Chromosome Evolution in Marsupials
Abstract
1. Introduction
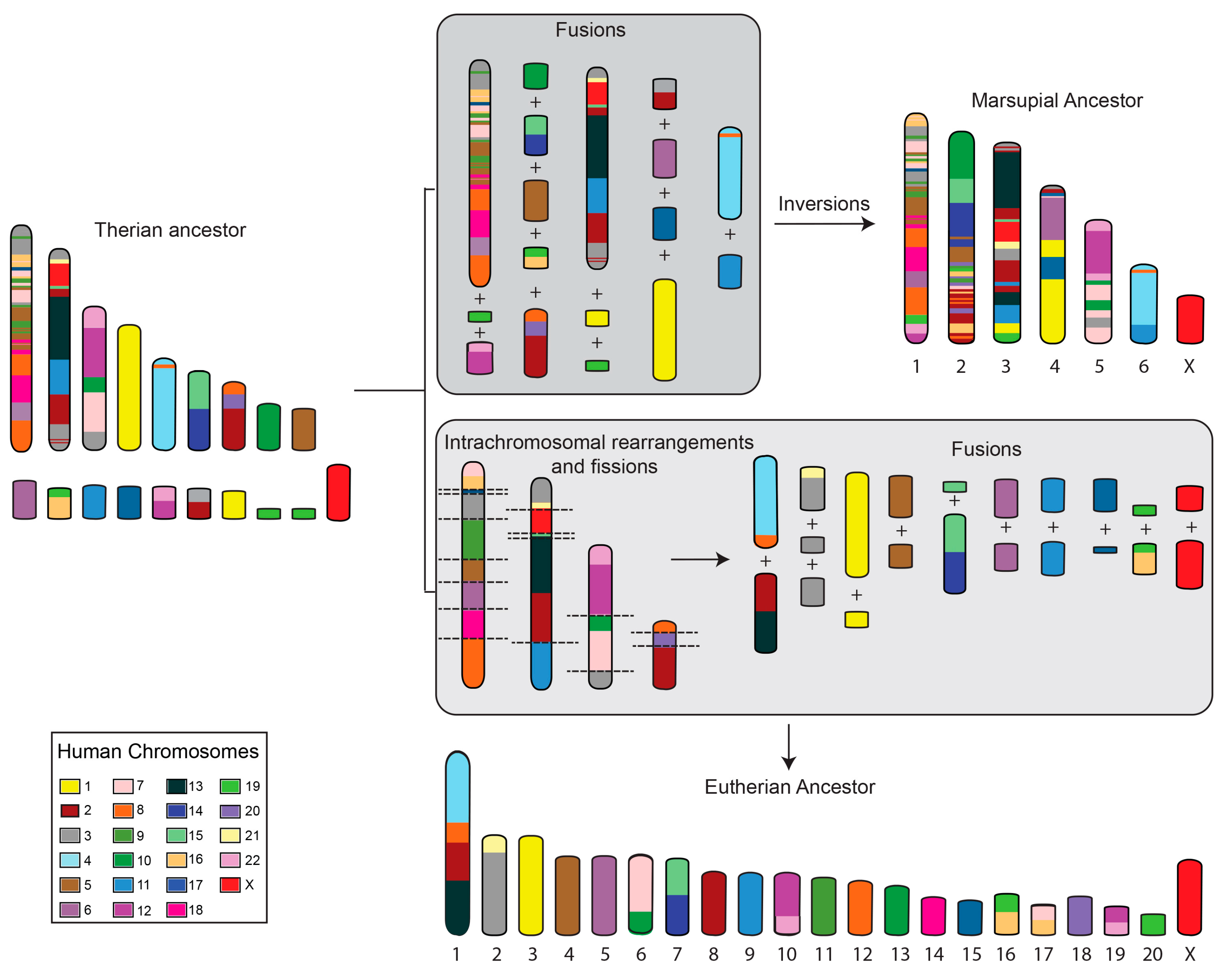
2. Marsupial Chromosome Evolution
2.1. Recombination in Marsupials
2.2. Marsupial Sex Chromosomes
2.3. Marsupial Genomics
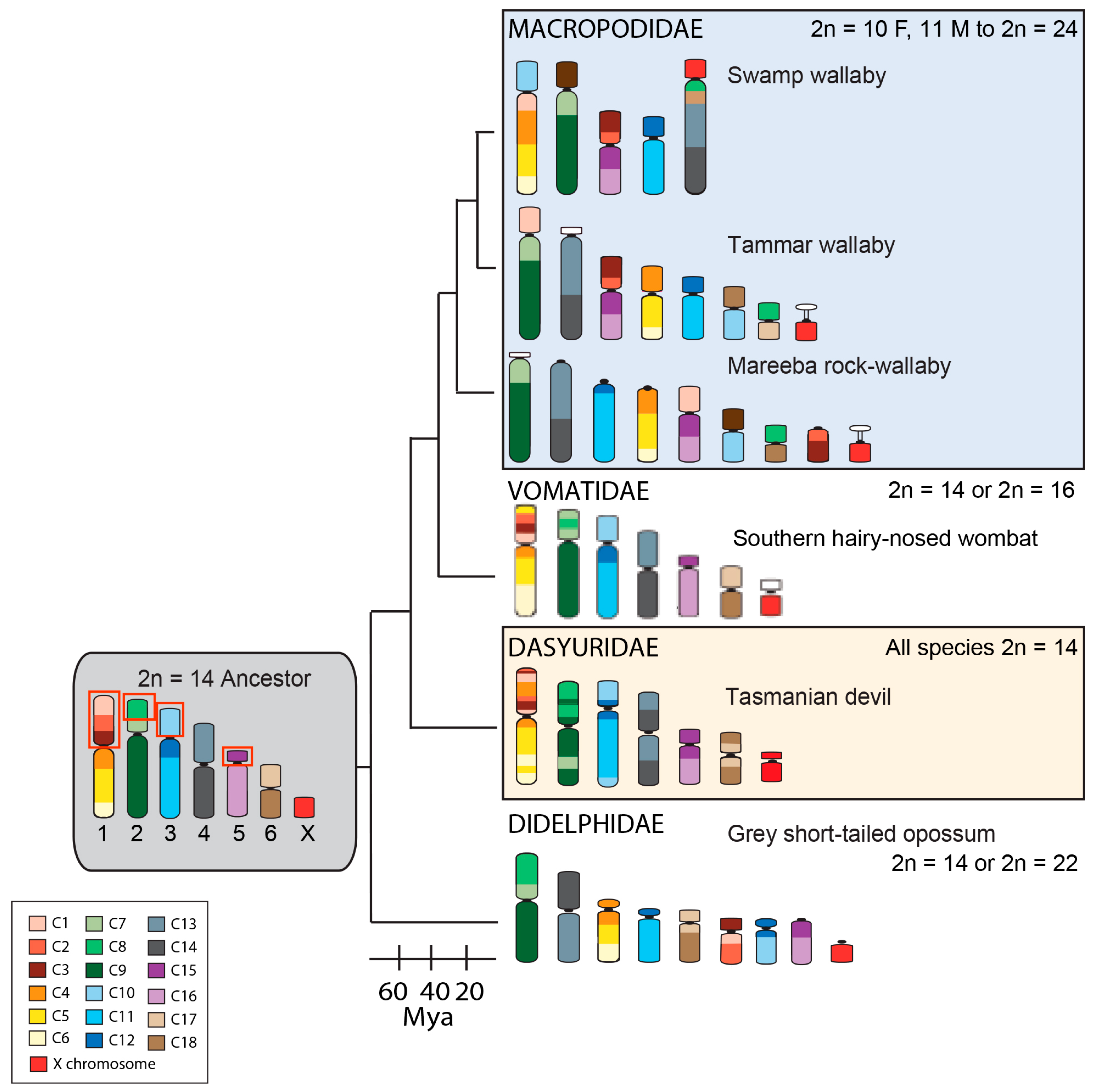
3. Dasyurid Chromosome Evolution and Unique Chromosome Features
Future Directions for Dasyurid Chromosome Research
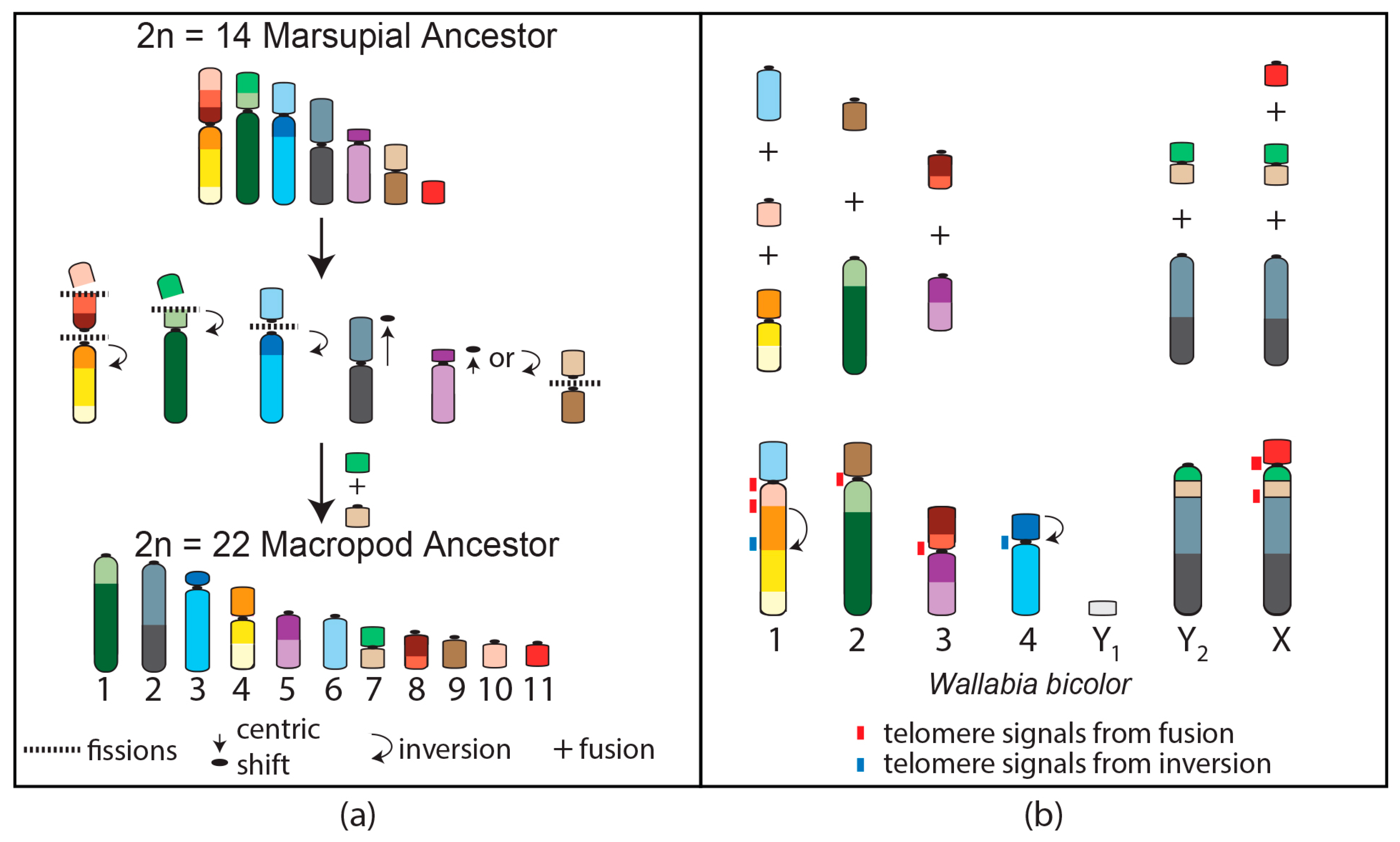
4. Macropod Chromosome Evolution
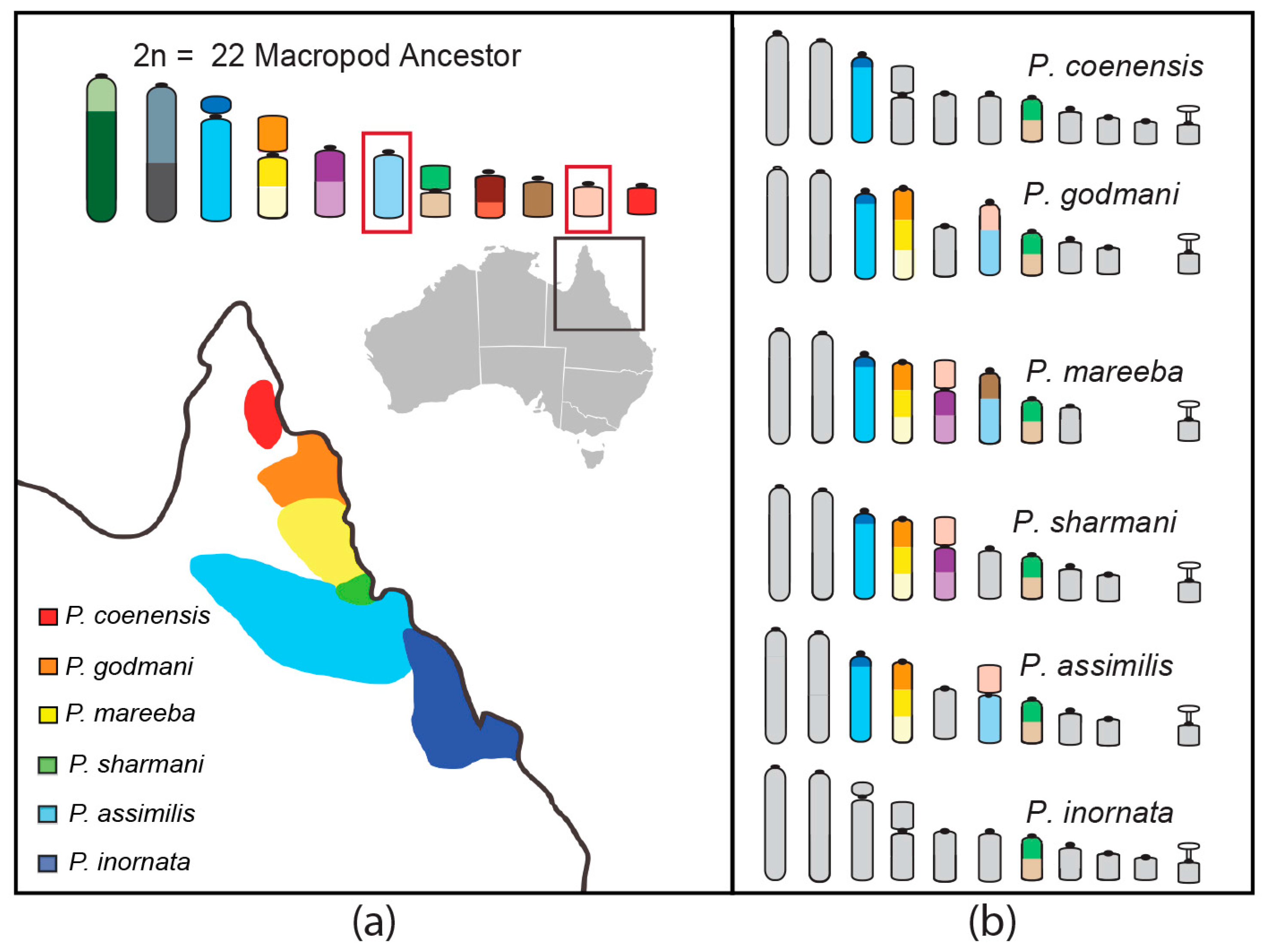
Rock-Wallabies as a Model Group for Studying Speciation
5. Conclusions
Acknowledgments
Conflicts of Interest
References
- King, M. Species Evolution: The Role of Chromosome Change; Cambridge University Press: Cambridge, UK, 1993; ISBN 0521484545. [Google Scholar]
- White, M.J.D. Modes of Speciation; W.H. Freeman and Co.: New York, NY, USA, 1978; ISBN 0716702843. [Google Scholar]
- Faria, R.; Navarro, A. Chromosomal speciation revisited: Rearranging theory with pieces of evidence. Trends Ecol. Evol. 2010, 25, 660–669. [Google Scholar] [CrossRef] [PubMed]
- Farré, M.; Robinson, T.J.; Ruiz-Herrera, A. An Integrative Breakage Model of genome architecture, reshuffling and evolution: The Integrative Breakage Model of genome evolution, a novel multidisciplinary hypothesis for the study of genome plasticity. BioEssays 2015, 37, 479–488. [Google Scholar] [CrossRef] [PubMed]
- Pevzner, P.; Tesler, G. Genome rearrangements in mammalian evolution: Lessons from human and mouse genomes. Genome Res. 2003, 13, 37–45. [Google Scholar] [CrossRef] [PubMed]
- Murphy, W.J.; Larkin, D.M.; Everts-van der Wind, A.; Bourque, G.; Tesler, G.; Auvil, L.; Beever, J.E.; Chowdhary, B.P.; Galibert, F.; Gatzke, L.; et al. Dynamics of mammalian chromosome evolution inferred from multispecies comparative maps. Science 2005, 309, 613–617. [Google Scholar] [CrossRef] [PubMed]
- Ruiz-Herrera, A.; Castresana, J.; Robinson, T.J. Is mammalian chromosomal evolution driven by regions of genome fragility? Genome Biol. 2006, 7, R115. [Google Scholar] [CrossRef] [PubMed]
- Capilla, L.; Sánchez-Guillén, R.A.; Farré, M.; Paytuví-Gallart, A.; Malinverni, R.; Ventura, J.; Larkin, D.M.; Ruiz-Herrera, A. Mammalian comparative genomics reveals genetic and epigenetic features associated with genome reshuffling in rodentia. Genome Biol. Evol. 2016, 8, 3703–3717. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Meredith, R.W.; Westerman, M.; Case, J.A.; Springer, M.S. A Phylogeny and timescale for marsupial evolution based on sequences for five nuclear genes. J. Mamm. Evol. 2008, 15, 1–36. [Google Scholar] [CrossRef]
- Sharman, G. The mitotic chromosomes of marsupials and their bearing on taxonomy and phylogeny. Aust. J. Zool. 1961, 9, 38–60. [Google Scholar] [CrossRef]
- Agar, W.E. The male meiotic phase in two genera of marsupials (Macropus and Petauroides). Q. J. Microsc. Sci. 1923, 67, 183–202. [Google Scholar]
- Painter, T.S. Studies in mammalian spermatogenesis. I. The spermatogenesis of the opossum. J. Exp. Zool. 1922, 35, 13–45. [Google Scholar] [CrossRef]
- Jordan, H.E. The microscopic anatomy of the epiphysis of the opossum. Anat. Rec. 1911, 5, 325–338. [Google Scholar] [CrossRef]
- Deakin, J.E.; Delbridge, M.L.; Koina, E.; Harley, N.; Alsop, A.E.; Wang, C.; Patel, V.S.; Graves, J.A.M. Reconstruction of the ancestral marsupial karyotype from comparative gene maps. BMC Evol. Biol. 2013, 13, 258. [Google Scholar] [CrossRef] [PubMed]
- Westerman, M.; Meredith, R.W.; Springer, M.S. Cytogenetics meets phylogenetics: A review of karyotype evolution in diprotodontian marsupials. J. Hered. 2010, 101, 690–702. [Google Scholar] [CrossRef] [PubMed]
- Rofe, R.; Hayman, D. G-banding evidence for a conserved complement in Marsupialia. Cytogenet. Cell Genet. 1985, 39, 40–50. [Google Scholar] [CrossRef] [PubMed]
- Rens, W.; Ferguson-Smith, M.A. The conserved marsupial karyotype: Chromosome painting and evolution. In Marsupial Genetics and Genomics; Deakin, J.E., Waters, P.D., Graves, J.A.M., Eds.; Springer: Dordrecht, The Netherlands, 2010; pp. 37–53. ISBN 9789048190225. [Google Scholar]
- Kim, J.; Farré, M.; Auvil, L.; Capitanu, B.; Larkin, D.M.; Ma, J.; Lewin, H.A. Reconstruction and evolutionary history of eutherian chromosomes. Proc. Natl. Acad. Sci. USA 2017, 114, E5379–E5388. [Google Scholar] [CrossRef] [PubMed]
- Hayman, D. Marsupial Cytogenetics. Aust. J. Zool. 1989, 37, 331. [Google Scholar] [CrossRef]
- Rens, W.; O’Brien, P.C.M.; Fairclough, H.; Harman, L.; Graves, J.A.M.; Ferguson-Smith, M.A. Reversal and convergence in marsupial chromosome evolution. Cytogenet. Genome Res. 2003, 102, 282–290. [Google Scholar] [CrossRef] [PubMed]
- Deakin, J.E.; Bender, H.S.; Pearse, A.M.; Rens, W.; O’Brien, P.C.M.; Ferguson-Smith, M.A.; Cheng, Y.; Morris, K.; Taylor, R.; Stuart, A.; et al. Genomic restructuring in the Tasmanian devil facial tumour: Chromosome painting and gene mapping provide clues to evolution of a transmissible tumour. PLoS Genet. 2012, 8, e1002483. [Google Scholar] [CrossRef] [PubMed]
- Bennett, J.H.; Hayman, D.L.; Hope, R.M. Novel sex differences in linkage values and meiotic chromosome behaviour in a marsupial. Nature 1986, 323, 59–60. [Google Scholar] [CrossRef] [PubMed]
- Hayman, D.L.; Moore, H.D.M.; Evans, E.P. Further evidence of novel sex differences in chiasma distribution in marsupials. Heredity 1988, 61, 455–458. [Google Scholar] [CrossRef]
- Hayman, D.L.; Rodger, J.C. Meiosis in male and female Trichosurus vulpecula (Marsupialia). Heredity 1990, 64, 251–254. [Google Scholar] [CrossRef]
- Samollow, P.B.; Gouin, N.; Miethke, P.; Mahaney, S.M.; Kenney, M.; VandeBerg, J.L.; Graves, J.A.M.; Kammerer, C.M. A microsatellite-based, physically anchored linkage map for the gray, short-tailed Opossum (Monodelphis domestica). Chromosome Res. 2007, 15, 269–281. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Webley, L.; Wei, K.; Wakefield, M.J.; Patel, H.R.; Deakin, J.E.; Alsop, A.; Marshall Graves, J.A.; Cooper, D.W.; Nicholas, F.W.; et al. A second-generation anchored genetic linkage map of the tammar wallaby (Macropus eugenii). BMC Genet. 2011, 12, 72. [Google Scholar] [CrossRef] [PubMed]
- Zenger, K.R.; McKenzie, L.M.; Cooper, D.W. The first comprehensive genetic linkage map of a marsupial: The tammar wallaby (Macropus eugenii). Genetics 2002, 162, 321–330. [Google Scholar] [PubMed]
- Samollow, P.B. Marsupial linkage maps. In Marsupial Genetics and Genomics; Deakin, J.E., Waters, P.D., Graves, J.A., Eds.; Springer: Dordrecht, The Netherlands; Heidelberg, Germany; London, UK; New York, NY, USA, 2010; pp. 75–100. ISBN 9789048190225. [Google Scholar]
- Hayman, D.L.; Smith, M.J.; Rodger, J.C. A comparative study of chiasmata in male and female Bettongia penicillata (Marsupialia). Genetica 1990, 83, 45–49. [Google Scholar] [CrossRef]
- Glas, R.; Graves, J.A.M.; Toder, R.; Ferguson-Smith, M.; O’Brien, P.C. Cross-species chromosome painting between human and marsupial directly demonstrates the ancient region of the mammalian X. Mamm. Genome 1999, 10, 1115–1116. [Google Scholar] [CrossRef] [PubMed]
- Deakin, J.E.; Koina, E.; Waters, P.D.; Doherty, R.; Patel, V.S.; Delbridge, M.L.; Dobson, B.; Fong, J.; Hu, Y.; Van Den Hurk, C.; et al. Physical map of two tammar wallaby chromosomes: A strategy for mapping in non-model mammals. Chromosome Res. 2008, 16, 1159–1175. [Google Scholar] [CrossRef] [PubMed]
- Graves, J.A. The origin and function of the mammalian Y chromosome and Y-borne genes—An evolving understanding. Bioessays 1995, 17, 311–320. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez Delgado, C.L.; Waters, P.D.; Gilbert, C.; Robinson, T.J.; Graves, J.A.M. Physical mapping of the elephant X chromosome: Conservation of gene order over 105 million years. Chromosome Res. 2009, 17, 917–926. [Google Scholar] [CrossRef] [PubMed]
- Proskuryakova, A.A.; Kulemzina, A.I.; Perelman, P.L.; Makunin, A.I.; Larkin, D.M.; Farré, M.; Kukekova, A.V.; Lynn Johnson, J.; Lemskaya, N.A.; Beklemisheva, V.R.; et al. X chromosome evolution in cetartiodactyla. Genes 2017, 8, 216. [Google Scholar] [CrossRef] [PubMed]
- Deakin, J.E. Marsupial genome sequences: Providing insight into evolution and disease. Scientifica 2012, 2012, 543176. [Google Scholar] [CrossRef] [PubMed]
- Rubtsov, N.B.; Rubtsova, N.V.; Anopriyenko, O.V.; Karamysheva, T.V.; Shevchenko, A.I.; Mazurok, N.A.; Nesterova, T.B.; Zakian, S.M. Reorganization of the X chromosome in voles of the genus microtus. Cytogenet. Genome Res. 2002, 99, 323–329. [Google Scholar] [CrossRef] [PubMed]
- Ohno, S. Sex Chromosomes and Sex Linked Genes; Springer Verlag: Berlin, Germany, 1967; ISBN 9783540039341. [Google Scholar]
- Vozdova, M.; Ruiz-Herrera, A.; Fernandez, J.; Cernohorska, H.; Frohlich, J.; Sebestova, H.; Kubickova, S.; Rubes, J. Meiotic behaviour of evolutionary sex-autosome translocations in Bovidae. Chromosome Res. 2016, 24, 325–338. [Google Scholar] [CrossRef] [PubMed]
- Toder, R.; O’Neill, R.J.W.; Wienberg, J.; O’Brien, P.C.M.; Voullaire, L.; Marshall-Graves, J.A. Comparative chromosome painting between two marsupials: Origins of an XX/XY1Y2 sex chromosome system. Mamm. Genome 1997, 8, 418–422. [Google Scholar] [CrossRef] [PubMed]
- Deakin, J.E. Marsupial X chromosome inactivation: Past, present and future. Aust. J. Zool. 2013, 61, 13–23. [Google Scholar] [CrossRef]
- Grant, J.; Mahadevaiah, S.K.; Khil, P.; Sangrithi, M.N.; Royo, H.; Duckworth, J.; McCarrey, J.R.; VandeBerg, J.L.; Renfree, M.B.; Taylor, W.; et al. Rsx is a metatherian RNA with Xist-like properties in X-chromosome inactivation. Nature 2012, 487, 254–258. [Google Scholar] [CrossRef] [PubMed]
- Toder, R.; Wakefield, M.J.; Graves, J.A. The minimal mammalian Y chromosome—The marsupial Y as a model system. Cytogenet. Cell Genet. 2000, 91, 285–292. [Google Scholar] [CrossRef] [PubMed]
- Foster, J.W.; Graves, J.A. An SRY-related sequence on the marsupial X chromosome: Implications for the evolution of the mammalian testis-determining gene. Proc. Natl. Acad. Sci. USA 1994, 91, 1927–1931. [Google Scholar] [CrossRef] [PubMed]
- Murtagh, V.J.; O’Meally, D.; Sankovic, N.; Delbridge, M.L.; Kuroki, Y.; Boore, J.L.; Toyoda, A.; Jordan, K.S.; Pask, A.J.; Renfree, M.B.; et al. Evolutionary history of novel genes on the tammar wallaby Y chromosome: Implications for sex chromosome evolution. Genome Res. 2012, 22, 498–507. [Google Scholar] [CrossRef] [PubMed]
- Bellott, D.W.; Hughes, J.F.; Skaletsky, H.; Brown, L.G.; Pyntikova, T.; Cho, T.-J.; Koutseva, N.; Zaghlul, S.; Graves, T.; Rock, S.; et al. Mammalian Y chromosomes retain widely expressed dosage-sensitive regulators. Nature 2014, 508, 494–499. [Google Scholar] [CrossRef] [PubMed]
- Sankovic, N.; Delbridge, M.L.; Grützner, F.; Ferguson-Smith, M.A.; O’Brien, P.C.M.; Marshall Graves, J.A. Construction of a highly enriched marsupial Y chromosome-specific BAC sub-library using isolated Y chromosomes. Chromosome Res. 2006, 14, 657–664. [Google Scholar] [CrossRef] [PubMed]
- Sharp, P. Sex chromosome pairing during male meiosis in marsupials. Chromosoma 1982, 86, 27–47. [Google Scholar] [CrossRef] [PubMed]
- Mikkelsen, T.S.; Wakefield, M.J.; Aken, B.; Amemiya, C.T.; Chang, J.L.; Duke, S.; Garber, M.; Gentles, A.J.; Goodstadt, L.; Heger, A.; et al. Genome of the marsupial Monodelphis domestica reveals innovation in non-coding sequences. Nature 2007, 447, 167–177. [Google Scholar] [CrossRef] [PubMed]
- Miller, W.; Hayes, V.M.; Ratan, A.; Petersen, D.C.; Wittekindt, N.E.; Miller, J.; Walenz, B.; Knight, J.; Qi, J.; Zhao, F.; et al. Genetic diversity and population structure of the endangered marsupial Sarcophilus harrisii (Tasmanian devil). Proc. Natl. Acad. Sci. USA 2011, 108, 12348–12353. [Google Scholar] [CrossRef] [PubMed]
- Murchison, E.P.; Schulz-Trieglaff, O.B.; Ning, Z.; Alexandrov, L.B.; Bauer, M.J.; Fu, B.; Hims, M.; Ding, Z.; Ivakhno, S.; Stewart, C.; et al. Genome sequencing and analysis of the Tasmanian devil and its transmissible cancer. Cell 2012, 148, 780–791. [Google Scholar] [CrossRef] [PubMed]
- Renfree, M.B.; Papenfuss, A.T.; Deakin, J.E.; Lindsay, J.; Heider, T.; Belov, K.; Rens, W.; Waters, P.D.; Pharo, E.A.; Shaw, G.; et al. Genome sequence of an Australian kangaroo, Macropus eugenii, provides insight into the evolution of mammalian reproduction and development. Genome Biol. 2011, 12, R81. [Google Scholar] [CrossRef] [PubMed]
- Koala Genome Consortium. Available online: https://koalagenome.org (accessed on 30 January 2018).
- Feigin, C.Y.; Newton, A.H.; Doronina, L.; Schmitz, J.; Hipsley, C.A.; Mitchell, K.J.; Gower, G.; Llamas, B.; Soubrier, J.; Heider, T.N.; et al. Genome of the Tasmanian tiger provides insights into the evolution and demography of an extinct marsupial carnivore. Nat. Ecol. Evol. 2018, 2, 182–192. [Google Scholar] [CrossRef] [PubMed]
- Oz Mammals Genomics Initiative. Available online: https://data.bioplatforms.com/organization/about/bpa-omg (accessed on 30 January 2018).
- Hayman, D.L.; Martin, P.G. Mammalia I: Monotremata and Marsupialia. In Animal Cytogenetics, Volume 4: Chordata; John, B., Ed.; Gebrüder Borntraeger: Berlin/Stuttgart, Germany, 1974. [Google Scholar]
- Young, G.J.; Graves, J.A.M.; Barbieri, I.; Woolley, P.A.; Cooper, D.W.; Westerman, M. The chromosomes of dasyurids (Masupialia). In Carnivorous Marsupials; Archer, M., Ed.; Royal Zoological Society: Sydney, NSW, Australia, 1982; pp. 783–795. ISBN 0959995137. [Google Scholar]
- Westerman, M.; Woolley, P.A. Cytogenetics of some New Guinean dasyurids and genome evolution in the Dasyuridae (Marsupialia). Aust. J. Zool. 1990, 37, 521–531. [Google Scholar] [CrossRef]
- Westerman, M.; Woolley, P.A. Chromosomes and the evolution of dasyurid marsupials: An overview. Sci. New Guin. 1993, 19, 123–130. [Google Scholar]
- Baverstock, P.R.; Adams, M.; Archer, M.; Mckenzie, N.L.; How, R. An Electrophoretic and Chromosomal Study of the Dasyurid Marsupial Genus Ningaui Archer. Aust. J. Zool. 1983, 31, 381–392. [Google Scholar] [CrossRef]
- Pyecroft, S.B.; Pearse, A.M.; Loh, R.; Swift, K.; Belov, K.; Fox, N.; Noonan, E.; Hayes, D.; Hyatt, A.; Wang, L.; et al. Towards a case definition for devil facial tumour disease: What is it? Ecohealth 2007, 4, 346–351. [Google Scholar] [CrossRef]
- Rocchi, M.; Archidiacono, N.; Schempp, W.; Capozzi, O.; Stanyon, R. Centromere repositioning in mammals. Heredity 2012, 108, 59–67. [Google Scholar] [CrossRef] [PubMed]
- Deakin, J.E.; Kruger-Andrezjewska, M. Marsupials as models for understanding the role of chromosome rearrangements in evolution and disease. Chromosoma 2016, 125, 633–644. [Google Scholar] [CrossRef] [PubMed]
- Bender, H.S.; Murchison, E.P.; Pickett, H.A.; Deakin, J.E.; Strong, M.A.; Conlan, C.; McMillan, D.A.; Neumann, A.A.; Greider, C.W.; Hannon, G.J.; et al. Extreme Telomere Length Dimorphism in the Tasmanian Devil and Related Marsupials Suggests Parental Control of Telomere Length. PLoS ONE 2012, 7. [Google Scholar] [CrossRef] [PubMed]
- Ingles, E.D. Telomeres, species differences, and unusual telomeres in vertebrates: Presenting challenges and opportunities to understanding telomere dynamics. AIMS Genet. 2016, 3, 1–24. [Google Scholar] [CrossRef]
- Griner, L.A. Neoplasms in Tasmanian Devils (Sarcophilus harrisii). J. Natl. Cancer Inst. 1979, 62, 589–595. [Google Scholar] [CrossRef] [PubMed]
- Canfield, P.J.; Cunningham, A.A. Disease and Mortality in Australasian Marsupials Held at London Zoo, 1872–1972. J. Zoo Wildl. Med. 1993, 24, 158–167. [Google Scholar]
- Canfield, P.J.; Hartley, W.J.; Reddacliff, G.L. Spontaneous proliferations in Australian marsupials—A survey and review. 2. Dasyurids and Bandicoots. J. Comp. Pathol. 1990, 103, 147–158. [Google Scholar] [CrossRef]
- Ingles, E.D.; Deakin, J.E. Global DNA Methylation patterns on marsupial and devil facial tumour chromosomes. Mol. Cytogenet. 2015, 8, 74. [Google Scholar] [CrossRef] [PubMed]
- Bennett, J.H.; Breed, W.G.; Hayman, D.L.; Hope, R.M. Reproductive and genetic studies with a laboratory colony of the dasyurid marsupial Sminthopsis crassicaudata. Aust. J. Zool. 1989, 37, 207–222. [Google Scholar] [CrossRef]
- Volker, M.; Backstrom, N.; Skinner, B.M.; Langley, E.J.; Bunzey, S.K.; Ellegren, H.; Griffin, D.K. Copy number variation, chromosome rearrangement, and their association with recombination during avian evolution. Genome Res. 2010, 20, 503–511. [Google Scholar] [CrossRef] [PubMed]
- Skinner, B.M.; Griffin, D.K. Intrachromosomal rearrangements in avian genome evolution: Evidence for regions prone to breakpoints. Heredity 2012, 108, 37–41. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Rofe, R. G-banded chromosomes and the evolution of Macropodidae. Aust. Mamm. 1978, 2, 53–63. [Google Scholar]
- Eldridge, M.D.B.; Close, R.L. Radiation of chromosome shuffles. Curr. Opin. Genet. Dev. 1993, 3, 915–922. [Google Scholar] [CrossRef]
- Rens, W.; O’Brien, P.C.M.; Yang, F.; Graves, J.A.M.; Ferguson-Smith, M.A. Karyotype relationships between four distantly related marsupials revealed by reciprocal chromosome painting. Chromosome Res. 1999, 7, 461–474. [Google Scholar] [CrossRef] [PubMed]
- Eldridge, M.D.B.; Metcalfe, C.J. Marsupialia. In Atlas of Mammalian Chromosomes; O’Brien, S.J., Menninger, J.C., Nash, W.G., Eds.; John Wiley & Sons, Inc.: Hoboken, NJ, USA, 2006; pp. 9–62. ISBN 10 0-471-35015-X. [Google Scholar]
- Eldridge, M.D.B.; Dollin, A.E.; Johnston, P.G.; Close, R.L.; Murray, J.D. Chromosomal rearrangements in rock wallabies, Petrogale (Marsupialia; Macropodidae). I. The Petrogale assimilis species complex. G-banding and synaptonemal complex analysis. Cytogenet. Cell Genet. 1988, 48, 228–232. [Google Scholar] [CrossRef]
- Eldridge, M.D.B.; Johnston, P.G.; Close, R.L.; Lowry, P.S. Chromosomal rearrangements in rock wallabies, Petrogale (Marsupialia, Macropodidae). 4. G-banding analysis of the Petrogale-lateralis complex. Genome 1989, 32, 935–940. [Google Scholar] [CrossRef] [PubMed]
- Eldridge, M.D.B.; Close, R.L.; Johnston, P.G. Chromosomal rearrangements in rock wallabies, Petrogale (Marsupialia, Macropodidae). III. G-banding analysis of Petrogale lateralis inornata and P. penicillata. Genome 1990, 33, 798–802. [Google Scholar] [CrossRef]
- Eldridge, M.D.B.; Close, R.L.; Johnston, P.G. Chromosomal rearrangements in rock wallabies, Petrogale (Marsupialia, Macropodidae). IV. G-banding analysis of the Petrogale lateralis complex. Aust. J. Zool. 1991, 39, 621–627. [Google Scholar] [CrossRef]
- Eldridge, M.D.B.; Johnston, P.G.; Lowry, P.S. Chromosomal Rearrangements in Rock Wallabies, Petrogale (Marsupialia, Macropodidae). VII. G-Banding Analysis of Petrogale brachyotis and P. concinna. Cytogenet. Cell Genet. 1992, 60, 34–39. [Google Scholar] [CrossRef]
- Eldridge, M.D.B.; Pearson, D.J. Chromosomal rearrangements in rock wallabies, Petrogale (Marsupialia: Macropodidae). IX. Further G-banding studies of the Petrogale lateralis complex: P. lateralis pearsoni, the West Kimberley race, and a population heterozygous for a centric fusion. Genome 1997, 40, 84–90. [Google Scholar] [CrossRef] [PubMed]
- Alsop, A.E.; Miethke, P.; Rofe, R.; Koina, E.; Sankovic, N.; Deakin, J.E.; Haines, H.; Rapkins, R.W.; Marshall Graves, J.A. Characterizing the chromosomes of the Australian model marsupial Macropus eugenii (tammar wallaby). Chromosome Res. 2005, 13, 627–636. [Google Scholar] [CrossRef] [PubMed]
- Glas, R.; De Leo, A.A.; Delbridge, M.L.; Reid, K.; Ferguson-Smith, M.A.; O’Brien, P.C.M.; Westerman, M.; Graves, J.A.M. Chromosome painting in marsupials: Genome conservation in the kangaroo family. Chromosome Res. 1999, 7, 167–176. [Google Scholar] [CrossRef] [PubMed]
- O’Neill, R.J.; Eldridge, M.D.B.; Toder, R.; Ferguson-Smith, M.A.; O’Brien, P.C.; Marshall Graves, J.A. Chromosome evolution in kangaroos (Marsupialia: Macropodidae): Cross species chromosome painting between the tammar wallaby and rock wallaby spp. with the 2n = 22 ancestral macropodid karyotype. Genome 1999, 42, 525–530. [Google Scholar] [CrossRef] [PubMed]
- Metcalfe, C.J.; Eldridge, M.D.B.; McQuade, L.R.; Johnston, P.G. Mapping the distribution of the telomeric sequence (T2AG3)n in rock-wallabies, Petrogale (Marsupialia: Macropodidae), by fluorescence in situ hybridization. I. The penicillata complex. Cytogenet. Genome Res. 1997, 78, 74–80. [Google Scholar] [CrossRef]
- Metcalfe, C.J.; Eldridge, M.D.B.; Toder, R.; Johnston, P.G. Mapping the distribution of the telomeric sequence (T2AG3)n in the Macropodoidea (Marsupialia), by fluorescence In situ hybridization. I. The swamp wallaby, Wallabia bicolor. Chromosome Res. 1998, 6, 603–610. [Google Scholar] [CrossRef] [PubMed]
- Metcalfe, C.J.; Eldridge, M.D.B.; Johnston, P.G. Mapping the distribution of the telomeric sequence (T2AG3)n in the Macropodoidea (Marsupialia) by fluorescence in situ hybridization. II. The ancestral 2n = 22 macropodid karyotype. Cytogenet. Genome Res. 2007, 116, 212–217. [Google Scholar] [CrossRef] [PubMed]
- Metcalfe, C.J.; Eldridge, M.D.B.; Johnston, P.G. Mapping the distribution of the telomeric sequence (T2AG3)n in rock wallabies, Petrogale (Marsupialia: Macropodidae), by fluorescence in situ hybridization. II. The lateralis complex. Cytogenet. Genome Res. 2002, 96, 169–175. [Google Scholar] [CrossRef] [PubMed]
- Eldridge, M.D.B.; Johnston, P.G. Chromosomal rearrangements in rock wallabies, Petrogale (Marsupialia: Macropodidae): VIII. An investigation of the nonrandom nature of karyotypic change. Genome 1993, 36, 524–534. [Google Scholar] [CrossRef] [PubMed]
- Bulazel, K.V.; Ferreri, G.C.; Eldridge, M.D.B.; O’Neill, R.J. Species-specific shifts in centromere sequence composition are coincident with breakpoint reuse in karyotypically divergent lineages. Genome Biol. 2007, 8, R170. [Google Scholar] [CrossRef] [PubMed]
- O’Neill, R.J.; O’Neill, M.J.; Graves, J.A. Undermethylation associated with retroelement activation and chromosome remodelling in an interspecific mammalian hybrid. Nature 1998, 393, 68–72. [Google Scholar] [CrossRef] [PubMed]
- Metcalfe, C.J.; Bulazel, K.V.; Ferreri, G.C.; Schroeder-Reiter, E.; Wanner, G.; Rens, W.; Obergfell, C.; Eldridge, M.D.B.; O’Neill, R.J. Genomic instability within centromeres of interspecific marsupial hybrids. Genetics 2007, 177, 2507–2517. [Google Scholar] [CrossRef] [PubMed]
- Close, R.L.; Lowry, P.S. Hybrids in Marsupial Research. Aust. J. Zool. 1990, 37, 259–267. [Google Scholar] [CrossRef]
- Close, R.L.; Bell, J.N. Fertile hybrids in two genera of wallabies: Petrogale and Thylogale. J. Hered. 1997, 88, 393–397. [Google Scholar] [CrossRef] [PubMed][Green Version]
- O’Neill, R.J.W.; Eldridge, M.D.B.; Graves, J.A.M. Chromosome heterozygosity and de novo chromosome rearrangements in mammalian interspecies hybrids. Mamm. Genome 2001, 12, 256–259. [Google Scholar] [CrossRef] [PubMed]
- Potter, S.; Bragg, J.G.; Blom, M.P.; Deakin, J.E.; Kirkpatrick, M.; Eldridge, M.D.; Moritz, C. Chromosomal speciation in the genomics era: Disentangling phylogenetic evolution of rock-wallabies. Front. Genet. 2017, 8, 1–18. [Google Scholar] [CrossRef] [PubMed]
- Eldridge, M.D.B.; Johnston, P.G.; Close, R.L. Chromosomal rearrangements in rock wallabies, Petrogale (Marsupialia, Macropodidae). VI. Determination of the plesiomorphic karyotype: G-Banding comparison of Thylogale with Petrogale persephone, P. xanthopus, and P. l. lateralis. Cytogenet. Cell Genet. 1992, 61, 29–33. [Google Scholar] [CrossRef] [PubMed]
- Potter, S.; Cooper, S.J.; Metcalfe, C.J.; Taggart, D.A.; Eldridge, M.D.B. Phylogenetic relationships of rock-wallabies Petrogale (Marsupialia: Macropodidae) and their biogeographic history within Australia. Mol. Ecol. 2012, 21, 2254–2269. [Google Scholar] [CrossRef] [PubMed]
- Eldridge, M.D.B.; Close, R.L. Taxonomy of rock wallabies, Petrogale (Marsupialia:Macropodidae). I. a revision of the eastern Petrogale with the description of three new species. Aust. J. Zool. 1992, 40, 563–571. [Google Scholar] [CrossRef]
- Close, R.L.; Bell, J.N.; Dollin, A.E.; Harding, H.R. Spermatogenesis and synaptonemal complexes of hybrid Petrogale (Marsupialia). J. Hered. 1996, 87, 96–107. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Potter, S.; Moritz, C.; Eldridge, M.D.B. Gene flow despite complex Robertsonian fusions among rock-wallaby (Petrogale) species. Biol. Lett. 2015, 11. [Google Scholar] [CrossRef] [PubMed]
© 2018 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Deakin, J.E. Chromosome Evolution in Marsupials. Genes 2018, 9, 72. https://doi.org/10.3390/genes9020072
Deakin JE. Chromosome Evolution in Marsupials. Genes. 2018; 9(2):72. https://doi.org/10.3390/genes9020072
Chicago/Turabian StyleDeakin, Janine E. 2018. "Chromosome Evolution in Marsupials" Genes 9, no. 2: 72. https://doi.org/10.3390/genes9020072
APA StyleDeakin, J. E. (2018). Chromosome Evolution in Marsupials. Genes, 9(2), 72. https://doi.org/10.3390/genes9020072




