagrass Restoration: What We Know and the Way Forwards. Water, 13(6), 829. https://doi.org/10.3390/w13060829









