Complemented Palindromic Small RNAs First Discovered from SARS Coronavirus
Abstract
:1. Introduction
2. Materials and Methods
2.1. Datasets and Data Analysis
2.2. RNAi and Cellular Experiments
3. Results
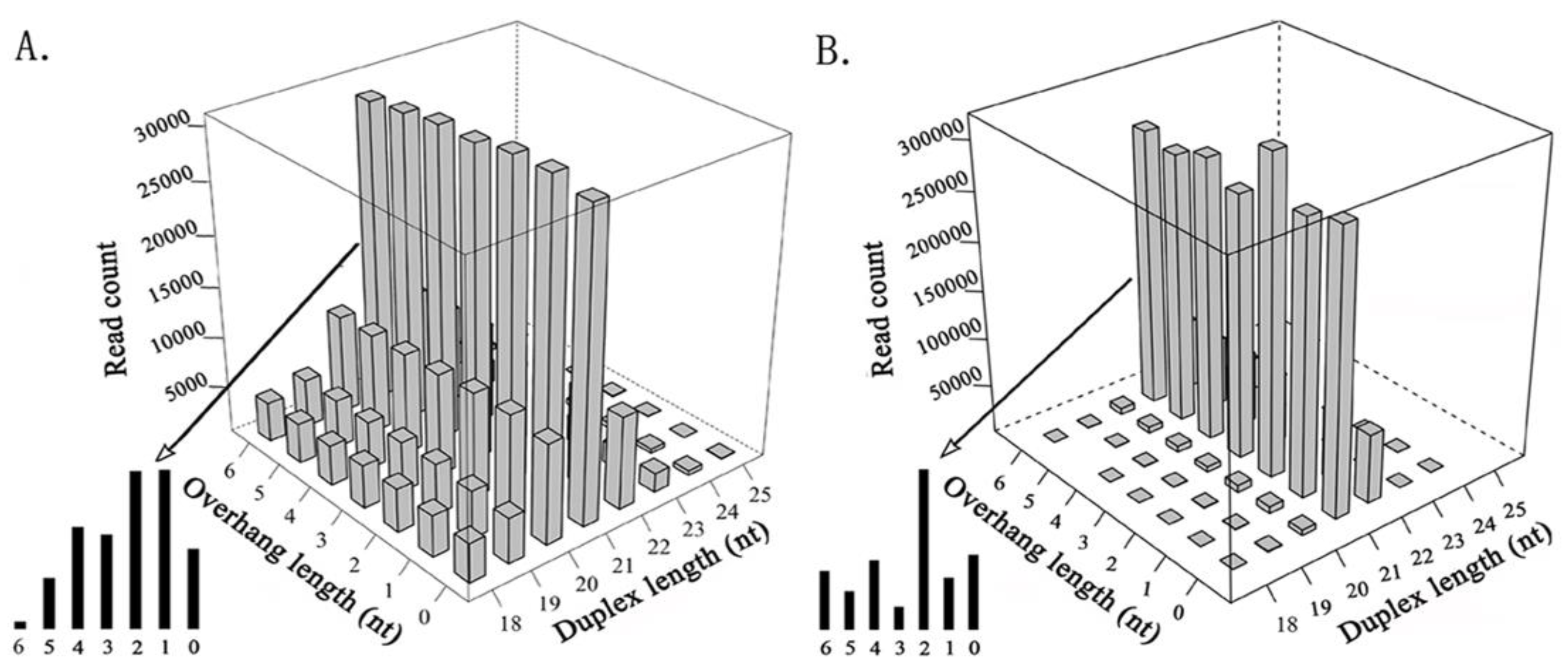
3.1. Comparison of siRNA Duplexes Induced by Plant, Invertebrate, and Mammalian Viruses
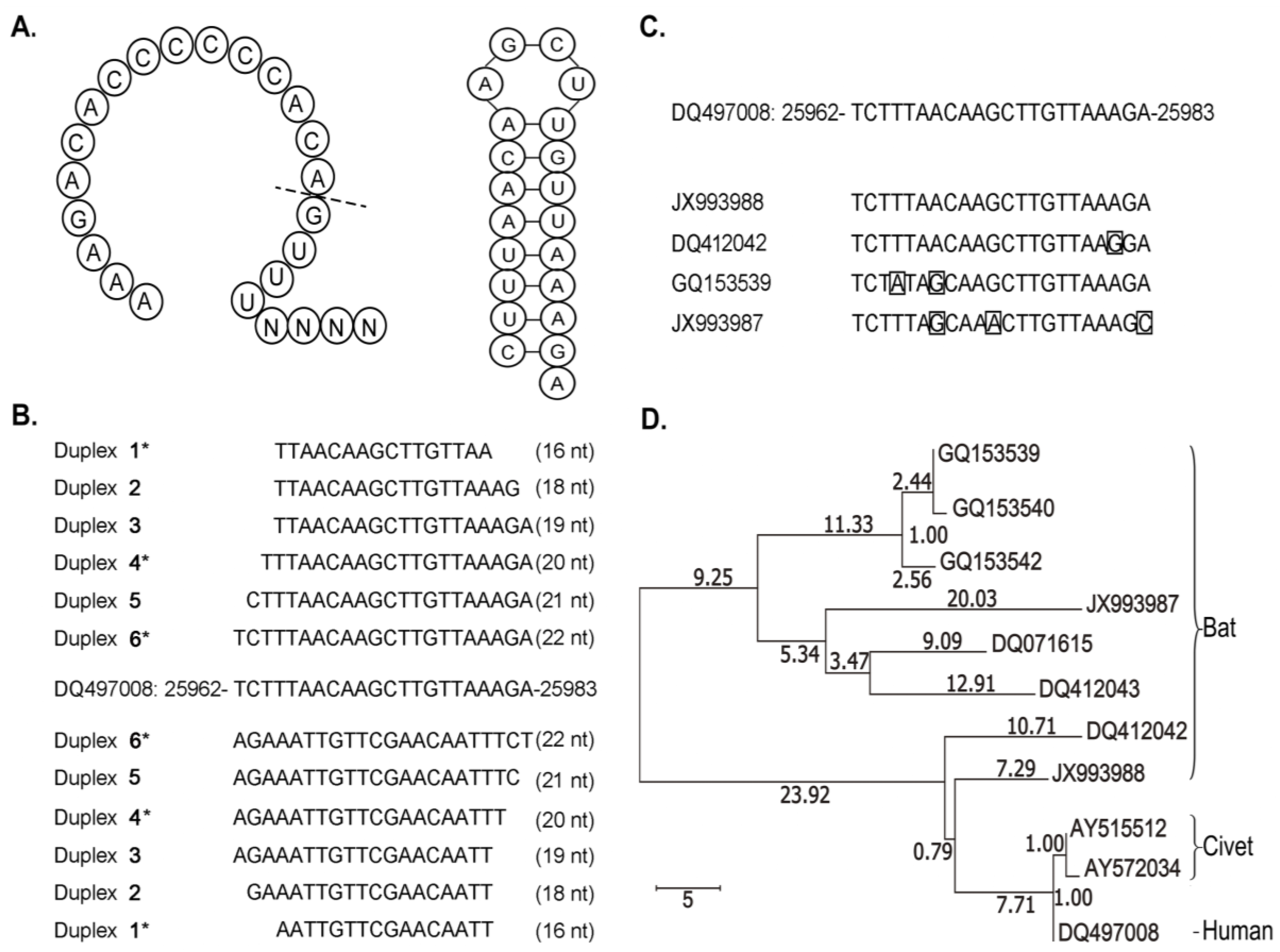
3.2. Discovery of psRNAs and cpsRNAs
3.3. Clues to Origins of SARS-CoV-cpsR-19
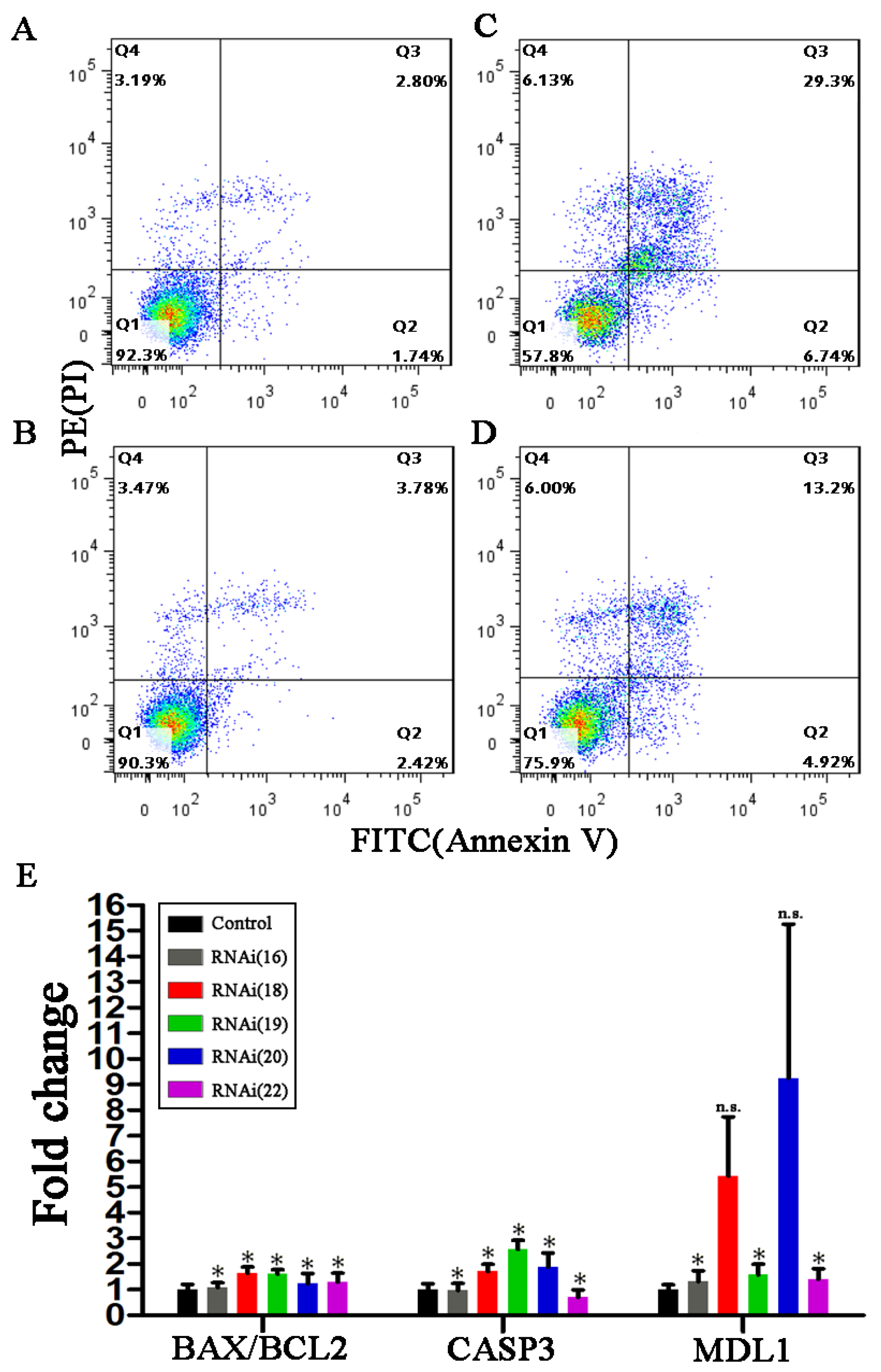
3.4. Preliminary Studies on Biological Functions of SARS-CoV-cpsR-19
4. Conclusions and Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Chen, Z.; Sun, Y.; Yang, X.; Wu, Z.; Guo, K.; Niu, X.; Wang, Q.; Ruan, J.; Bu, W.; Gao, S. Two featured series of rRNA-derived RNA fragments (rRFs) constitute a novel class of small RNAs. PLoS ONE 2017, 12, e0176458. [Google Scholar] [CrossRef] [PubMed]
- Kreuze, J.F.; Perez, A.; Untiveros, M.; Quispe, D.; Fuentes, S.; Barker, I.; Simon, R. Complete viral genome sequence and discovery of novel viruses by deep sequencing of small RNAs: A generic method for diagnosis, discovery and sequencing of viruses. Virology 2009, 388, 1–7. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, R.; Gao, S.; Hernandez, A.G.; Wechter, W.P.; Fei, Z.; Ling, K. Deep sequencing of small RNAs in tomato for virus and viroid identification and strain differentiation. PLoS ONE 2012, 7, e37127. [Google Scholar] [CrossRef] [PubMed]
- Zheng, Y.; Gao, S.; Chellappan, P.; Li, R.; Marco, G.; Dina, G.; Segundo, F.; Ling, K.; Jan, K.; Fei, Z. VirusDetect: An automated pipeline for efficient virus discovery using deep sequencing of small RNAs. Virology 2017, 500, 130–138. [Google Scholar] [CrossRef] [PubMed]
- Nayak, A.; Tassetto, M.; Kunitomi, M.; Andino, R. RNA Interference-Mediated Intrinsic Antiviral Immunity in Invertebrates; Springer: Berlin/Heidelberg, Germany, 2013; Volume 371, pp. 183–200. [Google Scholar]
- Wang, F.; Sun, Y.; Ruan, J.; Chen, R.; Chen, X.; Chen, C.; Kreuze, J.F.; Fei, Z.; Zhu, X.; Gao, S. Using small RNA deep sequencing to detect human viruses. BioMed Res. Int. 2016, 2016, 2596782. [Google Scholar] [CrossRef] [PubMed]
- Niu, X.; Sun, Y.; Chen, Z.; Li, R.; Padmanabhan, C.; Ruan, J.; Kreuze, J.F.; Ling, K.; Fei, Z.; Gao, S. Using small RNA-seq data to detect siRNA duplexes induced by plant viruses. Genes 2017, 8, 163. [Google Scholar] [CrossRef] [PubMed]
- Roberts, A.; Deming, D.; Paddock, C.D.; Cheng, A.; Yount, B.; Vogel, L.; Herman, B.D.; Sheahan, T.; Heise, M.; Genrich, G.L. A mouse-adapted SARS-coronavirus causes disease and mortality in BALB/c mice. PLoS Pathog. 2007, 3, e5. [Google Scholar] [CrossRef] [PubMed]
- Gao, S.; Tian, X.; Chang, H.; Sun, Y.; Wu, Z.; Cheng, Z.; Dong, P.; Zhao, Q.; Ruan, J.; Bu, W. Two novel lncRNAs discovered in human mitochondrial DNA using PacBio full-length transcriptome data. Mitochondrion 2017, 36. [Google Scholar] [CrossRef] [PubMed]
- Elbashir, S.M.; Harborth, J.; Weber, K.; Tuschl, T. Analysis of gene function in somatic mammalian cells using small interfering RNAs. Methods 2002, 26, 199–213. [Google Scholar] [CrossRef]
- Peng, X.; Gralinski, L.; Ferris, M.T.; Frieman, M.B.; Thomas, M.J.; Proll, S.; Korth, M.J.; Tisoncik, J.R.; Heise, M.; Luo, S. Integrative deep sequencing of the mouse lung transcriptome reveals differential expression of diverse classes of small RNAs in response to respiratory virus infection. Mbio 2011, 2, e00198-11. [Google Scholar] [CrossRef] [PubMed]
- Zhang, M.; Zhan, F.; Sun, H.; Gong, X.; Fei, Z.; Gao, S. Fastq_clean: An optimized pipeline to clean the Illumina sequencing data with quality control. In Proceedings of the 2014 IEEE International Conference on Bioinformatics and Biomedicine (BIBM), Belfast, UK, 2–5 November 2014. [Google Scholar]
- Langmead, B.; Trapnell, C.; Pop, M.; Salzberg, S.L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 2009, 10, R25. [Google Scholar] [CrossRef] [PubMed]
- Gao, S.; Ou, J.; Xiao, K. R Language and Bioconductor in Bioinformatics Applications, Chinese ed.; Tianjin Science and Technology Translation Publishing Co., Ltd.: Tianjin, China, 2014. [Google Scholar]
- Clustal Omega. Available online: http://www.ebi.ac.uk/Tools/msa/clustalo/ (accessed on 18th May 2014).
- Srivastava, S.K.; Robins, H.S. Palindromic nucleotide analysis in human T cell receptor rearrangements. PLoS ONE 2012, 7, e52250. [Google Scholar] [CrossRef] [PubMed]
- Enserink, M. Clues to the animal origins of SARS. Science 2003, 300, 1351. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Shi, Z.; Yu, M.; Ren, W.; Smith, C.; Epstein, J.H.; Wang, H.; Crameri, G.; Hu, Z.; Zhang, H. Bats are natural reservoirs of SARS-like coronaviruses. Science 2005, 310, 676–679. [Google Scholar] [CrossRef] [PubMed]
- Chew, D.S.H.; Choi, K.P.; Heidner, H.; Leung, M.Y. Palindromes in SARS and other coronaviruses. Informs J. Comput. 2004, 16, 331–340. [Google Scholar] [CrossRef] [PubMed]
- Morales, L.; Oliveros, J.C.; Fernandez-Delgado, R.; Tenoever, B.R.; Enjuanes, L.; Sola, I. SARS-CoV-encoded small RNAs contribute to infection-associated lung pathology. Cell Host Microbe 2017, 21, 344–355. [Google Scholar] [CrossRef] [PubMed]
| Gene Symbol | Forward Primer | Reverse Primer |
|---|---|---|
| GAPDH | ACATCGCTCAGACACCATG | TGTAGTTGAGGTCAATGAAGGG |
| BAX | AGTAACATGGAGCTGCAGAG | AGTAGAAAAGGGCGACAACC |
| BCL2 | GTGGATGACTGAGTACCTGAAC | GCCAGGAGAAATCAAACAGAGG |
| CASP3 | ACTGGACTGTGGCATTGAG | GAGCCATCCTTTGAATTTCGC |
| MDL1 * | CCCAATCCACATCAAAACCC | GGACGAGAAGGGATTTGACTG |
| DNA Complemented Palindrome | Start | End | Length | GC% | Tm | MFE |
|---|---|---|---|---|---|---|
| GGTAACTATAAAGTTACC | 1783 | 1800 | 18 | 33 | 20 | −6.9 |
| AATGTGAGAATCACATT | 2779 | 2795 | 17 | 29 | 18 | −4.4 |
| AAGAAACTAAGTTTCTT | 3923 | 3939 | 17 | 24 | 18 | −3.5 |
| ATGGTAAGCTTTACCAT | 3971 | 3987 | 17 | 35 | 18 | −4.5 |
| AAATGCAAATCTGCATTT | 4234 | 4251 | 18 | 28 | 18 | −4.9 |
| ATATGTCTATGACATAT | 4949 | 4965 | 17 | 24 | 18 | −4.3 |
| CCTCATGTAAATCATGAGG | 5020 | 5038 | 19 | 42 | 22 | −9.0 |
| ATAACAATTGTTAT | 5207 | 5220 | 14 | 14 | 12 | −0.9 |
| ACTTCAACAGCTTGAAGT | 5241 | 5258 | 18 | 39 | 18 | −4.7 |
| ACTTCAAATTCATTTGAAGT | 6256 | 6275 | 20 | 25 | 20 | −5.5 |
| GTACTTTTACTAAAAGTAC | 6734 | 6752 | 19 | 26 | 20 | −4.6 |
| ATCTACCAGTGGTAGAT | 9189 | 9205 | 17 | 41 | 20 | −6.5 |
| TTACCTTCCAAGGTAA | 10,892 | 10,907 | 16 | 38 | 16 | −4.0 |
| CCACTTATTAAGTGG | 14,160 | 14,174 | 15 | 40 | 18 | −4.7 |
| CCCATTTAATAAATGGG | 14,882 | 14,898 | 17 | 35 | 20 | −5.7 |
| CAGTGACAATGTCACTG | 16,463 | 16,479 | 17 | 47 | 22 | −7.7 |
| CACCTTTGAAAAAGGTG | 16,760 | 16,776 | 17 | 41 | 20 | −5.6 |
| TGTAAGAGAATTTCTTACA | 17,651 | 17,669 | 19 | 26 | 18 | −5.3 |
| TGAATATGACTATGTCATATTCA | 17,783 | 17,805 | 23 | 26 | 26 | −9.7 |
| CTACTTTAAGAAAGTAG | 20,081 | 20,097 | 17 | 29 | 18 | −3.6 |
| AGCATTCTTGGAATGCT | 21,106 | 21,122 | 17 | 41 | 20 | −6.1 |
| TTCCTCTTAAATTAAGAGGAA | 21,337 | 21,357 | 21 | 29 | 22 | −9.1 |
| GCATTACTACAGAAGTAATGC | 23,599 | 23,619 | 21 | 38 | 22 | −7.9 |
| AGCCCTTTATAAGGGCT | 25,480 | 25,496 | 17 | 47 | 22 | −8.7 |
| TCTTTAACAAGCTTGTTAAAGA * | 25,962 | 25,983 | 22 | 27 | 20 | −7.2 |
| CAACGGTACTATTACCGTTG | 26,406 | 26,425 | 20 | 45 | 24 | −9.0 |
| ACCTTCATGAAGGT | 28,048 | 28,061 | 14 | 43 | 14 | −2.6 |
| GCAAATTGCACAATTTGC * | 29,028 | 29,045 | 18 | 39 | 18 | −4.0 |
| TAAAATTAATTTTA | 29,668 | 29,681 | 14 | 0 | NA | NA |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, C.; Chen, Z.; Hu, Y.; Ji, H.; Yu, D.; Shen, W.; Li, S.; Ruan, J.; Bu, W.; Gao, S. Complemented Palindromic Small RNAs First Discovered from SARS Coronavirus. Genes 2018, 9, 442. https://doi.org/10.3390/genes9090442
Liu C, Chen Z, Hu Y, Ji H, Yu D, Shen W, Li S, Ruan J, Bu W, Gao S. Complemented Palindromic Small RNAs First Discovered from SARS Coronavirus. Genes. 2018; 9(9):442. https://doi.org/10.3390/genes9090442
Chicago/Turabian StyleLiu, Chang, Ze Chen, Yue Hu, Haishuo Ji, Deshui Yu, Wenyuan Shen, Siyu Li, Jishou Ruan, Wenjun Bu, and Shan Gao. 2018. "Complemented Palindromic Small RNAs First Discovered from SARS Coronavirus" Genes 9, no. 9: 442. https://doi.org/10.3390/genes9090442





