Bioinformatics Analysis of SNPs in IL-6 Gene Promoter of Jinghai Yellow Chickens
Abstract
:1. Introduction
2. Materials and Methods
2.1. Experimental Animals
2.2. Genomic DNA Extraction
2.3. Primer Design and Polymerase Chain Reaction Conditions
2.4. Gene Polymorphism Analysis
2.5. Bioinformatics Analysis of Interleukin-6 in Jinghai Yellow Chickens
2.5.1. Prediction of the Promoter Region
2.5.2. Transcription Factor Binding Site Prediction
2.5.3. Prediction of CpG Islands
2.5.4. Genetic Diversity Analysis
3. Results
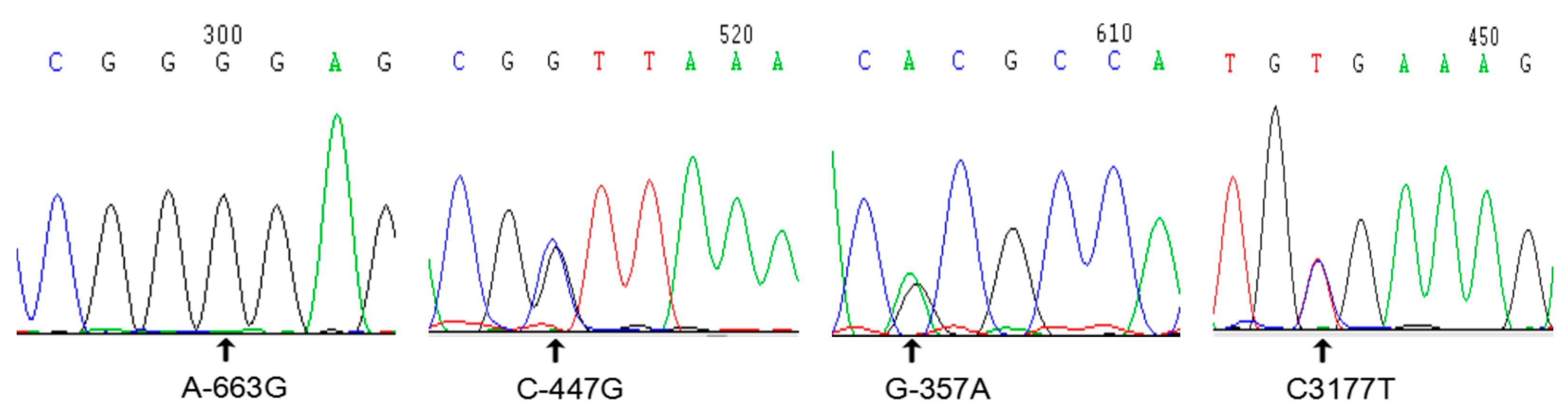
3.1. Single-Nucleotide Polymorphisms of Interleukin-6 in Jinghai Yellow Chickens
3.2. Population Genetic Variation of Newly Discovered Interleukin-6 Single-Nucleotide Polymorphisms
3.3. Results of Prediction of the Interleukin-6 Promoter Region and Screening of Promoter Region Single-Nucleotide Polymorphisms
3.4. Prediction of Putative Transcription Factor Binding in the Interleukin-6 Promoter by Single-Nucleotide Polymorphisms
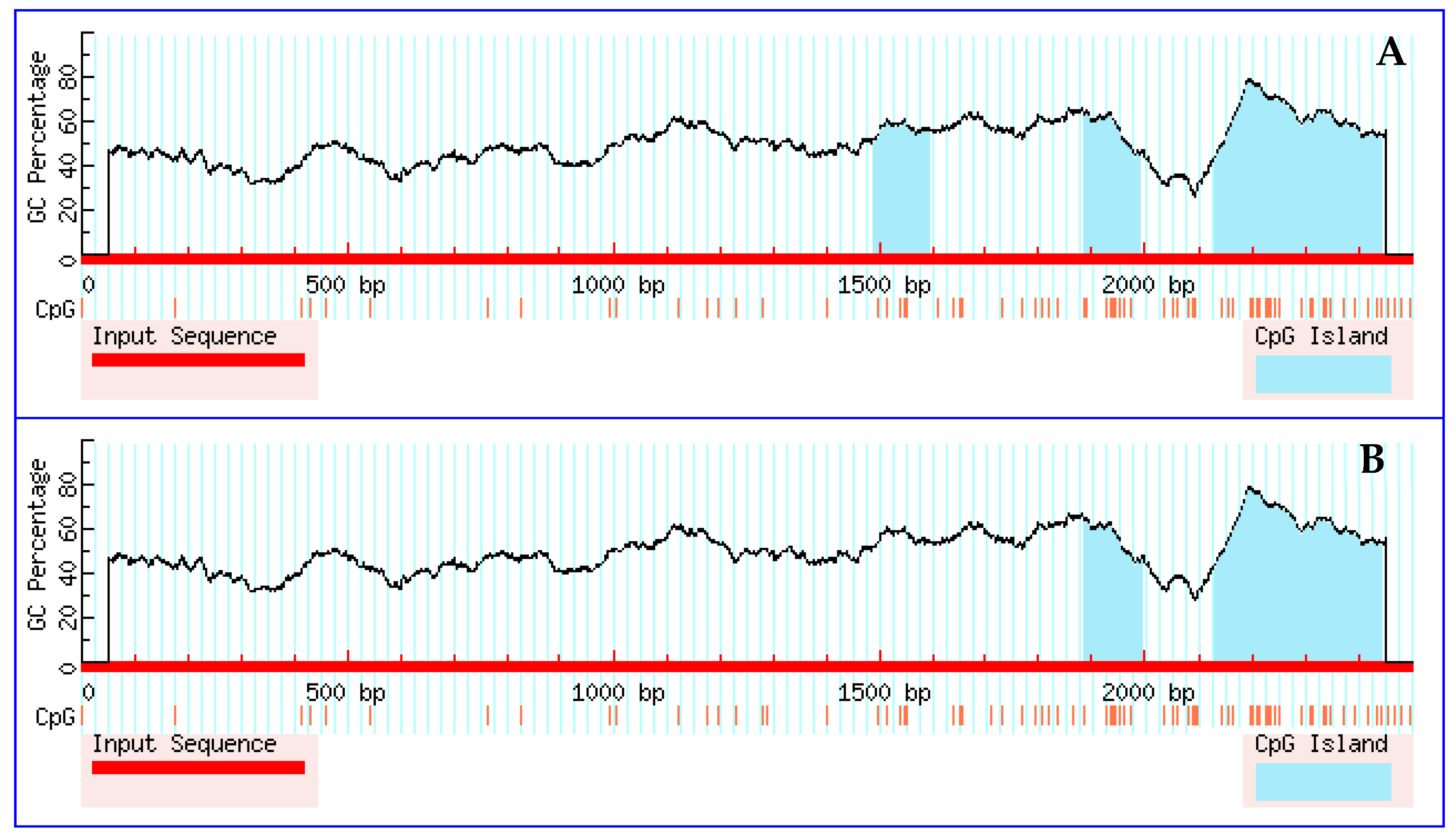
3.5. Prediction Results for CpG Islands in the Promoter Region
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Nishimichi, N.; Kawashima, T.; Hojyo, S.; Horiuchi, H.; Furusawa, S.; Matsuda, H. Characterization and expression analysis of a chicken interleukin-6 receptor alpha. Dev. Comp. Immunol. 2006, 30, 419–429. [Google Scholar] [CrossRef] [PubMed]
- Amrani, D.L.; Mauzy-Melitz, D.; Mosesson, M.W. Effect of hepatocyte-stimulating factor and glucocorticoids on plasma fibronectin levels. Biochem. J. 1986, 238, 365–371. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Schneider, K.; Klaas, R.; Kaspers, B.; Staeheli, P. Chicken interleukin-6. cDNA structure and biological properties. Eur. J. Biochem. 2001, 268, 4200–4206. [Google Scholar] [CrossRef] [PubMed]
- Horiuchi, H.; Tategaki, A.; Yamashita, Y.; Hisamatsu, H.; Ogawa, M.; Noguchi, T.; Aosasa, M.; Kawashima, T.; Akita, S.; Nishimichi, N.; et al. Chicken leukemia inhibitory factor maintains chicken embryonic stem cells in the undifferentiated state. J. Biol. Chem. 2004, 279, 24514–24520. [Google Scholar] [CrossRef] [PubMed]
- Sun, J.H.; Yan, Y.X.; Jiang, J.; Lu, P. DNA immunization against very virulent infectious bursal disease virus with VP2-4-3 gene and chicken IL-6 gene. J. Vet. Med. Ser. B 2005, 52, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Neurath, M.F.; Finotto, S. Il-6 signaling in autoimmunity, chronic inflammation and inflammation-associated cancer. Cytokine Growth Factor Rev. 2011, 22, 83–89. [Google Scholar] [CrossRef] [PubMed]
- Wu, M.; Gao, R.; Meng, M.; Li, J.; Tan, M.; Shen, Y.; Wang, L.; Yin, X.; Wu, X.; Xie, H.; et al. Regulating effects of porcine interleukin-6 gene and CpG motifs on immune responses to porcine trivalent vaccines in mice. Res. Vet. Sci. 2004, 77, 49–57. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.-H.; Tian, X.-S.; Guo, Y.; Zhou, F.-Z.; Meng, M.-J. Effect of transgenic expression of porcine interleukin-6 gene and CpG sequences on immune responses of newborn piglets inoculated with pseudorabies attenuated vaccine. Res. Vet. Sci. 2006, 80, 281–286. [Google Scholar] [CrossRef]
- Hui, G.; Hashimoto, C. Interleukin-6 has differential influence on the ability of adjuvant formulations to potentiate antibody reponses to a Plasmodium falciparum blood-stage vaccine. Vaccine 2007, 25, 6598–6603. [Google Scholar] [CrossRef] [PubMed]
- Su, B.; Wang, J.; Wang, X.; Jin, H.; Zhao, G.; Ding, Z.; Kang, Y.; Wang, B. The effects of IL-6 and TNF-α as molecular adjuvants on immune responses to FMDV and maturation of dendritic cells by DNA vaccination. Vaccine 2008, 26, 5111–5122. [Google Scholar] [CrossRef] [PubMed]
- Swaggerty, C.L.; Pevzner, I.Y.; Kogut, M.H. Selection for pro-inflammatory mediators produces chickens more resistant to Eimeria tenella. Poult. Sci. 2015, 94, 37–42. [Google Scholar] [CrossRef] [PubMed]
- Lin, Y.X. RNA Sequencing Analysis of Chicken Cecum Tissues Following E. tenella Infection and Coccidiosis Evaluation of Jinghai Yellow Chicken Cross-Breeding System Parents. Master’s Thesis, Yangzhou University, Yangzhou, China, 2015. [Google Scholar] [CrossRef]
- Liu, W.; Kong, Z.; Li, H.; Liu, Z.; Yang, S.; Huang, Y.; Zhang, X.; Zhang, L.; Wu, J. Comparison of host innate immune response of chicken (Gallus gallus) and duck (Anas platyrhynchos) infected with duck-origin newcastle disease virus. J. Agric. Biotechnol. 2017, 25, 307–315. [Google Scholar] [CrossRef]
- Tamura, K.; Stecher, G.; Peterson, D.; Filipski, A.; Kumar, S. MEGA6: Molecular evolutionary genetics analysis version 6.0. Mol. Biol. Evol. 2013, 30, 2725–2729. [Google Scholar] [CrossRef] [PubMed]
- Reddy, S.P.M.; Vuong, H.; Adiseshaiah, P. Interplay between proximal and distal promoter elements is required for squamous differentiation marker induction in the bronchial epithelium: Role for ESE-1, Sp1, and AP-1 proteins. J. Biol. Chem. 2003, 278, 21378–21387. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Hannenhalli, S. A mammalian promoter model links cis elements to genetic networks. Biochem. Biophys. Res. Commun. 2006, 347, 166–177. [Google Scholar] [CrossRef] [PubMed]
- Aneja, K.K.; Guha, P.; Shilpi, R.Y.; Chakraborty, S.; Schramm, L.M.; Haldar, D. The presence of distal and proximal promoters for rat mitochondrial glycerol-3-phosphate acyltransferase. Arch. Biochem. Biophys. 2008, 470, 35–43. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lee, T.I.; Young, R.A. Transcription of eukaryotic protein-coding genes. Annu. Rev. Genet. 2000, 34, 77–137. [Google Scholar] [CrossRef] [PubMed]
- Dan, S.P. Predicting Pol II promoter sequences using transcription factor binding sites. J. Mol. Bio. 1995, 249, 923–932. [Google Scholar] [CrossRef]
- Reese, M.G. Application of a time-delay neural network to promoter annotation in the Drosophila melanogaster genome. Comput. Chem. 2001, 26, 51–56. [Google Scholar] [CrossRef]
- Knudsen, S. Promoter2.0: For the recognition of PolII promoter sequences. Bioinformatics 1999, 15, 356–361. [Google Scholar] [CrossRef] [PubMed]
- Grabe, N. AliBaba2: Context specific identification of transcription factor binding sites. In Silico Biol. 2002, 2, S1–S15. [Google Scholar] [PubMed]
- McKeon, C.; Ohkubo, H.; Pastan, I.; de Crombrugghe, B. Unusual methylation pattern of the α2 (I) collagen gene. Cell 1982, 29, 203–210. [Google Scholar] [CrossRef]
- Li, L.C.; Dahiya, R. MethPrimer: Designing primers for methylation PCRs. Bioinformatics 2002, 18, 1427–1431. [Google Scholar] [CrossRef]
- Wigginton, J.E.; Cutler, D.J.; Abecasis, G.R. A note on exact tests of Hardy-Weinberg equilibrium. Am. J. Hum. Genet. 2005, 76, 887–893. [Google Scholar] [CrossRef] [PubMed]
- Sun, W.; Chen, H.; Lei, C.; Lei, X.; Zhang, Y. Study on population genetic characteristics of Qinchuan cows using microsatellite markers. J. Genet. Genom. 2007, 34, 17–25. [Google Scholar] [CrossRef]
- Sodhi, S.S.; Jeong, D.K.; Sharma, N.; Lee, J.H.; Kim, J.H.; Kim, S.H.; Kim, S.W.; Oh, S.J. Marker assisted selection-applications and evaluation for commercial poultry breeding. Korean J. Poult. Sci. 2013, 40, 223–234. [Google Scholar] [CrossRef]
- Toro, M.A.; Meuwissen, T.H.; Fernández, J.; Shaat, I.; Mäkitanila, A. Assessing the genetic diversity in small farm animal populations. Animal 2011, 5, 1669–1683. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cui, J.X.; Du, H.L.; Liang, Y.; Deng, X.M.; Li, N.; Zhang, X.Q. Association of polymorphisms in the promoter region of chicken prolactin with egg production. Poult. Sci. 2006, 85, 26–31. [Google Scholar] [CrossRef] [PubMed]
- Qiu, X.; Li, N.; Deng, X.; Zhao, X.; Meng, Q.; Wang, X. The single-nucleotide polymorphisms of chicken melanocortin-4 receptor (MC4R) gene and their association analysis with carcass traits. Sci. China Ser. C Life Sci. 2006, 49, 560–566. [Google Scholar] [CrossRef]
- Xu, M.; Sharma, P.; Pan, S.; Malik, S.; Roeder, R.G.; Martinez, E. Core promoter-selective function of HMGA1 and mediator in initiator-dependent transcription. Genes Dev. 2011, 25, 2513–2524. [Google Scholar] [CrossRef] [PubMed]
- Yan, Q.; Masson, R.; Pritz, M.M.; Rosati, B.; McKinnon, D. Evolution of KChIP2 gene function is localized within the core promoter and 5’ UTR. Biophys. J. 2011, 100, 573a. [Google Scholar] [CrossRef]
- Shir-Shapira, H.; Sharabany, J.; Filderman, M.; Ideses, D.; Ovadia-Shochat, A.; Mannervik, M.; Juven-Gershon, T. Structure-function analysis of the Drosophila melanogaster caudal transcription factor provides insights into core promoter-preferential activation. J. Biol. Chem. 2015, 290, 20747. [Google Scholar] [CrossRef] [PubMed]
- Kogut, M.H.; Genovese, K.J.; Haiqi, H.; Kaiser, P. Flagellin and lipopolysaccharide up-regulation of IL-6 and CXCLi2 gene expression in chicken heterophils is mediated by ERK1/2-dependent activation of AP-1 and NF-κB signaling pathways. Innate Immun. 2008, 14, 213–222. [Google Scholar] [CrossRef] [PubMed]
- Zhu, W.G.; Srinivasan, K.; Dai, Z.; Duan, W.; Druhan, L.J.; Ding, H.; Yee, L.; Villalona-Calero, M.A.; Plass, C.; Otterson, G.A. Methylation of adjacent CpG sites affects Sp1/Sp3 binding and activity in the p21Cip1 promoter. Mol. Cell Biol. 2003, 23, 4056–4065. [Google Scholar] [CrossRef] [PubMed]
- Isomura, H.; Stinski, M.F.; Kudoh, A.; Daikoku, T.; Shirata, N.; Tsurumi, T. Two Sp1/Sp3 binding sites in the major immediate-early proximal enhancer of human cytomegalovirus have a significant role in viral replication. J. Virol. 2005, 79, 9597–9607. [Google Scholar] [CrossRef] [PubMed]
- Tamura, G.; Yin, J.; Wang, S.; Fleisher, A.; Zou, T.; Abraham, J.; Kong, D.; Smolinski, K.; Wilson, K.; James, S.; et al. E-cadherin gene promoter hypermethylation in primary human gastric carcinomas. J. Natl. Cancer Inst. 2000, 92, 569–573. [Google Scholar] [CrossRef] [PubMed]
- Bird, A.P. CpG-rich islands and the function of DNA methylation. Nature 1986, 321, 209–213. [Google Scholar] [CrossRef] [PubMed]
- Yu, G.; Zhang, X.; Wang, H.; Rui, D.; Yin, A.; Qiu, G.; He, Y. CpG island methylation status in the EpCAM promoter region and gene expression. Oncol. Rep. 2008, 20, 1061–1067. [Google Scholar] [CrossRef] [PubMed]
- Macleod, D.; Charlton, J.; Mullins, J.; Bird, A.P. Sp1 sites in the mouse aprt gene promoter are required to prevent methylation of the CpG island. Genes Dev. 1994, 8, 2282–2292. [Google Scholar] [CrossRef] [PubMed]
- Uhm, T.G.; Lee, S.K.; Kim, B.S.; Kang, J.H.; Park, C.-S.; Rhim, T.Y.; Chang, H.S.; Kim, D.-J.; Chung, I.Y. CpG methylation at GATA elements in the regulatory region of CCR3 positively correlates with CCR3 transcription. Exp. Mol. Med. 2012, 44, 268–280. [Google Scholar] [CrossRef] [PubMed]
| Primer | Starting Position | Primer Sequence (5′ → 3′) | Annealing Temperature (°C) | Length (bp) |
|---|---|---|---|---|
| P1 | −2210 | F: AGAGAGGACTAACCCACAGAG | 58.5 | 698 |
| R: CCAGCTTCTCCAGTCTTGTC | ||||
| P2 | −1628 | F: AGGGACAGCAATGGCAGAAG | 60.5 | 717 |
| R: AAGAGCTGATCCTGGTTCTGG | ||||
| P3 | −994 | F: CAGAGGACGTCCTACCTCAA | 56.5 | 690 |
| R: GGTGAGCCTGGCAGCC | ||||
| P4 | −367 | F: AAGATAAGACGCGCCACACC | 58.5 | 803 |
| R: TTGAGGTTGTTCCGGACGAG | ||||
| P5 | 391 | F: CGTGTGCGAGAACAGCAT | 59.0 | 565 |
| R: AAATAGAAAGTTGGAAGGAGAGTACA | ||||
| P6 | 797 | F: GGTGTGCAGTCGGCAATAG | 57.5 | 803 |
| R: CTCATCATTCCACACAAGG | ||||
| P7 | 1388 | F: ACTGTGTGCTCCTAATGCCT | 60.5 | 770 |
| R: CTTCAGATTGGCGAGGAGGG | ||||
| P8 | 1989 | F: CGTACCTCAAAACCTACCTCAG | 59.0 | 768 |
| R: AGTGGAGTTCTTCAGCCTTATTTAA | ||||
| P9 | 2702 | F: CAAAACCAACCTGTCTAAGCTG | 59.5 | 773 |
| R: TTTTAAGAACTGTATGTTGATTGTGC |
| Mutation Site 1 | Chromosome Position | Serial Number | SNP Locus |
|---|---|---|---|
| 1 | 30947851 | rs315131240 | −1534 (T > C) |
| 2 | 30948105 | rs731482590 | −1280 (C > T) |
| 3 | 30948162 | rs15937151 | −1223 (G > A) |
| 4 | 30948165 | rs15937152 | −1220 (G > T) |
| 5 | 30948177 | rs15937153 | −1208 (A > G) |
| 6 | 30948446 | rs316873841 | −939 (C > G) |
| 7 | 30948496 | rs316713313 | −889 (G > A) |
| 8 | 30948520 | rs15937155 | −865 (G > A) |
| 9 | 30948594 | rs732353671 | −791 (G > C) |
| 10 | 30948651 | rs732784937 | −734 (G > A) |
| 11 | 30948722 | −663 (A > G) | |
| 12 | 30948751 | rs317547243 | −634 (A > G) |
| 13 | 30948775 | rs317511948 | −610 (G > A) |
| 14 | 30948874 | rs740913355 | −511 (C > T) |
| 15 | 30948902 | rs314710056 | −483 (A > G) |
| 16 | 30948927 | rs794072580 | −458 (T > G) |
| 17 | 30948938 | −447 (C > G) | |
| 18 | 30948985 | rs317677898 | −400 (A > G) |
| 19 | 30949028 | −357 (G > A) | |
| 20 | 30951133 | rs737499962 | 1749 (C > T) |
| 21 | 30951150 | rs736574918 | 1766 (G > A) |
| 22 | 30951621 | rs315658915 | 2237 (A > G) |
| 23 | 30951826 | rs794520257 | 2442 (T > C) |
| 24 | 30952410 | rs317567286 | 3026 (G > A) |
| 25 | 30952457 | rs794365857 | 3073 (C > T) |
| 26 | 30952561 | 3177 (C > T) | |
| 27 | 30952578 | rs316333207 | 3194 (C > T) |
| 28 | 30952600 | rs740039941 | 3216 (C > G) |
| No | Mutation Site | Sample Size | Genotype Frequency 1 | Gene Frequency | χ2 | p-Value | PIC | H | Ne | |||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | A-663G | 220 | AA: 0.000 (0) | GG: 1.000 (220) | AG: 0.000 (0) | A: 0.000 | G: 1.000 | 0.000 | 0.000 | 1.000 | ||
| 2 | C-447G | 220 | CC: 0.455 (100) | GG: 0.090 (20) | GC: 0.455 (100) | C: 0.680 | G: 0.320 | 0.500 | 0.799 | 0.340 | 0.434 | 1.766 |
| 3 | G-357A | 220 | GG: 0.290 (64) | AA: 0.255 (56) | GA: 0.455 (100) | G: 0.518 | A: 0.482 | 1.770 | 0.413 | 0.375 | 0.499 | 1.997 |
| 4 | C3177T | 220 | CC: 0.400 (88) | TT: 0.145 (32) | CT: 0.455 (100) | C: 0.627 | T: 0.373 | 0.170 | 0.918 | 0.358 | 0.468 | 1.878 |
| Prediction Software 1 | Start Site (bp) | Termination Site (bp) | Score |
|---|---|---|---|
| Promoter Scan [19] | −402 | −652 | 73.12 |
| −74 | −324 | 74.33 | |
| Promoter 2.0 Prediction Server [21] | −900 | - | 1.092 |
| Neural Network Promoter Prediction (NNPP version 2.2) [20] | −330 | −380 | 0.87 |
| −339 | −389 | 0.97 | |
| −519 | −569 | 0.99 | |
| −659 | −709 | 0.99 | |
| −1049 | −1099 | 0.92 | |
| −1192 | −1242 | 0.97 | |
| −1404 | −1454 | 0.91 | |
| −2179 | −2229 | 0.87 |
| Mutation Site | Base Group | Transcription Factor | Transcription Factor Binding Site Base Sequence | Transcription Factor Position |
|---|---|---|---|---|
| −357 | G | Sp1 | gacacgccacacc | −360–(−348) |
| A | ||||
| −447 | C | HB | cttaaaaacg | −447–(−438) |
| G | C/EBPα | aaacggttaa | −452–(−443) | |
| −458 | T | MEB-1 GLO | ttttttaaac ttttttaaac | −458(−449) −458–(−449) |
| G | Sp1 | ccgaggaggg | −467–(−458) | |
| −483 | A | PU.1 GATA-1 C/EBPα | aaaagagaac aaatgaaaag acaaatgaaa | −483–(−474) −488–(−479) −490–(−481) |
| G | PU.1 ISGF-3 TEC1 IRF-1 | gaaagagaac ggaaagagaa ggaaagagaa ggaaagagaa | −483–(−474) −484–(−475) −484–(−475) −484–(−475) | |
| −511 | C | TBP | tgcttataag | −513–(−504) |
| T | TBP ICS BP | atgtttataa agtttcatgt | −514–(−505) −520–(−511) | |
| −610 | G | Sp1 | ggagcgatgg | −615–(−606) |
| A | ||||
| −634 | A | GR Sp1 | cactgtgtgc cctccaccact | −635–(−626) −642–(−632) |
| G | ||||
| −663 | A | |||
| G | AP-2α Sp1 Sp1 | cccggggagc gccccggggag cctgctgccccggg | −668–(−659) −670–(−660) −676–(−663) | |
| −1208 | A | Sp1 | ggctcggggagccag | −1221–(−1207) |
| G | Sp1 | ggggagccgggc | −1216–(−1205) | |
| −1220 | G | Sp1 C/EBPβ | ggctcggggagccag gctgtggctc | −1221–(−1207) −1226–(−1217) |
| T | ||||
| −1223 | G | C/EBPβ C/EBPα | gctgtggctc ataattgctg | −1226–(−1217) −1232–(−1223) |
| A | ||||
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Xin, S.; Wang, X.; Dai, G.; Zhang, J.; An, T.; Zou, W.; Zhang, G.; Xie, K.; Wang, J. Bioinformatics Analysis of SNPs in IL-6 Gene Promoter of Jinghai Yellow Chickens. Genes 2018, 9, 446. https://doi.org/10.3390/genes9090446
Xin S, Wang X, Dai G, Zhang J, An T, Zou W, Zhang G, Xie K, Wang J. Bioinformatics Analysis of SNPs in IL-6 Gene Promoter of Jinghai Yellow Chickens. Genes. 2018; 9(9):446. https://doi.org/10.3390/genes9090446
Chicago/Turabian StyleXin, Shijie, Xiaohui Wang, Guojun Dai, Jingjing Zhang, Tingting An, Wenbin Zou, Genxi Zhang, Kaizhou Xie, and Jinyu Wang. 2018. "Bioinformatics Analysis of SNPs in IL-6 Gene Promoter of Jinghai Yellow Chickens" Genes 9, no. 9: 446. https://doi.org/10.3390/genes9090446





