Presence of Antibiotic-Resistant Escherichia coli in Wastewater Treatment Plant Effluents Utilized as Water Reuse for Irrigation
Abstract
1. Introduction
2. Materials and Methods
2.1. Sample Collection
2.2. Antibiotic Resistant Escherichia coli Culture Collection
2.2.1. Escherichia coli Isolation
2.2.2. Isolate Identification
2.3. Antibiotic Susceptibility Testing of the Isolates
2.4. Data Analysis
3. Results
3.1. Presumptive E. coli Growth on Plates Supplemented with Antibiotics
3.1.1. Growth on the Control Plates
3.1.2. E. coli Growth on Antibiotic Supplemented Media
3.1.3. Antibiotic Susceptibility in E. coli Isolates
4. Discussion
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Davies, J.; Davies, D. Origins and evolution of antibiotic resistance. Microbiol. Mol. Biol. Rev. 2010, 74, 417–433. [Google Scholar] [CrossRef] [PubMed]
- Stoll, C.; Sidhu, J.P.S.; Tiehm, A.; Toze, S. Prevalence of clinically relevant antibiotic resistance genes in surface water samples collected from Germany and Australia. Environ. Sci. Technol. 2012, 46, 9716–9726. [Google Scholar] [CrossRef] [PubMed]
- Burch, T.R.; Sadowsky, M.J.; LaPara, T.M. Fate of antibiotic resistance genes and class 1 integrons in soil microcosms following the application of treated residual municipal wastewater solids. Environ. Sci. Technol. 2014, 48, 5620–5627. [Google Scholar] [CrossRef] [PubMed]
- Ben, W.; Wang, J.; Cao, R.; Yang, M.; Zhang, Y.; Qiang, Z. Distribution of antibiotic resistance in the effluents of ten municipal wastewater treatment plants in China and the effect of treatment processes. Chemosphere 2017, 172, 392–398. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Watanabe, N.; Xiao, C.; Harter, T.; McCowan, B.; Liu, Y.; Atwill, E.R. Antibiotic-resistant E. coli in surface water and groundwater in dairy operations in Northern California. Environ. Monit. Assess. 2014, 186, 1253–1260. [Google Scholar] [CrossRef] [PubMed]
- Carnelli, A.; Mauri, F.; Demarta, A. Characterization of genetic determinants involved in antibiotic resistance in Aeromonas spp. and fecal coliforms isolated from different aquatic environments. Res. Microbiol. 2017, 168, 461–471. [Google Scholar] [CrossRef] [PubMed]
- Port, J.A.; Cullen, A.C.; Wallace, J.C.; Smith, M.N.; Faustman, E.M. Metagenomic frameworks for monitoring antibiotic resistance in aquatic environments. Environ. Health Perspect. 2014, 122, 222–228. [Google Scholar] [CrossRef] [PubMed]
- Manaia, C.M.; Macedo, G.; Fatta-Kassinos, D.; Nunes, O.C. Antibiotic resistance in urban aquatic environments: Can it be controlled? Appl. Microbiol. Biotechnol. 2016, 100, 1543–1557. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.; Han, B.; Gu, J.; Wang, C.; Wang, P.; Ma, Y.; Cao, J.; He, Z. Fate of antibiotic resistant cultivable heterotrophic bacteria and antibiotic resistance genes in wastewater treatment processes. Chemosphere 2015, 135, 138–145. [Google Scholar] [CrossRef] [PubMed]
- Frey, S.K.; Topp, E.; Khan, I.U.H.; Ball, B.R.; Edwards, M.; Gottschall, N.; Sunohara, M.; Lapen, D.R. Quantitative Campylobacter spp., antibiotic resistance genes, and veterinary antibiotics in surface and ground water following manure application: Influence of tile drainage control. Sci. Total Environ. 2015, 532, 138–153. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.-H.; Qiao, M.; Lv, Z.-E.; Guo, G.-X.; Jia, Y.; Su, Y.-H.; Zhu, Y.-G. Impact of reclaimed water irrigation on antibiotic resistance in public parks, Beijing, China. Environ. Pollut. 2014, 184, 247–253. [Google Scholar] [CrossRef] [PubMed]
- Blaustein, R.A.; Shelton, D.R.; Van Kessel, J.A.; Karns, J.S.; Stocker, M.D.; Pachepsky, Y.A. Irrigation waters and pipe-based biofilms as sources for antibiotic-resistant bacteria. Environ. Monit. Assess. 2016, 188, 56. [Google Scholar] [CrossRef] [PubMed]
- World Health Organization. Antimicrobial Resistance: An emerging Water, Sanitation and Hygiene Issue Briefing Note; WHO/FWC/WSH/14.07; World Health Organization: Geneva, Switzerland, 2014. [Google Scholar] [CrossRef]
- Xu, J.; Chen, W.; Wu, L.; Green, R.; Chang, A.C. Leachability of some emerging contaminants in reclaimed municipal wastewater-irrigated turf grass fields. Environ. Toxicol. Chem. 2009, 28, 1842–1850. [Google Scholar] [CrossRef] [PubMed]
- United States Environmental Protection Agency. Method 1604: Total Coliforms and Escherichia coli in Water by Membrane Filtration Using a Simultaneous Detection Technique (MI Medium); Standard Methods, EPA-821-R-02-024; United States Environmental Protection Agency: Washington, DC, USA, 2002. [Google Scholar]
- Brenner, K.P.; Rankin, C.C.; Roybal, Y.R.; Stelma, G.N.; Scarpino, P.V; Dufour, A.P. New medium for the simultaneous detection of total coliforms and Escherichia coli in water. Appl. Environ. Microbiol. 1993, 59, 3534–3544. [Google Scholar] [PubMed]
- Clinical and Laboratory Standards Institute. Performance Standards for Antimicrobial Susceptibility Testing, 27th ed.; CLSI Supplement M100; Clinical and Laboratory Standards Institute: Wayne, PA, USA, 2017. [Google Scholar]
- Watkinson, A.J.; Micalizzi, G.R.; Bates, J.R.; Costanzo, S.D. Novel method for rapid assessment of antibiotic resistance in Escherichia coli isolates from environmental waters by use of a modified chromogenic agar. Appl. Environ. Microbiol. 2007, 73, 2224–2229. [Google Scholar] [CrossRef] [PubMed]
- United States Environmental Protection Agency. Method 1609: Enterococci in Water by TaqMan® Quantitative Polymerase Chain Reaction (qPCR) with Internal Amplification Control (IAC) Assay; Standard Methods, EPA-820-R-15-099; U.S. Environmental Protection Agency: Washington, DC, USA, 2015. [Google Scholar]
- Chern, E.C.; Siefring, S.; Paar, J.; Doolittle, M.; Haugland, R.A. Comparison of quantitative PCR assays for Escherichia coli targeting ribosomal RNA and single copy genes. Lett. Appl. Microbiol. 2011, 52, 298–306. [Google Scholar] [CrossRef] [PubMed]
- Roe, M.T.; Vega, E.; Pillai, S.D. Antimicrobial resistance markers of Class 1 and Class 2 Integron-bearing Escherichia coli from irrigation water and sediments. Emerg. Infect. Dis. 2003, 9, 822–826. [Google Scholar] [CrossRef] [PubMed]
- Vital, P.G.; Zara, E.S.; Paraoan, C.E.M.; Dimasupil, M.A.Z.; Abello, J.J.M.; Santos, I.T.G.; Rivera, W.L. Antibiotic resistance and extended-spectrum beta-lactamase production of Escherichia coli isolated from irrigation waters in selected urban farms in metro Manila, Philippines. Water 2018, 10, 548. [Google Scholar] [CrossRef]
- Aijuka, M.; Charimba, G.; Hugo, C.J.; Buys, E.M. Characterization of bacterial pathogens in rural and urban irrigation water. J. Water Health 2015, 13, 103–117. [Google Scholar] [CrossRef] [PubMed]
- Ishii, S.; Sadowsky, M.J. Escherichia coli in the environment: Implications for water quality and human health. Microbes Environ. 2008, 23, 101–108. [Google Scholar] [CrossRef] [PubMed]
- Smith, J.E.; Perdek, J.M. Assessment and management of watershed microbial contaminants. Crit. Rev. Environ. Sci. Technol. 2004, 34, 109–139. [Google Scholar] [CrossRef]
- Al-Jassim, N.; Ansari, M.I.; Harb, M.; Hong, P.Y. Removal of bacterial contaminants and antibiotic resistance genes by conventional wastewater treatment processes in Saudi Arabia: Is the treated wastewater safe to reuse for agricultural irrigation? Water Res. 2015, 73, 277–290. [Google Scholar] [CrossRef] [PubMed]
- Kim, I.; Yamashita, N.; Tanaka, H. Performance of UV and UV/H2O2 processes for the removal of pharmaceuticals detected in secondary effluent of a sewage treatment plant in Japan. J. Hazard. Mater. 2009, 166, 1134–1140. [Google Scholar] [CrossRef] [PubMed]
- Xi, C.; Zhang, Y.; Marrs, C.F.; Ye, W.; Simon, C.; Foxman, B.; Nriagu, J. Prevalence of antibiotic resistance in drinking water treatment and distribution systems. Appl. Environ. Microbiol. 2009, 75, 5714–5718. [Google Scholar] [CrossRef] [PubMed]
- Srinivasan, S.; Aslan, A.; Xagoraraki, I.; Alocilja, E.; Rose, J.B. Escherichia coli, Enterococci, and Bacteroides thetaiotaomicron qPCR signals through wastewater and septage treatment. Water Res. 2011, 45, 2561–2572. [Google Scholar] [CrossRef] [PubMed]
- United States Environmental Protection Agency. Guidelines for Water Reuse; EPA/600/R-12/61; U.S. Environmental Protection Agency: Washington, DC, USA, 2012. [Google Scholar]
- Carducci, A.; Verani, M. Effects of bacterial, chemical, physical and meteorological variables on virus removal by a wastewater treatment plant. Food Environ. Virol. 2013, 5, 69–76. [Google Scholar] [CrossRef] [PubMed]
- McLellan, S.L.; Huse, S.M.; Mueller-Spitz, S.R.; Andreishcheva, E.N.; Sogin, M.L. Diversity and population structure of sewage-derived microorganisms in wastewater treatment plant influent. Environ. Microbiol. 2010, 12, 378–392. [Google Scholar] [CrossRef] [PubMed]
- Chigor, V.N.; Umoh, V.J.; Smith, S.I.; Igbinosa, E.O.; Okoh, A.I. Multidrug resistance and plasmid patterns of Escherichia coli O157 and other E. coli isolated from diarrhoeal stools and surface waters from some selected sources in Zaria, Nigeria. Int. J. Environ. Res. Public Health 2010, 7, 3831–3841. [Google Scholar] [CrossRef] [PubMed]
- Winfiel, M.D.; Groisman, E.A. Role of nonhost enviroments in the lifestyles of Salmonella and E. coli. Appl. Environ. Microbiol. 2003, 69, 3687–3694. [Google Scholar] [CrossRef]
- McLain, J.E.T.; Williams, C.F. Assessing environmental impacts of treated wastewater through monitoring of fecal indicator bacteria and salinity in irrigated soils. Environ. Monit. Assess. 2012, 184, 1559–1572. [Google Scholar] [CrossRef] [PubMed]
- Jiang, Y.; Xu, C.; Wu, X.; Chen, Y.; Han, W.; Gin, K.Y.H.; He, Y. Occurrence, seasonal variation and risk assessment of antibiotics in Qingcaosha reservoir. Water 2018, 10, 115. [Google Scholar] [CrossRef]
- Bhullar, K.; Waglechner, N.; Pawlowski, A.; Koteva, K.; Banks, E.D.; Johnston, M.D.; Barton, H.A.; Wright, G.D. Antibiotic resistance is prevalent in an isolated cave microbiome. PLoS ONE 2012, 7. [Google Scholar] [CrossRef] [PubMed]
- Dcosta, V.M.; King, C.E.; Kalan, L.; Morar, M.; Sung, W.W.L.; Schwarz, C.; Froese, D.; Zazula, G.; Calmels, F.; Debruyne, R.; et al. Antibiotic resistance is ancient. Nature 2011, 477, 457–461. [Google Scholar] [CrossRef] [PubMed]
- Jin, G.; Englande, A.J.; Bradford, H.; Englande, A.J. Comparison of E. coli, Enterococci, and fecal coliform as indicators for brackish water quality assessment. Water Environ. Res. 2004, 76, 245–255. [Google Scholar] [CrossRef] [PubMed]
- Dick, L.K.; Stelzer, E.A; Bertke, E.E.; Fong, D.L.; Stoeckel, D.M. Relative decay of Bacteroidales microbial source tracking markers and cultivated Escherichia coli in freshwater microcosms. Appl. Environ. Microbiol. 2010, 76, 3255–3262. [Google Scholar] [CrossRef] [PubMed]
- Martinez, J.L. Environmental pollution by antibiotics and by antibiotic resistance determinants. Environ. Pollut. 2009, 157, 2893–2902. [Google Scholar] [CrossRef] [PubMed]
- Li, D.; Zeng, S.; He, M.; Gu, A.Z. Water disinfection byproducts induce antibiotic resistance-role of environmental pollutants in resistance phenomena. Environ. Sci. Technol. 2016, 50, 3193–3201. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.-M.; Xu, L.-M.; Wang, X.-C.; Zhuang, K.; Liu, Q.-Q. Effects of ultraviolet disinfection on antibiotic-resistant Escherichia coli from wastewater: Inactivation, antibiotic resistance profiles and antibiotic resistance genes. J. Appl. Microbiol. 2017, 123, 295–306. [Google Scholar] [CrossRef] [PubMed]
- Zhang, K.; Farahbakhsh, K. Removal of native coliphages and coliform bacteria from municipal wastewater by various wastewater treatment processes: Implications to water reuse. Water Res. 2007, 41, 2816–2824. [Google Scholar] [CrossRef] [PubMed]
- Quach-Cu, J.; Herrera-Lynch, B.; Marciniak, C.; Adams, S.; Simmerman, A.; Reinke, R.A. The effect of primary, secondary, and tertiary wastewater treatment processes on antibiotic resistance gene (ARG) concentrations in solid and dissolved wastewater fractions. Water 2018, 10, 37. [Google Scholar] [CrossRef]
| Antibiotic | Abbreviation | Drug Family |
|---|---|---|
| Ampicillin | AM | β-lactam penicillin |
| Ciprofloxacin | CI | Fluoroquinolone |
| Sulfamethoxazole/Trimethoprim | ST | Folic acid synthesis inhibitor |
| Tetracycline | TC | Tetracycline |
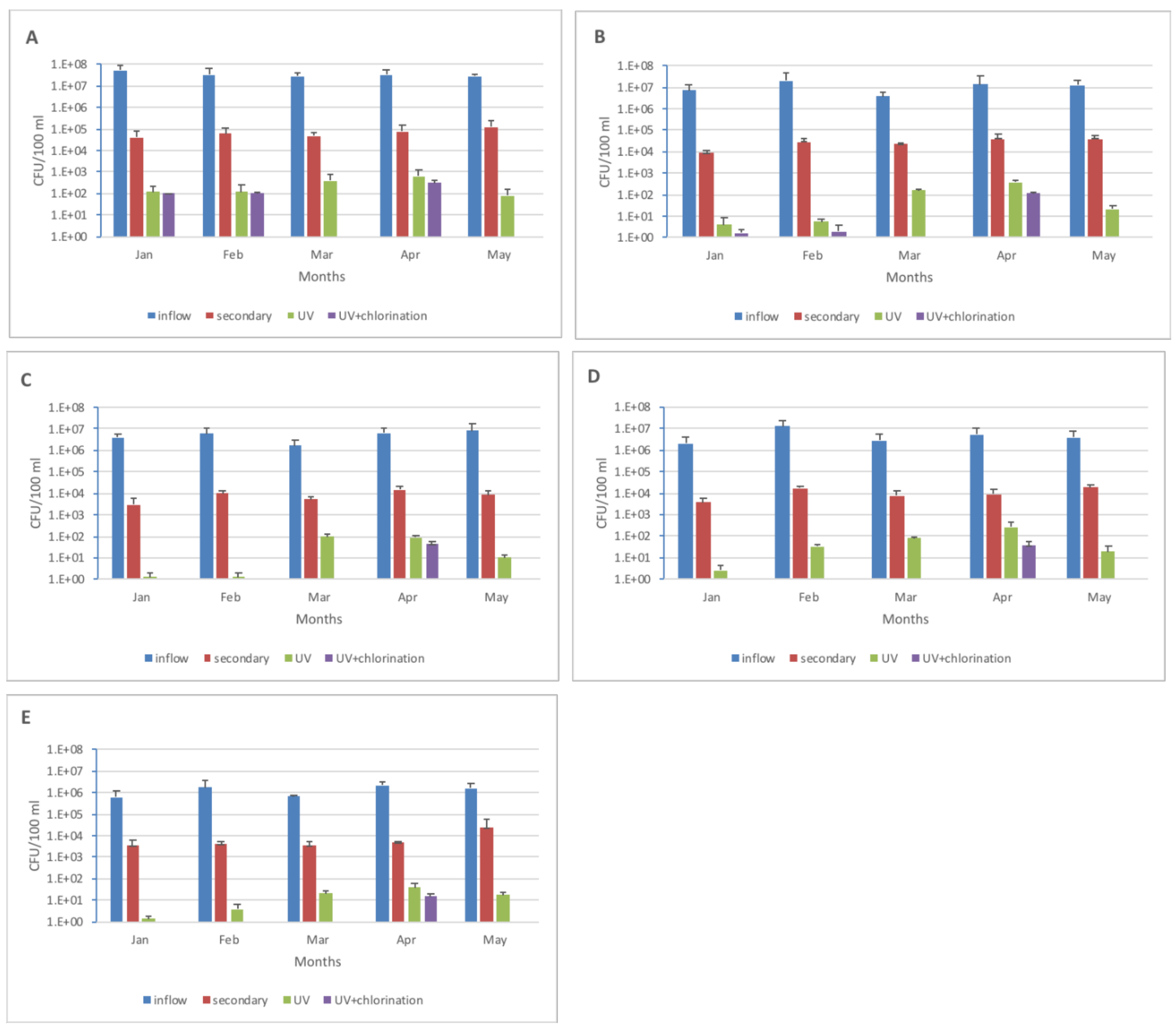
| Media | Influent to Secondary | Secondary to UV | Influent to UV | Influent to UV + Chlorination |
|---|---|---|---|---|
| Control | −2.68 | −2.56 | −5.24 | −6.33 |
| ST | −2.55 | −2.61 | −5.17 | −6.15 |
| CI | −3.11 | −2.17 | −5.29 | −5.95 |
| TC | −2.36 | −2.90 | −5.26 | −5.79 |
| AM | −2.50 | −2.83 | −5.33 | −6.26 |
| Stage | AM | ST | CI | TC |
|---|---|---|---|---|
| Inflow | 47 | 24 | 19 | 6 |
| Secondary | 42 | 17 | 12 | 12 |
| UV | 47 | 29 | 16 | 5 |
| UV + chlorination | 63 | 21 | 25 | 9 |
| Antibiotic | MIC Interpretive Criteria (μg/mL) | Percent Resistant † (n = 96) | ||
|---|---|---|---|---|
| S | I | R | ||
| AM | ≤8 | 16 | ≥32 | 95 |
| CI | ≤1 | 2 | ≥4 | 30 |
| ST | ≤2 | - | ≥4 | 70 |
| TC | ≤4 | 8 | ≥16 | 30 |
| Isolate | Location | AM | CI | ST | TC |
|---|---|---|---|---|---|
| EC1 | inflow | 64 | 4 | 6 | 2 |
| EC2 | secondary | 64 | 2 | 4 | 3 |
| EC3 | secondary | 48 | 3 | >32 | 0.5 |
| EC4 | inflow | 48 | 8 | 6 | 0.38 |
| EC5 | UV + chlorination | >256 | 0.5 | >32 | >256 |
| EC6 | UV + chlorination | 48 | 2 | >32 | 2 |
| EC7 | UV + chlorination | 64 | 24 | >32 | 4 |
| EC8 | UV | 48 | 3 | 4 | 1 |
| EC9 | UV + chlorination | >256 | 3 | 0.047 | >256 |
| EC10 | UV + chlorination | 128 | 6 | 4 | 4 |
| EC11 | secondary | >256 | 3 | 4 | 4 |
| EC12 | UV | 64 | 2 | 12 | 15 |
| EC13 | inflow | 64 | 4 | 0.38 | 24 |
| EC14 | UV + chlorination | >256 | 6 | >32 | 3 |
| EC15 | inflow | 48 | >32 | >32 | 4 |
| EC16 | inflow | 16 | 0.016 | 4 | 32 |
| EC17 | inflow | 24 | 4 | 0.047 | >256 |
| EC18 | UV | 24 | 3 | 24 | 0.5 |
| EC19 | UV | 48 | 4 | 6 | 2 |
| EC20 | secondary | 48 | 3 | >32 | 2 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Aslan, A.; Cole, Z.; Bhattacharya, A.; Oyibo, O. Presence of Antibiotic-Resistant Escherichia coli in Wastewater Treatment Plant Effluents Utilized as Water Reuse for Irrigation. Water 2018, 10, 805. https://doi.org/10.3390/w10060805
Aslan A, Cole Z, Bhattacharya A, Oyibo O. Presence of Antibiotic-Resistant Escherichia coli in Wastewater Treatment Plant Effluents Utilized as Water Reuse for Irrigation. Water. 2018; 10(6):805. https://doi.org/10.3390/w10060805
Chicago/Turabian StyleAslan, Asli, Zachariah Cole, Anunay Bhattacharya, and Oghenekpaobor Oyibo. 2018. "Presence of Antibiotic-Resistant Escherichia coli in Wastewater Treatment Plant Effluents Utilized as Water Reuse for Irrigation" Water 10, no. 6: 805. https://doi.org/10.3390/w10060805
APA StyleAslan, A., Cole, Z., Bhattacharya, A., & Oyibo, O. (2018). Presence of Antibiotic-Resistant Escherichia coli in Wastewater Treatment Plant Effluents Utilized as Water Reuse for Irrigation. Water, 10(6), 805. https://doi.org/10.3390/w10060805





