Optimisation of Various Physicochemical Variables Affecting Molybdenum Bioremediation Using Antarctic Bacterium, Arthrobacter sp. Strain AQ5-05
Abstract
:1. Introduction
2. Materials and Methods
2.1. Sample Collection and Maintenance
2.2. Screening the Bacterial Strain for Mo-Reducing Potential
2.3. Optimisation of Mo Reduction
2.3.1. One-Factor-at-a-Time (OFAT)
2.3.2. Response Surface Methodology (RSM)
3. Results and Discussion
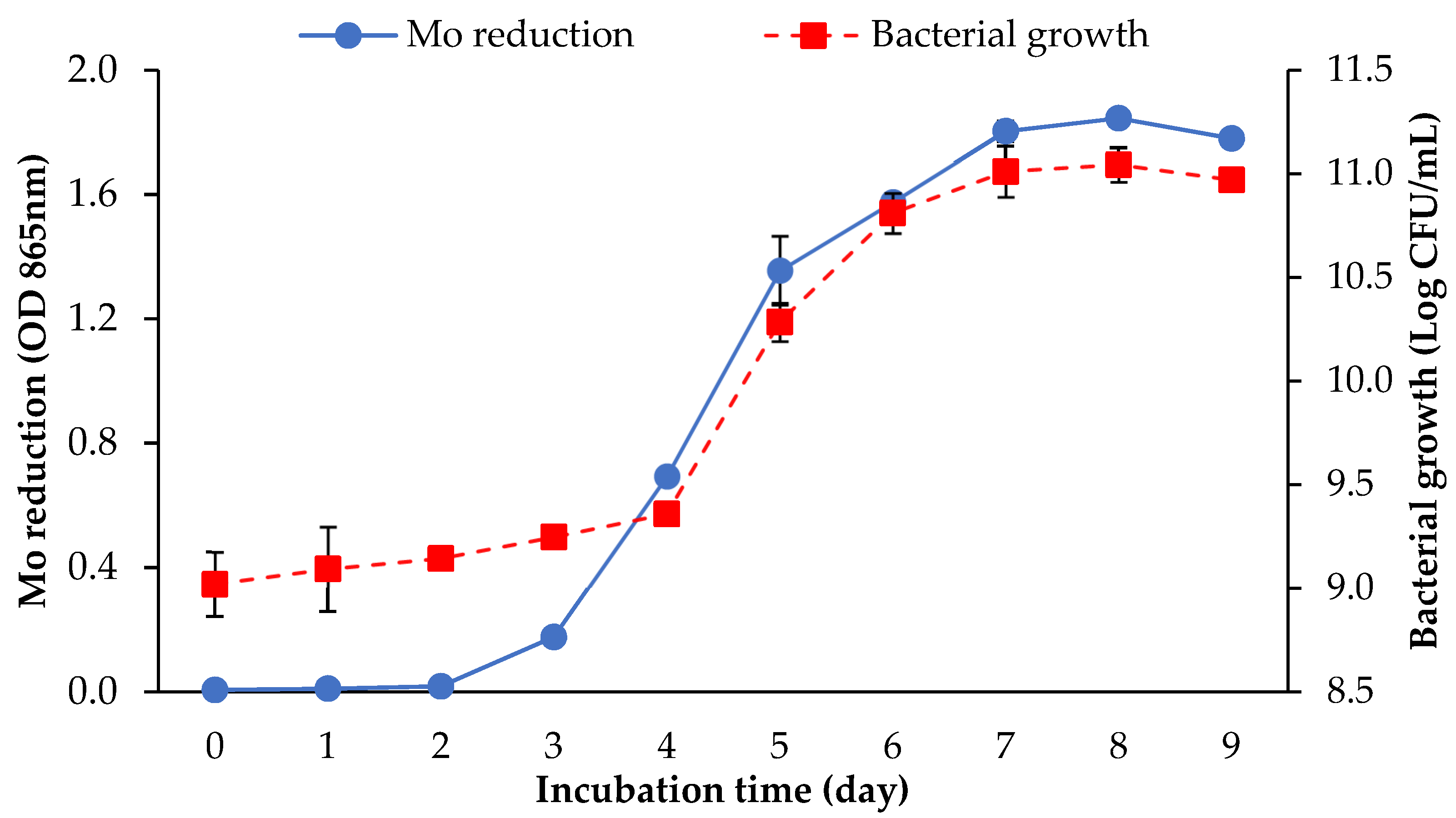
3.1. Screening of the Mo-Reducing Potential of Arthrobacter sp. Strain AQ5-05
3.2. Optimisation of Mo Reduction Using One-Factor-at-a-Time (OFAT)
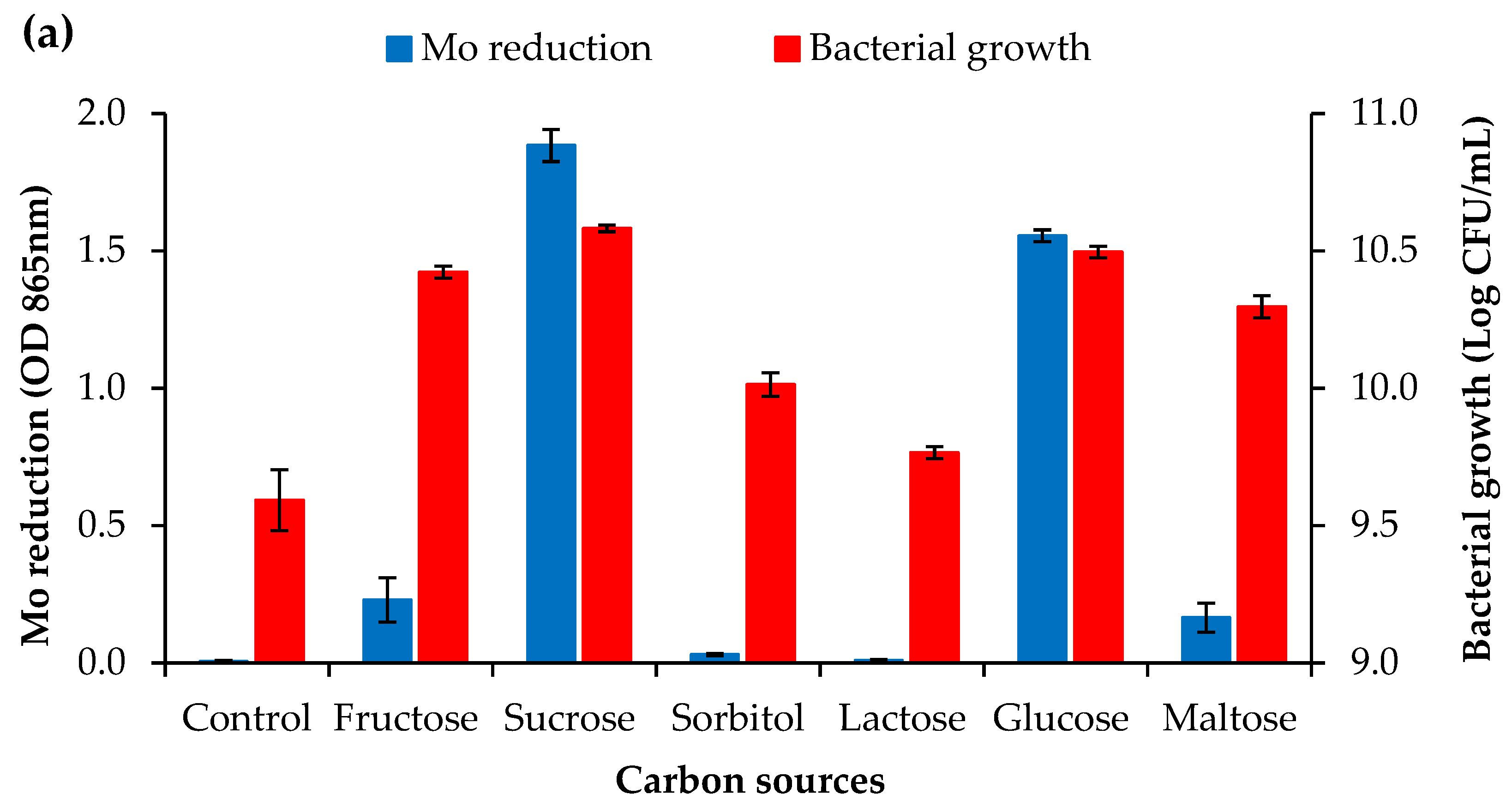
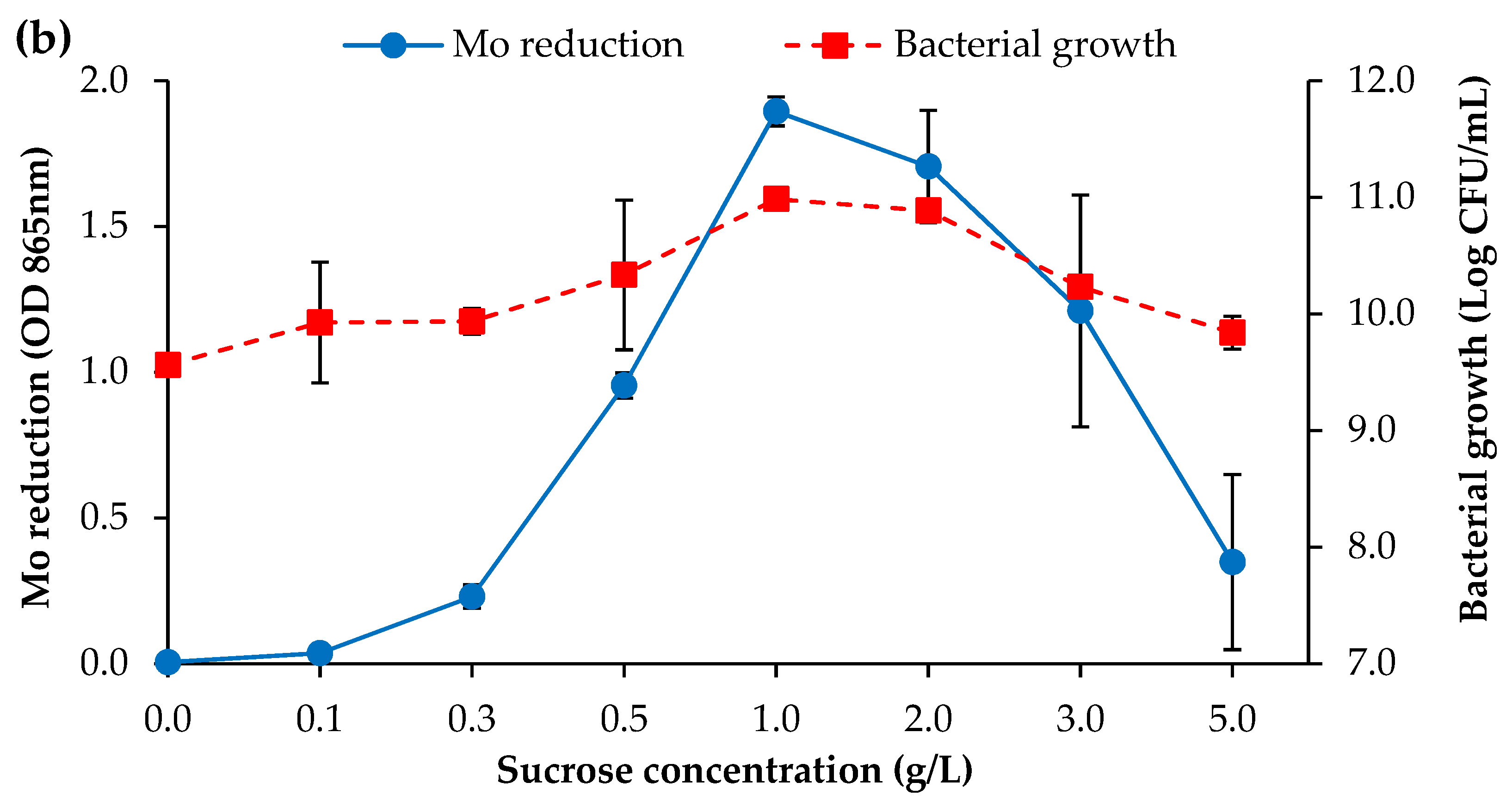
3.2.1. Carbon Source Concentration
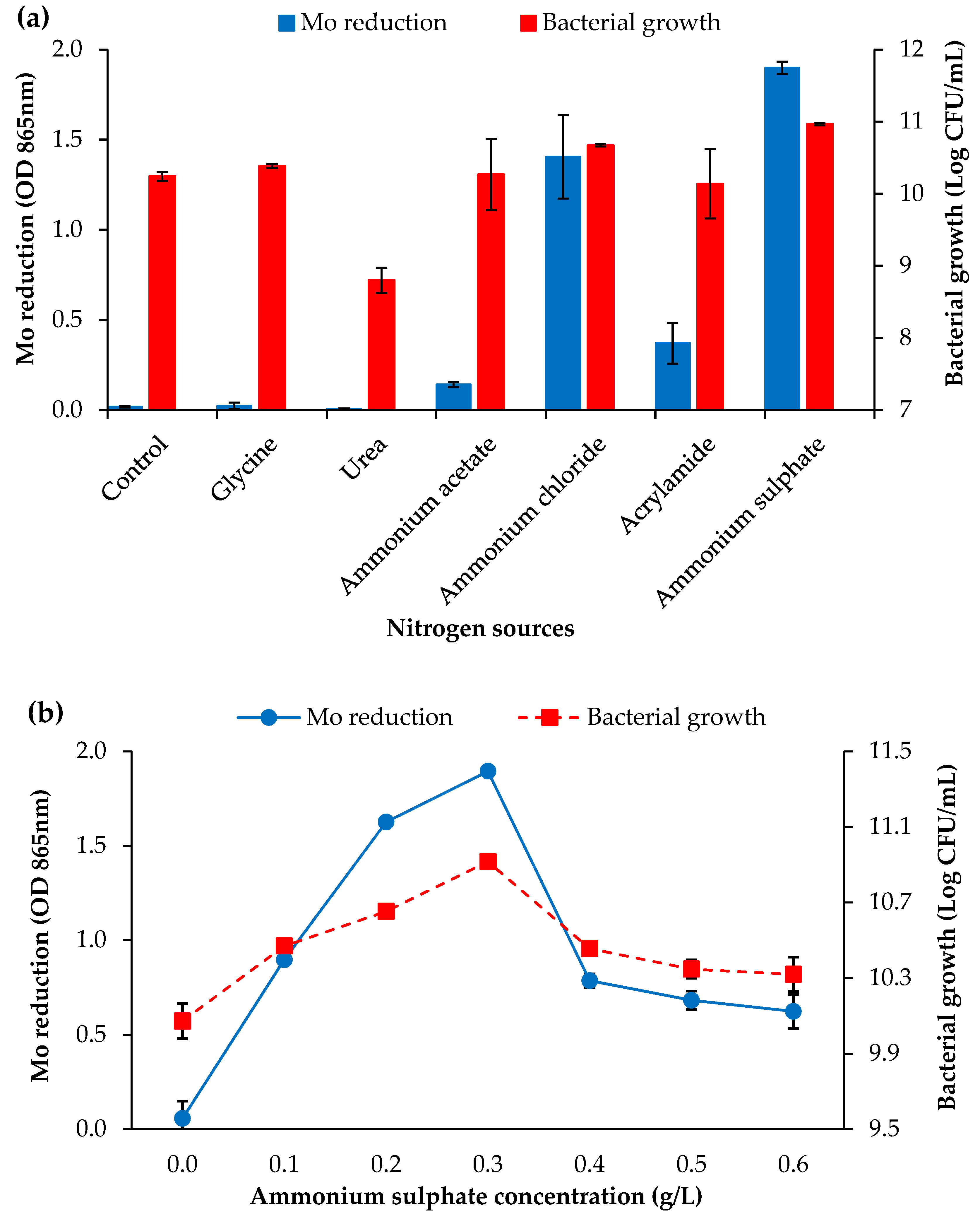
3.2.2. Nitrogen Sources and Concentration
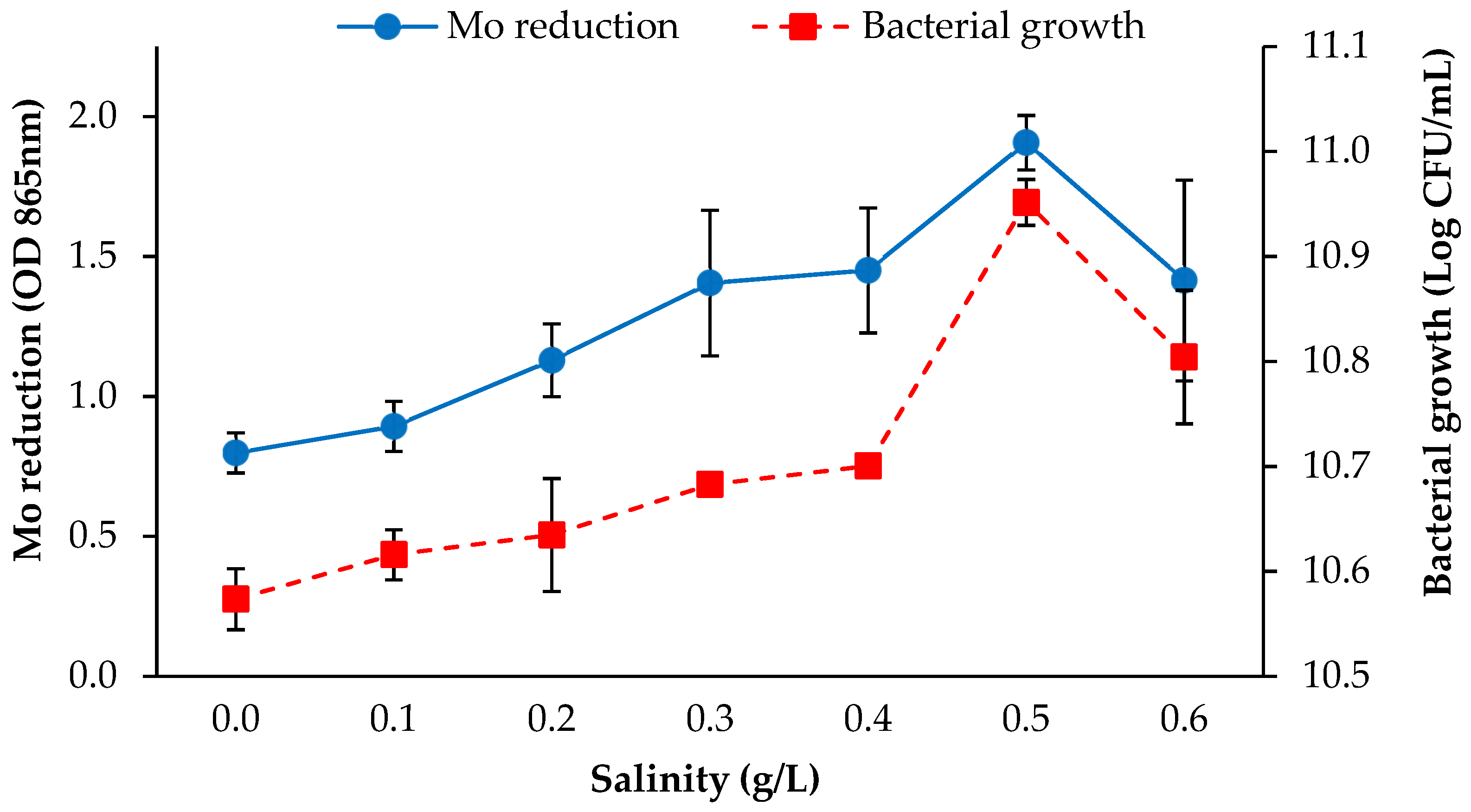
3.2.3. Salinity
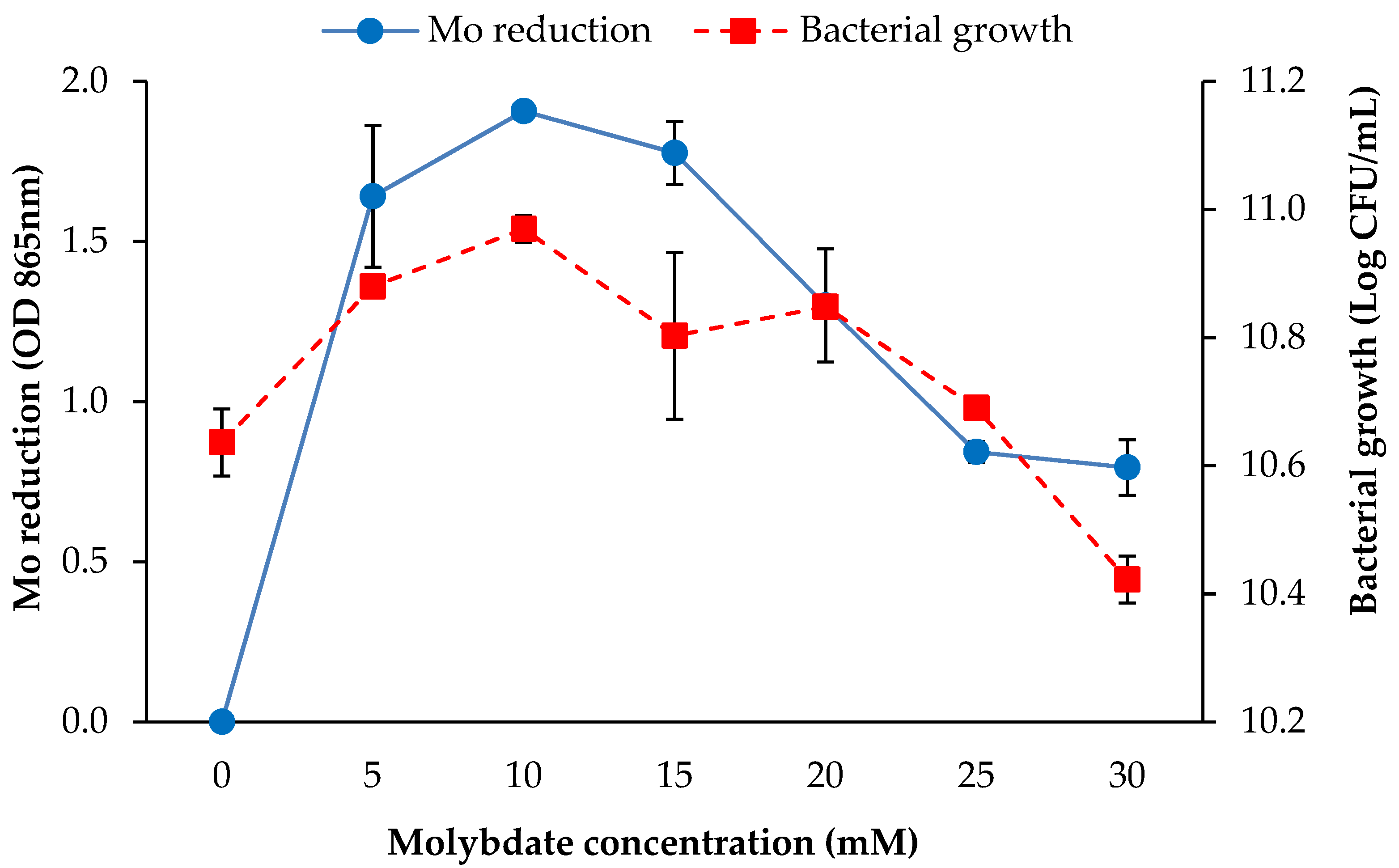
3.2.4. Molybdate Concentration
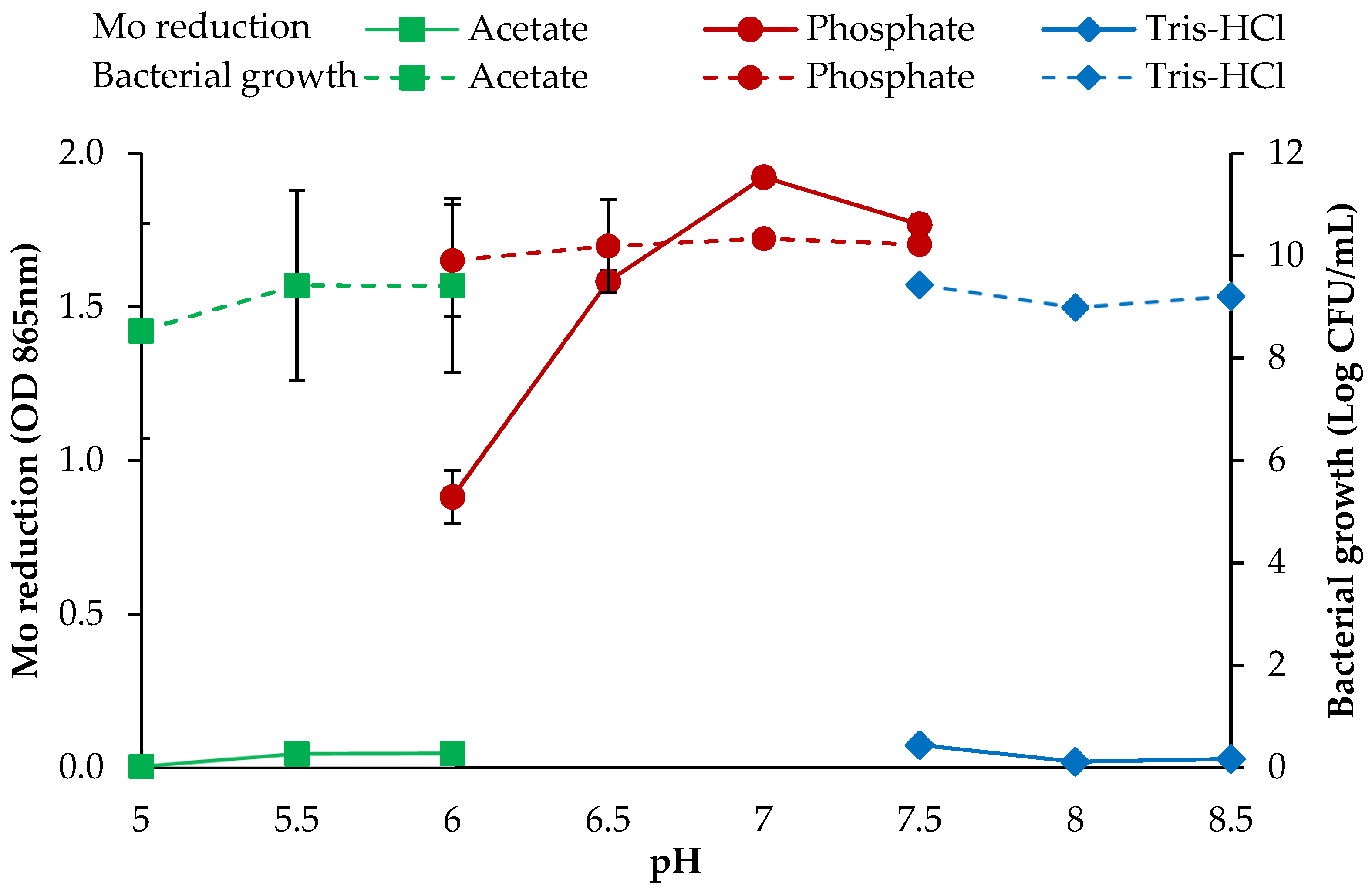
3.2.5. pH
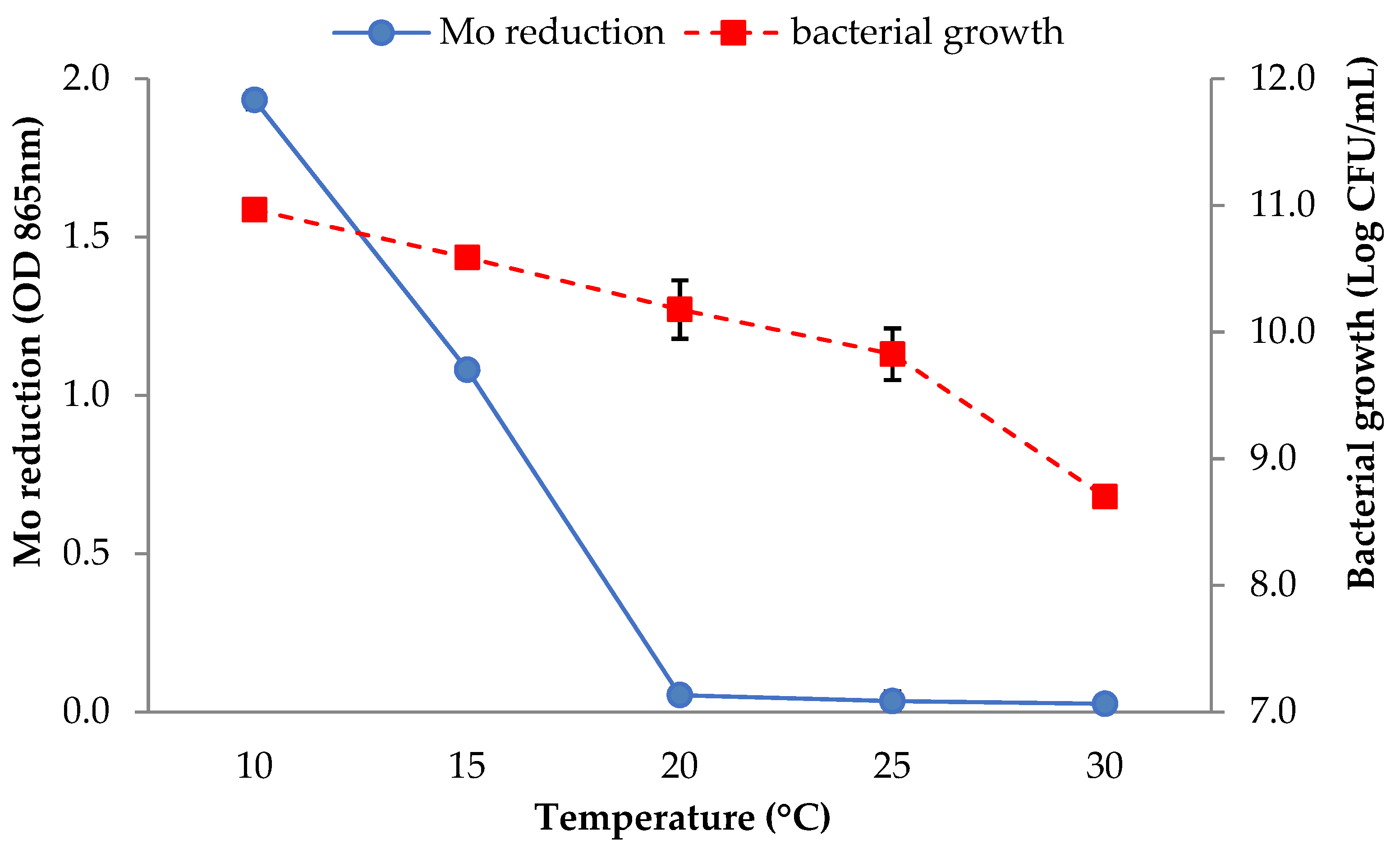
3.2.6. Temperature
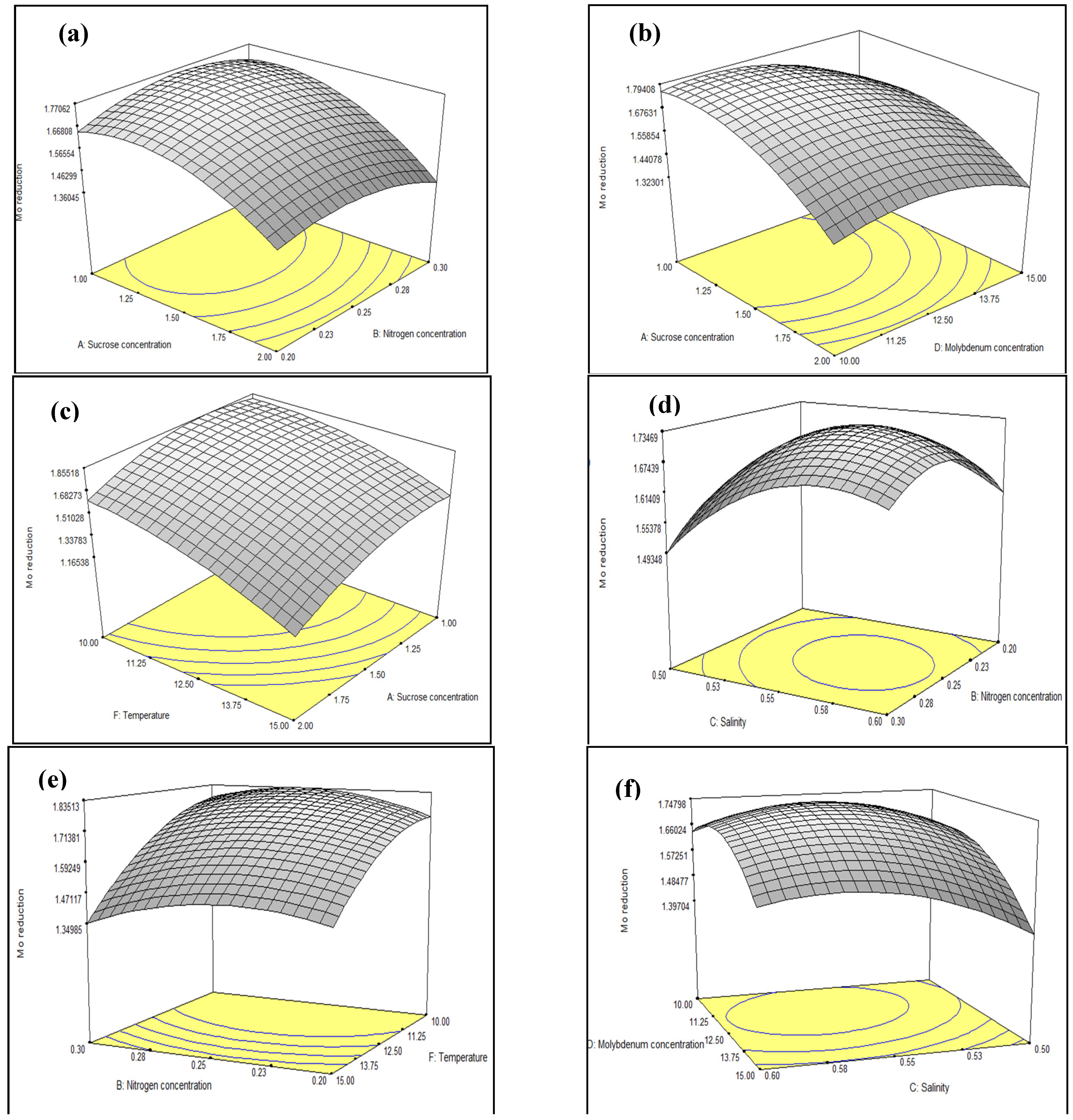
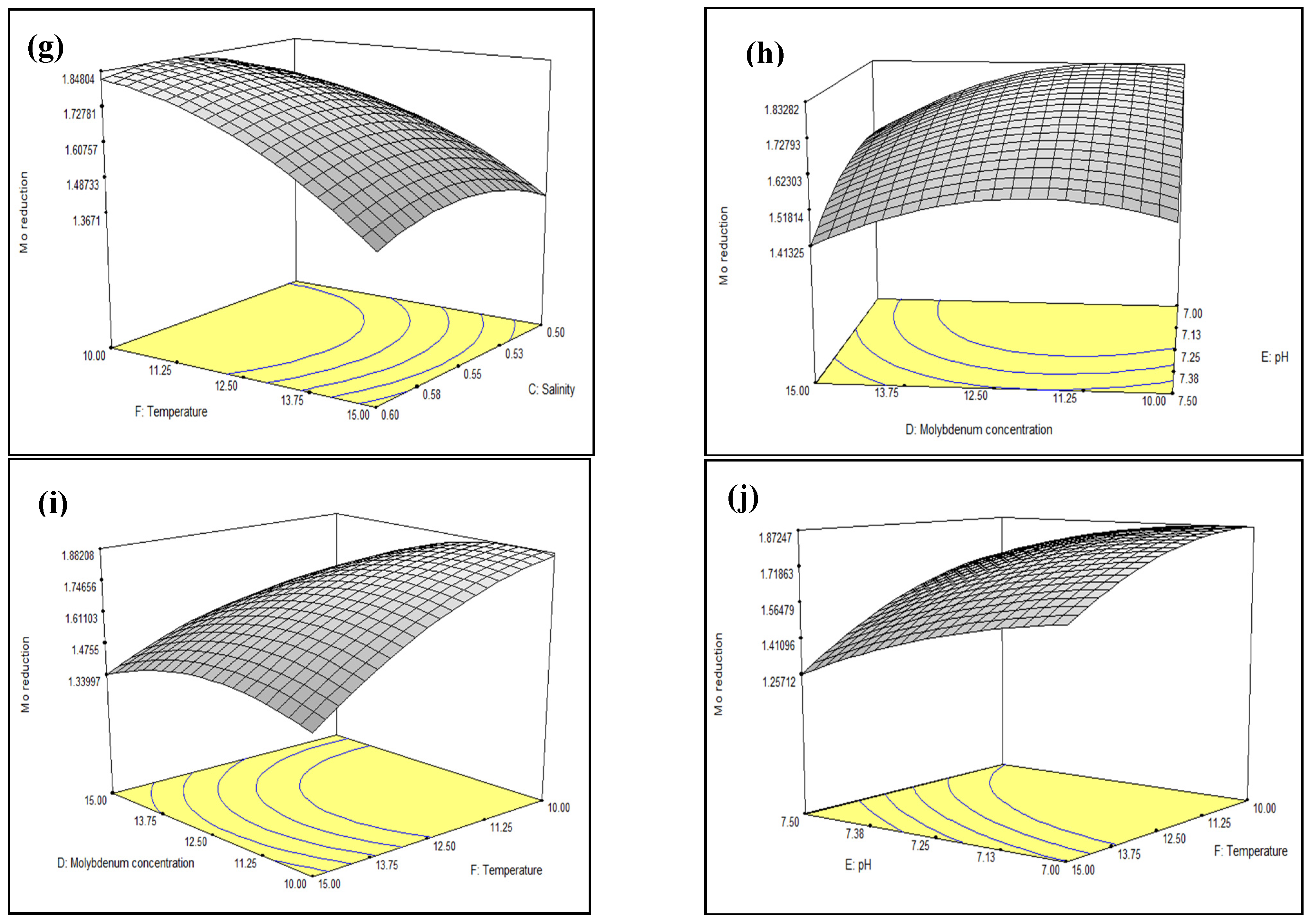
3.3. Optimisation of Mo Reduction Using Response Surface Methodology (RSM)
3.3.1. Central Composite Design (CCD)
3.3.2. Model Prediction and Validation
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Kulikova, O.I.; Fedorova, T.N.; Lopachev, A.V.; Orlova, V.S.; Grachev, V.A. Effects of antioxidants on the viability of the human neuroblastoma SH-SY5Y cell culture under the conditions of heavy-metal toxicity. Biol. Med. 2016, 8, 4–8. [Google Scholar] [CrossRef]
- Sabullah, M.K.; Rahman, M.F.; Ahmad, S.A.; Sulaiman, M.R.; Shukor, M.S.; Shamaan, N.A.; Shukor, M.Y. Assessing resistance and bioremediation ability of Enterobacter sp. strain Saw-1 on molybdenum in various heavy metals and pesticides. J. Math. Fundam. Sci. 2017, 49, 193–210. [Google Scholar] [CrossRef] [Green Version]
- Barra, F.; Alcota, H.; Rivera, S.; Valencia, V.; Munizaga, F.; Maksaev, V. Timing and formation of porphyry Cu-Mo mineralization in the Chuquicamata district, northern Chile: New constraints from the Toki cluster. Miner. Deposita. 2013, 48, 629–651. [Google Scholar] [CrossRef]
- Yang, N.; Welch, K.A.; Mohajerin, T.J.; Telfeyan, K.; Chevis, D.A.; Grimm, D.A.; Lyons, W.B.; White, C.D.; Johannesson, K.H. Comparison of arsenic and molybdenum geochemistry in meromictic lakes of the McMurdo Dry Valleys, Antarctica: Implications for oxyanion-forming trace element behavior in permanently stratified lakes. Chem. Geol. 2015, 404, 110–125. [Google Scholar] [CrossRef]
- Barakat, M.A. New trends in removing heavy metals from industrial wastewater. Arab. J. Chem. 2011, 4, 361–377. [Google Scholar] [CrossRef] [Green Version]
- Mansur, R.; Gusmanizar, N.; Roslan, M.A.; Ahmad, S.A.; Shukor, M.Y. Isolation and characterisation of a molybdenum-reducing and metanil yellow dye-decolourising Bacillus sp. strain neni-10 in soils from West Sumatera, Indonesia. Trop. Life Sci. Res. 2017, 28, 69–90. [Google Scholar] [CrossRef]
- Sabullah, M.K.; Rahman, M.F.; Ahmad, S.A.; Sulaiman, M.R.; Shukor, M.S.; Shamaan, N.A.; Shukor, M.Y. Isolation and characterization of a molybdenum-reducing and phenolic- and catechol-degrading Enterobacter sp. strain saw-2. Biotropia 2017, 24, 47–58. [Google Scholar] [CrossRef] [Green Version]
- Ahmad, S.A.; Shukor, M.Y.; Shamaan, N.A.; Mac Cormack, W.P.; Syed, M.A. Molybdate reduction to molybdenum blue by an Antarctic bacterium. BioMed Res. Int. 2013, 2013. [Google Scholar] [CrossRef]
- Lee, J.D. Concise Inorganic Chemistry; Van Reinhold Co.: New York, NY, USA, 1977; p. 325. [Google Scholar]
- Braithwaite, E.R. Molybdenum. In Specialty Inorganic Chemicals; Thompson, R., Ed.; The Royal Society of Chemistry, Burlington House: London, UK, 1981; pp. 350–351. [Google Scholar]
- Halmi, M.I.E.; Zuhainis, S.W.; Yusof, M.T.; Shaharuddin, N.A.; Helmi, W.; Shukor, Y.; Syed, M.A.; Ahmad, S.A. Hexavalent molybdenum reduction to Mo-blue by a sodium-dodecyl-sulfate-degrading Klebsiella oxytoca strain DRY14. BioMed Res. Int. 2013, 2013. [Google Scholar] [CrossRef]
- Komori, K.; Rivas, A.; Toda, K.; Ohtake, H. A method for removal of toxic chromium using dialysis-sac cultures of a chromate-reducing strain of Enterobacter cloacae. Appl. Microbiol. Biotechnol. 1990, 33, 117–119. [Google Scholar] [CrossRef]
- Halmi, M.I.E.; Wasoh, H.; Sukor, S.; Ahmad, S.A.; Yusof, M.T.; Shukor, M.Y. Bioremoval of molybdenum from aqueous solution. Int. J. Agric. Biol. 2014, 16, 848–850. [Google Scholar]
- Darham, S.; Gomez-Fuentes, C.; Zulkharnain, A.; Sabri, S.; Calisto-Ulloa, N.; Ramírez-Moreno, N.; Ahmad, S.A. Isolation and identification of molybdenum-reducing cold-adapted marine bacteria isolated from Bernardo O’Higgins Riquelme base station, Antarctica. Malays. J. Biochem. Mol. Biol. 2019, 22, 8–15. [Google Scholar]
- Abdulrasheed, M.; Zulkharnain, A.; Zakaria, N.N.; Roslee, A.F.A.; Khalil, K.A.; Napis, S.; Convey, P.; Gomez-Fuentes, C.; Ahmad, S.A. Response surface methodology optimization and kinetics of diesel degradation by a cold-adapted Antarctic bacterium, Arthrobacter sp. strain AQ5-05. Sustainability 2020, 12, 6966. [Google Scholar] [CrossRef]
- Lee, G.L.Y.; Ahmad, S.A.; Yasid, N.A.; Zulkharnain, A.; Convey, P.; Johari, W.L.W.; Alias, S.A.; Gonzalez-Rocha, G.; Shukor, M.Y. Biodegradation of phenol by cold-adapted bacteria from Antarctic soils. Polar. Biol. 2018, 41, 553–562. [Google Scholar] [CrossRef]
- Darham, S.; Zahri, K.N.M.; Zulkharnain, A.; Sabri, S.; Gomez-Fuentes, C.; Convey, P.; Khalil, K.A.; Ahmad, S.A. Statistical Optimisation and Kinetic Studies of Molybdenum Reduction Using a Psychrotolerant Marine Bacteria Isolated from Antarctica. J. Mar. Sci. Eng. 2021, 9, 648. [Google Scholar] [CrossRef]
- Campbell, A.M.; Campillo-Campbell, A.D.; Villaret, D.B. Molybdate reduction by Escherichia coli K-12 and its chl mutants. Proc. Natl. Acad. Sci. USA 1985, 82, 227–231. [Google Scholar] [CrossRef] [Green Version]
- Khuri, A.I.; Mukhopadhyay, S. Response surface methodology. Wiley Interdiscip. Rev. Comput. Stat. 2010, 2, 128–149. [Google Scholar] [CrossRef]
- Ghani, B.; Takai, M.; Hisham, N.Z.; Kishimoto, N.; Ismail, A.K.M.; Tano, T.; Sugio, T. Isolation and characterization of a Mo6+ reducing bacterium. Appl. Environ. Microbiol. 1993, 59, 1176–1180. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Adnan, A.S.M.; Zeid, I.M.A.; Ahmad, S.A.; Halmi, M.I.E.; Abdullah, S.R.S.; Masdor, N.A.; Shukor, M.S.; Shukor, M.Y. A molybdenum-reducing Bacillus sp. strain Zeid 14 in soils from Sudan that could grow on amides and acetonitrile. Malays. J. Soil Sci. 2016, 20, 111–134. [Google Scholar]
- Sabullah, M.K.; Rahman, M.F.; Ahmad, S.A.; Sulaiman, M.R.; Shukor, M.S.; Shamaan, N.A.; Shukor, M.Y. Isolation and characterization of a molybdenum-reducing and glyphosate-degrading Klebsiella oxytoca strain saw-5 in soils from Sarawak. J. Agric. Sci. 2016, 38, 1–13. [Google Scholar] [CrossRef]
- Manogaran, M.; Ahmad, S.A.; Yasid, N.A.; Yakasai, H.M.; Shukor, M.Y. Characterisation of the simultaneous molybdenum reduction and glyphosate degradation by Burkholderia vietnamiensis AQ5-12 and Burkholderia sp. AQ5-13. 3 Biotech 2018, 8, 1–8. [Google Scholar] [CrossRef]
- Mansur, R.; Gusmanizar, N.; Dahalan, F.A.; Masdor, N.A.; Ahmad, S.A.; Shukor, M.S.; Roslan, M.A.H.; Shukor, M.Y. Isolation and characterization of a molybdenum-reducing and amide-degrading Burkholderia sp. strain neni-11 in soils from West Sumatera, Indonesia. IIOAB J. 2016, 7, 28–40. [Google Scholar]
- Othman, A.R.; Bakar, N.A.; Halmi, M.I.E.; Johari, W.L.W.; Ahmad, S.A.; Jirangon, H.; Syed, M.A.; Shukor, M.Y. Kinetics of molybdenum reduction to molybdenum blue by Bacillus sp. strain A. rzi. BioMed Res. Int. 2013, 2013. [Google Scholar] [CrossRef] [Green Version]
- Underwood, E.J. Trace Elements in Human and Animal Nutrition; Academic Press: New York, NY, USA, 1971; pp. 356–357. [Google Scholar]
- Kubota, J. Areas of molybdenum toxicity to grazing animals in the western states. J. Range Manag. 1975, 28, 252–256. [Google Scholar] [CrossRef]
- Novotny, J.A.; Peterson, C.A. Molybdenum. Adv. Nutr. 2018, 9, 272–273. [Google Scholar] [CrossRef]
- Jin, Q.; Kirk, M.F. pH as a primary control in environmental microbiology: 1. Thermodynamic perspective. Front. Environ. Sci. 2018, 6, 21. [Google Scholar] [CrossRef]
- Glenn, J.L.; Crane, F.L. Studies on metalloflavoproteins: V. The action of silicomolybdate in the reduction of cytochrome c by aldehyde oxidase. Biochim. Biophys. Acta 1956, 22, 111–115. [Google Scholar] [CrossRef]
- Krishnan, C.V.; Garnett, M.; Chu, B. Influence of pH and acetate on the self-assembly process of (NH4)42[MoVI72MoV 60O372(CH3COO)30(H2O)72].ca.300H2O. Int. J. Electrochem. Sci. 2008, 3, 1299–1315. [Google Scholar]
- Khayat, M.E.; Rahman, M.F.A.; Shukor, M.S.; Ahmad, S.A.; Shamaan, N.A.; Shukor, M.Y. Characterization of a molybdenum-reducing Bacillus sp. strain khayat with the ability to grow on SDS and diesel. Rend. Fis. Acc. Lincei. 2016, 27, 547–556. [Google Scholar] [CrossRef]
- Roslee, A.F.A.; Gomez-Fuentes, C.; Zakaria, N.N.; Shaharuddin, N.A.; Zulkharnain, A.; Abdul Khalil, K.; Convey, P.; Ahmad, S.A. Growth optimisation and kinetic profiling of diesel biodegradation by a cold-adapted microbial consortium isolated from Trinity Peninsula, Antarctica. Biology 2021, 10, 493. [Google Scholar] [CrossRef] [PubMed]
- Zahri, K.N.M.; Zulkharnain, A.; Gomez-Fuentes, C.; Sabri, S.; Abdul Khalil, K.; Convey, P.; Ahmad, S.A. The use of response surface methodology as a statistical tool for the optimisation of waste and pure canola oil biodegradation by Antarctic soil bacteria. Life 2021, 11, 456. [Google Scholar] [CrossRef] [PubMed]
- Zakaria, N.N.; Gomez-Fuentes, C.; Abdul Khalil, K.; Convey, P.; Roslee, A.F.A.; Zulkharnain, A.; Sabri, S.; Shaharuddin, N.A.; Cárdenas, L.; Ahmad, S.A. Statistical optimisation of diesel biodegradation at low temperatures by an Antarctic marine bacterial consortium isolated from non-contaminated seawater. Microorganisms 2021, 9, 1213. [Google Scholar] [CrossRef] [PubMed]
- Lee, G.L.Y.; Zakaria, N.N.; Convey, P.; Futamata, H.; Zulkharnain, A.; Suzuki, K.; Khalil, K.A.; Shaharuddin, N.A.; Alias, S.A.; González-Rocha, G.; et al. Statistical optimisation of phenol degradation and pathway identification through whole genome sequencing of the cold-adapted Antarctic bacterium, Rhodococcus sp. strain AQ5-07. Int. J. Mol. Sci. 2020, 21, 9363. [Google Scholar] [CrossRef]
- Yakasai, H.M.; Karamba, K.I.; Yasid, N.A.; Halmi, M.I.E.; Rahman, M.F.; Ahmad, S.A.; Shukor, M.Y. Response surface-based optimization of a novel molybdenum-reducing cyanide-degrading Serratia sp. strain HMY1. Desalin. Water Treat. 2019, 145, 220–231. [Google Scholar] [CrossRef]
- Yusuf, I.; Ahmad, S.A.; Phang, L.Y.; Syed, M.A.; Shamaan, N.A.; Abdul Khalil, K.; Dahalan, F.A.; Shukor, M.Y. Keratinase production and biodegradation of polluted secondary chicken feather wastes by a newly isolated multi heavy metal tolerant bacterium-Alcaligenes sp. AQ05-001. J. Environ. Manag. 2016, 183, 182–195. [Google Scholar] [CrossRef] [PubMed]
| Variables | Symbol | Unit | Experimental Level | ||||
|---|---|---|---|---|---|---|---|
| −2 | −1 | 0 | +1 | +2 | |||
| Sucrose concentration | A | g/L | 0.09 | 1 | 1.5 | 2 | 2.91 |
| (NH4)2SO4 concentration | B | g/L | 0.11 | 0.2 | 0.25 | 0.3 | 0.39 |
| Salinity | C | g/L | 0.41 | 0.5 | 0.55 | 0.6 | 0.69 |
| Molybdate concentration | D | g/L | 5.43 | 10.0 | 12.5 | 15.0 | 19.57 |
| pH | E | - | 6.54 | 7.0 | 7.25 | 7.5 | 7.96 |
| Temperature | F | °C | 5.4 | 10.0 | 12.5 | 15.0 | 19.6 |
| Source | Sum of Squares | df | Mean Square | F Value | Prob > F |
|---|---|---|---|---|---|
| Model | 10.25654 | 27 | 0.379872 | 39.04075 | <0.0001 * |
| A | 1.533952 | 1 | 1.533952 | 157.6496 | <0.0001 * |
| B | 0.000928 | 1 | 0.000928 | 0.095385 | 0.7585 |
| C | 0.125828 | 1 | 0.125828 | 12.93182 | 0.0007 * |
| D | 0.506057 | 1 | 0.506057 | 52.00924 | <0.0001 * |
| E | 1.161845 | 1 | 1.161845 | 119.4068 | <0.0001 * |
| F | 2.791166 | 1 | 2.791166 | 286.8578 | <0.0001 * |
| A2 | 1.58817 | 1 | 1.58817 | 163.2217 | <0.0001 * |
| B2 | 0.684594 | 1 | 0.684594 | 70.35808 | <0.0001 * |
| C2 | 0.767384 | 1 | 0.767384 | 78.86666 | <0.0001 * |
| D2 | 1.128975 | 1 | 1.128975 | 116.0287 | <0.0001 * |
| E2 | 0.2677 | 1 | 0.2677 | 27.51244 | <0.0001 * |
| F2 | 0.769687 | 1 | 0.769687 | 79.10336 | <0.0001 * |
| AB | 0.054231 | 1 | 0.054231 | 5.573483 | 0.0216 * |
| AC | 0.033535 | 1 | 0.033535 | 3.446484 | 0.0685 |
| AD | 0.065344 | 1 | 0.065344 | 6.715643 | 0.0121 * |
| AE | 0.012183 | 1 | 0.012183 | 1.252052 | 0.2678 |
| AF | 0.08316 | 1 | 0.08316 | 8.546655 | 0.0049 * |
| BC | 0.063315 | 1 | 0.063315 | 6.507116 | 0.0134 * |
| BD | 0.011854 | 1 | 0.011854 | 1.218252 | 0.2743 |
| BE | 0.003525 | 1 | 0.003525 | 0.362317 | 0.5496 |
| BF | 0.049562 | 1 | 0.049562 | 5.093647 | 0.0278 * |
| CD | 0.055284 | 1 | 0.055284 | 5.681704 | 0.0204 * |
| CE | 0.005532 | 1 | 0.005532 | 0.568506 | 0.4539 |
| CF | 0.058746 | 1 | 0.058746 | 6.037493 | 0.0170 * |
| DE | 0.072025 | 1 | 0.072025 | 7.402273 | 0.0086 * |
| DF | 0.236561 | 1 | 0.236561 | 24.31215 | <0.0001 * |
| EF | 0.069498 | 1 | 0.069498 | 7.142564 | 0.0098 * |
| Residual | 0.564348 | 58 | 0.00973 | ||
| Lack of Fit | 0.469043 | 49 | 0.009572 | 0.903946 | 0.6238 |
| Pure Error | 0.095305 | 9 | 0.010589 | ||
| Cor Total | 10.82089 | 85 | |||
| Std. Dev. | 0.098641 | R-Squared | 0.948 | ||
| Mean | 1.250234 | Adj R-Squared | 0.9236 | ||
| C.V. | 7.889842 | Pred R-Squared | 0.87013 | ||
| PRESS | 1.405346 | Adeq Precision | 27.37344 | ||
| Symbol | Variable | Unit | Generated Value | Mo Reduction (OD865 nm) | |
|---|---|---|---|---|---|
| Predicted | Experimental | ||||
| A | Sucrose concentration | g/L | 1.0 | 2.1 | 2.19 |
| B | Nitrogen concentration | g/L | 0.25 | ||
| C | Salinity | g/L | 0.55 | ||
| D | Molybdate concentration | mM | 12.5 | ||
| E | pH | - | 7.0 | ||
| F | Temperature | °C | 10.0 | ||
| Symbol | Variable | Unit | Optimised Value | |
|---|---|---|---|---|
| OFAT | RSM | |||
| A | Sucrose concentration | g/L | 1.0 | 1.0 |
| B | Nitrogen concentration | g/L | 0.20 | 0.25 |
| C | Salinity | g/L | 0.50 | 0.55 |
| D | Molybdate concentration | mM | 10.0 | 12.5 |
| E | pH | - | 7.0 | 7.0 |
| F | Temperature | °C | 10.0 | 10.0 |
| Mo reduction (OD 865 nm) | 1.93 | 2.19 | ||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Darham, S.; Syed-Muhaimin, S.N.; Subramaniam, K.; Zulkharnain, A.; Shaharuddin, N.A.; Khalil, K.A.; Ahmad, S.A. Optimisation of Various Physicochemical Variables Affecting Molybdenum Bioremediation Using Antarctic Bacterium, Arthrobacter sp. Strain AQ5-05. Water 2021, 13, 2367. https://doi.org/10.3390/w13172367
Darham S, Syed-Muhaimin SN, Subramaniam K, Zulkharnain A, Shaharuddin NA, Khalil KA, Ahmad SA. Optimisation of Various Physicochemical Variables Affecting Molybdenum Bioremediation Using Antarctic Bacterium, Arthrobacter sp. Strain AQ5-05. Water. 2021; 13(17):2367. https://doi.org/10.3390/w13172367
Chicago/Turabian StyleDarham, Syazani, Sharifah Nabilah Syed-Muhaimin, Kavilasni Subramaniam, Azham Zulkharnain, Noor Azmi Shaharuddin, Khalilah Abdul Khalil, and Siti Aqlima Ahmad. 2021. "Optimisation of Various Physicochemical Variables Affecting Molybdenum Bioremediation Using Antarctic Bacterium, Arthrobacter sp. Strain AQ5-05" Water 13, no. 17: 2367. https://doi.org/10.3390/w13172367
APA StyleDarham, S., Syed-Muhaimin, S. N., Subramaniam, K., Zulkharnain, A., Shaharuddin, N. A., Khalil, K. A., & Ahmad, S. A. (2021). Optimisation of Various Physicochemical Variables Affecting Molybdenum Bioremediation Using Antarctic Bacterium, Arthrobacter sp. Strain AQ5-05. Water, 13(17), 2367. https://doi.org/10.3390/w13172367








