Vertebrate Genome Evolution in the Light of Fish Cytogenomics and rDNAomics
Abstract
1. Introduction
1.1. Introduction into Cytogenomics
1.2. How Can This Review Been Utilized?
1.3. The Complex Evolution of Vertebrates Began with Fish
2. The Importance of Fish Cytogenomic Research Demonstrated on Their Role in Medicine
2.1. Why Is the Zebrafish the Most Important Fish Model?
2.2. Other Fish Species Used as Medical Models Have Genomes Suited for Specific Human Diseases
2.3. Amazon Molly (Poecilia formosa) as a Cytogenomic and Epigenetic Model Species in Human Health Research
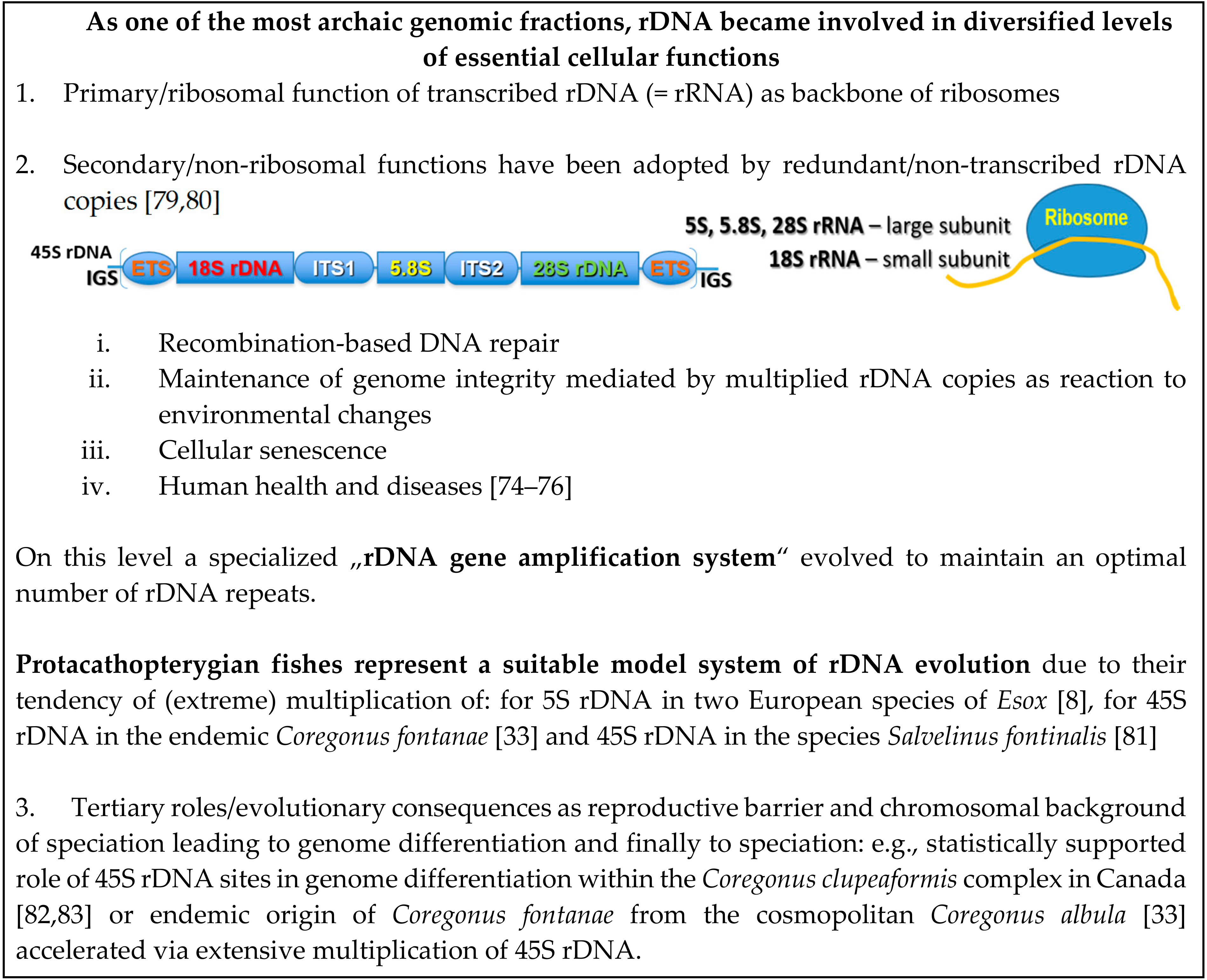
3. rDNAomics—Where Fish Ecological Cytogenomics Meets Human Cancer Genomics
4. State-of-the-Art in Fish (Cyto)Genomics—The Starting Point
5. Fish Cytogenomics—Practical Application of Integrated Cytogenetics and Genomics
5.1. Assignment of Linkage Groups to Specific Chromosomes Using FISH with Sequenced BACs
5.2. Quick Qualitative Analysis and Visualization of AT- vs. GC-Rich Chromosomal Regions
5.3. Cytogenetic Mapping of Repetitive Sequences on Conspecific Populations
5.4. Cytogenomics of Duplicated Genomes—Understanding Mechanisms of Genome Evolution in Vertebrates Which Have Undergone Whole-Genome Duplication(s)
6. Fish Cytogenetics and Cytogenomics in the Era of Digitalized Biology—The Boom of Databases
6.1. Databases for Fish Biology
6.2. History of the Zebrafish Reference Genome
6.3. Fish Genomes Available
6.4. Future of Fish Cytogenomics = Phylogenomics
7. Quantitative Eco-Evo Cytogenomics
8. Sex and B Chromosomes, Nuclear Architecture and Genome Evolution Research
8.1. Sex Chromosomes in Fish
8.2. B Chromosomes (Not Only) in Fish
8.3. Nuclear Architecture
9. Roots of (Population) rDNAomics in Fish Cytogenetics—rDNAomics as Another Dimension of Environmental Genomics
10. Conclusions
Acknowledgments
Conflicts of Interest
Appendix A. List of Fish Species with Sequenced Genomes Currently Available
| Species | Order a | Family a | Size (Mb) | C-val b | n/2n c | Assembly/Type d | GC% | Notes e |
|---|---|---|---|---|---|---|---|---|
| Acanthochromis polyacanthus | Ovalentaria | Pomacentridae | 991.585 | 0.94 | — | 1/S | 41.5 | |
| Amphilophus citrinellus | Cichliformes | Cichlasomat., Heroini | 844.903 | — | —/48 | 1/S | 41.4 | |
| Anguilla anguilla | Anguilliformes | Anguillidae | 1018.7 | 1.11–1.67 | —/38 | 1/S | 42.9 | |
| Anguilla japonica | Anguilliformes | Anguillidae | 1288.6 | 1.09 | —/38 | 1/S | 38.8 | |
| Anguilla rostrata | Anguilliformes | Anguillidae | 1413.03 | 1.01–1.66 | —/38 | 1/S | 41.0 | |
| Anoplopoma fimbria | Perciform × Scorpaeniform | Anoplomatidae | 699.326 | 0.71–0.84 | — | 1/C | 40.3 | |
| Astyanax mexicanus | Characiformes | Characidae | 1335.24 | — | 25/50 | 1/Ch | 38.4 | |
| Austrofundulus limnaeus | Cyprinodontiformes | Rivulidae | 866.963 | — | — | 1/S | 41.1 | |
| Boleophthalmus pectinirostris | Gobiiformes | Gobiid., Oxudercinae | 955.752 | — | —/46 | 1/S | 40.1 | |
| Branchiostoma belcheri | Cephalochordata | Branchiostomidae | 426.124 | — | —/36 | 2/S | 40.15 | |
| Branchiostoma floridae | Cephalochordata | Branchiostomidae | 521.895 | — | — | 1/S | 41.8 | |
| Callorhinchus milii | Chimaeriformes | Callorhinchidae | 974.499 | 1.94 | — | 1/S | 42.6 | |
| Channa argus | Perciformes | Channidae | 615.3 | 0.63–0.77 | —/48 | 1/D | — | |
| Clupea harengus | Clupeiformes | Clupeidae | 807.712 | — | —/50-52 | 1/S | 44.5 | |
| Cottus rhenanus | Perciformes | Cottidae | 563.609 | — | — | 1/S | 36.8 | |
| Cynoglossus semilaevis | Pleuronectiformes | Cynoglossidae | 470.199 | 0.62 C. bilineatus | 22 | 1/Ch | 41.27 | |
| Cyprinodon nevadensis | Cyprinodontiformes | Cyprinodontidae | 1011.85 | — | —/48 | 1/S | 39.0 | |
| Cyprinodon variegatus | Cyprinodontiformes | Cyprinodontidae | 1035.18 | 1.6 | —/48 | 1/S | 39.5 | |
| Cyprinus carpio | Cypriniformes | Cyprinidae | 1713.66 | 1.6–2.0 | 50/100 | 2/S, Ch | med 37.1 | |
| Danio rerio | Cypriniformes | Cyprinidae | 1679.2 | 1.6–2.3 | 25/48 | 4/3S, 1 Chr | med 36.7 | |
| Dicentrarchus labrax | Moroniformes | Moronidae | 675.917 | 0.78 | — | 2/S | 40.4 | |
| Eptatretus burgeri | Myxiniformes | Myxinidae/Eptatretinae | 2608.38 | 2.98 | /36 | 1/S | — | |
| Esox lucius | Esociformes | Esocidae | 904.497 | 0.8–1.4 | 25/50 | 3/Ch | 42.2 | PB |
| Fundulus heteroclitus | Cyprinodontiformes | Fundulidae | 1021.9 | 1.3–1.5 | -/48 | 1/S | 41.2 | |
| Gadus morhua | Gadiformes | Gadidae | 824.311 | 0.4–0.9 | -/46 | 2/S | 46.3 | 454, I, PB [124] |
| Gasterosteus aculeatus | Perciformes | Cottiodei, Gasterosteid. | 446.611 | 0.6–0.7 | —/42 | 2/S, Ch | 44.6 | LM [163] |
| Haplochromis burtoni | Perciformes × Cichliformes | Cichlidae (Afr.) | 831.412 | 0.97 | —/40 | 1/S | 41.9 | |
| Hippocampus comes | Syngnathiformes | Syngnathidae | 493.776 | — | — | 1/S | 43.7 | |
| Ictalurus punctatus | Siluriformes | Ictaluridae | 783.275 | 1.0 | 29/58 | 1/Ch | 39.8 | PB, I [164,165] |
| Kryptolebias marmoratus | Cyprinodontiformes | Rivulidae | 680.367 | — | —/48 | 2/S | med 39.5 | |
| Labeotropheus fuelleborni | Perciformes × Cichliformes | Cichlidae (Afr.) | 70.8584 | — | —/44 | 1/S | 42.1 | |
| Labrus bergylta | Labriformes | Labridae | 805.481 | — | — | 1/S | 40.9 | |
| Larimichthys crocea | Acanthuriformes | Sciaenidae | 678.938 | — | — | 2/S | med 41.4 | |
| Lates calcarifer | Perciformes | Centropomidae | 668.481 | 0.7 | —/48 | 2/S | med 40.6 | |
| Latimeria chalumnae | Ceolacanthiformes | Coelacanthidae | 2860.59 | 2.8–6.6 | —/48 | 2/S | med 42.5 | |
| Lepisosteus oculatus | Lepisosteiformes | Lepisosteidae | 945.878 | 1.4 | 29/58 | 1/Ch | 40.4 | |
| Lethenteron camtschaticum | Petromyzontiformes | Petromyzontidae | 1030.66 | ~1.4 | —/144-162 | 1/S | 48.1 | |
| Leuciscus waleckii | Cypriniformes | Cyprinidae | 752.539 | congeners 1.2–1.5 | —/50 | 1/S | 37.4 | |
| Leucoraja erinacea | Rajiformes | Rajidae | 1555.46 | 3.5–4.6 | — | 1/C | 40.3 | |
| Maccullochella peelii | Perciformes × | Percichthyidae | 633.241 | 0.83 | — | 1/S | — | |
| Maylandia zebra | Perciformes × Cichliformes | Cichlidae (Afr.) | 859.842 | — | — | 2/S | 41.4 | |
| Mchenga conophoros | Perciformes × Cichliformes | Cichlidae (Afr.) | 73.4256 | — | — | 1/S | 41.8 | |
| Megalobrama amblycephala | Cypriniformes | Cyprinidae, Cultrinae | 1116.0 | 1.12–1.35 | — | 1/D | 37.3 | [166] |
| Melanochromis auratus | Perciformes × Cichliformes | Cichlidae (Afr.) | 68.2386 | — | —/46 | 1/S | 41.5 | |
| Miichthys miiuy | Acanthuriformes | Sciaenidae | 619.301 | — | — | 1/S | 39.3 | |
| Mola mola | Tetraodontiformes | Molidae | 639.452 | 0.8–0.9 | —/46 | 1/S | 41.2 | [167] |
| Monopterus albus | Synbranchiformes | Synbranchidae | 684.144 | 0.6–0.9 | —/24 | 1/S | 41.5 | |
| Morone saxatilis | Moroniformes | Moronidae | 585.167 | 0.9 | —/48 | 1/S | 40.0 | |
| Neolamprologus brichardi | Perciformes × Cichliformes | Cichlidae (Afr.) | 847.91 | — | - | 1/S | 42 | |
| Nothobranchius furzeri | Cyprinodontiformes | Notobranchiidae | 1132.74 | 1.56 | 19/38 | 4/S, Chr | 43, 8 | |
| Nothobranchius kuhntae | Cyprinodontiformes | Notobranchiidae | 5.23461 | — | —/38 | 1/S | 44.8 | |
| Notothenia coriiceps | Perciformes | Nototheniidae | 636.614 | — | — | 1/S | 40.8 | |
| Oncorhynchus kisutch | Salmoniformes | Salmonidae | 2369.93 | 2.6–3.0 | 30/60 | 1/Ch | 43.6 | |
| Oncorhynchus mykiss | Salmoniformes | Salmonidae | 2179 | 1.9–2.9 | 29/60 | 2/Ch | med 43.7 | |
| Oreochromis niloticus | Perciformes × Cichliformes | Cichlidae (Afr.) | 1009.86 | 0.9–1.2 | 23/44 | 2/Ch | med 39.9 | PB, I [168,169] |
| Oryzias latipes | Beloniformes | Adrianichthyidae | 869.818 | 0.9–1.1 | 24/48 | 5/Ch | med 40.8 | |
| Pampus argenteus | Scombriformes | Stromateidae | 350.449 | — | — | 1/S | 38.2 | |
| Paralichthys olivaceus | Pleuronectiformes | Paralichthyidae | 643.911 | 0.7 | 24/46–48 | 2/S, Ch | med 42.4 | |
| Periophthalmodon schlosseri | Gobiiformes | Gobiid., Oxudercinae | 679.761 | 0.96 | — | 1/S | 40.2 | |
| Periophthalmus magnuspinnatus | Gobiiformes | Gobiid., Oxudercinae | 701.697 | — | — | 1/S | 40 | |
| Petromyzon marinus | Petromyzontiformes | Petromyzontidae | 885.535 | 1.6–2.4 | —/168 | 1/S | 46.8 | |
| Pimephales promelas | Cypriniformes | Cyprinidae | 1219.33 | 1.1 | —/50 | 2/S | med 40.6 | |
| Poecilia formosa | Cyprinodontiformes | Poeciliidae | 748.923 | 0.75–0.97 | —/46 | 1/S | 39.6 | |
| Poecilia latipinna | Cyprinodontiformes | Poeciliidae | 815.145 | 0.9–1.0 | —/46 | 1/S | 40.8 | |
| Poecilia mexicana | Cyprinodontiformes | Poeciliidae | 801.711 | 0.7–1.38 | —/46 | 1/S | 40.7 | |
| Poecilia reticulata | Cyprinodontiformes | Poeciliidae | 731.622 | 0.77–1.0 | 23/46 | 1/Ch | 40.3 | |
| Protosalanx hyalocranius | Osmeriformes | Salangidae | 525 | — | — | 1/D | — | |
| Pseudopleuronectes yokohamae | Pleuronectiformes | Pleuronectidae | 547.831 | 0.67 | —/48 | 1/C | 42 | |
| Pundamilia nyererei | Perciformes × Cichliformes | Cichlidae (Afr.) | 830.133 | — | — | 1/S | 41.9 | |
| Pygocentrus nattereri | Characiformes | Serrasalmidae | 1285.35 | — | — | 1/S | 40.6 | |
| Rhamphochromis esox | Perciformes × Cichliformes | Cichlidae (Afr.) | 71.2951 | — | — | 1/S | 42.3 | |
| Rhincodon typus | Orectolobiformes | Rhincodontidae | 2931.6 | — | — | 1/S | 41.8 | |
| Salmo salar | Salmoniformes | Salmonidae | 2966.89 | 3.0–3.3 | 29/60 | 1/Ch | 43.9 | PB, I, S [170] |
| Scartelaos histophorus | Gobiiformes | Gobiid., Oxudercinae | 695.009 | — | — | 1/S | 39.1 | |
| Scleropages formosus | Osteoglossiformes | Osteoglossidae | 777.359 | — | — | 4/S, Chr | med 43.9 | |
| Sebastes aleutianus | Scorpaeniformes | Sebastidae | 899.65 | — | — | 1/S | 40.9 | |
| Sebastes minor | Scorpaeniformes | Sebastidae | 681.653 | — | — | 1/S | 40.8 | |
| Sebastes nigrocinctus | Scorpaeniformes | Sebastidae | 746.045 | — | — | 1/S | 40.8 | |
| Sebastes rubrivinctus | Scorpaeniformes | Sebastidae | 756.297 | — | — | 1/S | 40.7 | |
| Sebastes steindachneri | Scorpaeniformes | Sebastidae | 648.011 | — | — | 1/S | 40.7 | |
| Seriola dumerili | Carangiformes | Carangidae | 677.67 | 0.74 | — | 1/S | 40.9 | |
| Seriola lalandi | Carangiformes | Carangidae | 685 | — | — | 1/S | - | PB, I |
| Seriola quinqueradiata | Carangiformes | Carangidae | med 750.45 | 0.83 | — | 2/S | 40.25 | |
| Sinocyclocheilus anshuiensis | Cypriniformes | Cyprinidae | 1632.72 | — | — | 1/S | 38 | |
| Sinocyclocheilus grahami | Cypriniformes | Cyprinidae | 1750.29 | 2.35 | —/96 | 1/S | 38.7 | |
| Sinocyclocheilus rhinocerous | Cypriniformes | Cyprinidae | 1655.79 | — | — | 1/S | 38.1 | |
| Squalius pyrenaicus | Cypriniformes | Cyprinidae | 48.1393 | 2.4 | —/50 | 1/C | 51.8 | |
| Stegastes partitus | Ovalentaria | Pomacentridae | 800.492 | — | — | 1/S | 42.1 | |
| Takifugu flavidus | Tetraodontiformes | Tetraodontidae | 378.032 | — | — | 1/S | 45.6 | |
| Takifugu rubripes | Tetraodontiformes | Tetraodontidae | 391.485 | 0.4 | 22/44 | 1/Ch | 45.8 | |
| Tetraodon nigroviridis | Tetraodontiformes | Tetraodontidae | 342.403 | 0.35–0.51 | 21/42 | 7/S, Ch | 46.6 | Genoscope |
| Thunnus orientalis | Scombriformes | Scombridae | 684.497 | — | —/48 | 1/C | 39.7 | |
| Xiphophorus couchianus | Cyprinodontiformes | Poeciliidae | 708.396 | 0.75 | — | 1/S | 40.9 | |
| Xiphophorus hellerii | Cyprinodontiformes | Poeciliidae | 733.802 | 0.7–1.0 | —/48 | 1/S | 41.2 | |
| Xiphophorus maculatus | Cyprinodontiformes | Poeciliidae | 729.664 | 0.8–1.0 | —/48 | 1/S | 39.8 |
References
- Bernheim, A. Cytogenomics of cancers: From chromosome to sequence. Mol. Oncol. 2010, 4, 309–322. [Google Scholar] [CrossRef] [PubMed]
- Xiang, B.; Leon, A.; Li, M.M.; Iqbal, A.M.; Li, P.; Li, S.; Papenhausen, P.R.; Schwartz, S.; Zhang, X.-X.; Geiersbach, K.B.; et al. Atlas of Cytogenomics in Oncology and Hematology: A Platform-Neutral Clinical Cancer Genomics Database. Cancer Genet. 2012, 205, 420. [Google Scholar] [CrossRef]
- McPherson, M.C.; Robinson, C.M.; Gehlen, L.P.; Delany, M.E. Comparative cytogenomics of poultry: Mapping of single gene and repeat loci in the Japanese quail (Coturnix japonica). Chromosome Res. 2014, 22, 71–83. [Google Scholar] [CrossRef] [PubMed]
- Barh, D.; Khan, M.S.; Davies, E. PlantOmics: The Omics of Plant Science; Springer: New Delhi, India, 2015; ISBN 978-81-322-2172-2. [Google Scholar]
- Kapusta, A.; Suh, A. Evolution of bird genomes-a transposon’s-eye view. Ann. N. Y. Acad. Sci. 2017, 1389, 164–185. [Google Scholar] [CrossRef] [PubMed]
- Nakajima, R.T.; Cabral-de-Mello, D.C.; Valente, G.T.; Venere, P.C.; Martins, C. Evolutionary dynamics of rRNA gene clusters in cichlid fish. BMC Evol. Biol. 2012, 12, 198. [Google Scholar] [CrossRef] [PubMed]
- Symonová, R.; Majtánová, Z.; Arias-Rodriguez, L.; Mořkovský, L.; Kořínková, T.; Cavin, L.; Pokorná, M.J.; Doležálková, M.; Flajšhans, M.; Normandeau, E.; et al. Genome Compositional Organization in Gars Shows More Similarities to Mammals than to Other Ray-Finned Fish: Cytogenomics of Gars. J. Exp. Zool. B Mol. Dev. Evol. 2017, 328, 607–619. [Google Scholar] [CrossRef] [PubMed]
- Symonová, R.; Ocalewicz, K.; Kirtiklis, L.; Delmastro, G.B.; Pelikánová, Š.; Garcia, S.; Kovařík, A. Higher-order organisation of extremely amplified, potentially functional and massively methylated 5S rDNA in European pikes (Esox sp.). BMC Genom. 2017, 18, 391. [Google Scholar] [CrossRef] [PubMed]
- Cioffi, M.B.; Bertollo, L.A.C. Chromosomal distribution and evolution of repetitive DNAs in fish. Genome Dyn. 2012, 7, 197–221. [Google Scholar] [CrossRef] [PubMed]
- Betancur-R, R.; Wiley, E.O.; Arratia, G.; Acero, A.; Bailly, N.; Miya, M.; Lecointre, G.; Ortí, G. Phylogenetic classification of bony fishes. BMC Evol. Biol. 2017, 17, 162. [Google Scholar] [CrossRef] [PubMed]
- Saha, N.R.; Smith, J.; Amemiya, C.T. Evolution of adaptive immune recognition in jawless vertebrates. Semin. Immunol. 2010, 22, 25–33. [Google Scholar] [CrossRef] [PubMed]
- Gregory, T.R. Animal Genome Size Database. Available online: http://genomesize.com (accessed on 9 November 2017).
- Caputo Barucchi, V.; Giovannotti, M.; Nisi Cerioni, P.; Splendiani, A. Genome Duplication in Early Vertebrates: Insights from Agnathan Cytogenetics. Cytogenet. Genome Res. 2013, 141, 80–89. [Google Scholar] [CrossRef] [PubMed]
- Smith, J.J.; Saha, N.R.; Amemiya, C.T. Genome biology of the cyclostomes and insights into the evolutionary biology of vertebrate genomes. Integr. Comp. Biol. 2010, 50, 130–137. [Google Scholar] [CrossRef] [PubMed]
- Stingo, V.; Rocco, L. Selachian cytogenetics: A review. Genetica 2001, 111, 329–347. [Google Scholar] [CrossRef] [PubMed]
- Rock, J.; Eldridge, M.; Champion, A.; Johnston, P.; Joss, J. Karyotype and nuclear DNA content of the Australian lungfish, Neoceratodus forsteri (Ceratodidae: Dipnoi). Cytogenet. Cell Genet. 1996, 73, 187–189. [Google Scholar] [CrossRef] [PubMed]
- Koch, J.; Lüdemann, J.; Spies, R.; Last, M.; Amemiya, C.T.; Burmester, T. Unusual Diversity of Myoglobin Genes in the Lungfish. Mol. Biol. Evol. 2016, 33, 3033–3041. [Google Scholar] [CrossRef] [PubMed]
- Bogart, J.P.; Balon, E.K.; Bruton, M.N. The chromosomes of the living coelacanth and their remarkable similarity to those of one of the most ancient frogs. J. Hered. 1994, 85, 322–325. [Google Scholar] [CrossRef] [PubMed]
- Andreyushkova, D.A.; Makunin, A.I.; Beklemisheva, V.R.; Romanenko, S.A.; Druzhkova, A.S.; Biltueva, L.B.; Serdyukova, N.A.; Graphodatsky, A.S.; Trifonov, V.A. Next Generation Sequencing of Chromosome-Specific Libraries Sheds Light on Genome Evolution in Paleotetraploid Sterlet (Acipenser ruthenus). Genes 2017, 8, 318. [Google Scholar] [CrossRef] [PubMed]
- Romanenko, S.A.; Biltueva, L.S.; Serdyukova, N.A.; Kulemzina, A.I.; Beklemisheva, V.R.; Gladkikh, O.L.; Lemskaya, N.A.; Interesova, E.A.; Korentovich, M.A.; Vorobieva, N.V.; et al. Segmental paleotetraploidy revealed in sterlet (Acipenser ruthenus) genome by chromosome painting. Mol. Cytogenet. 2015, 8, 90. [Google Scholar] [CrossRef] [PubMed]
- Helfman, G.S.; Collette, B.B.; Facey, D.E.; Bowen, B.W. The Diversity of Fishes: Biology, Evolution, and Ecology, 2nd ed.; Wiley-Blackwell: Oxford, UK, 2009; ISBN 978-1-4051-2494-2. [Google Scholar]
- Nelson, J.S.; Grande, T.; Wilson, M.V.H. Fishes of the World, 5th ed.; John Wiley & Sons: Hoboken, NJ, USA, 2016; ISBN 978-1-118-34233-6. [Google Scholar]
- Cavin, L. Freshwater Fishes: 250 Million Years of Evolutionary History; ISTE Press/Elsevier: London, UK, 2017. [Google Scholar]
- Sallan, L.C. Major issues in the origins of ray-finned fish (Actinopterygii) biodiversity. Biol. Rev. Camb. Philos. Soc. 2014, 89, 950–971. [Google Scholar] [CrossRef] [PubMed]
- Majtánová, Z.; Symonová, R.; Arias-Rodriguez, L.; Sallan, L.; Ráb, P. “Holostei versus Halecostomi” Problem: Insight from Cytogenetics of Ancient Nonteleost Actinopterygian Fish, Bowfin Amia calva. J. Exp. Zool. B Mol. Dev. Evol. 2017, 328, 620–628. [Google Scholar] [CrossRef] [PubMed]
- Vega, C.G.; Wiens, J.J. Why are there so few fish in the sea? Proc. R. Soc. B Biol. Sci. 2012, 279, 2323–2329. [Google Scholar] [CrossRef] [PubMed]
- Comber, S.C.L.; Smith, C. Polyploidy in fishes: Patterns and processes: Polyploidy in fishes. Biol. J. Linn. Soc. 2004, 82, 431–442. [Google Scholar] [CrossRef]
- Smith, E.M.; Gregory, T.R. Patterns of genome size diversity in the ray-finned fishes. Hydrobiologia 2009, 625, 1–25. [Google Scholar] [CrossRef]
- Mank, J.E.; Avise, J.C. Phylogenetic conservation of chromosome numbers in Actinopterygiian fishes. Genetica 2006, 127, 321–327. [Google Scholar] [CrossRef] [PubMed]
- Francis, W.R.; Wörheide, G. Similar Ratios of Introns to Intergenic Sequence across Animal Genomes. Genome Biol. Evol. 2017, 9, 1582–1598. [Google Scholar] [CrossRef] [PubMed]
- Sarropoulou, E.; Fernandes, J.M.O. Comparative genomics in teleost species: Knowledge transfer by linking the genomes of model and non-model fish species. Comp. Biochem. Physiol. Part D Genom. Proteom. 2011, 6, 92–102. [Google Scholar] [CrossRef] [PubMed]
- Howe, K.; Clark, M.D.; Torroja, C.F.; Torrance, J.; Berthelot, C.; Muffato, M.; Collins, J.E.; Humphray, S.; McLaren, K.; Matthews, L.; et al. The zebrafish reference genome sequence and its relationship to the human genome. Nature 2013, 496, 498–503. [Google Scholar] [CrossRef] [PubMed]
- Santoriello, C.; Zon, L.I. Hooked! Modeling human disease in zebrafish. J. Clin. Investig. 2012, 122, 2337–2343. [Google Scholar] [CrossRef] [PubMed]
- Gibbons, J.G.; Branco, A.T.; Godinho, S.A.; Yu, S.; Lemos, B. Concerted copy number variation balances ribosomal DNA dosage in human and mouse genomes. Proc. Natl. Acad. Sci. USA 2015, 112, 2485–2490. [Google Scholar] [CrossRef] [PubMed]
- Symonová, R.; Majtánová, Z.; Sember, A.; Staaks, G.B.; Bohlen, J.; Freyhof, J.; Rábová, M.; Ráb, P. Genome differentiation in a species pair of coregonine fishes: An extremely rapid speciation driven by stress-activated retrotransposons mediating extensive ribosomal DNA multiplications. BMC Evol. Biol. 2013, 13, 42. [Google Scholar] [CrossRef] [PubMed]
- Xu, B.; Li, H.; Perry, J.M.; Singh, V.P.; Unruh, J.; Yu, Z.; Zakari, M.; McDowell, W.; Li, L.; Gerton, J.L. Ribosomal DNA copy number loss and sequence variation in cancer. PLOS Genet. 2017, 13, e1006771. [Google Scholar] [CrossRef] [PubMed]
- Cioffi, M.B.; Martins, C.; Bertollo, L.A.C. Chromosome spreading of associated transposable elements and ribosomal DNA in the fish Erythrinus erythrinus. Implications for genome change and karyoevolution in fish. BMC Evol. Biol. 2010, 10, 271. [Google Scholar] [CrossRef] [PubMed]
- Animal rDNA Database. Available online: www.animalrdnadatabase.com (accessed on 9 November 2017).
- Sochorová, J.; Garcia, S.; Gálvez, F.; Symonová, R.; Kovařík, A. Evolutionary trends in animal ribosomal DNA loci: Introduction to a new online database. Chromosoma 2017. [Google Scholar] [CrossRef] [PubMed]
- Reed, K.M.; Phillips, R.B. Structure and organization of the rDNA intergenic spacer in lake trout (Salvelinus namaycush). Chromosome Res. Int. J. Mol. Supramol. Evol. Asp. Chromosome Biol. 2000, 8, 5–16. [Google Scholar] [CrossRef]
- Reed, K.M.; Hackett, J.D.; Phillips, R.B. Comparative analysis of intra-individual and inter-species DNA sequence variation in salmonid ribosomal DNA cistrons. Gene 2000, 249, 115–125. [Google Scholar] [CrossRef]
- Mazzuchelli, J.; Kocher, T.D.; Yang, F.; Martins, C. Integrating cytogenetics and genomics in comparative evolutionary studies of cichlid fish. BMC Genom. 2012, 13, 463. [Google Scholar] [CrossRef] [PubMed]
- Chitramuthu, B. Modeling Human Disease and Development in Zebrafish. Hum. Genet. Embryol. 2013, 3, e108. [Google Scholar] [CrossRef]
- Liu, S.; Leach, S.D. Zebrafish Models for Cancer. Annu. Rev. Pathol. Mech. Dis. 2011, 6, 71–93. [Google Scholar] [CrossRef] [PubMed]
- Schartl, M. Beyond the zebrafish: Diverse fish species for modeling human disease. Dis. Model. Mech. 2014, 7, 181–192. [Google Scholar] [CrossRef] [PubMed]
- Kari, G.; Rodeck, U.; Dicker, A.P. Zebrafish: An emerging model system for human disease and drug discovery. Clin. Pharmacol. Ther. 2007, 82, 70–80. [Google Scholar] [CrossRef] [PubMed]
- Amores, A.; Force, A.; Yan, Y.L.; Joly, L.; Amemiya, C.; Fritz, A.; Ho, R.K.; Langeland, J.; Prince, V.; Wang, Y.L.; et al. Zebrafish HOX clusters and vertebrate genome evolution. Science 1998, 282, 1711–1714. [Google Scholar] [CrossRef] [PubMed]
- Braasch, I.; Postlethwait, J.H. Polyploidy in Fish and the Teleost Genome Duplication. In Polyploidy and Genome Evolution; Soltis, P.S., Soltis, D.E., Eds.; Springer: Berlin/Heidelberg, Germany, 2012; pp. 341–383. ISBN 978-3-642-31441-4. [Google Scholar]
- Meyer, A.; Schartl, M. Gene and genome duplications in vertebrates: The one-to-four (-to-eight in fish) rule and the evolution of novel gene functions. Curr. Opin. Cell Biol. 1999, 11, 699–704. [Google Scholar] [CrossRef]
- Postlethwait, J.H.; Woods, I.G.; Ngo-Hazelett, P.; Yan, Y.L.; Kelly, P.D.; Chu, F.; Huang, H.; Hill-Force, A.; Talbot, W.S. Zebrafish comparative genomics and the origins of vertebrate chromosomes. Genome Res. 2000, 10, 1890–1902. [Google Scholar] [CrossRef] [PubMed]
- Ceol, C.J.; Houvras, Y.; Jane-Valbuena, J.; Bilodeau, S.; Orlando, D.A.; Battisti, V.; Fritsch, L.; Lin, W.M.; Hollmann, T.J.; Ferré, F.; et al. The histone methyltransferase SETDB1 is recurrently amplified in melanoma and accelerates its onset. Nature 2011, 471, 513–517. [Google Scholar] [CrossRef] [PubMed]
- Golzio, C.; Willer, J.; Talkowski, M.E.; Oh, E.C.; Taniguchi, Y.; Jacquemont, S.; Reymond, A.; Sun, M.; Sawa, A.; Gusella, J.F.; et al. KCTD13 is a major driver of mirrored neuroanatomical phenotypes of the 16p11.2 copy number variant. Nature 2012, 485, 363–367. [Google Scholar] [CrossRef] [PubMed]
- Panizzi, J.R.; Becker-Heck, A.; Castleman, V.H.; Al-Mutairi, D.A.; Liu, Y.; Loges, N.T.; Pathak, N.; Austin-Tse, C.; Sheridan, E.; Schmidts, M.; et al. CCDC103 mutations cause primary ciliary dyskinesia by disrupting assembly of ciliary dynein arms. Nat. Genet. 2012, 44, 714–719. [Google Scholar] [CrossRef] [PubMed]
- Roscioli, T.; Kamsteeg, E.-J.; Buysse, K.; Maystadt, I.; van Reeuwijk, J.; van den Elzen, C.; van Beusekom, E.; Riemersma, M.; Pfundt, R.; Vissers, L.E.L.M.; et al. Mutations in ISPD cause Walker-Warburg syndrome and defective glycosylation of α-dystroglycan. Nat. Genet. 2012, 44, 581–585. [Google Scholar] [CrossRef] [PubMed]
- Patton, E.E.; Widlund, H.R.; Kutok, J.L.; Kopani, K.R.; Amatruda, J.F.; Murphey, R.D.; Berghmans, S.; Mayhall, E.A.; Traver, D.; Fletcher, C.D.M.; et al. BRAF mutations are sufficient to promote nevi formation and cooperate with p53 in the genesis of melanoma. Curr. Biol. 2005, 15, 249–254. [Google Scholar] [CrossRef] [PubMed]
- Wittbrodt, J.; Shima, A.; Schartl, M. Medaka—A model organism from the far East. Nat. Rev. Genet. 2002, 3, 53–64. [Google Scholar] [CrossRef] [PubMed]
- Albertson, R.C.; Cresko, W.; Detrich, H.W.; Postlethwait, J.H. Evolutionary mutant models for human disease. Trends Genet. 2009, 25, 74–81. [Google Scholar] [CrossRef] [PubMed]
- McGaugh, S.E.; Gross, J.B.; Aken, B.; Blin, M.; Borowsky, R.; Chalopin, D.; Hinaux, H.; Jeffery, W.R.; Keene, A.; Ma, L.; et al. The cavefish genome reveals candidate genes for eye loss. Nat. Commun. 2014, 5, 5307. [Google Scholar] [CrossRef] [PubMed]
- Protas, M.E.; Hersey, C.; Kochanek, D.; Zhou, Y.; Wilkens, H.; Jeffery, W.R.; Zon, L.I.; Borowsky, R.; Tabin, C.J. Genetic analysis of cavefish reveals molecular convergence in the evolution of albinism. Nat. Genet. 2006, 38, 107–111. [Google Scholar] [CrossRef] [PubMed]
- Terzibasi, E.; Valenzano, D.R.; Cellerino, A. The short-lived fish Nothobranchius furzeri as a new model system for aging studies. Exp. Gerontol. 2007, 42, 81–89. [Google Scholar] [CrossRef] [PubMed]
- Hárosi, F.I.; von Herbing, I.H.; Van Keuren, J.R. Sickling of anoxic red blood cells in fish. Biol. Bull. 1998, 195, 5–11. [Google Scholar] [CrossRef] [PubMed]
- Meierjohann, S.; Schartl, M. From Mendelian to molecular genetics: The Xiphophorus melanoma model. Trends Genet. 2006, 22, 654–661. [Google Scholar] [CrossRef] [PubMed]
- Schartl, M.; Walter, R.B.; Shen, Y.; Garcia, T.; Catchen, J.; Amores, A.; Braasch, I.; Chalopin, D.; Volff, J.-N.; Lesch, K.-P.; et al. The genome of the platyfish, Xiphophorus maculatus, provides insights into evolutionary adaptation and several complex traits. Nat. Genet. 2013, 45, 567–572. [Google Scholar] [CrossRef] [PubMed]
- Schmale, M.C.; Hensley, G.T.; Udey, L.R. Neurofibromatosis in the bicolor damselfish (Pomacentrus partitus) as a model of von Recklinghausen neurofibromatosis. Ann. N. Y. Acad. Sci. 1986, 486, 386–402. [Google Scholar] [CrossRef] [PubMed]
- Williams, D.E. The rainbow trout liver cancer model: Response to environmental chemicals and studies on promotion and chemoprevention. Comp. Biochem. Physiol. Part C Toxicol. Pharmacol. 2012, 155, 121–127. [Google Scholar] [CrossRef] [PubMed]
- Burnett, K.G.; Bain, L.J.; Baldwin, W.S.; Callard, G.V.; Cohen, S.; Di Giulio, R.T.; Evans, D.H.; Gómez-Chiarri, M.; Hahn, M.E.; Hoover, C.A.; et al. Fundulus as the premier teleost model in environmental biology: Opportunities for new insights using genomics. Comp. Biochem. Physiol. Part D Genom. Proteom. 2007, 2, 257–286. [Google Scholar] [CrossRef] [PubMed]
- Lampert, K.; Schartl, M. The origin and evolution of a unisexual hybrid: Poecilia formosa. Philos. Trans. R. Soc. B Biol. Sci. 2008, 363, 2901–2909. [Google Scholar] [CrossRef] [PubMed]
- Schartl, M.; Nanda, I.; Schlupp, I.; Wilde, B.; Epplen, J.T.; Schmid, M.; Parzefall, J. Incorporation of subgenomic amounts of DNA as compensation for mutational load in a gynogenetic fish. Nature 1995, 373, 68–71. [Google Scholar] [CrossRef]
- Schartl, A.; Hornung, U.; Nanda, I.; Wacker, R.; Müller-Hermelink, H.K.; Schlupp, I.; Parzefall, J.; Schmid, M.; Schartl, M. Susceptibility to the development of pigment cell tumors in a clone of the Amazon molly, Poecilia formosa, introduced through a microchromosome. Cancer Res. 1997, 57, 2993–3000. [Google Scholar] [PubMed]
- Tobler, M.; Schlupp, I. Parasites in sexual and asexual mollies (Poecilia, Poeciliidae, Teleostei): A case for the Red Queen? Biol. Lett. 2005, 1, 166–168. [Google Scholar] [CrossRef] [PubMed]
- Woodhead, A.D.; Setlow, R.B.; Pond, V. The Amazon molly, Poecilia formosa, as a test animal in carcinogenicity studies: Chronic exposures to physical agents. Natl. Cancer Inst. Monogr. 1984, 65, 45–52. [Google Scholar] [PubMed]
- Scahill, C.M.; Digby, Z.; Sealy, I.M.; Wojciechowska, S.; White, R.J.; Collins, J.E.; Stemple, D.L.; Bartke, T.; Mathers, M.E.; Patton, E.E.; et al. Loss of the chromatin modifier Kdm2aa causes BrafV600E-independent spontaneous melanoma in zebrafish. PLoS Genet. 2017, 13, e1006959. [Google Scholar] [CrossRef] [PubMed]
- Cossins, A.R.; Crawford, D.L. Fish as models for environmental genomics. Nat. Rev. Genet. 2005, 6, 324–333. [Google Scholar] [CrossRef] [PubMed]
- Aquaculture Genomics, Genetics and Breeding Workshop; Abdelrahman, H.; ElHady, M.; Alcivar-Warren, A.; Allen, S.; Al-Tobasei, R.; Bao, L.; Beck, B.; Blackburn, H.; Bosworth, B.; et al. Aquaculture genomics, genetics and breeding in the United States: Current status, challenges and priorities for future research. BMC Genom. 2017, 18, 191. [Google Scholar] [CrossRef]
- Kobayashi, T. Genome Instability of Repetitive Sequence: Lesson from the Ribosomal RNA Gene Repeat. In DNA Replication, Recombination and Repair; Hanaoka, F., Sugasawa, K., Eds.; Springer: Tokyo, Japan, 2016; pp. 235–247. ISBN 978-4-431-55871-2. [Google Scholar]
- Wang, M.; Lemos, B. Ribosomal DNA copy number amplification and loss in human cancers is linked to tumor genetic context, nucleolus activity and proliferation. PLoS Genet. 2017, 13, e1006994. [Google Scholar] [CrossRef] [PubMed]
- Boisvert, F.-M.; van Koningsbruggen, S.; Navascués, J.; Lamond, A.I. The multifunctional nucleolus. Nat. Rev. Mol. Cell Biol. 2007, 8, 574–585. [Google Scholar] [CrossRef] [PubMed]
- Guetg, C.; Santoro, R. Formation of nuclear heterochromatin: The nucleolar point of view. Epigenetics 2012, 7, 811–814. [Google Scholar] [CrossRef] [PubMed]
- Makunin, A.I.; Dementyeva, P.V.; Graphodatsky, A.S.; Volobouev, V.T.; Kukekova, A.V.; Trifonov, V.A. Genes on B chromosomes of vertebrates. Mol. Cytogenet. 2014, 7, 99. [Google Scholar] [CrossRef] [PubMed]
- Terencio, M.L.; Schneider, C.H.; Gross, M.C.; do Carmo, E.J.; Nogaroto, V.; de Almeida, M.C.; Artoni, R.F.; Vicari, M.R.; Feldberg, E. Repetitive sequences: The hidden diversity of heterochromatin in prochilodontid fish. Comp. Cytogenet. 2015, 9, 465–481. [Google Scholar] [CrossRef] [PubMed]
- Tsekrekou, M.; Stratigi, K.; Chatzinikolaou, G. The Nucleolus: In Genome Maintenance and Repair. Int. J. Mol. Sci. 2017, 18, 1411. [Google Scholar] [CrossRef] [PubMed]
- Braasch, I.; Gehrke, A.R.; Smith, J.J.; Kawasaki, K.; Manousaki, T.; Pasquier, J.; Amores, A.; Desvignes, T.; Batzel, P.; Catchen, J.; et al. The spotted gar genome illuminates vertebrate evolution and facilitates human-teleost comparisons. Nat. Genet. 2016, 48, 427–437. [Google Scholar] [CrossRef] [PubMed]
- Dion- Côté, A.-M.; Symonová, R.; Ráb, P.; Bernatchez, L. Reproductive isolation in a nascent species pair is associated with aneuploidy in hybrid offspring. Proc. R. Soc. B Biol. Sci. 2015, 282, 20142862. [Google Scholar] [CrossRef] [PubMed]
- Clark, M.S. Genomics and Mapping of Teleostei (Bony Fish). Comp. Funct. Genom. 2003, 4, 182–193. [Google Scholar] [CrossRef] [PubMed]
- Carvalho, G.R.; Hauser, L.; Martinsohn, J.; Naish, K. Fish, genes and genomes: Contributions to ecology, evolution and management. J. Fish Biol. 2016, 89, 2471–2478. [Google Scholar] [CrossRef] [PubMed]
- Arai, R. Fish Karyotypes; Springer: Tokyo, Japan, 2011; ISBN 978-4-431-53876-9. [Google Scholar]
- Majtánová, Z.; Choleva, L.; Symonová, R.; Ráb, P.; Kotusz, J.; Pekárik, L.; Janko, K. Asexual Reproduction Does Not Apparently Increase the Rate of Chromosomal Evolution: Karyotype Stability in Diploid and Triploid Clonal Hybrid Fish (Cobitis, Cypriniformes, Teleostei). PLoS ONE 2016, 11, e0146872. [Google Scholar] [CrossRef] [PubMed]
- Rampin, M.; Bi, K.; Bogart, J.P.; Collares-Pereira, M.J. Identifying parental chromosomes and genomic rearrangements in animal hybrid complexes of species with small genome size using Genomic In Situ Hybridization (GISH). Comp. Cytogenet. 2012, 6, 287–300. [Google Scholar] [CrossRef] [PubMed]
- Havelka, M.; Bytyutskyy, D.; Symonová, R.; Ráb, P.; Flajšhans, M. The second highest chromosome count among vertebrates is observed in cultured sturgeon and is associated with genome plasticity. Genet. Sel. Evol. 2016, 48, 12. [Google Scholar] [CrossRef] [PubMed]
- Jaillon, O.; Aury, J.-M.; Brunet, F.; Petit, J.-L.; Stange-Thomann, N.; Mauceli, E.; Bouneau, L.; Fischer, C.; Ozouf-Costaz, C.; Bernot, A.; et al. Genome duplication in the teleost fish Tetraodon nigroviridis reveals the early vertebrate proto-karyotype. Nature 2004, 431, 946–957. [Google Scholar] [CrossRef] [PubMed]
- Roest Crollius, H.; Weissenbach, J. Fish genomics and biology. Genome Res. 2005, 15, 1675–1682. [Google Scholar] [CrossRef] [PubMed]
- Phillips, R.B.; Amores, A.; Morasch, M.R.; Wilson, C.; Postlethwait, J.H. Assignment of zebrafish genetic linkage groups to chromosomes. Cytogenet. Genome Res. 2006, 114, 155–162. [Google Scholar] [CrossRef] [PubMed]
- Phillips, R.B.; Nichols, K.M.; DeKoning, J.J.; Morasch, M.R.; Keatley, K.A.; Rexroad, C.; Gahr, S.A.; Danzmann, R.G.; Drew, R.E.; Thorgaard, G.H. Assignment of rainbow trout linkage groups to specific chromosomes. Genetics 2006, 174, 1661–1670. [Google Scholar] [CrossRef] [PubMed]
- Phillips, R.B.; Keatley, K.A.; Morasch, M.R.; Ventura, A.B.; Lubieniecki, K.P.; Koop, B.F.; Danzmann, R.G.; Davidson, W.S. Assignment of Atlantic salmon (Salmo salar) linkage groups to specific chromosomes: Conservation of large syntenic blocks corresponding to whole chromosome arms in rainbow trout (Oncorhynchus mykiss). BMC Genet. 2009, 10, 46. [Google Scholar] [CrossRef] [PubMed]
- Guyon, R.; Rakotomanga, M.; Azzouzi, N.; Coutanceau, J.P.; Bonillo, C.; D’Cotta, H.; Pepey, E.; Soler, L.; Rodier-Goud, M.; D’Hont, A.; et al. A high-resolution map of the Nile Tilapia genome: A resource for studying cichlids and other percomorphs. BMC Genom. 2012, 13, 222. [Google Scholar] [CrossRef] [PubMed]
- Dion-Côté, A.-M.; Symonová, R.; Lamaze, F.C.; Pelikánová, Š.; Ráb, P.; Bernatchez, L. Standing chromosomal variation in Lake Whitefish species pairs: The role of historical contingency and relevance for speciation. Mol. Ecol. 2017, 26, 178–192. [Google Scholar] [CrossRef] [PubMed]
- Rondeau, E.B.; Minkley, D.R.; Leong, J.S.; Messmer, A.M.; Jantzen, J.R.; von Schalburg, K.R.; Lemon, C.; Bird, N.H.; Koop, B.F. The genome and linkage map of the northern pike (Esox lucius): Conserved synteny revealed between the salmonid sister group and the Neoteleostei. PLoS ONE 2014, 9, e102089. [Google Scholar] [CrossRef] [PubMed]
- Sutherland, B.J.G.; Gosselin, T.; Normandeau, E.; Lamothe, M.; Isabel, N.; Audet, C.; Bernatchez, L. Salmonid Chromosome Evolution as Revealed by a Novel Method for Comparing RADseq Linkage Maps. Genome Biol. Evol. 2016, 8, 3600–3617. [Google Scholar] [CrossRef] [PubMed]
- Symonová, R.; Sutherland, B.J.G.; Bernatchez, L. Residually tetrasomic sites in Coregonus clupeaformis. Under preparation.
- Cozzi, P.; Milanesi, L.; Bernardi, G. Segmenting the Human Genome into Isochores. Evol. Bioinform. 2015, 11, 253–261. [Google Scholar] [CrossRef] [PubMed]
- Bernardi, G. Structural and Evolutionary Genomics: Natural Selection in Genome Evolution; New Comprehensive Biochemistry; Elsevier: Amsterdam, The Netherlands, 2004; ISBN 978-0-444-51255-0. [Google Scholar]
- Daniel-Silva, M.F.Z.; Almeida-Toledo, L.F. Chromosome evolution in fish: BrdU replication patterns demonstrate chromosome homeologies in two species of the genus Astyanax. Cytogenet. Genome Res. 2005, 109, 497–501. [Google Scholar] [CrossRef] [PubMed]
- Zerbino, D.R.; Achuthan, P.; Akanni, W.; Amode, M.R.; Barrell, D.; Bhai, J.; Billis, K.; Cummins, C.; Gall, A.; Girón, C.G.; et al. Ensembl 2018. Nucleic Acids Res. 2018, 46, D754–D761. [Google Scholar] [CrossRef] [PubMed]
- Cioffi, M.B.; Yano, C.F.; Sember, A.; Bertollo, L.A.C. Chromosomal Evolution in Lower Vertebrates: Sex Chromosomes in Neotropical Fishes. Genes 2017, 8, 258. [Google Scholar] [CrossRef] [PubMed]
- Cioffi, M.B.; Franco, W.; Ferreira, R.; Carlos Bertollo, L.A. Chromosomes as Tools for Discovering Biodiversity—The Case of Erythrinidae Fish Family. In Recent Trends in Cytogenetic Studies—Methodologies and Applications; Tirunilai, P., Ed.; InTech: Rijeka, Croatia, 2012; ISBN 978-953-51-0178-9. [Google Scholar]
- Glasauer, S.M.K.; Neuhauss, S.C.F. Whole-genome duplication in teleost fishes and its evolutionary consequences. Mol. Genet. Genom. 2014, 289, 1045–1060. [Google Scholar] [CrossRef] [PubMed]
- Mable, B.; Alexandrou, M.; Taylor, M. Genome duplication in amphibians and fish: An extended synthesis. J. Zool. 2011, 284, 151–182. [Google Scholar] [CrossRef]
- Symonová, R.; Havelka, M.; Amemiya, C.T.; Howell, W.M.; Kořínková, T.; Flajšhans, M.; Gela, D.; Ráb, P. Molecular cytogenetic differentiation of paralogs of HOX paralogs in duplicated and re-diploidized genome of the North American paddlefish (Polyodon spathula). BMC Genet. 2017, 18, 19. [Google Scholar] [CrossRef] [PubMed]
- Nagpure, N.S.; Rashid, I.; Pathak, A.K.; Singh, M.; Singh, S.P.; Sarkar, U.K. FBIS: A regional DNA barcode archival and analysis system for Indian fishes. Bioinformation 2012, 8, 483–488. [Google Scholar] [CrossRef] [PubMed]
- Nagpure, N.S.; Rashid, I.; Pathak, A.K.; Singh, M.; Pati, R.; Singh, S.P.; Sarkar, U.K. FMiR: A Curated Resource of Mitochondrial DNA Information for Fish. PLoS ONE 2015, 10, e0136711. [Google Scholar] [CrossRef] [PubMed]
- Nagpure, N.S.; Rashid, I.; Pati, R.; Pathak, A.K.; Singh, M.; Singh, S.P.; Sarkar, U.K. FishMicrosat: A microsatellite database of commercially important fishes and shellfishes of the Indian subcontinent. BMC Genom. 2013, 14, 630. [Google Scholar] [CrossRef] [PubMed]
- Avvaru, A.K.; Saxena, S.; Sowpati, D.T.; Mishra, R.K. MSDB: A Comprehensive Database of Simple Sequence Repeats. Genome Biol. Evol. 2017, 9, 1797–1802. [Google Scholar] [CrossRef] [PubMed]
- Nagpure, N.S.; Pathak, A.K.; Pati, R.; Rashid, I.; Sharma, J.; Singh, S.P.; Singh, M.; Sarkar, U.K.; Kushwaha, B.; Kumar, R.; et al. Fish Karyome version 2.1: A chromosome database of fishes and other aquatic organisms. Database 2016, 2016, baw012. [Google Scholar] [CrossRef] [PubMed]
- Froese, R.; Pauly, D. FishBase. World Wide Web Electronic Publication. 2017. Available online: www.fishbase.org (accessed on 9 November 2017).
- Bhartiya, D.; Maini, J.; Sharma, M.; Joshi, P.; Laddha, S.V.; Jalali, S.; Patowary, A.; Purkanti, R.; Lalwani, M.; Singh, A.R.; et al. FishMap Zv8 update—A genomic regulatory map of zebrafish. Zebrafish 2010, 7, 179–180. [Google Scholar] [CrossRef] [PubMed]
- Amores, A.; Postlethwait, J.H. Banded Chromosomes and the Zebrafish Karyotype. In Methods in Cell Biology; Elsevier: Amsterdam, The Netherlands, 1998; Volume 60, pp. 323–338. ISBN 978-0-12-544162-9. [Google Scholar]
- Gornung, E.; De Innocentiis, S.; Annesi, F.; Sola, L. Zebrafish 5S rRNA genes map to the long arms of chromosome 3. Chromosome Res. Int. J. Mol. Supramol. Evol. Asp. Chromosome Biol. 2000, 8, 362. [Google Scholar] [CrossRef]
- Phillips, R.B.; Reed, K.M. Localization of repetitive DNAs to zebrafish (Danio rerio) chromosomes by fluorescence in situ hybridization (FISH). Chromosome Res. Int. J. Mol. Supramol. Evol. Asp. Chromosome Biol. 2000, 8, 27–35. [Google Scholar] [CrossRef]
- Sola, L.; Gornung, E. Classical and molecular cytogenetics of the zebrafish, Danio rerio (Cyprinidae, Cypriniformes): An overview. Genetica 2001, 111, 397–412. [Google Scholar] [CrossRef] [PubMed]
- National Center for Biotechnology Information (NCBI). Genome Browser. Available online: www.ncbi.nlm.nih.gov/genome/browse (accessed on 9 November 2017).
- Personal Webpage of Radka Symonová. Available online: http://lide.uhk.cz/Symonra1 (accessed on 9 November 2017).
- Kitts, P.A.; Church, D.M.; Thibaud-Nissen, F.; Choi, J.; Hem, V.; Sapojnikov, V.; Smith, R.G.; Tatusova, T.; Xiang, C.; Zherikov, A.; et al. Assembly: A resource for assembled genomes at NCBI. Nucleic Acids Res. 2016, 44, D73–D80. [Google Scholar] [CrossRef] [PubMed]
- European Nucleotide Archive (ENA). Genome Assembly Database. Available online: www.ebi.ac.uk/ena/browse/genome-assembly-database (accessed on 9 November 2017).
- Tørresen, O.K.; Star, B.; Jentoft, S.; Reinar, W.B.; Grove, H.; Miller, J.R.; Walenz, B.P.; Knight, J.; Ekholm, J.M.; Peluso, P.; et al. An improved genome assembly uncovers prolific tandem repeats in Atlantic cod. BMC Genom. 2017, 18, 95. [Google Scholar] [CrossRef] [PubMed]
- Malmstrøm, M.; Matschiner, M.; Tørresen, O.K.; Jakobsen, K.S.; Jentoft, S. Whole genome sequencing data and de novo draft assemblies for 66 teleost species. Sci. Data 2017, 4, 160132. [Google Scholar] [CrossRef] [PubMed]
- Koepfli, K.-P.; Paten, B. Genome 10K Community of Scientists; O’Brien, S.J. The Genome 10K Project: A way forward. Annu. Rev. Anim. Biosci. 2015, 3, 57–111. [Google Scholar] [CrossRef] [PubMed]
- Pasquier, J.; Cabau, C.; Nguyen, T.; Jouanno, E.; Severac, D.; Braasch, I.; Journot, L.; Pontarotti, P.; Klopp, C.; Postlethwait, J.H.; et al. Gene evolution and gene expression after whole genome duplication in fish: The PhyloFish database. BMC Genom. 2016, 17, 368. [Google Scholar] [CrossRef] [PubMed]
- China National Genebank. FishT1K. Available online: https://db.cngb.org/fisht1k/status (accessed on 9 November 2017).
- Garamszegi, L. Z. Modern Phylogenetic Comparative Methods and Their Application in Evolutionary Biology: Concepts and Practice; Springer: Berlin, Germany, 2014; ISBN 978-3-662-43550-2. [Google Scholar]
- Hardie, D.C.; Hebert, P.D.N. The nucleotypic effects of cellular DNA content in cartilaginous and ray-finned fishes. Genome 2003, 46, 683–706. [Google Scholar] [CrossRef] [PubMed]
- Lefébure, T.; Morvan, C.; Malard, F.; François, C.; Konecny-Dupré, L.; Guéguen, L.; Weiss-Gayet, M.; Seguin-Orlando, A.; Ermini, L.; Sarkissian, C.D.; et al. Less effective selection leads to larger genomes. Genome Res. 2017, 27, 1016–1028. [Google Scholar] [CrossRef] [PubMed]
- Gregory, T.R.; Witt, J.D.S. Population size and genome size in fishes: A closer look. Genome 2008, 51, 309–313. [Google Scholar] [CrossRef] [PubMed]
- Elliott, T.A.; Gregory, T.R. Do larger genomes contain more diverse transposable elements? BMC Evol. Biol. 2015, 15, 69. [Google Scholar] [CrossRef] [PubMed]
- Tarallo, A.; Angelini, C.; Sanges, R.; Yagi, M.; Agnisola, C.; D’Onofrio, G. On the genome base composition of teleosts: The effect of environment and lifestyle. BMC Genom. 2016, 17, 173. [Google Scholar] [CrossRef] [PubMed]
- Hardie, D.C.; Hebert, P.D. Genome-size evolution in fishes. Can. J. Fish. Aquat. Sci. 2004, 61, 1636–1646. [Google Scholar] [CrossRef]
- Yi, S.; Streelman, J.T. Genome size is negatively correlated with effective population size in ray-finned fish. Trends Genet. 2005, 21, 643–646. [Google Scholar] [CrossRef] [PubMed]
- Vervoort, A. Tetraploidy in Protopterus (Dipnoi). Experientia 1980, 36, 294–296. [Google Scholar] [CrossRef]
- Paim, F.G.; da Hora Almeida, L.A.; de Mell Affonso, P.R.A.; Sobrinho-Scudeler, P.E.; Oliveira, C.; Diniz, D. Chromosomal stasis in distinct families of marine Percomorpharia from South Atlantic. Comp. Cytogenet. 2017, 11, 299–307. [Google Scholar] [CrossRef] [PubMed]
- Camacho, J.P.M.; Sharbel, T.F.; Beukeboom, L.W. B-chromosome evolution. Philos. Trans. R. Soc. B Biol. Sci. 2000, 355, 163–178. [Google Scholar] [CrossRef] [PubMed]
- Valente, G.T.; Nakajima, R.T.; Fantinatti, B.E.A.; Marques, D.F.; Almeida, R.O.; Simões, R.P.; Martins, C. B chromosomes: From cytogenetics to systems biology. Chromosoma 2017, 126, 73–81. [Google Scholar] [CrossRef] [PubMed]
- Anderson, J.L.; Rodríguez Marí, A.; Braasch, I.; Amores, A.; Hohenlohe, P.; Batzel, P.; Postlethwait, J.H. Multiple sex-associated regions and a putative sex chromosome in zebrafish revealed by RAD mapping and population genomics. PLoS ONE 2012, 7, e40701. [Google Scholar] [CrossRef] [PubMed]
- Bradley, K.M.; Breyer, J.P.; Melville, D.B.; Broman, K.W.; Knapik, E.W.; Smith, J.R. An SNP-Based Linkage Map for Zebrafish Reveals Sex Determination Loci. G3 2011, 1, 3–9. [Google Scholar] [CrossRef] [PubMed]
- Nagabhushana, A.; Mishra, R.K. Finding clues to the riddle of sex determination in zebrafish. J. Biosci. 2016, 41, 145–155. [Google Scholar] [CrossRef] [PubMed]
- Matsuda, M.; Nagahama, Y.; Shinomiya, A.; Sato, T.; Matsuda, C.; Kobayashi, T.; Morrey, C.E.; Shibata, N.; Asakawa, S.; Shimizu, N.; et al. DMY is a Y-specific DM-domain gene required for male development in the Medaka fish. Nature 2002, 417, 559–563. [Google Scholar] [CrossRef] [PubMed]
- De Andrade Silva, D.M.Z.; Utsunomia, R.; Ruiz-Ruano, F.J.; Daniel, S.N.; Porto-Foresti, F.; Hashimoto, D.T.; Oliveira, C.; Camacho, J.P.M.; Foresti, F. High-throughput analysis unveils a highly shared satellite DNA library among three species of fish genus Astyanax. Sci. Rep. 2017, 7, 12726. [Google Scholar] [CrossRef] [PubMed]
- Ruiz-Estévez, M.; López-León, M.D.; Cabrero, J.; Camacho, J.P.M. B-chromosome ribosomal DNA is functional in the grasshopper Eyprepocnemis plorans. PLoS ONE 2012, 7, e36600. [Google Scholar] [CrossRef] [PubMed]
- Utsunomia, R.; de Andrade Silva, D.M.Z.; Ruiz-Ruano, F.J.; Araya-Jaime, C.; Pansonato-Alves, J.C.; Scacchetti, P.C.; Hashimoto, D.T.; Oliveira, C.; Trifonov, V.A.; Porto-Foresti, F.; et al. Uncovering the Ancestry of B Chromosomes in Moenkhausia sanctaefilomenae (Teleostei, Characidae). PLoS ONE 2016, 11, e0150573. [Google Scholar] [CrossRef] [PubMed]
- Lamatsch, D.K.; Trifonov, V.; Schories, S.; Epplen, J. T.; Schmid, M.; Schartl, M. Isolation of a cancer-associated microchromosome in the sperm-dependent parthenogen Poecilia formosa. Cytogenet. Genome Res. 2011, 135, 135–142. [Google Scholar] [CrossRef] [PubMed]
- Schmid, M.; Ziegler, C.G.; Steinlein, C.; Nanda, I.; Schartl, M. Cytogenetics of the bleak (Alburnus alburnus), with special emphasis on the B chromosomes. Chromosome Res. Int. J. Mol. Supramol. Evol. Asp. Chromosome Biol. 2006, 14, 231–242. [Google Scholar] [CrossRef] [PubMed]
- Takagui, F.H.; Dias, A.L.; Birindelli, J.L.O.; Swarça, A.C.; da Rosa, R.; Lui, R.L.; Fenocchio, A.S.; Giuliano-Caetano, L. First report of B chromosomes in three neotropical thorny catfishes (Siluriformes, Doradidae). Comp. Cytogenet. 2017, 11, 55–64. [Google Scholar] [CrossRef] [PubMed]
- Jones, R.N.; Diez, M. The B chromosome database. Cytog. Gen. Res. 2004, 106, 149–150. [Google Scholar] [CrossRef] [PubMed]
- D’Ambrosio, U.; Alonso-Lifante, M.P.; Barros, K.; Kovařík, A.; Mas de Xaxars, G.; Garcia, S. B-chrom: A database on B-chromosomes of plants, animals and fungi. New Phytol. 2017, 216, 635–642. [Google Scholar] [CrossRef] [PubMed]
- Grummt, I. The nucleolus—Guardian of cellular homeostasis and genome integrity. Chromosoma 2013, 122, 487–497. [Google Scholar] [CrossRef] [PubMed]
- Ide, S.; Miyazaki, T.; Maki, H.; Kobayashi, T. Abundance of ribosomal RNA gene copies maintains genome integrity. Science 2010, 327, 693–696. [Google Scholar] [CrossRef] [PubMed]
- Charlesworth, B.; Sniegowski, P.; Stephan, W. The evolutionary dynamics of repetitive DNA in eukaryotes. Nature 1994, 371, 215–220. [Google Scholar] [CrossRef] [PubMed]
- Federico, C.; Scavo, C.; Cantarella, C.D.; Motta, S.; Saccone, S.; Bernardi, G. Gene-rich and gene-poor chromosomal regions have different locations in the interphase nuclei of cold-blooded vertebrates. Chromosoma 2006, 115, 123–128. [Google Scholar] [CrossRef] [PubMed]
- Kirubakaran, T.G.; Grove, H.; Kent, M.P.; Sandve, S.R.; Baranski, M.; Nome, T.; De Rosa, M.C.; Righino, B.; Johansen, T.; Otterå, H.; et al. Two adjacent inversions maintain genomic differentiation between migratory and stationary ecotypes of Atlantic cod. Mol. Ecol. 2016, 25, 2130–2143. [Google Scholar] [CrossRef] [PubMed]
- Fujiwara, A.; Abe, S.; Yamaha, E.; Yamazaki, F.; Yoshida, M.C. Chromosomal localization and heterochromatin association of ribosomal RNA gene loci and silver-stained nucleolar organizer regions in salmonid fishes. Chromosome Res. Int. J. Mol. Supramol. Evol. Asp. Chromosome Biol. 1998, 6, 463–471. [Google Scholar] [CrossRef]
- Costa, G.W.W.F.; Cioffi, M.B.; Bertollo, L.A.C.; Molina, W.F. Unusual dispersion of histone repeats on the whole chromosomal complement and their colocalization with ribosomal genes in Rachycentron canadum (Rachycentridae, Perciformes). Cytogenet. Genome Res. 2014, 144, 62–67. [Google Scholar] [CrossRef] [PubMed]
- Costa, G.W.W.F.; Cioffi, M.B.; Bertollo, L.A.C.; Molina, W.F. The Evolutionary Dynamics of Ribosomal Genes, Histone H3 and Transposable Rex Elements in the Genome of Atlantic Snappers. J. Hered. 2016, 107, 173–180. [Google Scholar] [CrossRef] [PubMed]
- Mehner, T.; Pohlmann, K.; Elkin, C.; Monaghan, M.T.; Nitz, B.; Freyhof, J. Genetic population structure of sympatric and allopatric populations of Baltic ciscoes (Coregonus albula complex, Teleostei, Coregonidae). BMC Evol. Biol. 2010, 10, 85. [Google Scholar] [CrossRef] [PubMed]
- Kobayashi, T. Ribosomal RNA gene repeats, their stability and cellular senescence. Proc. Jpn. Acad. Ser. B Phys. Biol. Sci. 2014, 90, 119–129. [Google Scholar] [CrossRef] [PubMed]
- Glazer, A.M.; Killingbeck, E.E.; Mitros, T.; Rokhsar, D.S.; Miller, C.T. Genome Assembly Improvement and Mapping Convergently Evolved Skeletal Traits in Sticklebacks with Genotyping-by-Sequencing. G3 2015, 5, 1463–1472. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Zhong, L.; Bian, C.; Xu, P.; Qiu, Y.; You, X.; Zhang, S.; Huang, Y.; Li, J.; Wang, M.; et al. High-quality genome assembly of channel catfish, Ictalurus punctatus. GigaScience 2016, 5. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Liu, S.; Yao, J.; Bao, L.; Zhang, J.; Li, Y.; Jiang, C.; Sun, L.; Wang, R.; Zhang, Y.; et al. The channel catfish genome sequence provides insights into the evolution of scale formation in teleosts. Nat. Commun. 2016, 7, 11757. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Chen, C.; Gao, Z.; Min, J.; Gu, Y.; Jian, J.; Jiang, X.; Cai, H.; Ebersberger, I.; Xu, M.; et al. The draft genome of blunt snout bream (Megalobrama amblycephala) reveals the development of intermuscular bone and adaptation to herbivorous diet. GigaScience 2017, 6, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Pan, H.; Yu, H.; Ravi, V.; Li, C.; Lee, A. P.; Lian, M.M.; Tay, B.-H.; Brenner, S.; Wang, J.; Yang, H.; et al. The genome of the largest bony fish, ocean sunfish (Mola mola), provides insights into its fast growth rate. GigaScience 2016, 5. [Google Scholar] [CrossRef] [PubMed]
- Brawand, D.; Wagner, C.E.; Li, Y.I.; Malinsky, M.; Keller, I.; Fan, S.; Simakov, O.; Ng, A.Y.; Lim, Z.W.; Bezault, E.; et al. The genomic substrate for adaptive radiation in African cichlid fish. Nature 2014, 513, 375–381. [Google Scholar] [CrossRef] [PubMed]
- Conte, M.A.; Gammerdinger, W.J.; Bartie, K.L.; Penman, D.J.; Kocher, T.D. A high quality assembly of the Nile Tilapia (Oreochromis niloticus) genome reveals the structure of two sex determination regions. BMC Genom. 2017, 18. [Google Scholar] [CrossRef] [PubMed]
- Lien, S.; Koop, B.F.; Sandve, S.R.; Miller, J.R.; Kent, M.P.; Nome, T.; Hvidsten, T.R.; Leong, J.S.; Minkley, D.R.; Zimin, A.; et al. The Atlantic salmon genome provides insights into rediploidization. Nature 2016, 533, 200–205. [Google Scholar] [CrossRef] [PubMed]

| Group | 2n Chromosome Counts (Basic Features) | Micro- Chromosomes | CG Heterogeneity | WGD after the First Two Basal Vertebrates´ WGDs | C-value/Haploid DNA Content (pg) [12] | Specific Features in the Genome History and Chromosomal Evolution |
|---|---|---|---|---|---|---|
| Myxiniformes (hagfishes) | 14–48 | NO | unknown | not observed | Myxinidae 2.5–4.59 | chromatin diminution, programmed genome rearrengement [13,14] |
| Petromyzontiformes (lampreys) | 76–178 | NO | GC-rich DNA repeats | not observed | Petromyzontidae 1.29–2.5 | programmed genome rearrengement [14] |
| Chondrichthyes (cartilaginous fishes) | 54–102 | YES | observed, presumably satellite DNA | not observed | Chimeriformes 1.5-2 Selachimorpha ~3–17 Rajimorphii 2.7–17 | AT/GC heterogeneity positively correlated with genome size [15] |
| Ceratodontiformes (lungfishes) | 34–68 | Only N. forsteri [15] otherwise not | unknown | not observed | Neoceradotus 52.75–74.86 Lepidosiren 80.55–123.9 Protopterus 40–132.8 | “genomic obesity” without WGD documented [16,17] |
| Coelacanthiformes (lobe-finned fishes) | 48 | YES | unknown | not observed | Latimeria 2.8–6.6 | chromosomes similar to ancient frogs [18] |
| Acipenseriformes (sturgeons, paddlefish) | ~ 120–240–360 | YES ~ 50% | NORs and GC-rich microchromosomes [7] andG-banding [19,20] | multiple in sturgeons, one in paddlefish | Acipenser 1.8–9.3 Polyodon 1.6–2.4 | multiple WGD, ploidy diversity |
| Lepisosteiformes (gars) | 56–58 | small sized chromosomes | in both genera | not observed | Atractosteus 1.2 Lepisosteus 1.4 | regionally high recombination rate |
| Amiiformes (bowfin) | 46 | NO | only NORs | not observed | Amia calva 1.2 | convergent evolution with teleosts? |
| Polypteriformes (bichirs) | 36–38 biarmed, extremelly large | NO | only NORs | not observed | Erpetoichthys 4.5 Polypterus 3.6-7.2 | not investigated |
| Teleostei | ~ 50 (exceptions up to 100–150 or more) | Micro B-chromosomes | only NORs | TGD and lineage specific WGDs | mostly 0.4- ~ 1.0 | from genome compaction to lineage specific WGD |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Symonová, R.; Howell, W.M. Vertebrate Genome Evolution in the Light of Fish Cytogenomics and rDNAomics. Genes 2018, 9, 96. https://doi.org/10.3390/genes9020096
Symonová R, Howell WM. Vertebrate Genome Evolution in the Light of Fish Cytogenomics and rDNAomics. Genes. 2018; 9(2):96. https://doi.org/10.3390/genes9020096
Chicago/Turabian StyleSymonová, Radka, and W. Mike Howell. 2018. "Vertebrate Genome Evolution in the Light of Fish Cytogenomics and rDNAomics" Genes 9, no. 2: 96. https://doi.org/10.3390/genes9020096
APA StyleSymonová, R., & Howell, W. M. (2018). Vertebrate Genome Evolution in the Light of Fish Cytogenomics and rDNAomics. Genes, 9(2), 96. https://doi.org/10.3390/genes9020096






