Solar Septic Tank: Next Generation Sequencing Reveals Effluent Microbial Community Composition as a Useful Index of System Performance
Abstract
1. Introduction
- Qualitatively assess microbial community composition and persistence in the sludge and effluent from each of the CT and SST, operated under near-identical conditions, over a four-month time period;
- Associate microbial community composition in the CT and SST with system operation and performance;
- Assess the sampling locations in each system that most usefully explain performance variability. This will help guide the sampling strategy for an extensive study.
2. Materials and Methods
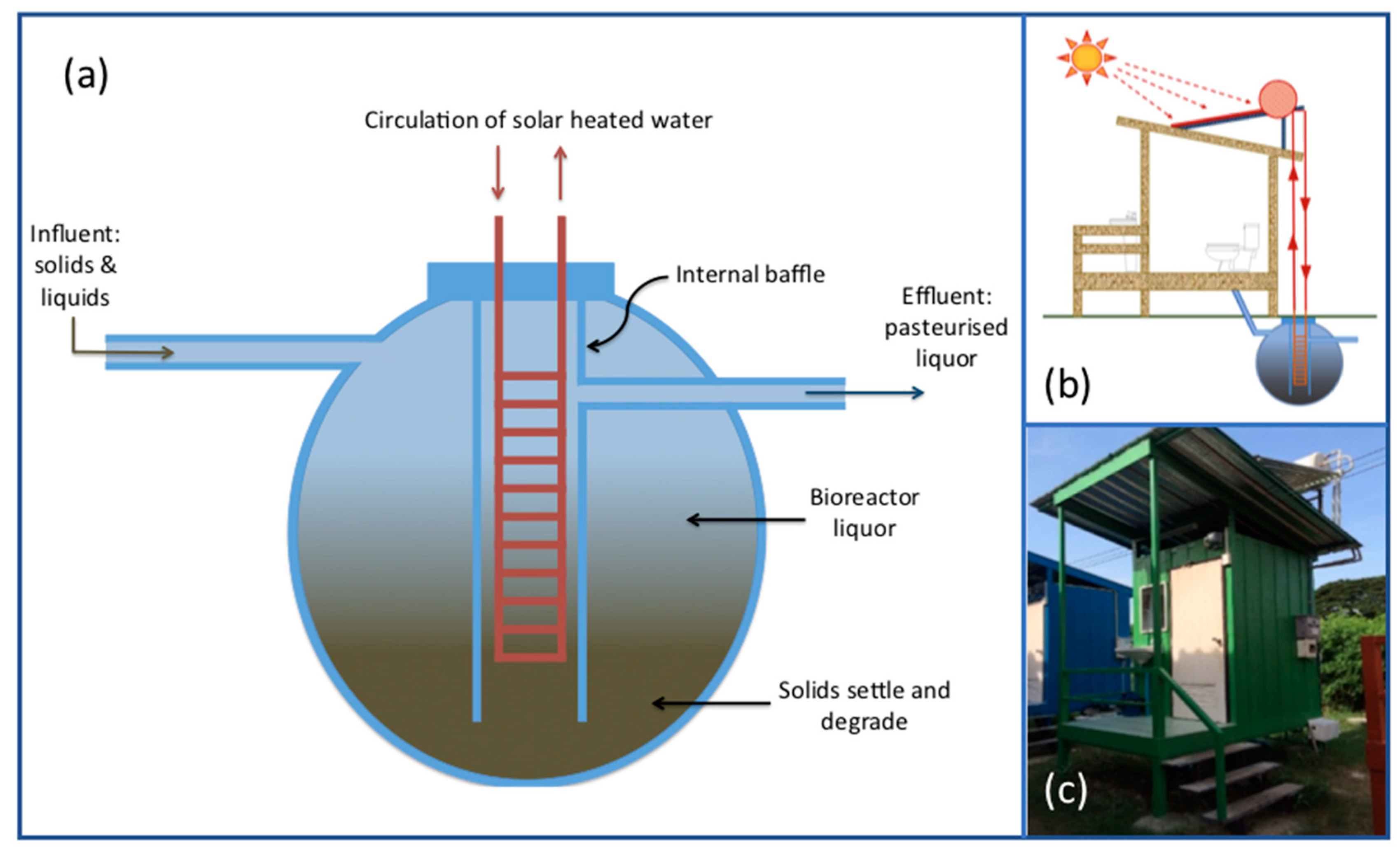
2.1. SST and CT System Design
2.2. Sampling
2.3. Environmental Operation and Performance Monitoring
2.4. DNA Extraction
2.5. Amplicon Library Preparation
2.6. Bioinformatics and Statistical Analysis
3. Results
3.1. Operation and Performance
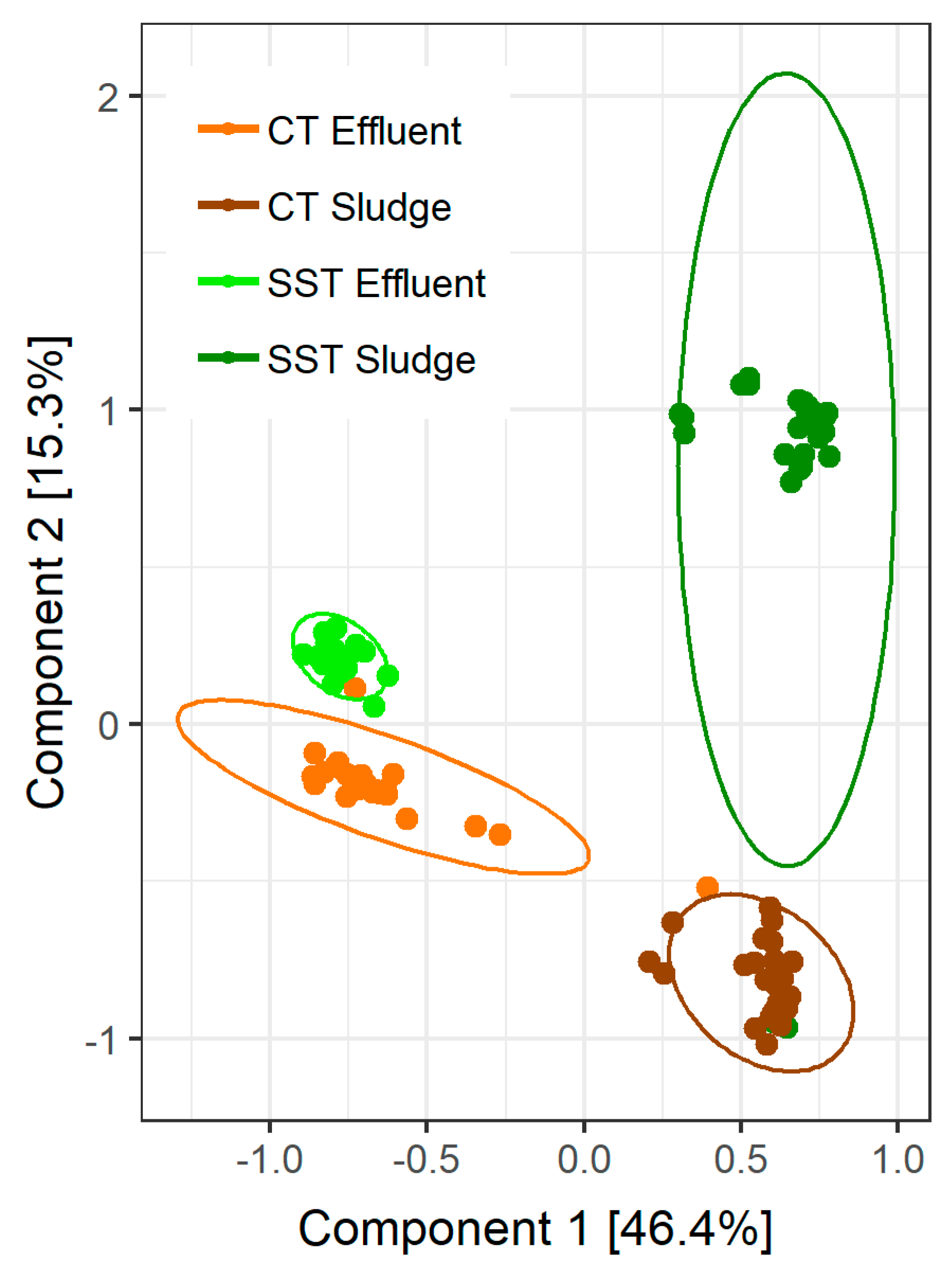
3.2. Microbial Community Composition and Sampling Variability
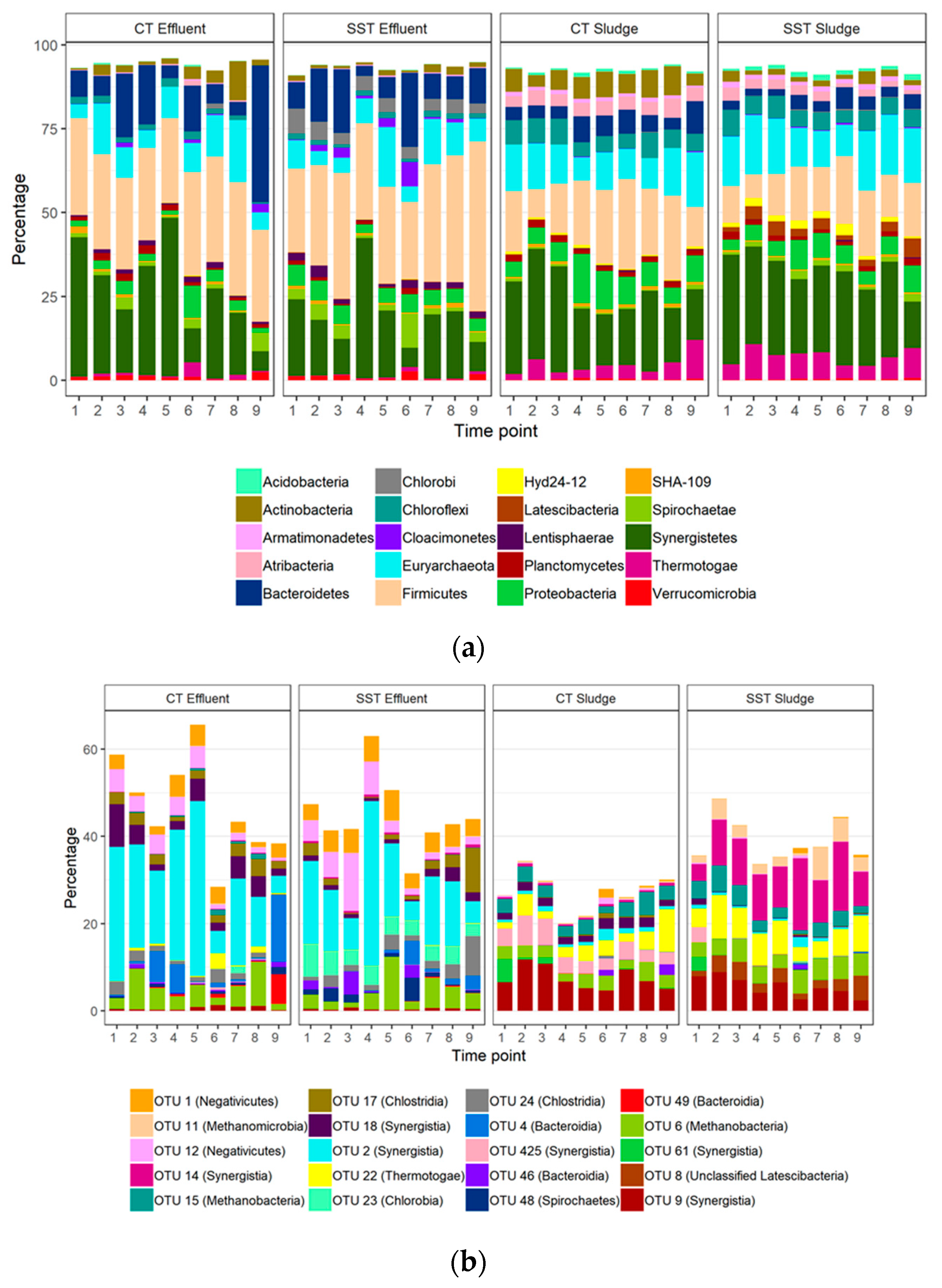
3.3. Microbial Community Composition within and between Samples
3.4. Correlating System Operation and Microbial Composition in Sludge and Effluent Samples
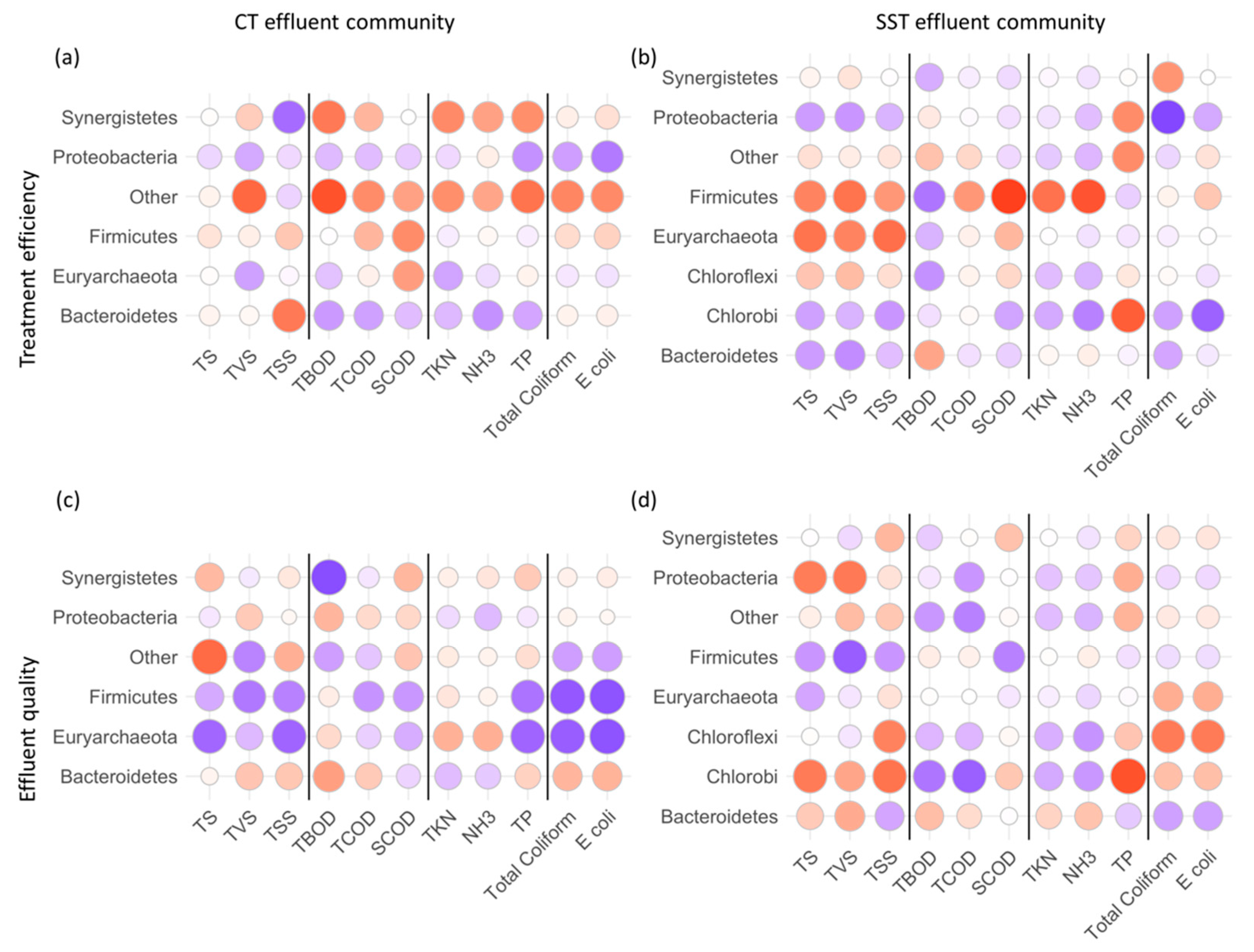
3.5. Microbial Taxa in Effluent Indexing Treatment Efficiency and Effluent Water Quality
4. Discussion
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Withers, P.J.A.; Jordan, P.; May, L.; Jarvie, H.P.; Deal, N.E. Do septic tank systems pose a hidden threat to water quality? Front. Ecol. Environ. 2014, 12, 123–130. [Google Scholar] [CrossRef]
- Manual, D. Onsite Wastewater Treatment and Disposal Systems; EPA: Washington, DC, USA, 1980.
- May, L.; Place, C.; O’Malley, M.; Spears, B. The Impact of Phosphorus Inputs From Small Discharges on Designated Freshwater Sites; Natural England Commissioned Reports, Number 170, Natural England - GOV.UK: London, UK, 2015. [Google Scholar]
- Meuler, S.; Paris, S.; Hackner, T. Membrane bio-reactors for decentralized wastewater treatment and reuse. Water Sci. Technol. 2008, 58, 285–294. [Google Scholar] [CrossRef] [PubMed]
- Appling, D.; Habteselassie, M.; Radcliffe, D.; Bradshaw, J. Preliminary study on the effect of wastewater storage in septic tank on E. coli concentration in summer. Water 2013, 5, 1141–1151. [Google Scholar] [CrossRef]
- Polpraserta, C.; Koottatepb, T.; Pussayanavinb, T. Solar septic tanks: A new sanitation paradigm for Thailand 4.0. Sci. Asia 2018, 44, 39–43. [Google Scholar] [CrossRef]
- US Environmental Protection Agency. Onsite Wastewater Treatment Systems Manual; US EPA: Washington, DC, USA, 2002.
- Kjellén, M.; Pensulo, C.; Nordqvist, P.; Fogde, M. Global review of sanitation system trends and interactions with menstrual management practices. In Report for the Menstrual Management and Sanitation Systems Project; Proj Report-2011; Stock Environ Institute: Stockholm, Sweden, 2011. [Google Scholar]
- Liu, C.; Li, H.; Zhang, Y.; Si, D.; Chen, Q. Evolution of microbial community along with increasing solid concentration during high-solids anaerobic digestion of sewage sludge. Bioresour. Technol. 2016, 216, 87–94. [Google Scholar] [CrossRef]
- Fitamo, T.; Treu, L.; Boldrin, A.; Sartori, C.; Angelidaki, I.; Scheutz, C. Microbial population dynamics in urban organic waste anaerobic co-digestion with mixed sludge during a change in feedstock composition and different hydraulic retention times. Water Res. 2017, 118, 261–271. [Google Scholar] [CrossRef]
- Lee, C.; Kim, J.; Hwang, K.; O’Flaherty, V.; Hwang, S. Quantitative analysis of methanogenic community dynamics in three anaerobic batch digesters treating different wastewaters. Water Res. 2009, 43, 157–165. [Google Scholar] [CrossRef]
- Ziganshin, A.M.; Schmidt, T.; Lv, Z.; Liebetrau, J.; Richnow, H.H.; Kleinsteuber, S.; Nikolausz, M. Reduction of the hydraulic retention time at constant high organic loading rate to reach the microbial limits of anaerobic digestion in various reactor systems. Bioresour. Technol. 2016, 217, 62–71. [Google Scholar] [CrossRef]
- Schmidt, T.; Ziganshin, A.M.; Nikolausz, M.; Scholwin, F.; Nelles, M.; Kleinsteuber, S.; Pröter, J. Effects of the reduction of the hydraulic retention time to 1.5 days at constant organic loading in CSTR, ASBR, and fixed-bed reactors—Performance and methanogenic community composition. Biomass Bioenergy 2014, 69, 241–248. [Google Scholar] [CrossRef]
- Kundu, K.; Sharma, S.; Sreekrishnan, T.R. Changes in microbial communities in a hybrid anaerobic reactor with organic loading rate and temperature. Bioresour. Technol. 2013, 129, 538–547. [Google Scholar] [CrossRef]
- Rincón, B.; Borja, R.; González, J.M.; Portillo, M.C.; Sáiz-Jiménez, C. Influence of organic loading rate and hydraulic retention time on the performance, stability and microbial communities of one-stage anaerobic digestion of two-phase olive mill solid residue. Biochem. Eng. J. 2008, 40, 253–261. [Google Scholar] [CrossRef]
- Goux, X.; Calusinska, M.; Lemaigre, S.; Marynowska, M.; Klocke, M.; Udelhoven, T.; Benizri, E.; Delfosse, P. Microbial community dynamics in replicate anaerobic digesters exposed sequentially to increasing organic loading rate, acidosis, and process recovery. Biotechnol. Biofuels 2015, 8, 122. [Google Scholar] [CrossRef] [PubMed]
- Koster, I.W.; Lettinga, G. The influence of ammonium-nitrogen on the specific activity of pelletized methanogenic sludge. Agric. Wastes 1984, 9, 205–216. [Google Scholar] [CrossRef]
- Belostotskiy, D.E.; Ziganshina, E.E.; Siniagina, M.; Boulygina, E.A.; Miluykov, V.A.; Ziganshin, A.M. Impact of the substrate loading regime and phosphoric acid supplementation on performance of biogas reactors and microbial community dynamics during anaerobic digestion of chicken wastes. Bioresour. Technol. 2015, 193, 42–52. [Google Scholar] [CrossRef]
- Karlsson, A.; Einarsson, P.; Schnürer, A.; Sundberg, C.; Ejlertsson, J.; Svensson, B.H. Impact of trace element addition on degradation efficiency of volatile fatty acids, oleic acid and phenyl acetate and on microbial populations in a biogas digester. J. Biosci. Bioeng. 2012, 114, 446–452. [Google Scholar] [CrossRef]
- Westerholm, M.; Müller, B.; Isaksson, S.; Schnürer, A. Trace element and temperature effects on microbial communities and links to biogas digester performance at high ammonia levels. Biotechnol. Biofuels 2015, 8, 154. [Google Scholar] [CrossRef]
- Cai, Y.; Hua, B.; Gao, L.; Hu, Y.; Yuan, X.; Cui, Z.; Zhu, W.; Wang, X. Effects of adding trace elements on rice straw anaerobic mono-digestion: Focus on changes in microbial communities using high-throughput sequencing. Bioresour. Technol. 2017, 239, 454–463. [Google Scholar] [CrossRef]
- Carballa, M.; Regueiro, L.; Lema, J.M. Microbial management of anaerobic digestion: Exploiting the microbiome-functionality nexus. Curr. Opin. Biotechnol. 2015, 33, 103–111. [Google Scholar] [CrossRef]
- Griffiths, R.I.; Whiteley, A.S.; O’Donnell, A.G.; Bailey, M.J. Rapid method for coextraction of DNA and RNA from natural environments for analysis of ribosomal DNA-and rRNA-based microbial community composition. Appl. Environ. Microbiol. 2000, 66, 5488–5491. [Google Scholar] [CrossRef]
- Keating, C.; Chin, J.P.; Hughes, D.; Manesiotis, P.; Cysneiros, D.; Mahony, T.; Smith, C.J.; McGrath, J.W.; O’Flaherty, V. Biological phosphorus removal during high-rate, low-temperature, anaerobic digestion of wastewater. Front. Microbiol. 2016, 7, 226. [Google Scholar] [CrossRef]
- Caporaso, J.G.; Lauber, C.L.; Walters, W.A.; Berg-Lyons, D.; Huntley, J.; Fierer, N.; Owens, S.M.; Betley, J.; Fraser, L.; Bauer, M.; et al. Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. ISME J. 2012, 6, 1621. [Google Scholar] [CrossRef] [PubMed]
- Connelly, S.; Shin, S.G.; Dillon, R.J.; Ijaz, U.Z.; Quince, C.; Sloan, W.T.; Collins, G. Bioreactor Scalability: Laboratory-Scale Bioreactor Design Influences Performance, Ecology, and Community Physiology in Expanded Granular Sludge Bed Bioreactors. Front. Microbiol. 2017, 8, 664. [Google Scholar] [CrossRef] [PubMed]
- Masella, A.P.; Bartram, A.K.; Truszkowski, J.M.; Brown, D.G.; Neufeld, J.D. PANDAseq: Paired-end assembler for illumina sequences. BMC Bioinform. 2012, 13, 31. [Google Scholar] [CrossRef] [PubMed]
- Rognes, T.; Flouri, T.; Nichols, B.; Quince, C.; Mahé, F. VSEARCH: A versatile open source tool for metagenomics. Peer J. 2016, 4, e2584. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R.C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 2010, 26, 2460–2461. [Google Scholar] [CrossRef]
- Aßhauer, K.P.; Wemheuer, B.; Daniel, R.; Meinicke, P. Tax4Fun: Predicting functional profiles from metagenomic 16S rRNA data. Bioinformatics 2015, 31, 2882–2884. [Google Scholar] [CrossRef]
- Zabranska, J.; Pokorna, D. Bioconversion of carbon dioxide to methane using hydrogen and hydrogenotrophic methanogens. Biotechnol. Adv. 2018, 36, 707–720. [Google Scholar] [CrossRef]
- Liu, C.; Wachemo, A.C.; Tong, H.; Shi, S.; Zhang, L.; Yuan, H.; Li, X. Biogas production and microbial community properties during anaerobic digestion of corn stover at different temperatures. Bioresour. Technol. 2018, 261, 93–103. [Google Scholar] [CrossRef]
- Azman, S.; Khadem, A.F.; van Lier, J.B.; Zeeman, G.; Plugge, C.M. Presence and Role of Anaerobic Hydrolytic Microbes in Conversion of Lignocellulosic Biomass for Biogas Production. Crit. Rev. Environ. Sci. Technol. 2015, 45, 2523–2564. [Google Scholar] [CrossRef]
- Vanwonterghem, I.; Jensen, P.D.; Ho, D.P.; Batstone, D.J.; Tyson, G.W. Linking microbial community structure, interactions and function in anaerobic digesters using new molecular techniques. Curr. Opin. Biotechnol. 2014, 27, 55–64. [Google Scholar] [CrossRef]
- Koch, C.; Müller, S.; Harms, H.; Harnisch, F. Microbiomes in bioenergy production: From analysis to management. Curr. Opin. Biotechnol. 2014, 27, 65–72. [Google Scholar] [CrossRef] [PubMed]
- Sboner, A.; Mu, X.J.; Greenbaum, D.; Auerbach, R.K.; Gerstein, M.B. The real cost of sequencing: Higher than you think! Genome Biol. 2011, 12, 125. [Google Scholar] [CrossRef] [PubMed]
- Muir, P.; Li, S.; Lou, S.; Wang, D.; Spakowicz, D.J.; Salichos, L.; Zhang, J.; Weinstock, G.M.; Isaacs, F.; Rozowsky, J.; et al. The real cost of sequencing: Scaling computation to keep pace with data generation. Genome Biol. 2016, 17, 53. [Google Scholar] [CrossRef] [PubMed]
- Rivière, D.; Desvignes, V.; Pelletier, E.; Chaussonnerie, S.; Guermazi, S.; Weissenbach, J.; Li, T.; Camacho, P.; Sghir, A. Towards the definition of a core of microorganisms involved in anaerobic digestion of sludge. ISME J. 2009, 3, 700. [Google Scholar] [CrossRef] [PubMed]
| Parameter | CT Influent | SST Influent | P-Value |
|---|---|---|---|
| Temperature (°C) * | 37.2 ± 4.5 | 42.7 ± 4.9 | <0.001 |
| Wastewater flowrate (L/d) | 72.1 ± 32.1 | 93.7 ± 48.1 | <0.001 |
| Organic loading rate (kg/d) | 0.14 ± 0.06 | 0.47 ± 0.53 | <0.001 |
| Solid loading rate (kg/d) | 0.12 ± 0.06 | 0.24 ± 0.14 | <0.001 |
| TS (mg/L) | 1895 ± 414 | 2550 ± 1015 | 0.153 |
| TVS (mg/L) | 1210 ± 294 | 2088 ± 933 | 0.048 |
| TSS (mg/L) | 1032 ± 545 | 1834 ± 953 | 0.083 |
| TBOD (mg/L) | 734 ± 345 | 800 ± 369 | 0.719 |
| TCOD (mg/L) | 1823 ± 582 | 6093 ± 6731 | 0.116 |
| SCOD (mg/L) | 671 ± 253 | 683 ± 187 | 0.917 |
| TKN (mg/L as N) | 394 ± 86 | 375 ± 97 | 0.690 |
| NH3 (mg/L as N) | 312 ± 73 | 196 ± 77 | 0.008 |
| TP (mg/L as P) | 40 ± 17 | 34 ± 25 | 0.604 |
| Total coliform (MPN per 100mL) | 3.40 × 107 ± 5.45 × 107 | 1.03 × 107 ± 1.94 × 107 | 0.279 |
| E.coli (MPN per 100mL) | 1.03 × 107 ± 1.847 | 1.26 × 107 ± 1.90 × 107 | 0.814 |
| Parameter | CT Effluent | CT Removal Efficiency (%) | SST Effluent | SST Removal Efficiency (%) | P-Value |
|---|---|---|---|---|---|
| TS (mg/L) | 967 ± 169 | 48 ± 5 | 564 ± 160 | 73 ± 16 | 0.006 |
| TVS (mg/L) | 433 ± 106 | 62 ± 17 | 172 ± 62 | 89 ± 7 | 0.004 |
| TSS (mg/L) | 158 ± 92 | 84 ± 5 | 63 ± 18 | 96 ± 3 | <0.001 |
| TBOD (mg/L) | 181 ± 84 | 70 ± 22 | 138 ± 112 | 75 ± 28 | 0.660 |
| TCOD (mg/L) | 534 ± 254 | 70 ± 11 | 359 ± 275 | 89 ± 9 | 0.002 |
| SCOD (mg/L) | 223 ± 146 | 64 ± 18 | 171 ± 74 | 73 ± 15 | 0.329 |
| TKN (mg/L as N) | 366 ± 49 | 3 ± 22 | 296 ± 29 | 17 ± 23 | 0.261 |
| NH3 (mg/L as N) | 341 ± 34 | −14 ± 26 | 287 ± 32 | −66 ± 65 | 0.065 |
| TP (mg/L as P) | 34 ± 7 | 4 ± 29 | 25 ± 6 | −62 ± 199 | 0.378 |
| Total coliform *,* | 9.11 × 104 ± 1.11 × 105 | 2.33 ± 1.09 * | 1.73 × 104 ± 2.84 × 104 | 2.79 ± 0.95 * | 0.399 |
| E. coli *,* | 8.47 × 104 ± 1.14 × 105 | 2.01 ± 1.11 * | 1.73 × 104 ± 2.84 × 104 | 3.02 ± 0.69 * | 0.054 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Connelly, S.; Pussayanavin, T.; J. Randle-Boggis, R.; Wicheansan, A.; Jampathong, S.; Keating, C.; Z. Ijaz, U.; T. Sloan, W.; Koottatep, T. Solar Septic Tank: Next Generation Sequencing Reveals Effluent Microbial Community Composition as a Useful Index of System Performance. Water 2019, 11, 2660. https://doi.org/10.3390/w11122660
Connelly S, Pussayanavin T, J. Randle-Boggis R, Wicheansan A, Jampathong S, Keating C, Z. Ijaz U, T. Sloan W, Koottatep T. Solar Septic Tank: Next Generation Sequencing Reveals Effluent Microbial Community Composition as a Useful Index of System Performance. Water. 2019; 11(12):2660. https://doi.org/10.3390/w11122660
Chicago/Turabian StyleConnelly, Stephanie, Tatchai Pussayanavin, Richard J. Randle-Boggis, Araya Wicheansan, Suparat Jampathong, Ciara Keating, Umer Z. Ijaz, Willian T. Sloan, and Thammarat Koottatep. 2019. "Solar Septic Tank: Next Generation Sequencing Reveals Effluent Microbial Community Composition as a Useful Index of System Performance" Water 11, no. 12: 2660. https://doi.org/10.3390/w11122660
APA StyleConnelly, S., Pussayanavin, T., J. Randle-Boggis, R., Wicheansan, A., Jampathong, S., Keating, C., Z. Ijaz, U., T. Sloan, W., & Koottatep, T. (2019). Solar Septic Tank: Next Generation Sequencing Reveals Effluent Microbial Community Composition as a Useful Index of System Performance. Water, 11(12), 2660. https://doi.org/10.3390/w11122660





