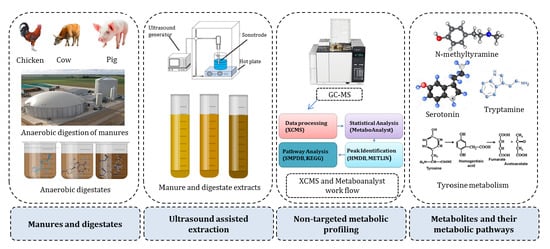
Untargeted Metabolite Profiling for Screening Bioactive Compounds in Digestate of Manure under Anaerobic Digestion
Abstract
1. Introduction
2. Materials and Methods
2.1. Materials
2.2. Extraction and GCMS Detection
2.3. Data Analysis
3. Results and Discussion
3.1. Primary Metabolite Profiling
3.2. Primary Metabolite Derivative Pathways
3.3. Characterization of Bioactive Compounds
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Duan, N.; Lin, C.; Wang, P.; Meng, J.; Chen, H.; Li, X.L. Ecological analysis of a typical farm-scale biogas plant in China. Front. Earth Sci. 2014, 8, 375–384. [Google Scholar] [CrossRef]
- Cao, Y.; Wang, J.; Wu, H.; Yan, S.; Guo, D.; Wang, G.; Ma, Y. Soil chemical and microbial responses to biogas slurry amendment and its effect on Fusarium wilt suppression. Appl. Soil Ecol. 2016, 107, 116–123. [Google Scholar] [CrossRef]
- Wentzel, S.; Joergensen, R.G. Effects of biogas and raw slurries on grass growth and soil microbial indices. J. Plant Nutr. Soil Sci. 2016, 179, 215–222. [Google Scholar] [CrossRef]
- Zheng, X.; Fan, J.; Cui, J.; Wang, Y.; Zhou, J.; Ye, M.; Sun, M. Effects of biogas slurry application on peanut yield, soil nutrients, carbon storage, and microbial activity in an Ultisol soil in southern China. J. Soils Sediments 2016, 16, 449–460. [Google Scholar] [CrossRef]
- Zhang, S.; Hua, Y.; Deng, L. Nutrient Status and Contamination Risks from Digested Pig Slurry Applied on a Vegetable Crops Field. Int. J. Environ. Res. Public Health 2016, 13, 406. [Google Scholar] [CrossRef] [PubMed]
- Bougnom, B.P.; Niederkofler, C.; Knapp, B.A.; Stimpfl, E.; Insam, H. Residues from renewable energy production: Their value for fertilizing pastures. Biomass Bioenergy 2012, 39, 290–295. [Google Scholar] [CrossRef]
- Ahmad, M.Z.; Zahir, Z.A.; Jamil, M.; Nazli, F.; Latif, M. Integrated use of plant growth promoting rhizobacteria, biogas slurry and chemical nitrogen for sustainable production of maize under salt-affected conditions. Pak. J. Bot. 2014, 46, 375–382. [Google Scholar]
- Cao, Y.; Chang, Z.; Wang, J.; Ma, Y.; Fu, G. The fate of antagonistic microorganisms and antimicrobial substances during anaerobic digestion of pig and dairy manure. Bioresour. Technol. 2013, 136, 664–671. [Google Scholar] [CrossRef]
- Heuer, H.; Schmitt, H.; Smalla, K. Antibiotic resistance gene spread due to manure application on agricultural fields. Curr. Opin. Microbiol. 2011, 14, 236–243. [Google Scholar] [CrossRef]
- Tambone, F.; Adani, F.; Gigliotti, G.; Volpe, D.; Fabbri, C.; Provenzano, M.R. Organic matter characterization during the anaerobic digestion of different biomasses by means of CPMAS 13C NMR spectroscopy. Biomass Bioenergy 2013, 48, 111–120. [Google Scholar] [CrossRef]
- Lorenz, K.; Lal, R.; Preston, C.M.; Nierop, K.G.J. Strengthening the soil organic carbon pool by increasing contributions from recalcitrant aliphatic bio(macro)molecules. Geoderma 2007, 142, 1–10. [Google Scholar] [CrossRef]
- Popp, D.; Harms, H.; Sträuber, H. The alkaloid gramine in the anaerobic digestion process—Inhibition and adaptation of the methanogenic community. Appl. Microbiol. Biotechnol. 2016, 100, 7311–7322. [Google Scholar] [CrossRef]
- Han, J.S.; Lee, S.; Kim, H.Y.; Lee, C.H. MS-Based Metabolite Profiling of Aboveground and Root Components of Zingiber mioga and Officinale. Molecules 2015, 20, 16170–16185. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Nagele, T.; Doerfler, H.; Fragner, L.; Chaturvedi, P.; Nukarinen, E.; Bellaire, A.; Huber, W.; Weiszmann, J.; Engelmeier, D.; et al. System level analysis of cacao seed ripening reveals a sequential interplay of primary and secondary metabolism leading to polyphenol accumulation and preparation of stress resistance. Plant. J. 2016, 87, 318–332. [Google Scholar] [CrossRef] [PubMed]
- Scaglia, B.; Pognani, M.; Adani, F. Evaluation of hormone-like activity of the dissolved organic matter fraction (DOM) of compost and digestate. Sci. Total Environ. 2015, 514, 314–321. [Google Scholar] [CrossRef]
- Caroline, H.J.; Julijana, I.; Gary, S. Metabolomics: Beyond biomarkers and towards mechanisms. Nat. Rev. Mol. Cell Biol. 2016, 17, 451–459. [Google Scholar]
- Hites, R.A.; Jobst, K.J. Is Nontargeted Screening Reproducible? Environ. Sci. Technol. 2018, 52, 11975–11976. [Google Scholar] [CrossRef]
- Nomura, D.K.; Long, J.Z.; Niessen, S.; Hoover, H.S.; Ng, S.W.; Cravatt, B.F. Monoacylglycerol lipase regulates a fatty acid network that promotes cancer pathogenesis. Cell 2010, 140, 49–61. [Google Scholar] [CrossRef]
- Patti, G.J.; Yanes, O.; Siuzdak, G. Metabolomics implicates altered sphingolipids in chronic pain of neuropathic origin. Nat. Chem. Biol. 2012, 8, 232–234. [Google Scholar] [CrossRef]
- Yanes, O.; Clark, J.; Wong, D.M.; Patti, G.J.; Sánchez-Ruiz, A.; Benton, H.P.; Trauger, S.A.; Desponts, C.; Ding, S.; Siuzdak, G. Metabolic oxidation regulates embryonic stem cell differentiation. Nat. Chem. Biol. 2010, 6, 411–417. [Google Scholar] [CrossRef]
- Lu, J.; Muhmood, A.; Liu, H.; Dong, R.; Pang, S.; Wu, S. Exploring Bioactive Compounds in Anaerobically Digested Slurry: Extraction, Characterization, and Assessment of Antifungal Activity. Waste Biomass Valorization 2018, 1–10. [Google Scholar] [CrossRef]
- Tautenhahn, R.; Patti, G.J.; Rinehart, D.; Siuzdak, G. XCMS Online: A web-based platform to process untargeted metabolomic data. Anal. Chem. 2012, 84, 5035–5039. [Google Scholar] [CrossRef] [PubMed]
- Gowda, H.; Ivanisevic, J.; Johnson, C.H.; Kurczy, M.E.; Benton, H.P.; Rinehart, D.; Nguyen, T.; Ray, J.; Kuehl, J.; Arevalo, B.; et al. Interactive XCMS Online: Simplifying advanced metabolomic data processing and subsequent statistical analyses. Anal. Chem. 2014, 86, 6931–6939. [Google Scholar] [CrossRef] [PubMed]
- Xia, J.; Wishart, D.S. Web-based inference of biological patterns, functions and pathways from metabolomic data using MetaboAnalyst. Nat. Protoc. 2011, 6, 743–760. [Google Scholar] [CrossRef] [PubMed]
- Aittokallio, T.; Schwikowski, B. Graph-based methods for analysing networks in cell biology. Brief. Bioinform. 2006, 7, 243–255. [Google Scholar] [CrossRef]
- Patti, G.J.; Yanes, O.; Siuzdak, G. Innovation: Metabolomics: The apogee of the omics trilogy. Nat. Rev. Mol. Cell Biol. 2012, 13, 263–269. [Google Scholar] [CrossRef]
- Wang, K.; Li, X.; He, C.; Chen, C.L.; Bai, J.; Ren, N.; Wang, J.Y. Transformation of dissolved organic matters in swine, cow and chicken manures during composting. Bioresour. Technol. 2014, 168, 222–228. [Google Scholar] [CrossRef]
- Huang, J.; Yu, Z.; Gao, H.; Yan, X.; Chang, J.; Wang, C.; Hu, J.; Zhang, L. Chemical structures and characteristics of animal manures and composts during composting and assessment of maturity indices. PLoS ONE 2017, 12, e0178110. [Google Scholar] [CrossRef]
- Nie, H.; Jacobi, H.F.; Strach, K.; Xu, C.; Zhou, H.; Liebetrau, J. Mono-fermentation of chicken manure: Ammonia inhibition and recirculation of the digestate. Bioresour. Technol. 2015, 178, 238–246. [Google Scholar] [CrossRef]
- Tesso, T.A.; Liu, G. Indole Degradation in a Model System and in Poultry Manure by Acinetobacter spp. Appl. Sci. 2019, 9, 1622. [Google Scholar] [CrossRef]
- Yan, Q.; Liu, X.; Wang, Y.; Li, H.; Li, Z.; Zhou, L.; Qu, Y.; Li, Z.; Bao, X. Cow manure as a lignocellulosic substrate for fungal cellulase expression and bioethanol production. AMB Express 2018, 8, 190. [Google Scholar] [CrossRef] [PubMed]
- Sawatdeenarunat, C.; Surendra, K.C.; Takara, D.; Oechsner, H.; Khanal, S.K. Anaerobic digestion of lignocellulosic biomass: Challenges and opportunities. Bioresour. Technol. 2015, 178, 178–186. [Google Scholar] [CrossRef] [PubMed]
- Sakar, S.; Yetilmezsoy, K.; Kocak, E. Anaerobic digestion technology in poultry and livestock waste treatment—A literature review. Waste Manag. Res. 2009, 27, 3–18. [Google Scholar] [CrossRef] [PubMed]
- Yang, D.; Fan, X.; Shi, X.; Lian, S.; Qiao, J.; Guo, R. Metabolomics reveals stage-specific metabolic pathways of microbial communities in two-stage anaerobic fermentation of corn-stalk. Biotechnol. Lett. 2014, 36, 1461–1468. [Google Scholar] [CrossRef]
- Li, P.; Tang, H.; Shi, C.; Xie, Y.; Zhou, H.; Xia, B.; Zhang, C.; Chen, L.; Jiang, L. Untargeted metabolomics analysis of Mucor racemosus Douchi fermentation process by gas chromatography with time-of-flight mass spectrometry. Food Sci. Nutr. 2019, 7, 1865–1874. [Google Scholar] [CrossRef]
- Xia, J.; Wishart, D.S. The efficiency of shredded and briquetted wheat straw in anaerobic co-digestion with dairy cattle manure. Biosyst. Eng. 2015, 139, 16–24. [Google Scholar]
- Gadde, U.D.; Oh, S.; Lillehoj, H.S.; Lillehoj, E.P. Antibiotic growth promoters virginiamycin and bacitracin methylene disalicylate alter the chicken intestinal metabolome. Sci. Rep. 2018, 8, 3592. [Google Scholar] [CrossRef]
- Granot, D.; David-Schwartz, R.; Kelly, G. Hexose kinases and their role in sugar-sensing and plant development. Front. Plant. Sci. 2013, 4, 44. [Google Scholar] [CrossRef]
- Dove, C.R. Vitamins: Water Soluble: Pantothenic Acid, Folic Acid, and B12. In Encyclopedia of Animal Science, 2nd ed.; CRC Press: Boca Raton, FL, USA, 2011; pp. 1076–1078. [Google Scholar]
- Ragaller, V.; Lebzien, P.; Sudekum, K.H.; Huther, L.; Flachowsky, G. Pantothenic acid in ruminant nutrition: A review. J. Anim. Physiol. Anim. Nutr. 2011, 95, 6–16. [Google Scholar] [CrossRef]
- Muller, S.; Kappes, B. Vitamin and cofactor biosynthesis pathways in Plasmodium and other apicomplexan parasites. Trends Parasitol. 2007, 23, 112–121. [Google Scholar] [CrossRef][Green Version]
- Angelidaki, I.; Ahring, B.K. Methods for increasing the biogas potential from the recalcitrant organic matter contained in manure. Water Sci. Technol. 2000, 41, 189–194. [Google Scholar] [CrossRef] [PubMed]
- Vanholme, R.; Ralph, J.; Akiyama, T.; Lu, F.; Pazo, J.R.; Kim, H.; Christensen, J.H.; Van Reusel, B.; Storme, V.; De Rycke, R.; et al. Engineering traditional monolignols out of lignin by concomitant up-regulation of F5H1 and down-regulation of COMT in Arabidopsis. Plant. J. 2010, 64, 885–897. [Google Scholar] [CrossRef] [PubMed]
- Mackie, R.I.; Stroot, P.G.; Varel, V.H. Biochemical identification and biological origin of key odor components in livestock waste. J. Anim. Sci. 1998, 76, 1331–1342. [Google Scholar] [CrossRef] [PubMed]
- Marhuenda-Egea, F.C.; Martinez-Sabater, E.; Jorda, J.; Moral, R.; Bustamante, M.A.; Paredes, C.; Pérez-Murcia, M.D. Dissolved organic matter fractions formed during composting of winery and distillery residues: Evaluation of the process by fluorescence excitation-emission matrix. Chemosphere 2007, 68, 301–309. [Google Scholar] [CrossRef] [PubMed]
- Shrestha, S.; Fonoll, X.; Khanal, S.K.; Raskin, L. Biological strategies for enhanced hydrolysis of lignocellulosic biomass during anaerobic digestion: Current status and future perspectives. Bioresour. Technol. 2017, 245, 1245–1257. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Guo, J.; Dong, R.; Zhang, W. Indolic Derivatives Metabolism in the Anaerobic Reactor Treating Animal Manure: Pathways and Regulation. ACS Sustain. Chem. Eng. 2018, 6, 11511–11518. [Google Scholar] [CrossRef]
- Xu, J.; Hu, F.L.; Wang, W.; Wan, X.C.; Bao, G.H. Investigation on biochemical compositional changes during the microbial fermentation process of Fu brick tea by LC-MS based metabolomics. Food Chem. 2015, 186, 176–184. [Google Scholar] [CrossRef]
- Combalbert, S.; Bellet, V.; Dabert, P.; Bernet, N.; Balaguer, P.; Hernandez-Raquet, G. Fate of steroid hormones and endocrine activities in swine manure disposal and treatment facilities. Water Res. 2012, 46, 895–906. [Google Scholar] [CrossRef]
- Gao, J.; Xu, K.; Liu, H.; Liu, G.; Bai, M.; Peng, C.; Li, T.; Yin, Y. Impact of the Gut Microbiota on Intestinal Immunity Mediated by Tryptophan Metabolism. Front. Cell. Infect. Microbiol. 2018, 8, 13. [Google Scholar] [CrossRef]
- Jensen, M.T.; Cox, R.P.; Jensen, B.B. 3-Methylindole (Skatole) and Indole Production by Mixed Populations of Pig Fecal Bacteria. Appl. Environ. Microbiol. 1995, 61, 3180–3184. [Google Scholar]




| Biogas Plant | Doudian | Donghuashan | Liu Minying | |||
|---|---|---|---|---|---|---|
| Model | CSTR | USR | USR | |||
| Temperature (°C) | 33–35 | 32–35 | 30 | |||
| HRT (d) | 30 | 20 | 15 | |||
| OLR (kg m−³d⁻¹) | 3–4 | 4.0–4.7 | 3–4 | |||
| Parameters | Cow Manure | Digestate | Swine Manure | Digestate | Chicken Manure | Digestate |
| pH | 6.80 ± 0.18 | 7.75 ± 0.23 | 6.55 ± 0.20 | 7.41 ± 0.18 | 7.78 ± 0.23 | 8.38 ± 0.26 |
| VS (%) | 7.20 ± 0.27 | 6.03 ± 0.02 | 7.64 ± 0.22 | 6.33 ± 0.33 | 7.10 ± 0.59 | 4.02 ± 0.68 |
| COD (g L−1) | 3.20 ± 1.03 | 8.8 ± 4.32 | 9.4 ± 2.23 | 18.6 ± 3.34 | 78.7 ± 3.56 | 67.5 ± 6.89 |
| NH₄⁺-N (mg L−1) | 333.4 ± 62 | 1959 ± 79 | 979.3 ± 29.3 | 1938 ± 7.80 | 8199 ± 53.3 | 7032 ± 34.6 |
| NO₃−-N (mg L−1) | 1.70 ± 0.27 | 0.89 ± 0.13 | 2.02 ± 0.29 | 0.96 ± 0.14 | 6.71 ± 0.45 | 4.71 ± 0.92 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lu, J.; Muhmood, A.; Czekała, W.; Mazurkiewicz, J.; Dach, J.; Dong, R. Untargeted Metabolite Profiling for Screening Bioactive Compounds in Digestate of Manure under Anaerobic Digestion. Water 2019, 11, 2420. https://doi.org/10.3390/w11112420
Lu J, Muhmood A, Czekała W, Mazurkiewicz J, Dach J, Dong R. Untargeted Metabolite Profiling for Screening Bioactive Compounds in Digestate of Manure under Anaerobic Digestion. Water. 2019; 11(11):2420. https://doi.org/10.3390/w11112420
Chicago/Turabian StyleLu, Jiaxin, Atif Muhmood, Wojciech Czekała, Jakub Mazurkiewicz, Jacek Dach, and Renjie Dong. 2019. "Untargeted Metabolite Profiling for Screening Bioactive Compounds in Digestate of Manure under Anaerobic Digestion" Water 11, no. 11: 2420. https://doi.org/10.3390/w11112420
APA StyleLu, J., Muhmood, A., Czekała, W., Mazurkiewicz, J., Dach, J., & Dong, R. (2019). Untargeted Metabolite Profiling for Screening Bioactive Compounds in Digestate of Manure under Anaerobic Digestion. Water, 11(11), 2420. https://doi.org/10.3390/w11112420