Pathogenicity Islands Distribution in Non-O157 Shiga Toxin-Producing Escherichia coli (STEC)
Abstract
:1. Introduction
2. Materials and Methods
2.1. Bacterial Strains
2.2. Detection of Non-Locus of the Enterocyte Effacement Effector Genes Encoded in Pathogenicity Islands
2.3. Detection of Particular Genes for the Presence of OI-122
2.4. Cluster Analysis
3. Results
4. Discussion
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Karmali, M.A.; Gannon, V.; Sargeant, J.M. Verocytotoxin-producing Escherichia coli (VTEC). Vet. Microbiol. 2010, 140, 360–370. [Google Scholar] [CrossRef] [PubMed]
- Gould, L.H.; Mody, R.K.; Ong, K.L.; Clogher, P.; Cronquist, A.B.; Garman, K.N.; Lathrop, S.; Medus, C.; Spina, N.L.; Webb, T.H.; et al. Increased recognition of non-O157 Shiga toxin-producing Escherichia coli infections in the United States during 2000-2010: Epidemiologic features and comparison with E. coli O157 infections. Foodborne Pathog. Dis. 2013, 10, 453–460. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, H.; Hensel, M. Pathogenicity islands in bacterial pathogenesis. Clinical Microbiology Reviews 2004, 17, 14–56. [Google Scholar] [CrossRef] [PubMed]
- Karmali, M.A.; Mascarenhas, M.; Shen, S.; Ziebell, K.; Johnson, S.; Reid-Smith, R.; Isaac-Renton, J.; Clark, C.; Rahn, K.; Kaper, J.B. Association of genomic O Island 122 of Escherichia coli EDL 933 with Verocytotoxin-Producing Escherichia coli seropathotypes that are linked to epidemic and/or serious disease. J. Clin. Microbiol. 2003, 41, 4930–4940. [Google Scholar] [CrossRef] [PubMed]
- Ju, W.; Shen, J.; Toro, M.; Zhao, S.; Menga, J. Distribution of pathogenicity islands OI-122, OI-43/48, and OI-57 and a high-pathogenicity island in Shiga toxin-producing Escherichia coli. Appl. Environ. Microbiol. 2013, 79, 3406–3412. [Google Scholar] [CrossRef] [PubMed]
- García-Angulo, V.A.; Martínez-Santos, V.I.; Villaseñor, T.; Santana, F.J.; Huerta-Saquero, A.; Martínez, L.C.; Jiménez, R.; Lara-Ochoa, C.; Téllez-Sosa, J.; Bustamante, V.H.; et al. A distinct regulatory sequence is essential for the expression of a subset of nle genes in attaching and effacing Escherichia coli. J. Bacteriol. 2012, 194, 5589–5603. [Google Scholar] [CrossRef]
- Coombes, B.K.; Wickham, M.E.; Mascarenhas, M.; Gruenheid, S.; Finlay, B.B.; Karmali, M.A. Molecular analysis as an aid to assess the public health risk of non-O157 shiga toxin-producing Escherichia coli strains. Appl. Environ. Microbiol. 2008, 74, 2153–2160. [Google Scholar] [CrossRef] [PubMed]
- Konczy, P.; Ziebell, K.; Mascarenhas, M.; Choi, A.; Michaud, C.; Kropinski, A.M.; Whittam, T.S.; Wickham, M.; Finlay, B.; Karmali, M.A. Genomic O island 122, locus for enterocyte effacement, and the evolution of virulent verocytotoxin-producing Escherichia coli. J. Bacteriol. 2008, 190, 5832–5840. [Google Scholar] [CrossRef] [PubMed]
- Bugarel, M.; Beutin, L.; Martin, A.; Gill, A.; Fach, P. Micro-array for the identification of Shiga toxin-producing Escherichia coli (STEC) seropathotypes associated with Hemorrhagic Colitis and Hemolytic Uremic Syndrome in humans. Int. J. Food Microbiol. 2010, 142, 318–329. [Google Scholar] [CrossRef] [PubMed]
- Bugarel, M.; Beutin, L.; Fach, P. Low-density macroarray targeting non-locus of enterocyte effacement effectors (nle genes) and major virulence factors of Shiga toxin-producing Escherichia coli (STEC): A new approach for molecular risk assessment of STEC isolates. Appl. Environ. Microbiol. 2010, 76, 203–211. [Google Scholar] [CrossRef] [PubMed]
- Imamovic, L.; Tozzoli, R.; Michelacci, V.; Minelli, F.; Marziano, M.L.; Caprioli, A.; Morabito, S. OI-57, a genomic island of Escherichia coli O157, is present in other seropathotypes of Shiga toxin-producing E. coli associated with severe human disease. Infect. Immun. 2010, 78, 4697–4704. [Google Scholar] [CrossRef] [PubMed]
- Wu, B.; Skarina, T.; Yee, A.; Jobin, M.C.; Dileo, R.; Semesi, A.; Fares, C.; Lemak, A.; Coombes, B.K.; Arrowsmith, C.H.; et al. NleG type 3 effectors from enterohaemorrhagic Escherichia coli are U-Box E3 ubiquitin ligases. PLoS Pathog. 2010, 6, e1000960. [Google Scholar] [CrossRef] [PubMed]
- Morabito, S.; Tozzoli, R.; Oswald, E.; Caprioli, A. A mosaic pathogenicity island made up of the locus of enterocyte effacement and a pathogenicity island of Escherichia coli O157:H7 is frequently present in attaching and effacing E. coli. Infect. Immun. 2003, 71, 3343–3348. [Google Scholar] [CrossRef] [PubMed]
- Wickham, M.E.; Lupp, C.; Mascarenhas, M.; Vazquez, A.; Coombes, B.K.; Brown, N.F.; Coburn, B.A.; Deng, W.; Puente, J.L.; Karmali, M.A.; Finlay, B.B. Bacterial genetic determinants of non-O157 STEC outbreaks and hemolytic-uremic syndrome after infection. J. Infect. Dis. 2006, 194, 819–827. [Google Scholar] [CrossRef] [PubMed]
- Buvens, G.; Piérard, D. virulence profiling and disease association of verocytotoxin-producing Escherichia coli O157 and Non-O157 isolates in Belgium. Foodborne Pathog. Dis. 2012, 9, 530–535. [Google Scholar] [CrossRef] [PubMed]
- Parma, A.E.; Sanz, M.E.; Blanco, J.E.; Blanco, J.; Viñas, M.R.; Blanco, M.; Padola, N.L.; Etcheverria, A.I. Virulence genotypes and serotypes of verotoxigenic Escherichia coli isolated from cattle and foods in Argentina: Importance in public health. Eur. J. Epidemiol. 2000, 16, 757–762. [Google Scholar] [CrossRef] [PubMed]
- Blanco, M.; Padola, N.L.; Krüger, A.; Sanz, M.E.; Blanco, J.E.; Gonzalez, E.A.; Dahbi, G.; Mora, A.; Bernardez, M.I.; Etcheverria, A.I.; et al. Virulence genes and intimin types of Shiga-toxin-producing Escherichia coli isolated from cattle and beef products in Argentina. Int. Microbiol. 2004, 7, 269–276. [Google Scholar] [PubMed]
- Sanz, M.E.; Cristina, V.; Elida, E.; Arroyo, G. Prevalencia de Escherichia coli verotoxigénico en productos cárnicos de la ciudad de Tandil. La Ind. Cárnica Latinoam. 2007, 146, 56–58. [Google Scholar]
- Rivero, M.A.; Passucci, J.A.; Rodriguez, E.M.; Parma, A.E. Role and clinical course of verotoxigenic Escherichia coli infections in childhood acute diarrhoea in Argentina. J. Med. Microbiol. 2010, 59, 345–352. [Google Scholar] [CrossRef] [PubMed]
- González, J.; Cadona, J.S.; Sanz, M.; Bustamante, A.V.; Sanso, A.M. Molecular characterization of diarrheagenic Escherichia coli isolated from vegetables in Argentina. Int. J. Food Microbiol. 2017, 261, 57–61. [Google Scholar] [CrossRef] [PubMed]
- Toma, C.; Espinosa, E.M.; Song, T.; Miliwebsky, E.; Chinen, I.; Iyoda, S.; Iwanaga, M.; Rivas, M. Distribution of putative adhesins in different seropathotypes of Shiga toxin-producing Escherichia coli. J. Clin. Microbiol. 2004, 42, 4937–4946. [Google Scholar] [CrossRef] [PubMed]
- Franz, E.; Van Hoek, A.H.A.M.; Wuite, M.; Van Der Wal, F.J.; De Boer, A.G.; Bouw, E.I.; Aarts, H.J.M. Molecular hazard identification of non-O157 Shiga toxin-producing Escherichia coli (STEC). PLoS ONE 2015, 10, 1–21. [Google Scholar] [CrossRef] [PubMed]
- Ju, W.; Rump, L.; Toro, M.; Shen, J.; Cao, G.; Zhao, S.; Meng, J. Pathogenicity islands in Shiga toxin-producing Escherichia coli O26, O103, and O111 isolates from humans and animals. Foodborne Pathog. Dis. 2014, 11, 342–345. [Google Scholar] [CrossRef] [PubMed]
- Chui, L.; Li, V.; Fach, P.; Delannoy, S.; Malejczyk, K.; Patterson-Fortin, L.; Poon, A.; King, R.; Simmonds, K.; Scott, A.N.; Lee, M.C. Molecular profiling of Escherichia coli O157:H7 and non-O157 strains isolated from humans and cattle in Alberta, Canada. J. Clin. Microbiol. 2015, 53, 986–990. [Google Scholar] [CrossRef] [PubMed]
- Rivas, M.; Miliwebsky, E.; Chinen, I.; Deza, N.; Leotta, G.A. Epidemiología del síndrome urémico hemolítico en Argentina. Diagnóstico del agente etiológico, reservorios y vías de transmisión. Medicina 2006, 66, 27–32. [Google Scholar] [PubMed]
- Newton, H.J.; Pearson, J.S.; Badea, L.; Kelly, M.; Lucas, M.; Holloway, G.; Wagstaff, K.M.; Dunstone, M.A.; Sloan, J.; Whisstock, J.C.; et al. The type III effectors NleE and NleB from enteropathogenic E. coli and Ospz from Shigella block nuclear translocation of NF-κB p65. PLoS Pathog. 2010, 6, e1000898. [Google Scholar] [CrossRef] [PubMed]
- Cadona, J.S.; Bustamante, A.V.; González, J.; Sanso, A.M. Genetic relatedness and novel sequence types of Non-O157 Shiga toxin-producing Escherichia coli strains isolated in Argentina. Front. Cell. Infect. Microbiol. 2016, 6, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Montero, D.A.; Velasco, J.; Del Canto, F.; Puente, J.L.; Padola, N.L.; Rasko, D.A.; Farfán, M.; Salazar, J.C.; Vidal, R. Locus of adhesion and autoaggregation (LAA), a pathogenicity island present in emerging Shiga Toxin-producing Escherichia coli strains. Sci. Rep. 2017, 7, 7011. [Google Scholar] [CrossRef] [PubMed]
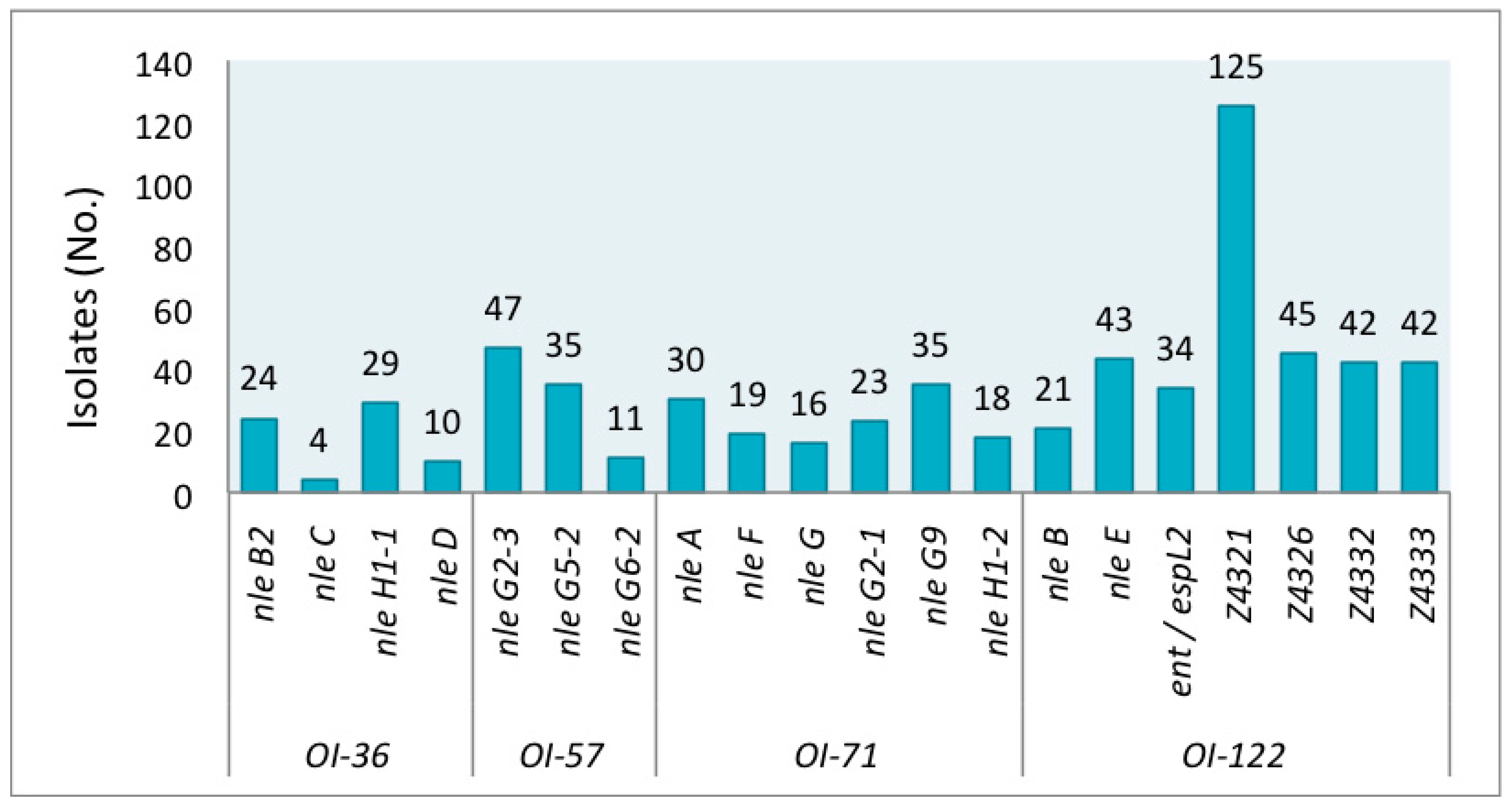
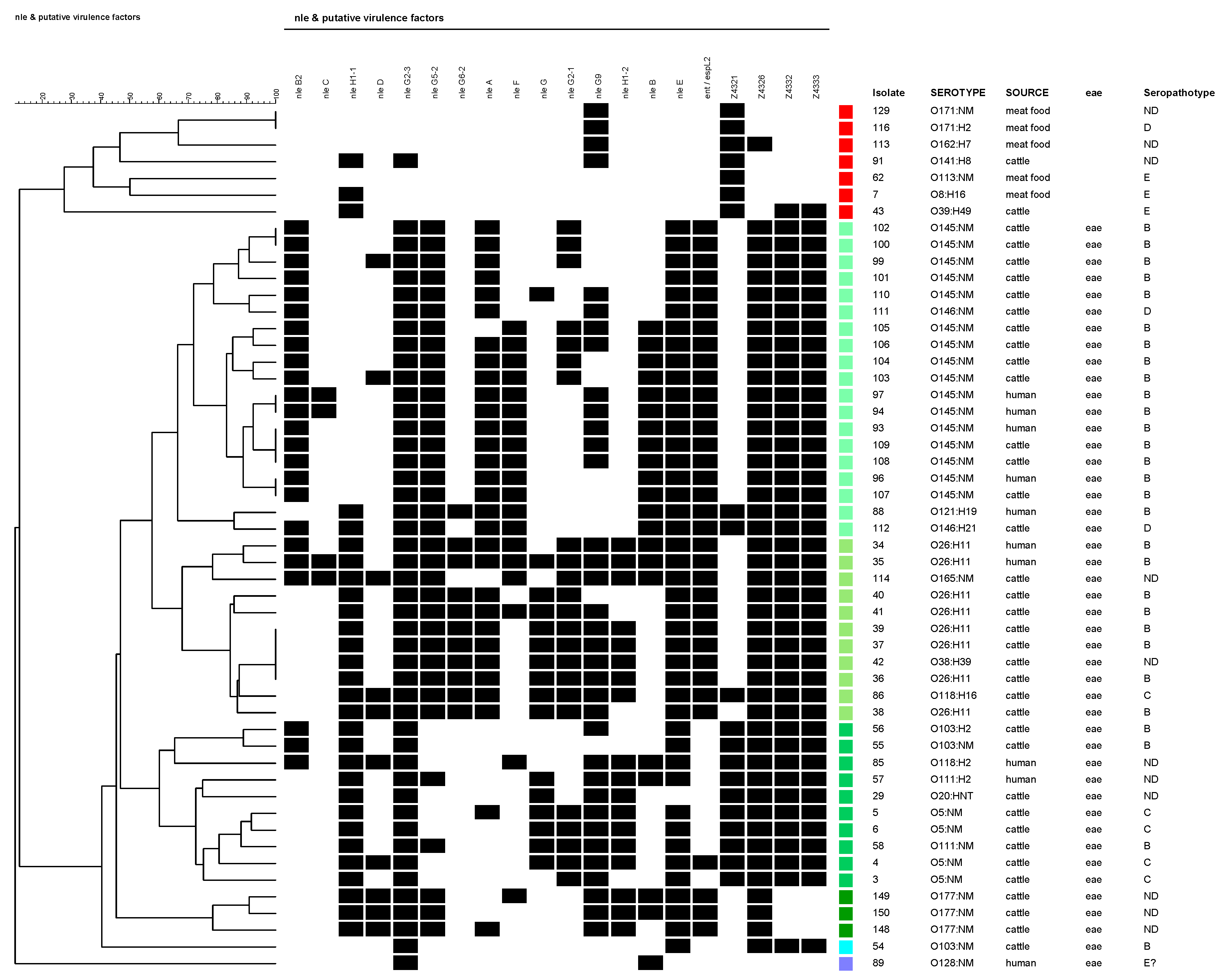


| PAIs | Target | Encoded Protein or Family Effector 1 |
|---|---|---|
| O-Island 36 | nle B2 | Non-LEE encoded type III effector |
| nle C | Immunomodulation, zinc-metalloprotease | |
| nle H1-1 | Immunomodulation | |
| nle D | Immunomodulation, zinc-metalloprotease | |
| O-Island 57 | nle G2-3 | Ubiquitin ligase |
| nle G5-2 | Ubiquitin ligase | |
| nle G6-2 | Ubiquitin ligase | |
| O-Island 71 | nle A | Disruption tight junctions and protein trafficking |
| nle F | Disruption protein trafficking | |
| nle G | Ubiquitin ligase | |
| nle G2-1 | Ubiquitin ligase | |
| nle G9 | Ubiquitin ligase | |
| nle H1-2 | Immunomodulation | |
| O-Island 122 | nle B | Immunomodulation |
| nle E | Immunomodulation | |
| ent/espL2 | Microcolony formation and F-actin aggregation | |
| Z4321 (pagC) | Similarity to Salmonella enterica serovar Typhimurium PhoP-activated gene C | |
| Z4326 (sen) | Similarity to Shigella flexneri enterotoxin 2 | |
| Z4332 (efa1) | EHEC factor for adherence | |
| Z4333 (efa2) | EHEC factor for adherence |
| % Isolates Positive for Gene | ||||||
|---|---|---|---|---|---|---|
| PAI | Gene | O5:NM (n = 4) | O26:H11 (n = 8) | O103 (n = 3) | O145:NM (n = 18) | O177:NM (n = 3) |
| O-Island 36 | nle B2 | 0 | 25 | 66.7 | 88.9 | 0 |
| nle C | 0 | 12.5 | 0 | 11.1 | 0 | |
| nle H1-1 | 100 | 100 | 66.7 | 0 | 100 | |
| nle D | 25 | 12.5 | 0 | 11.1 | 100 | |
| O-Island 57 | nle G2-3 | 100 | 100 | 100 | 94.4 | 100 |
| nle G5-2 | 0 | 100 | 0 | 88.9 | 100 | |
| nle G6-2 | 0 | 100 | 0 | 0 | 0 | |
| O-Island 71 | nle A | 25 | 100 | 0 | 83.3 | 33.3 |
| nle F | 0 | 37.5 | 0 | 61.1 | 33.3 | |
| nle G | 75 | 87.5 | 0 | 5.5 | 0 | |
| nle G2-1 | 100 | 87.5 | 0 | 38.9 | 0 | |
| nle G9 | 100 | 87.5 | 33.3 | 44.4 | 100 | |
| nle H1-2 | 75 | 62.5 | 0 | 0 | 100 | |
| O-Island 122 | nle B | 0 | 25 | 0 | 61.1 | 66.7 |
| nle E | 100 | 100 | 100 | 88.9 | 100 | |
| ent/espL2 | 25 | 100 | 0 | 88.9 | 100 | |
| Z4321 (pagC) | 100 | 0 | 66.7 | 0 | 0 | |
| Z4326 (sen) | 100 | 100 | 100 | 88.9 | 100 | |
| Z4332 (efa1) | 100 | 100 | 100 | 88.9 | 0 | |
| Z4333 (efa2) | 100 | 100 | 100 | 88.9 | 0 | |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cadona, J.S.; Bustamante, A.V.; González, J.; Sanso, A.M. Pathogenicity Islands Distribution in Non-O157 Shiga Toxin-Producing Escherichia coli (STEC). Genes 2018, 9, 81. https://doi.org/10.3390/genes9020081
Cadona JS, Bustamante AV, González J, Sanso AM. Pathogenicity Islands Distribution in Non-O157 Shiga Toxin-Producing Escherichia coli (STEC). Genes. 2018; 9(2):81. https://doi.org/10.3390/genes9020081
Chicago/Turabian StyleCadona, Jimena Soledad, Ana Victoria Bustamante, Juliana González, and Andrea Mariel Sanso. 2018. "Pathogenicity Islands Distribution in Non-O157 Shiga Toxin-Producing Escherichia coli (STEC)" Genes 9, no. 2: 81. https://doi.org/10.3390/genes9020081
APA StyleCadona, J. S., Bustamante, A. V., González, J., & Sanso, A. M. (2018). Pathogenicity Islands Distribution in Non-O157 Shiga Toxin-Producing Escherichia coli (STEC). Genes, 9(2), 81. https://doi.org/10.3390/genes9020081





