Design of a Smart System for Rapid Bacterial Test
Abstract
1. Introduction
2. Materials and Methods
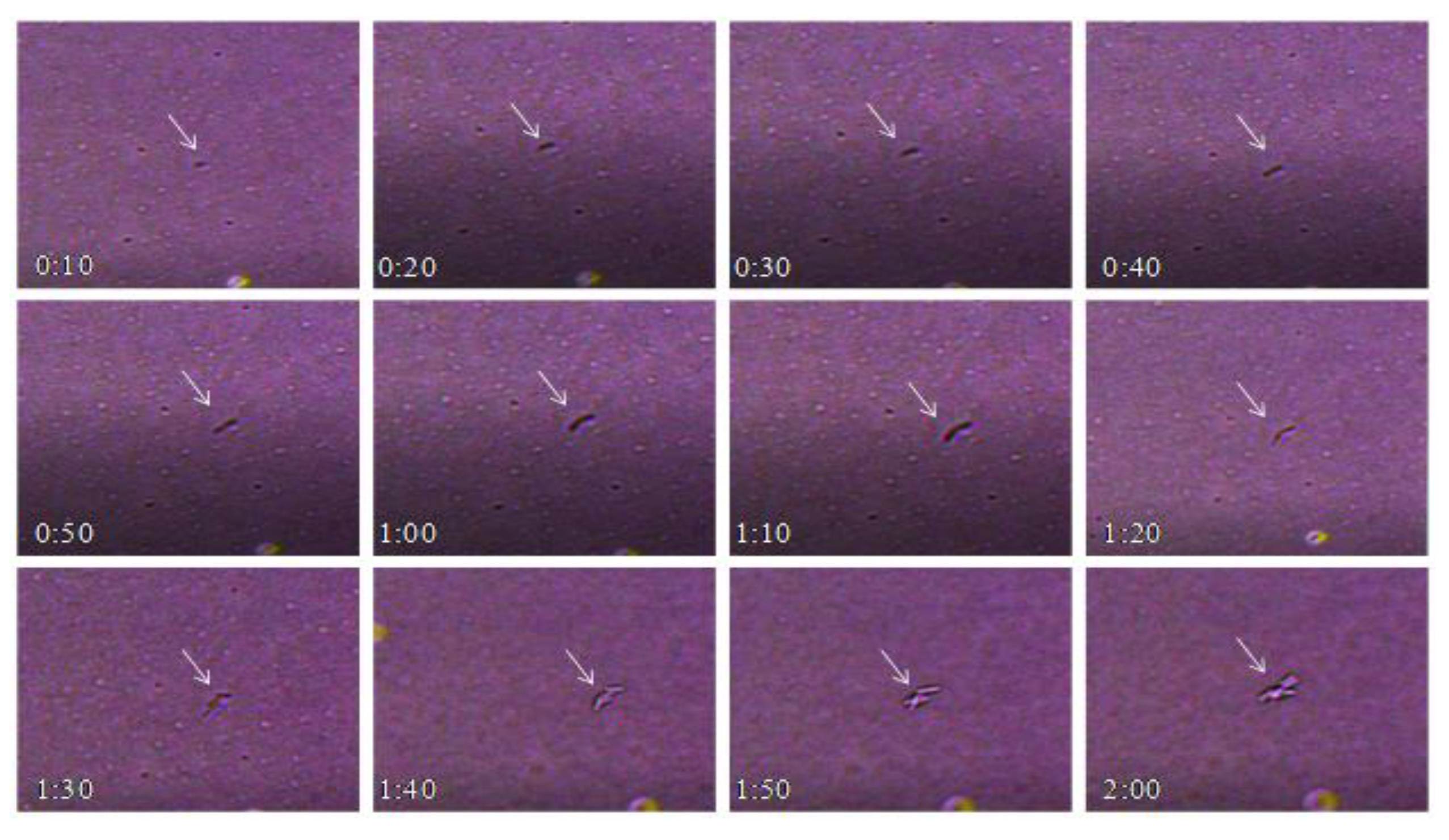
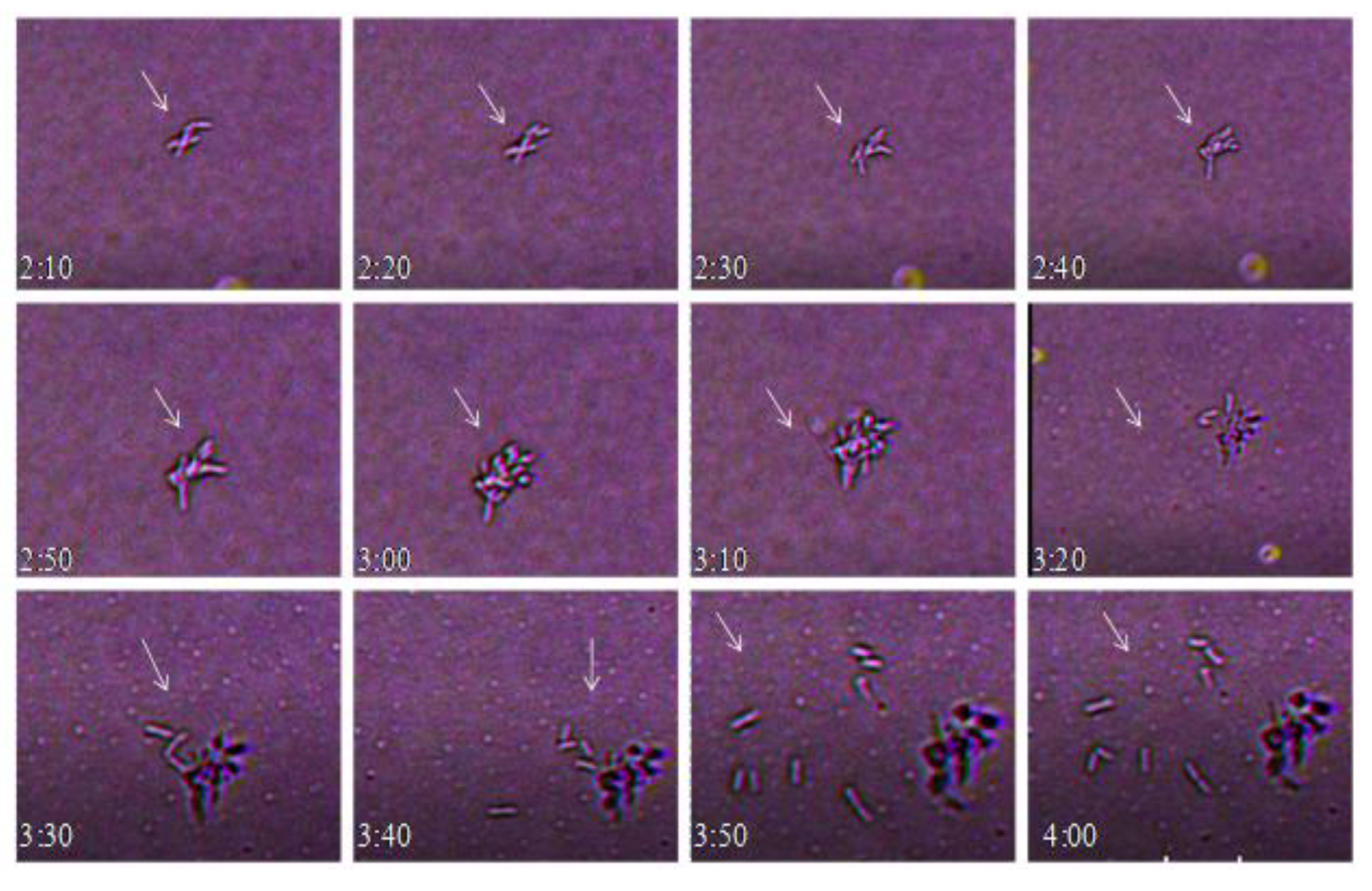
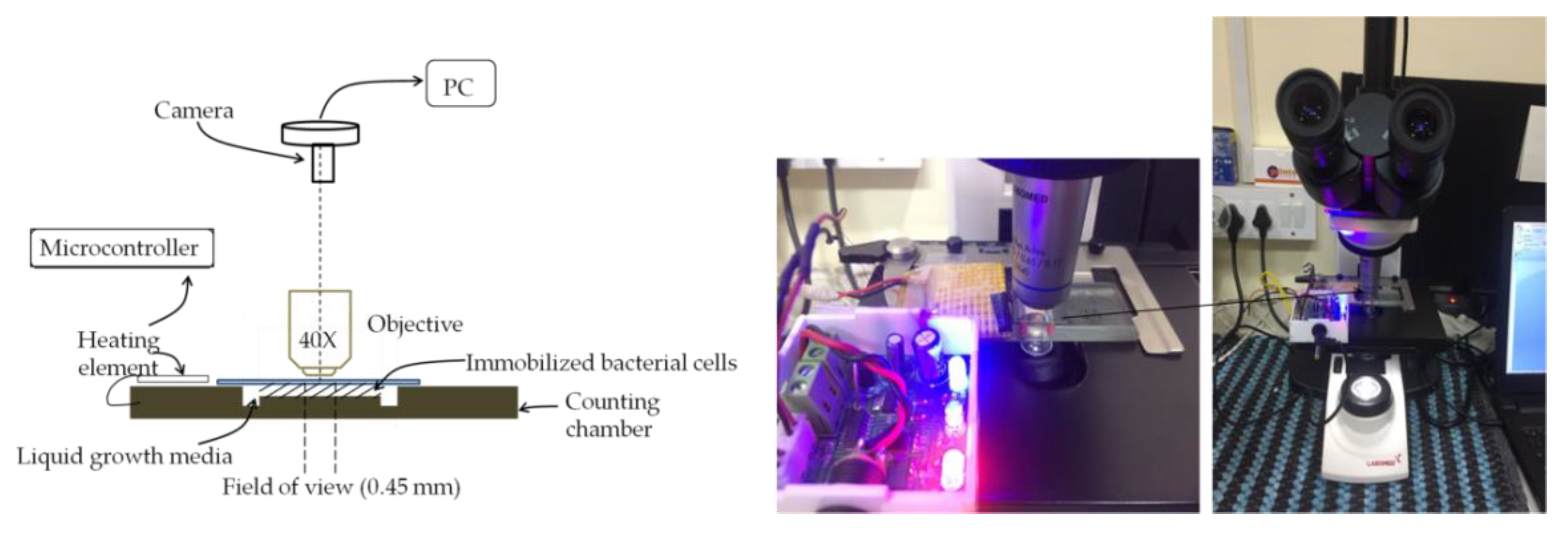
Live-Cell Imaging of Bacteria
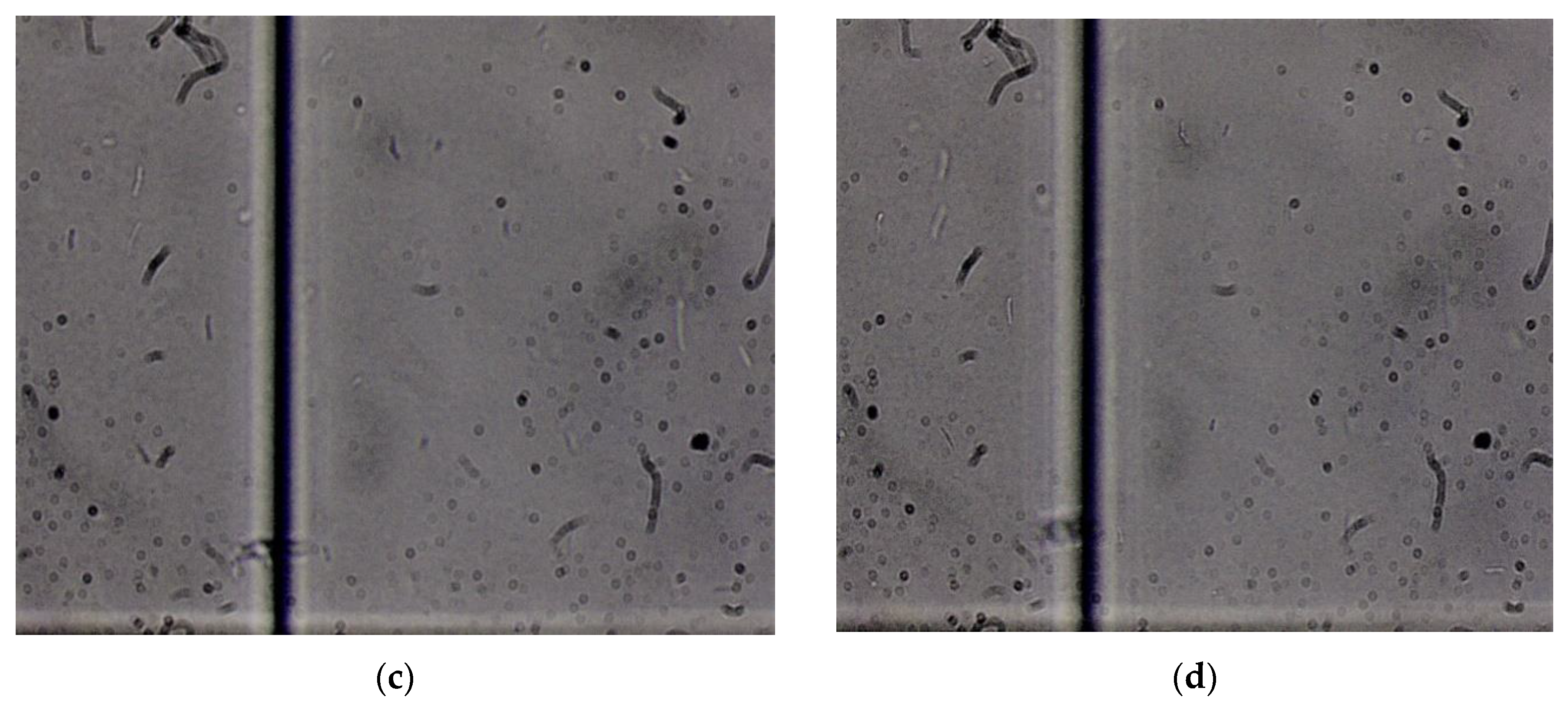
3. Results and Discussion
4. Configuration of the Proposed Device
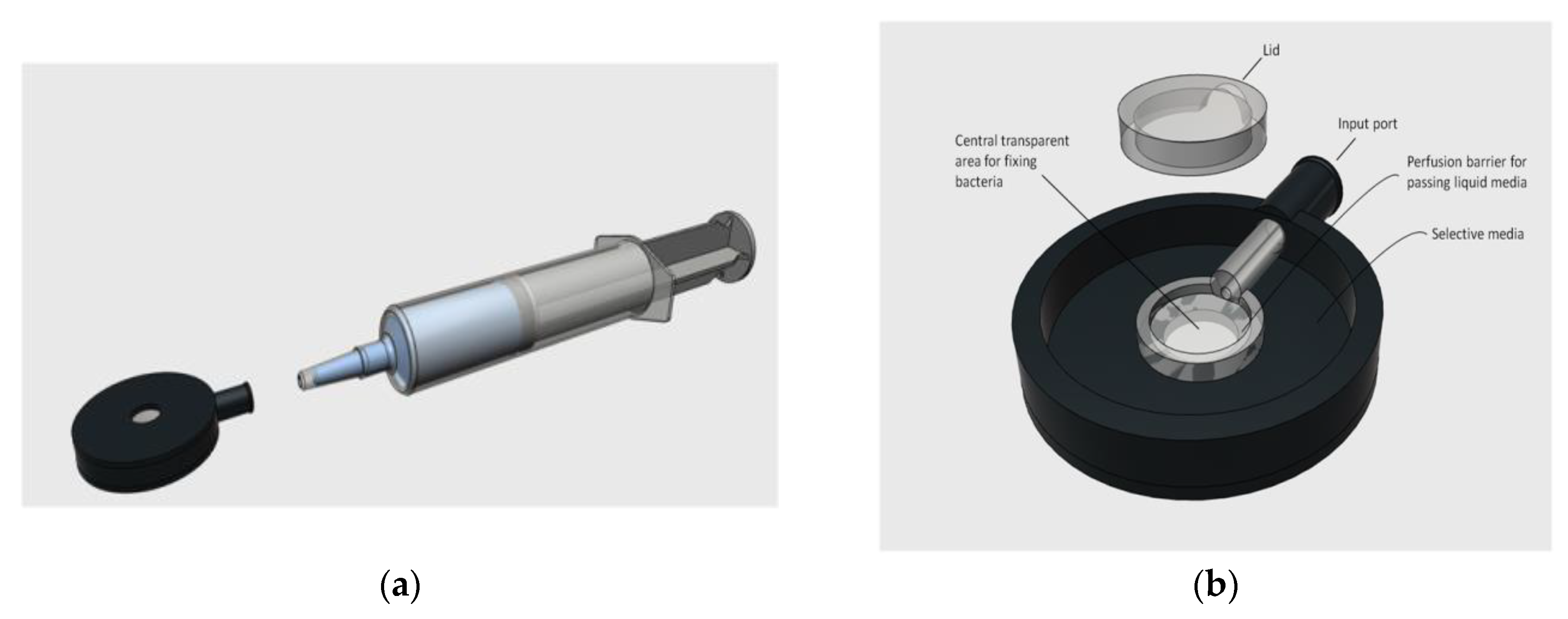
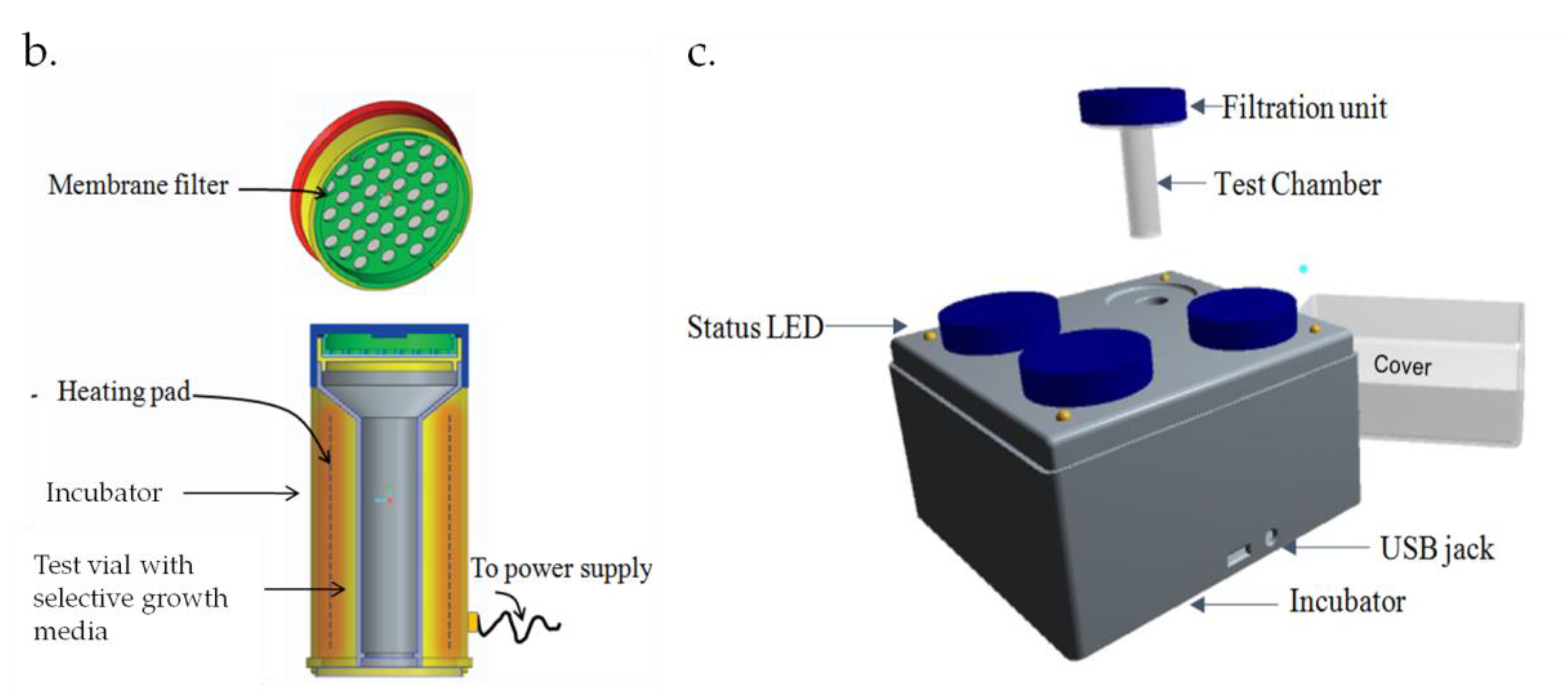
4.1. Test Chamber
4.2. Sample Pre-Enrichment
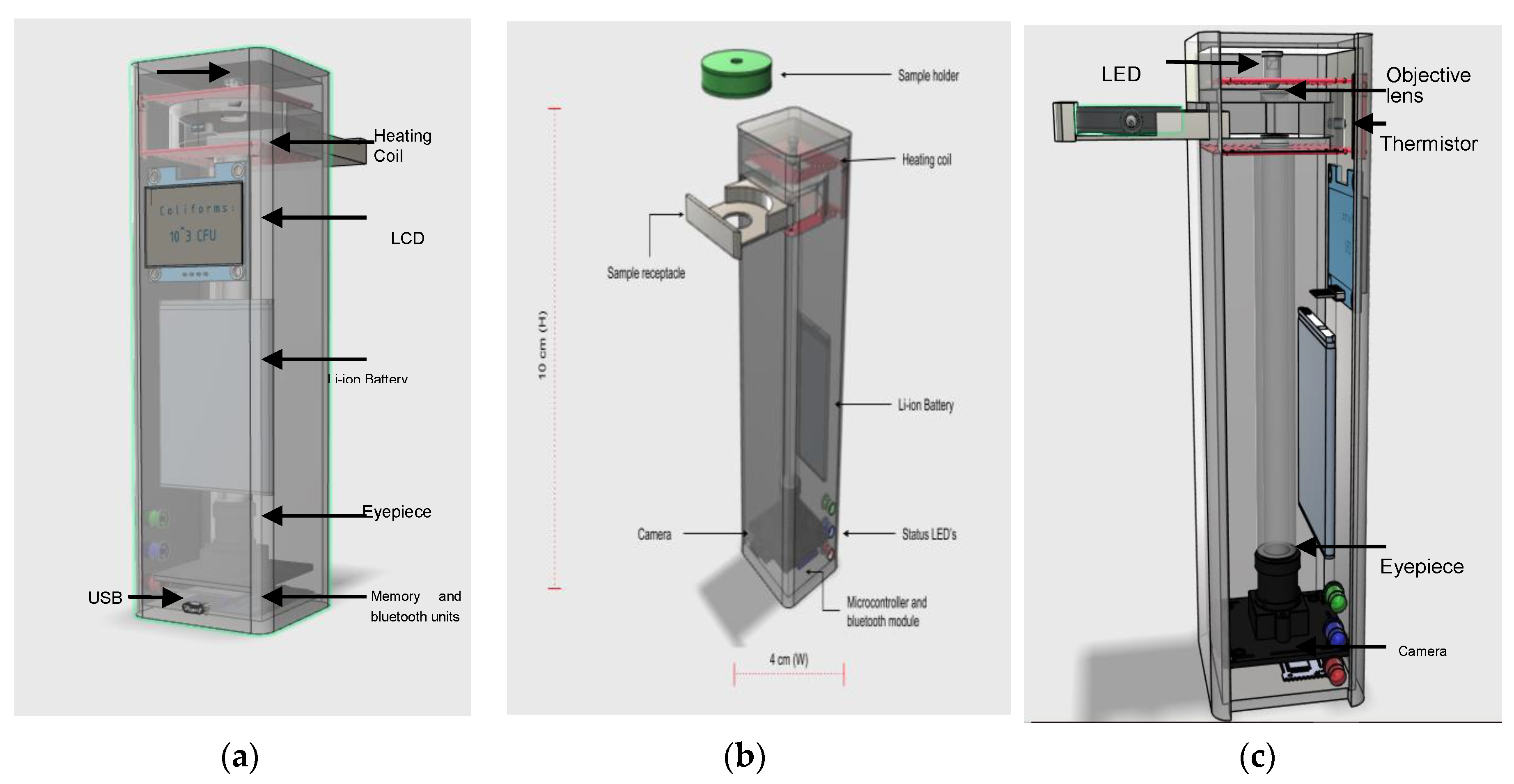
4.3. The Complete System
5. The Decision System
6. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Ashbolt, N.J. Microbial contamination of drinking water and human health from community water systems. Curr. Environ. Health Rep. 2015, 2, 95–106. [Google Scholar] [CrossRef] [PubMed]
- Parsai, A.; Rokade, V. Water Quality Monitoring Infrastructure for Tackling Water-Borne Diseasesin the State of Madhya Pradesh, India, and Its Implication on the Sustainable Development Goals (SDGs). In The Relevance of Hygiene to Health in Developing Countries; Potgieter, N., Hoffman, A.T., Eds.; IntechOpen: London, UK, 2019. [Google Scholar] [CrossRef]
- Holm, R.; Singini, W.; Gwayi, S. Comparative evaluation of the cost of water in northern Malawi: From rural water wells to science education. Appl. Econ. 2016, 48, 4573–4583. [Google Scholar] [CrossRef]
- Tortajada, C.; Biswas, A.K. Achieving universal access to clean water and sanitation in an era of water scarcity: Strengthening contributions from academia. Curr. Opin. Environ. Sustain. 2018, 34, 21–25. [Google Scholar] [CrossRef]
- Kuylenstierna, J.L.; Björklund, G.; Najlis, P. Sustainable water future with global implications: everyone’s responsibility. In Natural Resources Forum; Blackwell Publishing Ltd.: Oxford, UK, 1997; Volume 21, pp. 181–190. [Google Scholar]
- Batchelor, C.H.; Rama Mohan Rao, M.S.; Manohar Rao, S. Watershed development: A solution to water shortages in semi-arid India or part of the problem? Land Use Water Resour. Res. 2003, 3, 1–10. [Google Scholar] [CrossRef]
- Hanchett, S.; Akhter, S.; Khan, M.H.; Mezulianik, S.; Blagbrough, V. Water, sanitation and hygiene in Bangladeshi slums: An evaluation of the WaterAid–Bangladesh urban programme. Environ. Urban. 2003, 15, 43–56. [Google Scholar]
- Osborn, D.; Cutter, A.; Ullah, F.; Universal Sustainable Development Goals. Understanding the Transformational Challenge for Developed Countries. Report of a Study by Stakeholder Forum. May 2015. Available online: https://sustainabledevelopment.un.org/content/documents/1684SF_-_SDG_Universality_Report_-_May_2015.pdf (accessed on 10 December 2019).
- Martin, N.H.; Trmčić, A.; Hsieh, T.H.; Boor, K.J.; Wiedmann, M. The evolving role of coliforms as indicators of unhygienic processing conditions in dairy foods. Front. Microbiol. 2016, 7, 1–8. [Google Scholar] [CrossRef]
- Odonkor, S.T.; Ampofo, J.K. Escherichia coli as an indicator of bacteriological quality of water: An overview. Microbiol. Res. 2013, 4. [Google Scholar] [CrossRef]
- Tallon, P.; Magajna, B.; Lofranco, C.; Leung, K.T. Microbial indicators of faecal contamination in water: A current perspective. Water Air Soil Pollut. 2005, 166, 139–166. [Google Scholar] [CrossRef]
- Bain, R.B.; Elliott, M.; Matthews, R.; Mcmahan, L.; Tung, R.; Chuang, P.; Gundry, S. A summary catalogue of microbial drinking water tests for low and medium resource settings. Int. J. Environ. Res. Pub. Health 2012, 9, 1609–1625. [Google Scholar] [CrossRef]
- Gunda, N.S.K.; Naicker, S.; Shinde, S.; Kimbahune, S.; Shrivastava, S.; Mitra, S. Mobile water kit (MWK): A smartphone compatible low-cost water monitoring system for rapid detection of total coliform and E. coli. Anal. Methods 2014, 6, 6236–6246. [Google Scholar] [CrossRef]
- Rompré, A.; Servais, P.; Baudart, J.; De-Roubin, M.R.; Laurent, P. Detection and enumeration of coliforms in drinking water: Current methods and emerging approaches. J. Microbiol. Methods 2002, 49, 31–54. [Google Scholar] [CrossRef]
- Warren, L.S.; Benoit, R.E.; Jessee, J.A. Rapid enumeration of Fecal Coliforms in water by a colorimetric beta-galactosidase assay. Appl. Environ. Microbiol. 1978, 35, 136–141. [Google Scholar] [PubMed]
- Lewis, C.L.; Craig, C.C.; Senecal, A.G. Mass and density measurements of live and dead Gram-negative and Gram-positive bacterial populations. Appl. Environ. Microbiol. 2014, 80, 3622–3631. [Google Scholar] [CrossRef] [PubMed]
- Law, J.W.; Ab Mutalib, N.S.; Chan, K.G.; Lee, L.H. Rapid methods for the detection of foodborne bacterial pathogens: Principles, applications, advantages and limitations. Front. Microbiol. 2015, 5, 770. [Google Scholar] [CrossRef] [PubMed]
- Molina, F.; López-Acedo, E.; Tabla, R.; Roa, I.; Gómez, A.; Rebollo, J.E. Improved detection of Escherichia coli and coliform bacteria by multiplex PCR. BMC Biotechnol. 2015, 15, 48. [Google Scholar] [CrossRef]
- Priest, D.G.; Tanaka, N.; Tanaka, Y.; Taniguchi, Y. Micro-patterned agarose gel devices for single-cell high-throughput microscopy of E. coli cells. Sci. Rep. 2017, 7, 17750. [Google Scholar] [CrossRef]
- Joyce, G.; Robertson, B.D.; Williams, K.J. A modified agar pad method for mycobacterial live-cell imaging. BMC Res. Notes 2011, 4, 73. [Google Scholar] [CrossRef]
- Fernández-Miranda, E.; Majada, J.; Casares, A. Efficacy of propidium iodide and FUN-1 stains for assessing viability in basidiospores of Rhizopogonroseolus. Mycologia 2017, 109, 350–358. [Google Scholar] [CrossRef]
- Gao, P.; Sun, C.; Li, Y.; Zou, X.; Wu, X.; Ling, Y.; Luan, C.; Chen, H. Vital staining of bacteria by sunset yellow pigment. Pol. J. Microbiol. 2017, 66, 113–117. [Google Scholar] [CrossRef]
- Haines, A.M.; Tobe, S.S.; Kobus, H.; Linacre, A. Finding DNA: Using fluorescent in situ detection. Forensic Sci. Int. Genet. Suppl. Ser. 2015, 5, e501–e502. [Google Scholar] [CrossRef]
- Haines, A.M.; Tobe, S.S.; Kobus, H.; Linacre, A. Duration of in situ fluorescent signals within hairs follicles. Forensic Sci. Int. Genet. Suppl. Ser. 2015, 5, 175–176. [Google Scholar] [CrossRef]
- Mishra, M.; Chauhan, P. Applications of Microscopy in Bacteriology. Microsc. Res. 2016, 4, 1–9. [Google Scholar] [CrossRef][Green Version]
- World Health Organization (WHO). Guidelines for Drinking-Water Quality; World Health Organization: Geneva, Switzerland, 2011. [Google Scholar]
- Pettipher, G.L.; Mansell, R.; McKinnon, C.H.; Cousins, C.M. Rapid membrane filtration-epifluorescent microscopy technique for direct enumeration of bacteria in raw milk. Appl. Environ. Microbiol. 1980, 39, 423–429. [Google Scholar] [PubMed]
- Patil, R.; Levin, S.; Halery, N.; Gupta, I.; Rajkumar, S. A smartphone-based early alert system for screening of coliform contamination in drinking water. J. Microbiol. Biotechnol. Food Sci. 2019, 539–547. [Google Scholar] [CrossRef][Green Version]
- Mekterović, I.; Mekterović, D. BactImAS: A platform for processing and analysis of bacterial time-lapse microscopy movies. BMC Bioinform. 2014, 15, 251. [Google Scholar] [CrossRef]






© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Patil, R.; Levin, S.; Rajkumar, S.; Ajmal, T. Design of a Smart System for Rapid Bacterial Test. Water 2020, 12, 15. https://doi.org/10.3390/w12010015
Patil R, Levin S, Rajkumar S, Ajmal T. Design of a Smart System for Rapid Bacterial Test. Water. 2020; 12(1):15. https://doi.org/10.3390/w12010015
Chicago/Turabian StylePatil, Rajshree, Saurabh Levin, Samuel Rajkumar, and Tahmina Ajmal. 2020. "Design of a Smart System for Rapid Bacterial Test" Water 12, no. 1: 15. https://doi.org/10.3390/w12010015
APA StylePatil, R., Levin, S., Rajkumar, S., & Ajmal, T. (2020). Design of a Smart System for Rapid Bacterial Test. Water, 12(1), 15. https://doi.org/10.3390/w12010015




