Classification and Prediction of Fecal Coliform in Stream Waters Using Decision Trees (DTs) for Upper Green River Watershed, Kentucky, USA
Abstract
1. Introduction
2. Description of Study Area and Data
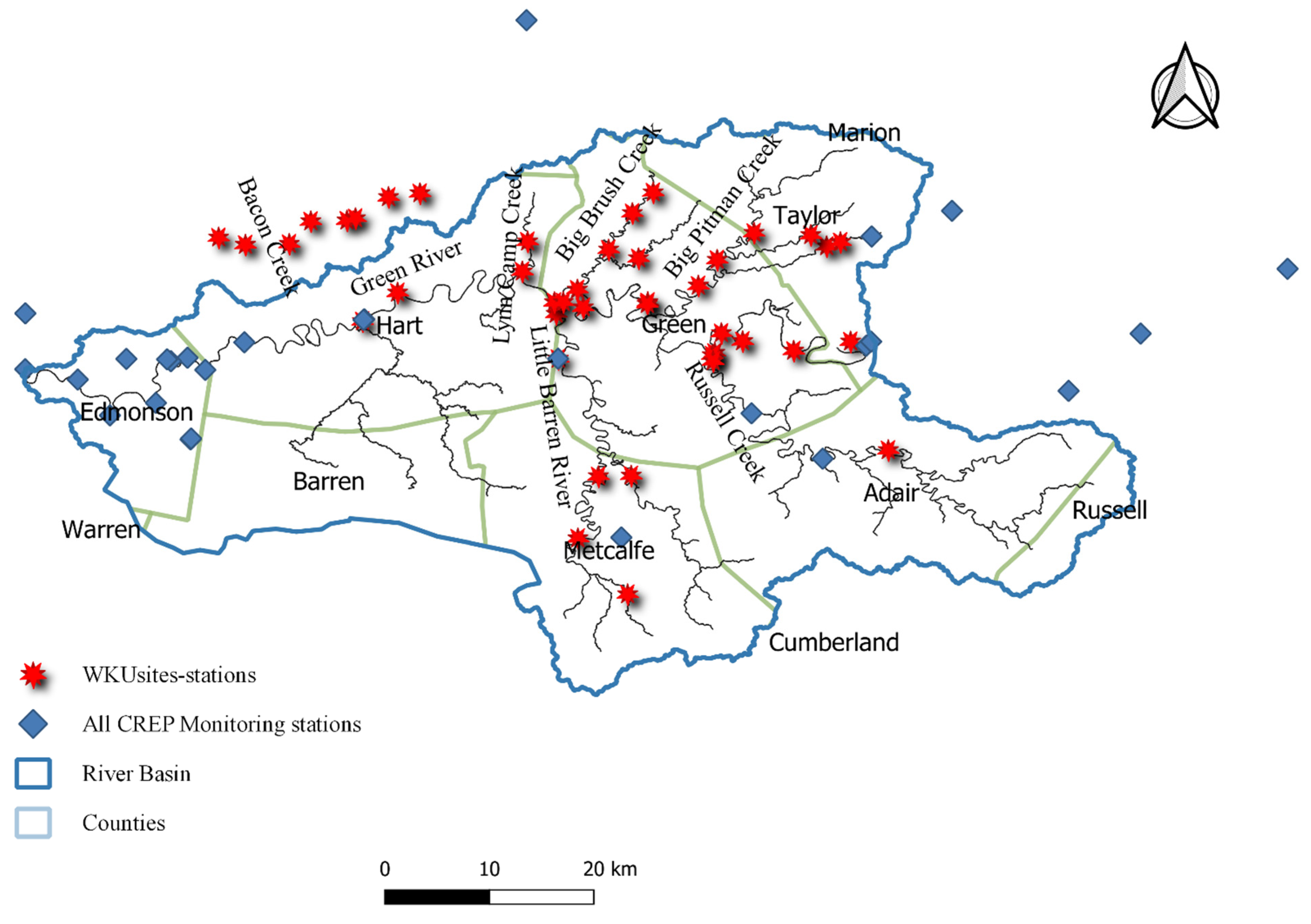
2.1. Study Area
2.2. Data
3. Methodology
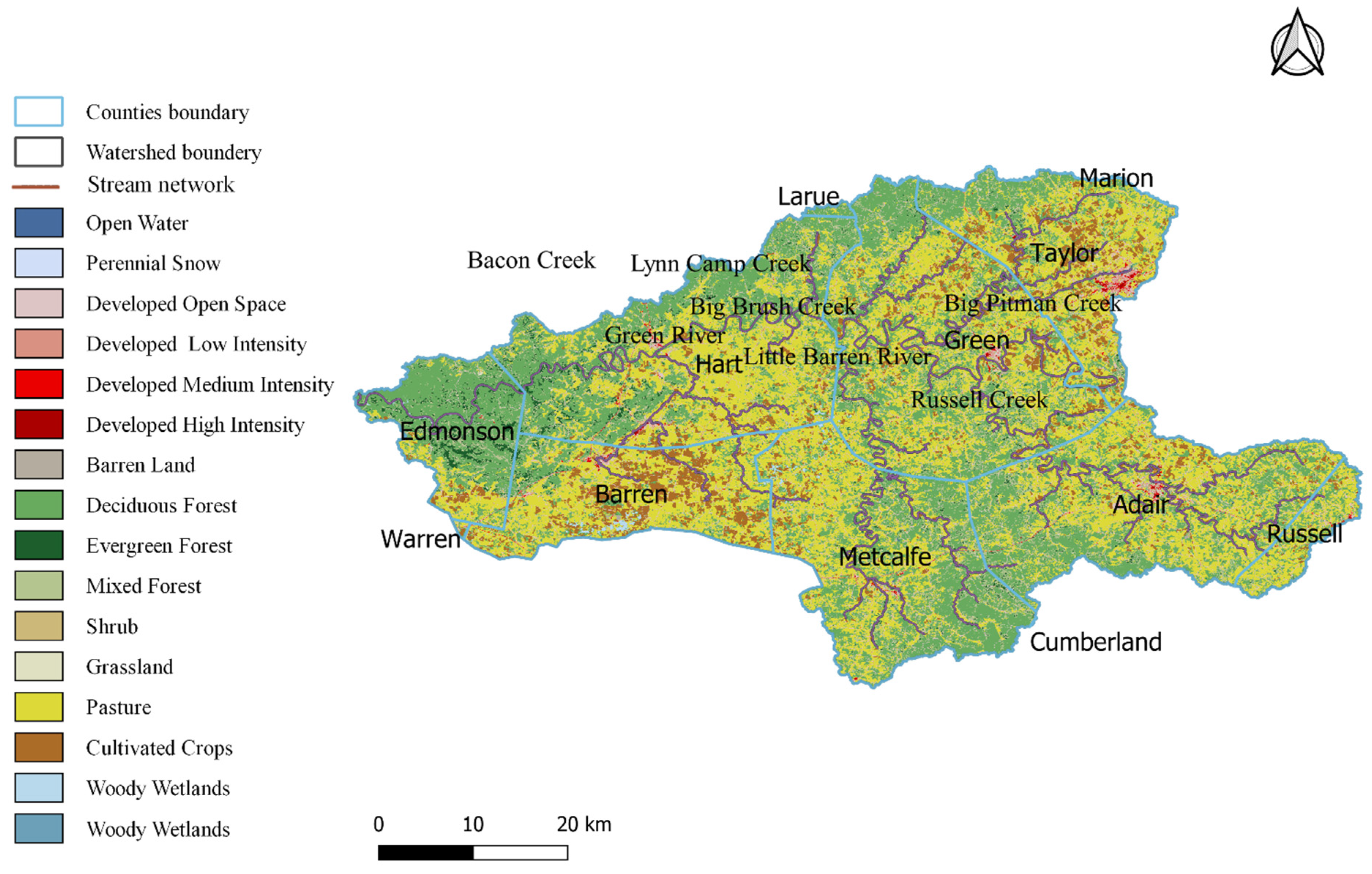
3.1. GIS Landuse Analysis
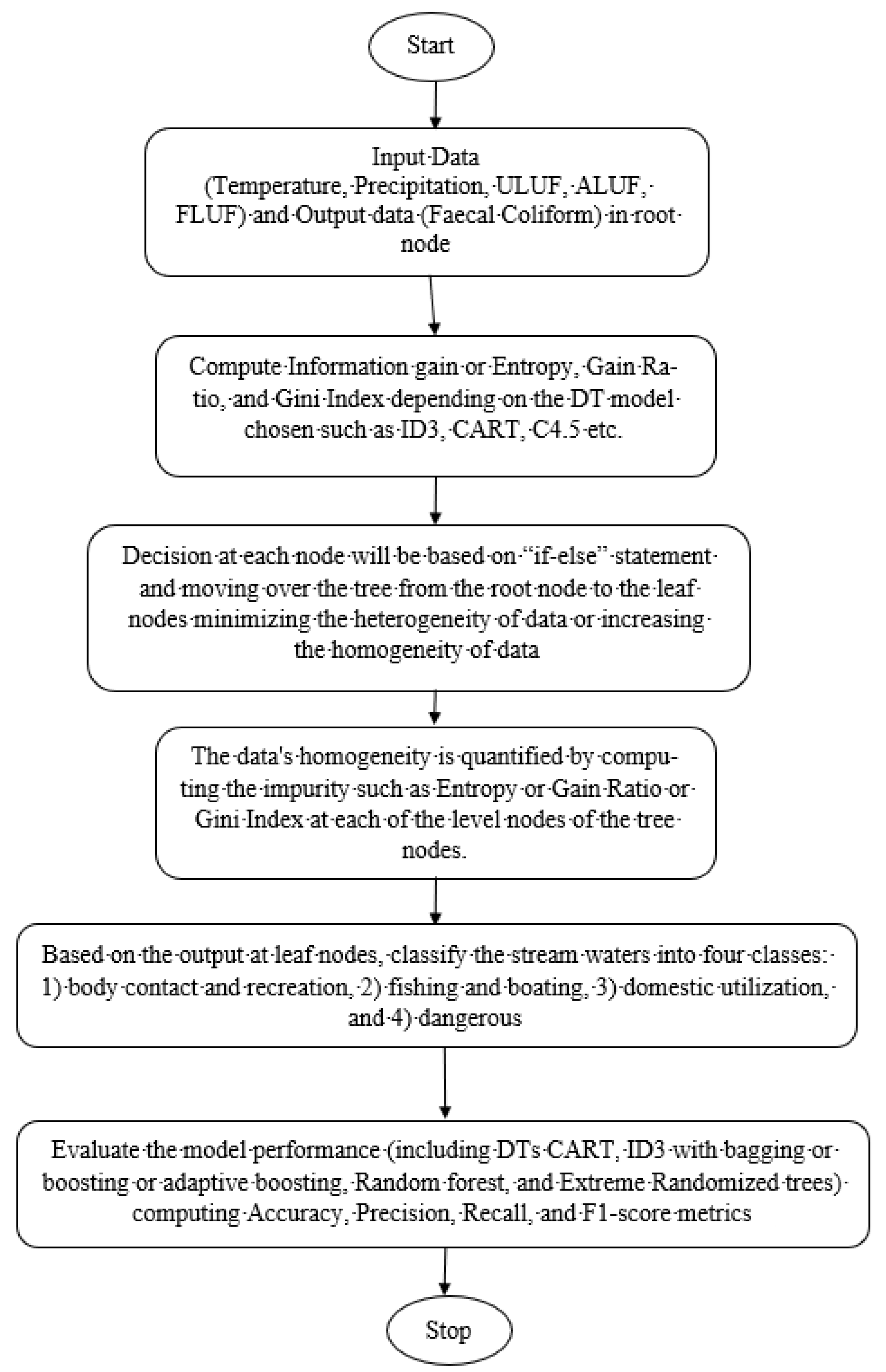
3.2. Decision Trees
3.3. Attribute Selection Measures
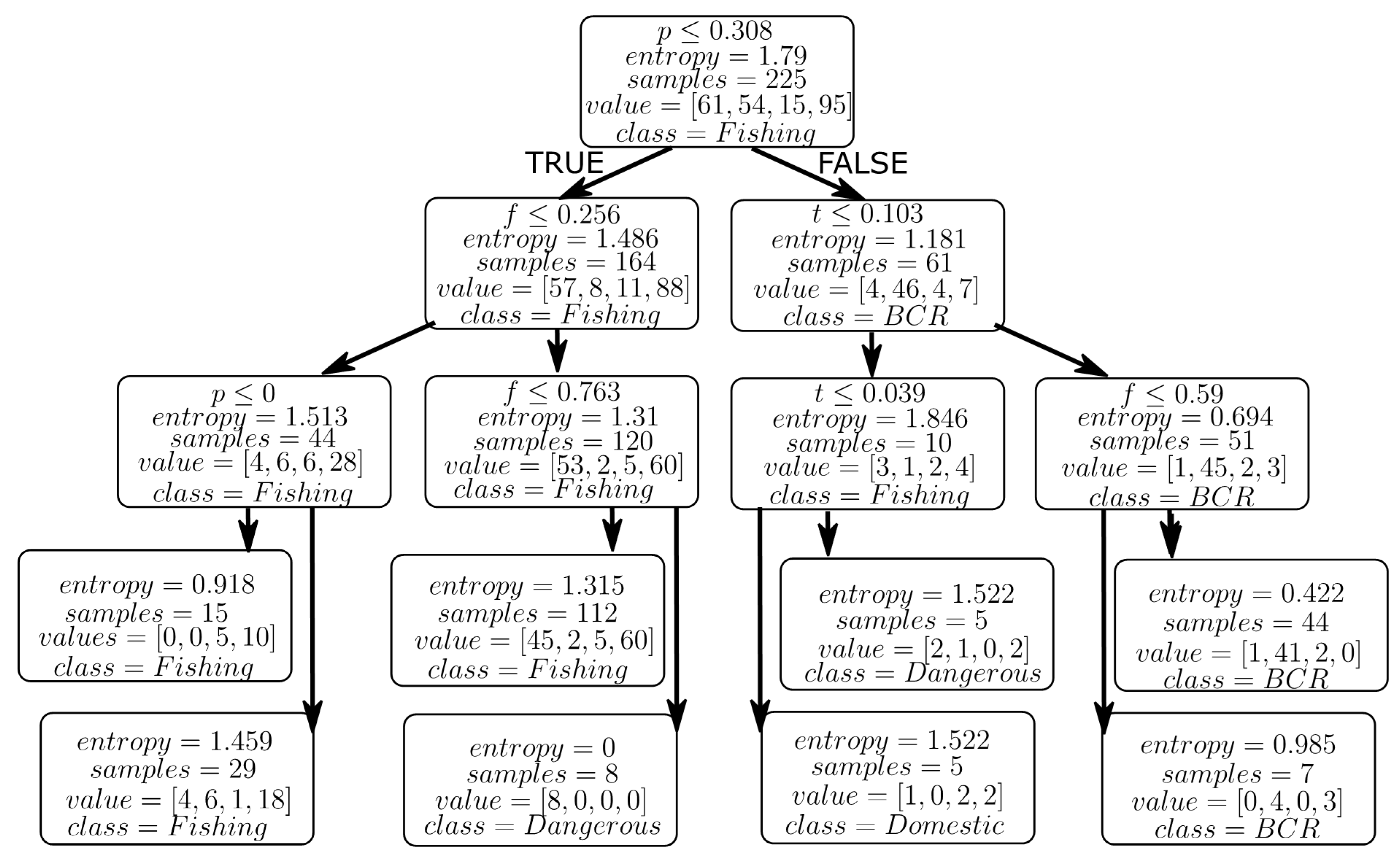
3.3.1. Entropy
3.3.2. Information Gain
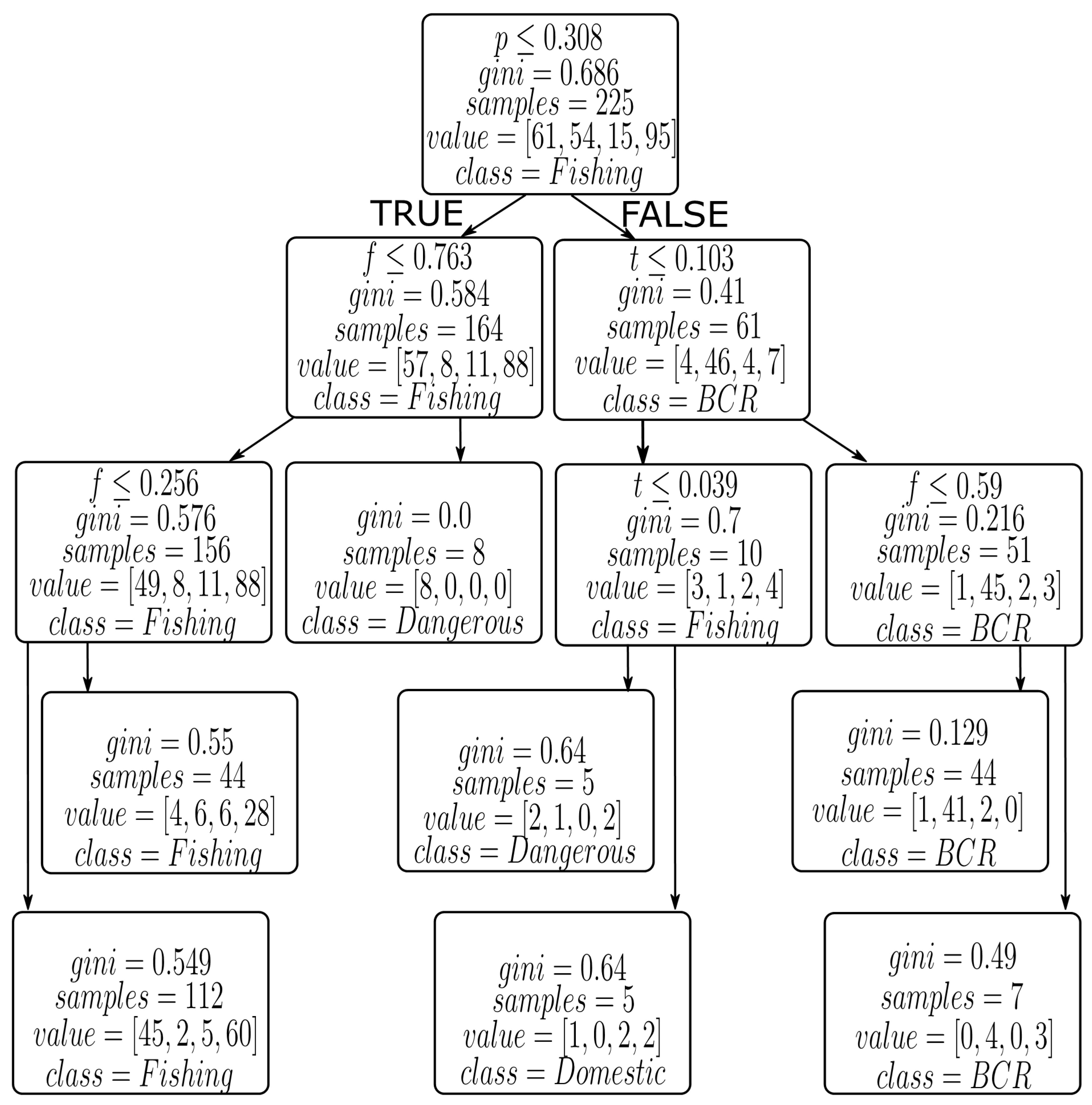
3.3.3. Gini Index
3.3.4. Gain Ratio
3.4. Bagging and Boosting
4. Results and Discussion
4.1. Overview
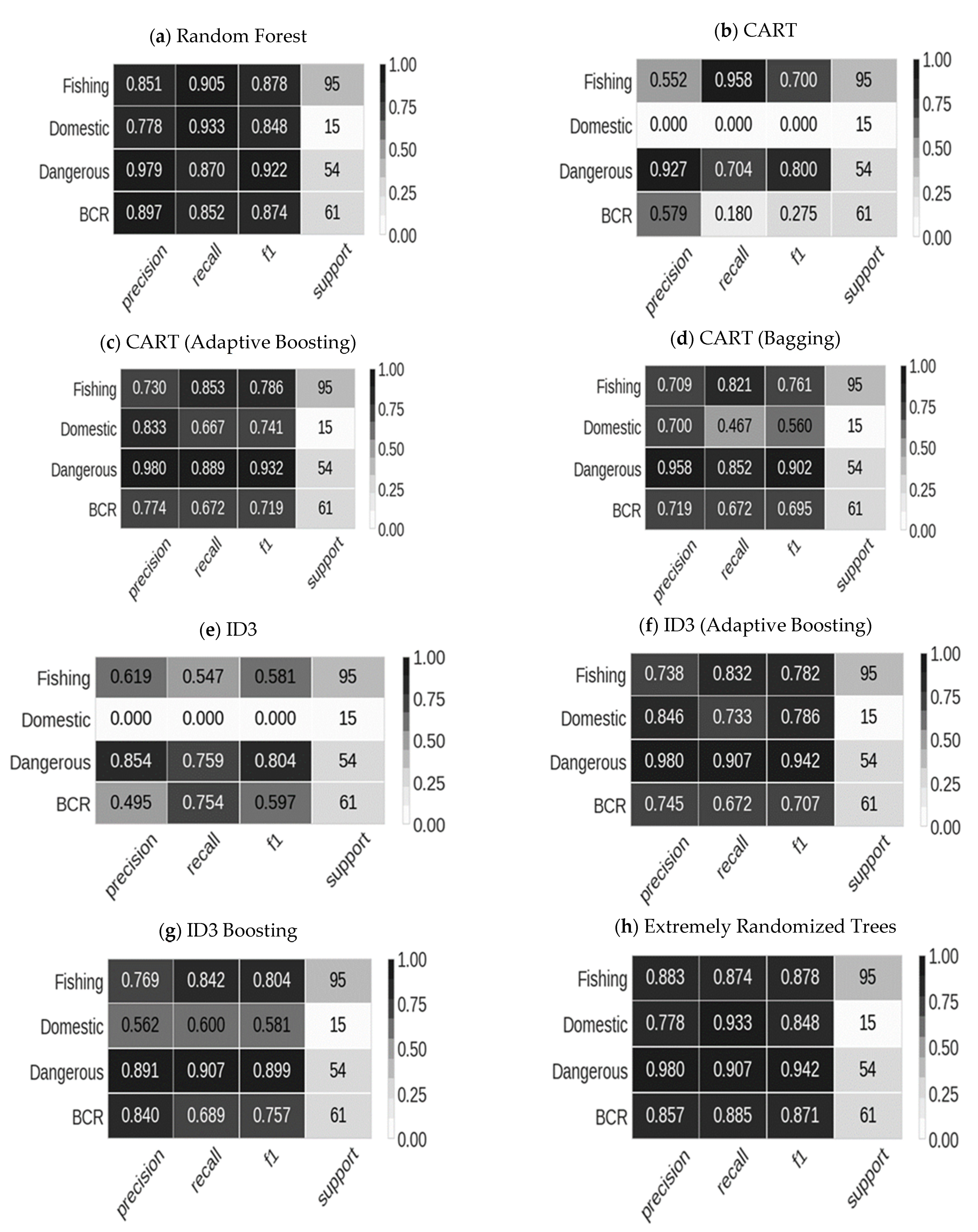
4.2. Results from Decision Tree Models
4.3. CART with Bagging and Adaptive Boosting
4.4. ID3 with Bagging and Adaptive Boosting
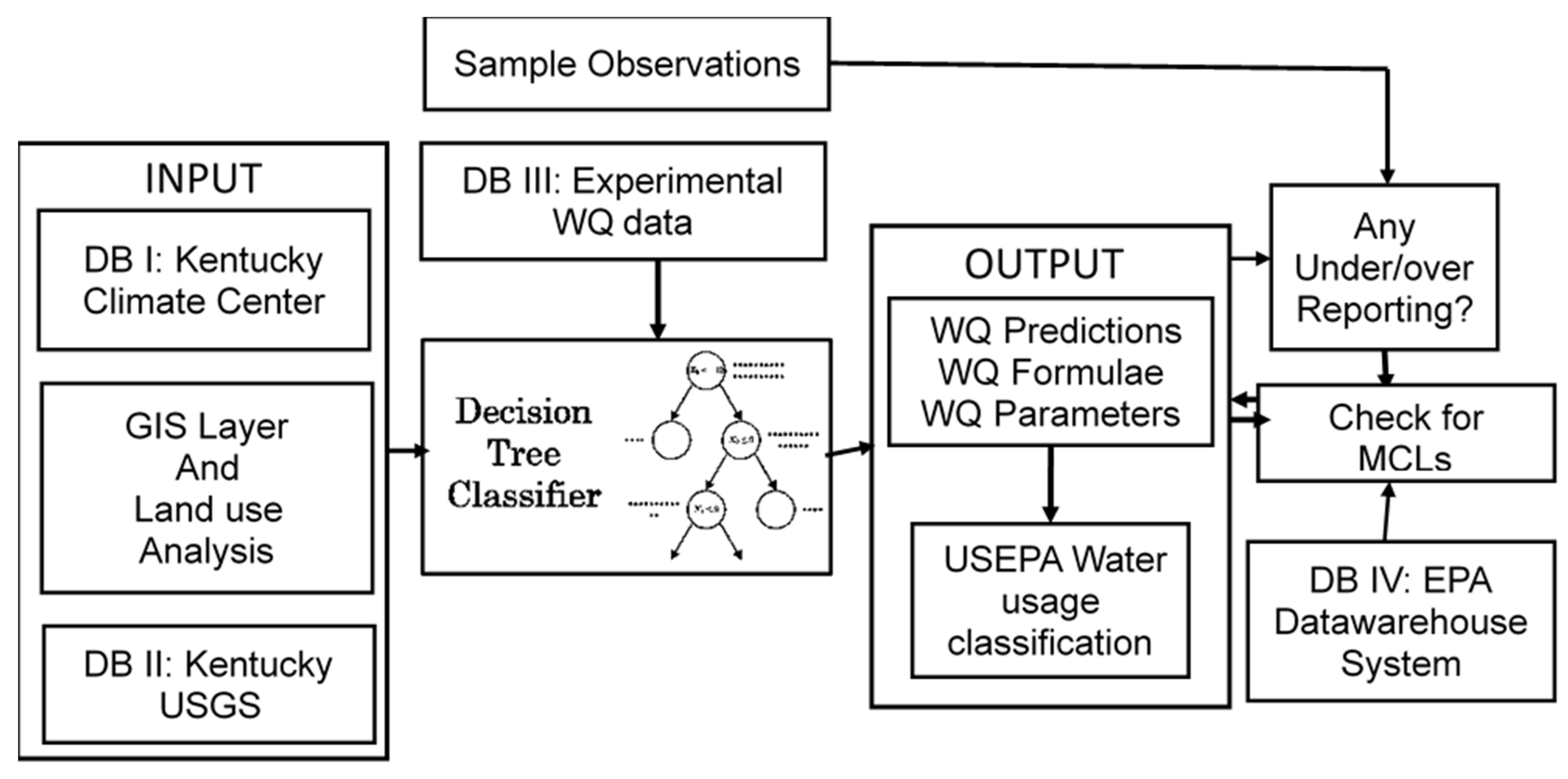
4.5. Decision Tree Classifier Based Decision Support System (DTCDSS)
5. Conclusions
- (i)
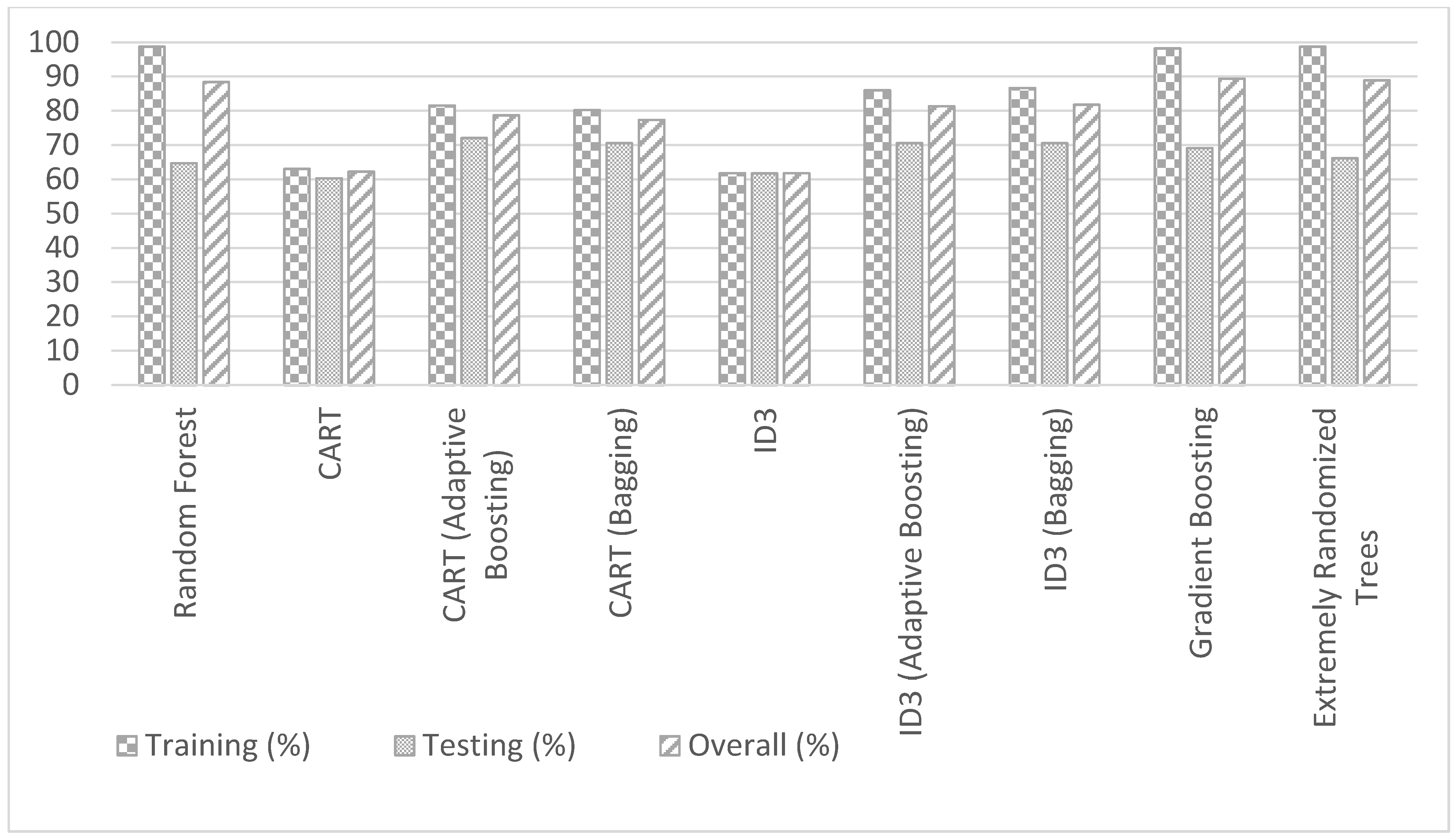
- The Decision Trees of Gradient Boosting (GB), Extremely Randomized Trees (ERT), and RF perform better than simple (without bagging and boosting) ID3, and CART models in training, testing, and overall.
- (ii)
- The bagging and adaptive boosting Decision Trees of CART, and ID3 significantly improve the performance over simple (without bagging and boosting) CART, and ID3 models.
- (iii)
- The performances of bagging and adaptive boosting Decision Trees of CART, and ID3 are slightly better than GB, ERT, and RF in testing. However, the training and overall accuracies of GB, ERT, and RF are better than all the models (including bagging and adaptive boosting) of CART and ID3.
- (iv)
- The Decision Tree models of GB, ERT, and RF are more consistent than other models in training, testing, and overall accuracies.
- (v)
- Overtraining the trees increases training accuracy at the expense of testing accuracy. A judicious choice need to be made in cutting down the trees, so that an optimal performance of training, testing, and overall accuracies is obtained.
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Ormsbee, L.J.; Anmala, S.E.M. Total Maximum Daily Load (TMDL) Development for Eagle Creek; Report submitted to Kentucky Department for Environmental Protection, Division of Water, Frankfort; University of Kentucky: Lexington, Kentucky, 2002. [Google Scholar]
- D’Agostino, V.; Greene, E.A.; Passarella, G.; Vurro, M. Spatial and temporal study of nitrate concentration in groundwater by means of coregionalization. Environ. Earth Sci. 1998, 36, 285–295. [Google Scholar] [CrossRef]
- Gaus, I.; Kinniburgh, D.G.; Talbot, J.C.; Webster, R. Geostatistical analysis of arsenic concentration in groundwater in Bangladesh using disjunctive kriging. Environ. Earth Sci. 2003, 44, 939–948. [Google Scholar] [CrossRef][Green Version]
- Arslan, H. Spatial and temporal mapping of groundwater salinity using ordinary kriging and indicator kriging: The case of Bafra Plain, Turkey. Agric. Water Manag. 2012, 113, 57–63. [Google Scholar] [CrossRef]
- Ahn, H.; Chon, H. Assessment of groundwater contamination using geographic information systems. Environ. Geochem. Health 1999, 21, 273–289. [Google Scholar] [CrossRef]
- Bae, H.-K.; Olson, B.H.; Hsu, K.-L.; Sorooshian, S. Classification and regression tree (CART) analysis for indicator bacterial concentration prediction for a Californian coastal area. Water Sci. Technol. 2010, 61, 545–553. [Google Scholar] [CrossRef]
- Liao, H.; Sun, W. Forecasting and Evaluating Water Quality of Chao Lake based on an Improved Decision Tree Method. Procedia Environ. Sci. 2010, 2, 970–979. [Google Scholar] [CrossRef]
- Nikoo, M.R.; Karimi, A.; Kerachian, R.; Poorsepahy-Samian, H.; Daneshmand, F. Rules for Optimal Operation of Reservoir-River-Groundwater Systems Considering Water Quality Targets: Application of M5P Model. Water Resour. Manag. 2013, 27, 2771–2784. [Google Scholar] [CrossRef]
- Azam, M.; Aslam, M.; Khan, K.; Mughal, A.; Inayat, A. Comparisons of decision tree methods using water data. Commun. Stat. -Simul. Comput. 2016, 46, 2924–2934. [Google Scholar] [CrossRef]
- Maier, P.M.; Keller, S. Machine Learning Regression on Hyperspectral Data to Estimate Multiple Water Parameters. In 2018 9th Workshop on Hyperspectral Image and Signal Processing: Evolution in Remote Sensing (WHISPERS); IEEE: Piscataway, NJ, USA, 2018; pp. 1–5. [Google Scholar]
- Jerves-Cobo, R.; Córdova-Vela, G.; Iñiguez-Vela, X.; Díaz-Granda, C.; Van Echelpoel, W.; Cisneros, F.; Nopens, I.; Goethals, P.L.M. Model-Based Analysis of the Potential of Macroinvertebrates as Indicators for Microbial Pathogens in Rivers. Water 2018, 10, 375. [Google Scholar] [CrossRef]
- Geetha Jenifel, M.; Jemila Rose, R. Recursive partitioning algorithm in water quality prediction. Int. J. Environ. Sci. Technol. 2020, 17, 745–754. [Google Scholar] [CrossRef]
- Ho, J.Y.; Afan, H.A.; El-Shafie, A.H.; Koting, S.B.; Mohd, N.S.; Jaafar, W.Z.B.; Sai, H.L.; Malek, M.A.; Ahmed, A.N.; Mohtar, W.H.M.W.; et al. Towards a time and cost effective approach to water quality index class prediction. J. Hydrol. 2019, 575, 148–165. [Google Scholar] [CrossRef]
- Sepahvand, A.; Singh, B.; Sihag, P.; Samani, A.N.; Ahmadi, H.; Nia, S.F. Assessment of the various soft computing techniques to predict sodium absorption ratio (SAR). ISH J. Hydraul. Eng. 2019, 1–12. [Google Scholar] [CrossRef]
- Lu, H.; Ma, X. Hybrid decision tree-based machine learning models for short-term water quality prediction. Chemosphere 2020, 249, 126169. [Google Scholar] [CrossRef]
- Shin, Y.; Kim, T.; Hong, S.; Lee, S.; Lee, E.; Hong, S.; Lee, C.; Kim, T.; Park, M.S.; Park, J.; et al. Prediction of Chlorophyll-a Concentrations in the Nakdong River Using Machine Learning Methods. Water 2020, 12, 1822. [Google Scholar] [CrossRef]
- Mosavi, A.; Hosseini, F.S.; Choubin, B.; Abdolshahnejad, M.; Gharechaee, H.; Lahijanzadeh, A.; Dineva, A.A. Susceptibility Prediction of Groundwater Hardness Using Ensemble Machine Learning Models. Water 2020, 12, 2770. [Google Scholar] [CrossRef]
- Naloufi, M.; Lucas, F.S.; Souihi, S.; Servais, P.; Janne, A.; De Abreu, T.W.M. Evaluating the Performance of Machine Learning Approaches to Predict the Microbial Quality of Surface Waters and to Optimize the Sampling Effort. Water 2021, 13, 2457. [Google Scholar] [CrossRef]
- Chen, K.; Chen, H.; Zhou, C.; Huang, Y.; Qi, X.; Shen, R.; Liu, F.; Zuo, M.; Zou, X.; Wang, J.; et al. Comparative analysis of surface water quality prediction performance and identification of key water parameters using different machine learning models based on big data. Water Res. 2020, 171, 115454. [Google Scholar] [CrossRef]
- Alizamir, M.; Heddam, S.; Kim, S.; Mehr, A.D. On the implementation of a novel data-intelligence model based on extreme learning machine optimized by bat algorithm for estimating daily chlorophyll-a concentration: Case studies of river and lake in USA. J. Clean. Prod. 2021, 285, 124868. [Google Scholar] [CrossRef]
- Khullar, S.; Singh, N. Machine learning techniques in river water quality modelling: A research travelogue. Water Supply 2021, 21, 1–13. [Google Scholar] [CrossRef]
- Asadollah, S.B.H.S.; Sharafati, A.; Motta, D.; Yaseen, Z.M. River water quality index prediction and uncertainty analysis: A comparative study of machine learning models. J. Environ. Chem. Eng. 2021, 9, 104599. [Google Scholar] [CrossRef]
- Zounemat-Kermani, M.; Alizamir, M.; Fadaee, M.; Namboothiri, A.S.; Shiri, J. Online sequential extreme learning machine in river water quality (turbidity) prediction: A comparative study on different data mining approaches. Water Environ. J. 2021, 35, 335–348. [Google Scholar] [CrossRef]
- Bui, D.T.; Khosravi, K.; Tiefenbacher, J.; Nguyen, H.; Kazakis, N. Improving prediction of water quality indices using novel hybrid machine-learning algorithms. Sci. Total. Environ. 2020, 721, 137612. [Google Scholar] [CrossRef]
- Jeihouni, M.; Toomanian, A.; Mansourian, A. Decision Tree-Based Data Mining and Rule Induction for Identifying High Quality Groundwater Zones to Water Supply Management: A Novel Hybrid Use of Data Mining and GIS. Water Resour. Manag. 2020, 34, 139–154. [Google Scholar] [CrossRef]
- Saghebian, S.M.; Sattari, M.T.; Mirabbasi, R.; Pal, M. Ground water quality classification by decision tree method in Ardebil region, Iran. Arab. J. Geosci. 2014, 7, 4767–4777. [Google Scholar] [CrossRef]
- Anmala, J.; Turuganti, V. Comparison of the performance of decision tree (DT) algorithms and extreme learning machine (ELM) model in the prediction of water quality of the Upper Green River watershed. Water Environ. Res. 2021. [Google Scholar] [CrossRef] [PubMed]
- Status Report, Green and Tradewater Basins; Kentucky Division of Water: Frankfort, KY, USA, 2001.
- Anmala, J.; Meier, O.W.; Meier, A.J.; Grubbs, S. A GIS and an artificial neural network based water quality model for a stream network in Upper Green River Basin, Kentucky, USA. ASCE J. Environm. Eng. 2015, 141, 04014082. [Google Scholar] [CrossRef]
- Kaplan, C.; Pasternack, B.; Shah, H.; Gallo, G. Age-related incidence of sclerotic glomeruli in human kidneys. Am. J. Pathol. 1975, 80, 227–234. [Google Scholar]
- Sokal, R.R.; Rohlf, J.F. Biometry: The Principle and Practice of Statistics in Biological Research, 2nd ed.; W.H. Freeman and Company: San Francisco, CA, USA, 1981. [Google Scholar]
- Snedecor, G.W.; Cochran, W.G. Statistical Methods, 8th ed.; Iowa University Press: Ames, IA, USA, 1989. [Google Scholar] [CrossRef]
- Rao, P.V. Statistical Research Methods in the Life Sciences; Duxbury Press: Austin, TX, USA, 1998; 889p. [Google Scholar]
- Sutton, C.D. Classification and Regression Trees, Bagging, and Boosting. Hand. Stat. 2005, 24, 303–329. [Google Scholar] [CrossRef]
- Bramer, M. Principles of Data Mining; Springer: London, UK, 2007; 343p, ISBN 978-1-84628-765-7. [Google Scholar]
- Quinlan, J.R. Induction of Decision Trees. Mach. Learn. 1986, 1, 81–106. [Google Scholar] [CrossRef]
- Breiman, L.; Firedman, J.H.; Olshen, R.A.; Stone, C.J. Classification and Regression Trees; Chapman & Hall/CRC: Boca Raton, FL, USA, 1984; ISBN 978-0-412-04841-8. [Google Scholar]
- Breiman, L. Bagging predictors. Mach. Learn. 1996, 24, 123–140. [Google Scholar] [CrossRef]
- Tan, P.-N.; Steinbach, M. Introduction to Data Mining; Pearson India Education Services Pvt. Ltd: Bengaluru, India, 2016; p. 760. [Google Scholar]
- Shehane, S.; Harwood, V.; Whitlock, J.; Rose, J. The influence of rainfall on the incidence of microbial faecal indicators and the dominant sources of faecal pollution in a Florida river. J. Appl. Microbiol. 2005, 98, 1127–1136. [Google Scholar] [CrossRef]
- Santiago-Rodriguez, T.M.; Tremblay, R.L.; Toledo-Hernandez, C.; Gonzalez-Nieves, J.E.; Ryu, H.; Domingo, J.W.S.; Toranzos, G.A. Microbial Quality of Tropical Inland Waters and Effects of Rainfall Events. Appl. Environ. Microbiol. 2012, 78, 5160–5169. [Google Scholar] [CrossRef]
- Islam, M.M.M.; Hofstra, N.; Islam, A. The Impact of Environmental Variables on Faecal Indicator Bacteria in the Betna River Basin, Bangladesh. Environ. Process. 2017, 4, 319–332. [Google Scholar] [CrossRef]
- Leight, A.K.; Crump, B.C.; Hood, R. Assessment of Fecal Indicator Bacteria and Potential Pathogen Co-Occurrence at a Shellfish Growing Area. Front. Microbiol. 2018, 9, 384. [Google Scholar] [CrossRef]
- Seo, M.; Lee, H.; Kim, Y. Relationship between Coliform Bacteria and Water Quality Factors at Weir Stations in the Nakdong River, South Korea. Water 2019, 11, 1171. [Google Scholar] [CrossRef]
- Kagalou, I.; Tsimarakis, G.; Bezirtzoglou, E. Inter-relationships between Bacteriological and Chemical Variations in Lake Pamvotis—Greece. Microb. Ecol. Health Dis. 2002, 14, 37–41. [Google Scholar] [CrossRef]
- Jin, G.; Englande, A.; Bradford, H.; Jeng, H.-W. Comparison of E.Coli, Enterococci, and Fecal Coliform as Indicators for Brackish Water Quality Assessment. Water Environ. Res. 2004, 76, 245–255. [Google Scholar] [CrossRef] [PubMed]
- Venkateswarlu, T.; Anmala, J.; Dharwa, M. PCA, CCA, and ANN modeling of climate and land-use effects on stream water quality of Karst watershed in Upper Green River, Kentucky, USA. ASCE J. Hydrol. Eng. 2020, 25, 05020008. [Google Scholar] [CrossRef]
- Breiman, L. Random forests. Mach. Learn. 2001, 45, 5–32. [Google Scholar] [CrossRef]
- Geurts, P.; Ernst, D.; Wehenkel, L. Extremely randomized trees. Mach. Learn. 2006, 63, 3–42. [Google Scholar] [CrossRef]
- Breiman, L. Arcing the Edge; Technical Report 486, Ann. Prob; Statistics Department, University of California: Berkeley, CA, USA, 1997; Volume 26, pp. 1683–1702. [Google Scholar]
- Freund, Y.; Schapire, R.E. A short introduction to Boosting. J. Jpn. Soc. Artif. Intell. 1999, 14, 771–780. [Google Scholar]
- Mitchell, T.M. Machine Learning, 1st ed.; Mc-Graw Hill Education: New York, NY, USA, 1997. [Google Scholar]

| Water Quality Parameter | Variable | Sum | Average | Standard Deviation | Input/Output |
|---|---|---|---|---|---|
| Precipitation (in cm) | P | 817.0 | 3.6 | 4.21 | Input |
| Temperature (°C) | T | 4250.7 | 18.9 | 4.36 | Input |
| Urban land-use factor | U | 35.3 | 0.16 | 0.12 | Input |
| Forest land-use factor | F | 165.2 | 0.73 | 0.12 | Input |
| Agricultural land-use factor | A | 178.0 | 0.79 | 0.11 | Input |
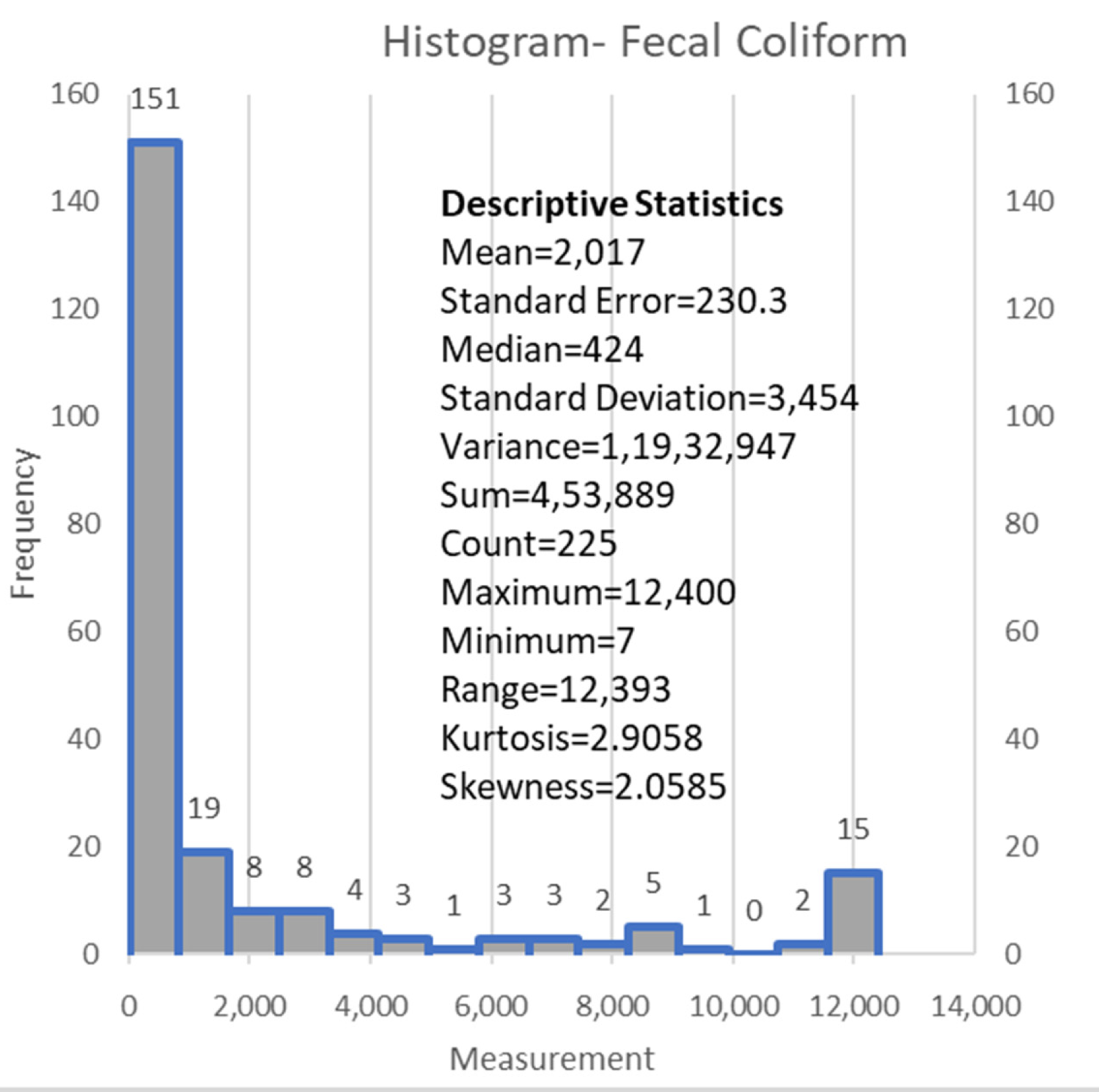
| Fecal Coliform (#colonies/100 mL) | FC | 453,889 | 2017.3 | 3454 | Output |
| Model | Training (%) | Testing (%) | Overall (%) |
|---|---|---|---|
| CART (Adaptive Boosting) | 81.53 | 72.06 | 78.67 |
| ID3-Bagg | 86.62 | 70.58 | 81.78 |
| ID3-AB | 85.98 | 70.58 | 81.33 |
| CART (Bagging) | 80.25 | 70.58 | 77.33 |
| Gradient Boosting (GBM) | 98.19 | 69.12 | 89.33 |
| Extremely Randomized Trees (ERT) | 98.72 | 66.17 | 88.89 |
| Random Forest (RF) | 98.70 | 64.70 | 88.40 |
| ID3 | 61.78 | 61.76 | 61.77 |
| CART | 63.05 | 60.29 | 62.22 |
| Class | Fecal Coliform (FC) Range (cfu/100 mL) |
|---|---|
| Body contact and recreation (BCR) | 0 < FC ≤ 200 |
| Fishing and boating | 200 < FC ≤ 1000 |
| Domestic utilization | 1000 < FC ≤ 2000 |
| Dangerous | FC > 2000 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hannan, A.; Anmala, J. Classification and Prediction of Fecal Coliform in Stream Waters Using Decision Trees (DTs) for Upper Green River Watershed, Kentucky, USA. Water 2021, 13, 2790. https://doi.org/10.3390/w13192790
Hannan A, Anmala J. Classification and Prediction of Fecal Coliform in Stream Waters Using Decision Trees (DTs) for Upper Green River Watershed, Kentucky, USA. Water. 2021; 13(19):2790. https://doi.org/10.3390/w13192790
Chicago/Turabian StyleHannan, Abdul, and Jagadeesh Anmala. 2021. "Classification and Prediction of Fecal Coliform in Stream Waters Using Decision Trees (DTs) for Upper Green River Watershed, Kentucky, USA" Water 13, no. 19: 2790. https://doi.org/10.3390/w13192790
APA StyleHannan, A., & Anmala, J. (2021). Classification and Prediction of Fecal Coliform in Stream Waters Using Decision Trees (DTs) for Upper Green River Watershed, Kentucky, USA. Water, 13(19), 2790. https://doi.org/10.3390/w13192790






