Transcriptomic Analysis Provides Insights into Grafting Union Development in Pecan (Carya illinoinensis)
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Material and Grafting Procedures
2.2. RNA Extraction, Library Construction, and Sequencing
2.3. De Novo Assembly and Functional Annotation
2.4. Analysis of Differentially Expressed Genes (DEGs)
2.5. Validation of RNA-Seq Data by Quantitative Real-Time PCR (qRT-PCR)
2.6. Detection of Hormones Content by ELISA
3. Results and Discussion
3.1. De Novo Assembly and Functional Annotation
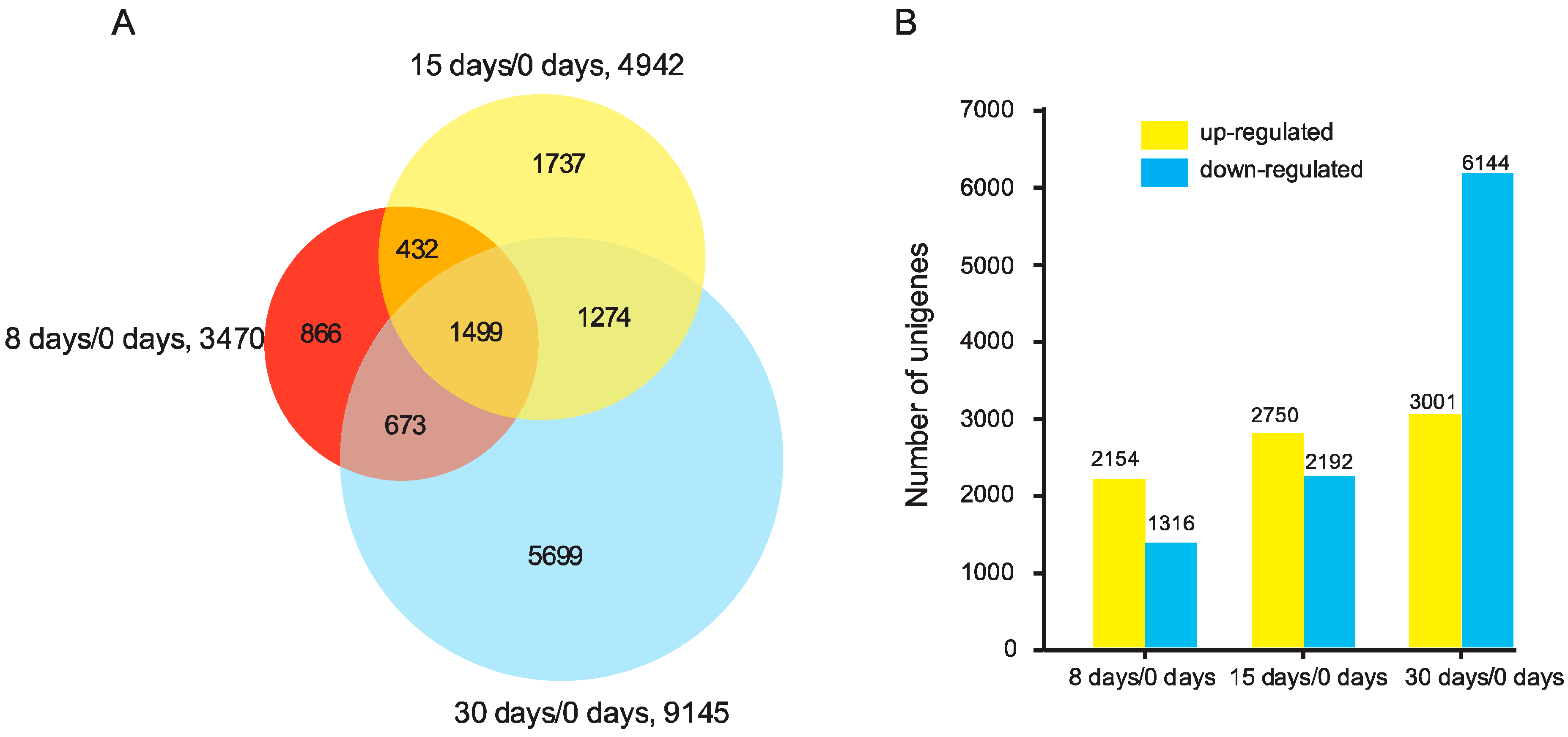
3.2. Analysis of DEGs in the Graft Process of Pecan
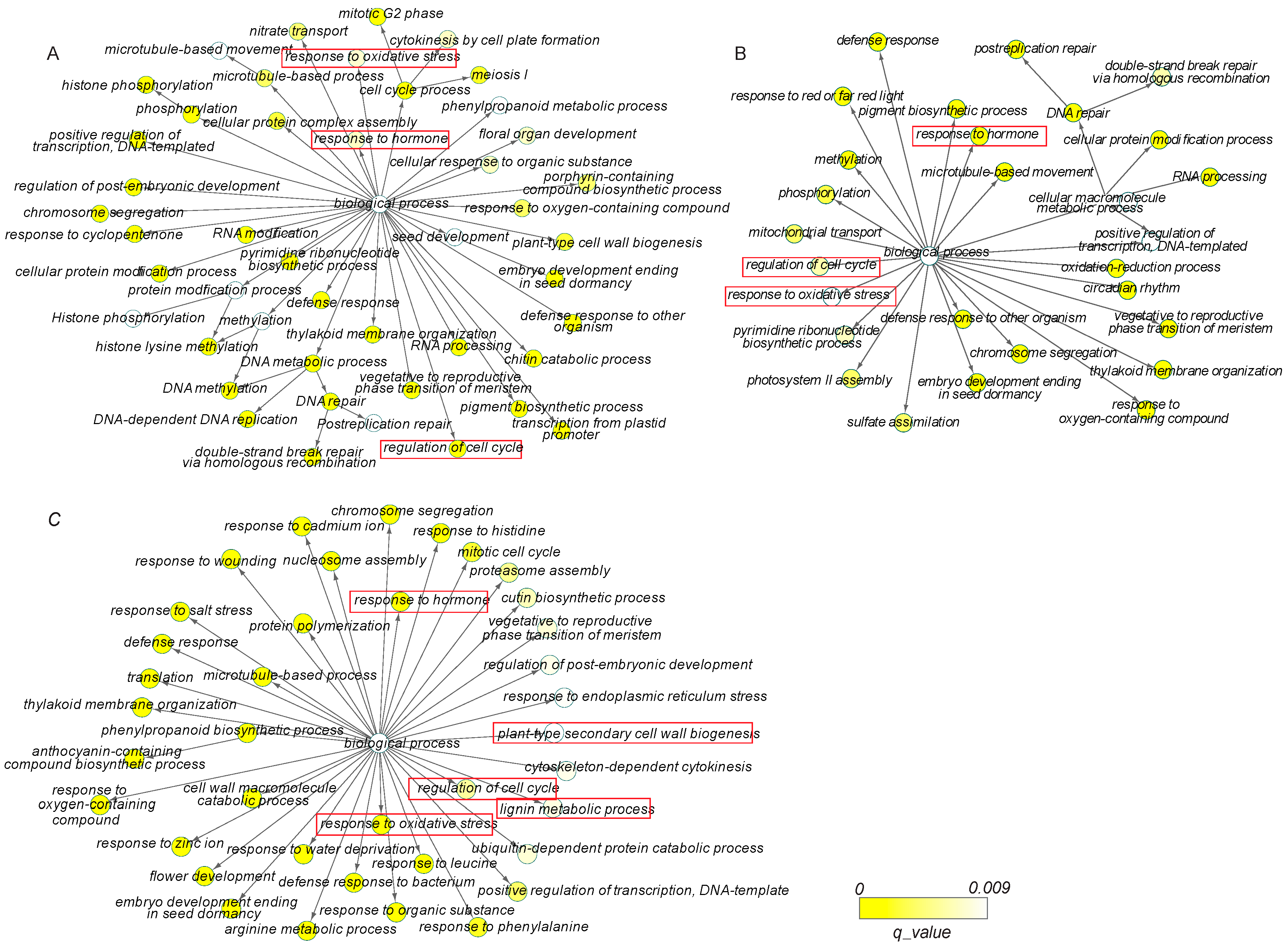
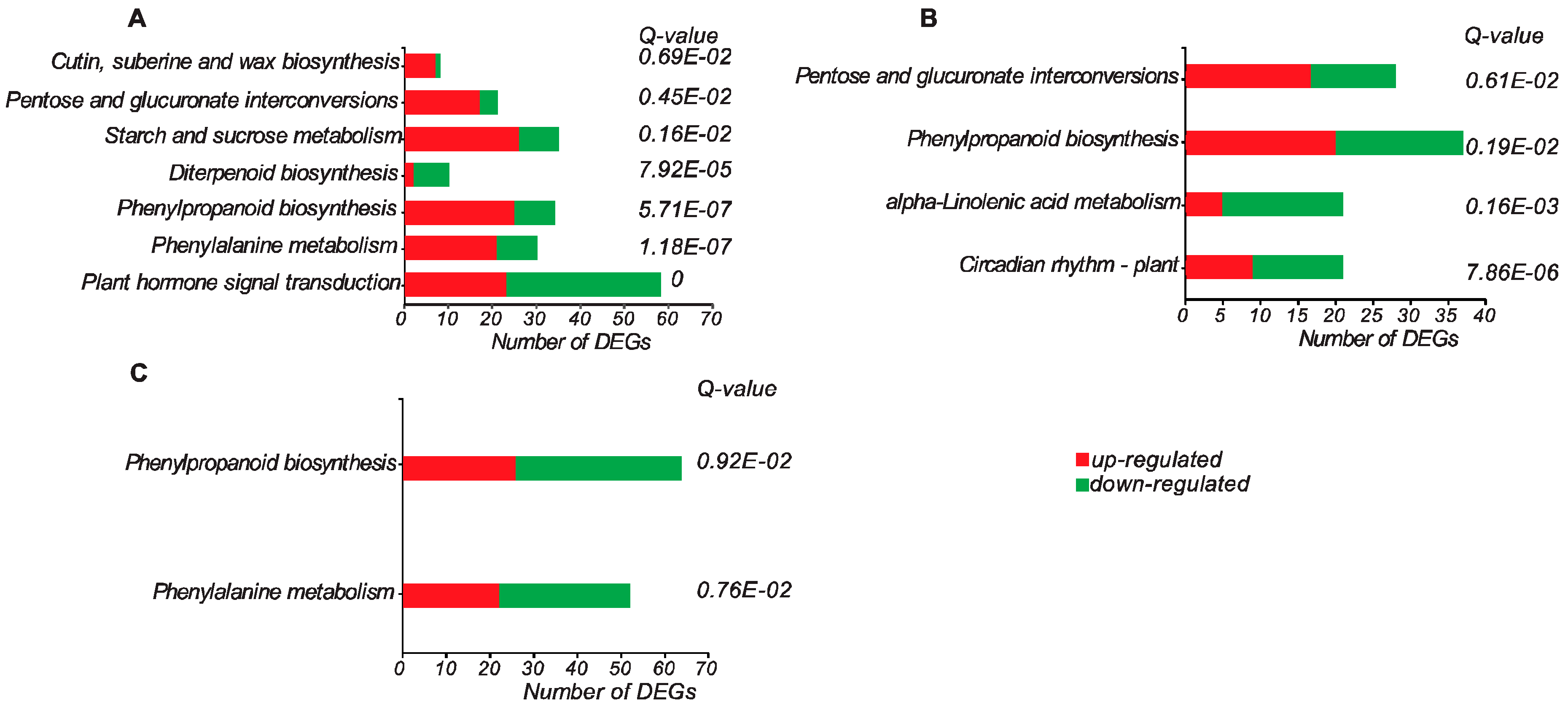
3.3. Gene Ontology and Pathway Enrichment Analyses of DEGs
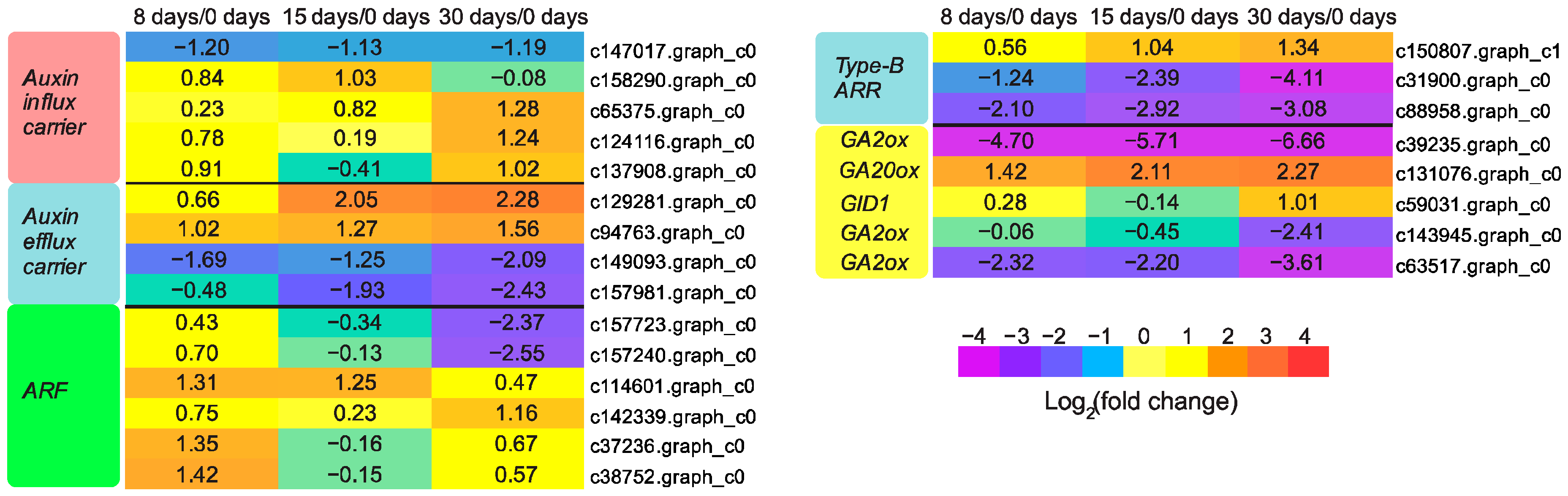
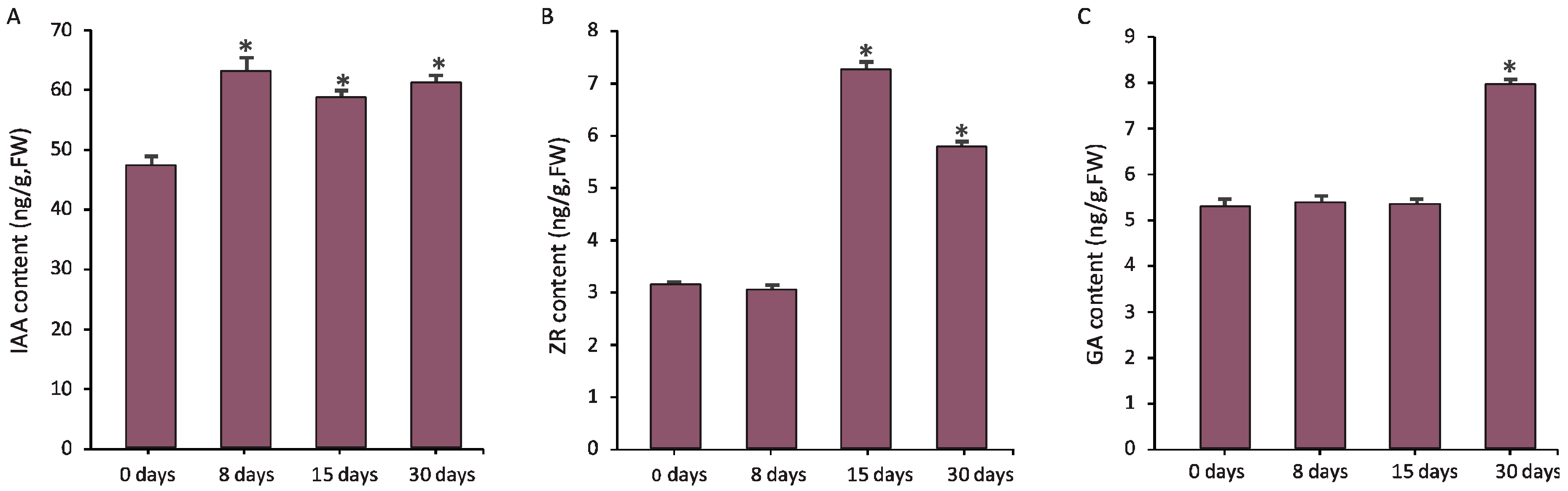
3.4. Hormones Were Critical Regulators for Graft Union Development
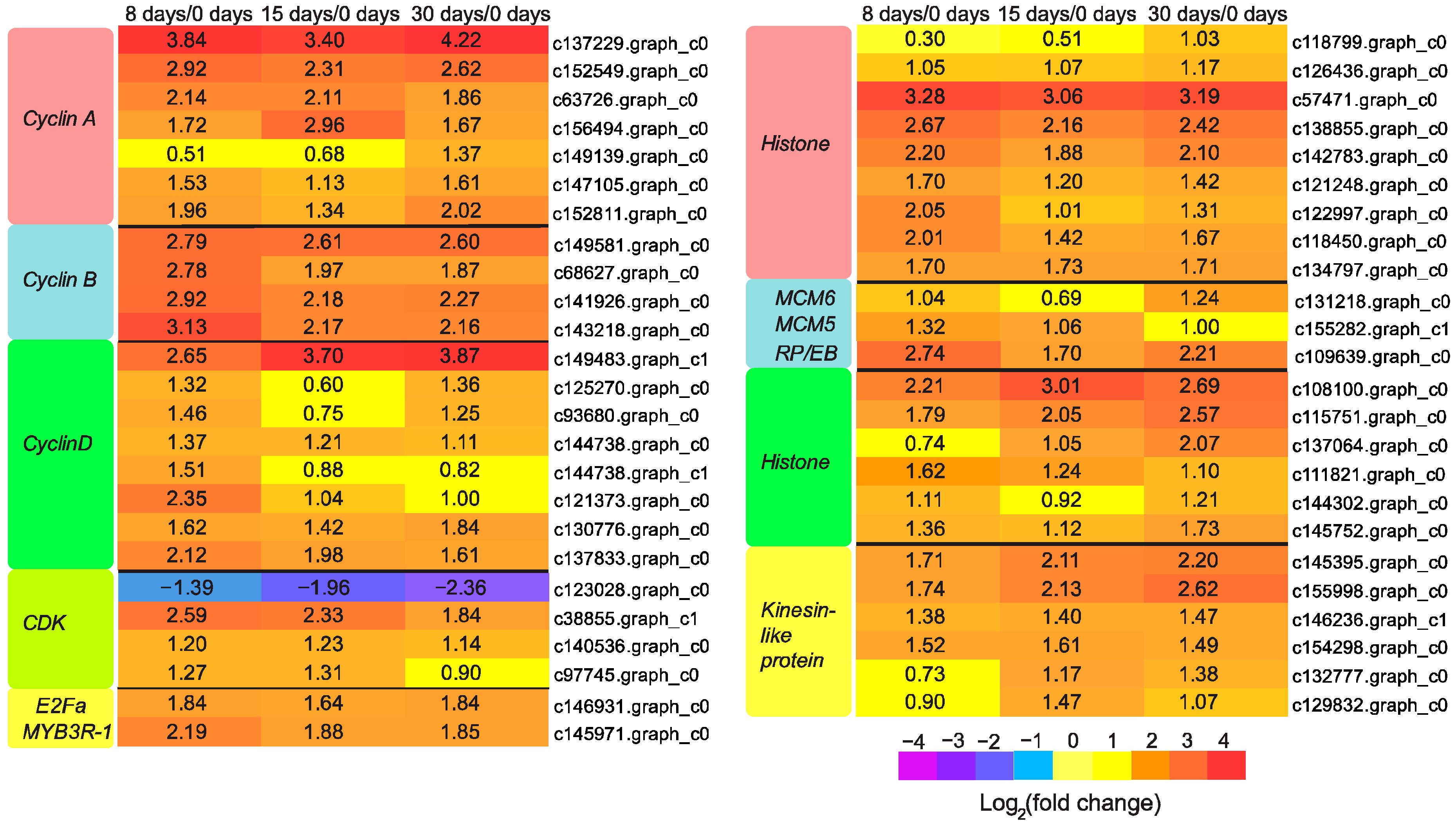
3.5. Genes Responsible for Callus Formation
3.6. Genes Participated in Vascular Bundle Formation
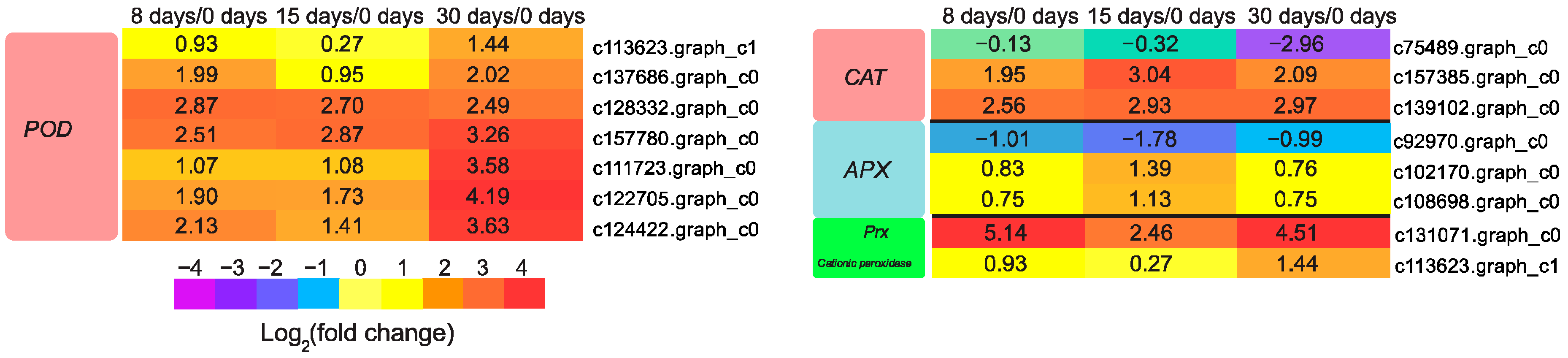
3.7. Genes Involved in Reactive Oxygen Species (ROS) Scavenging
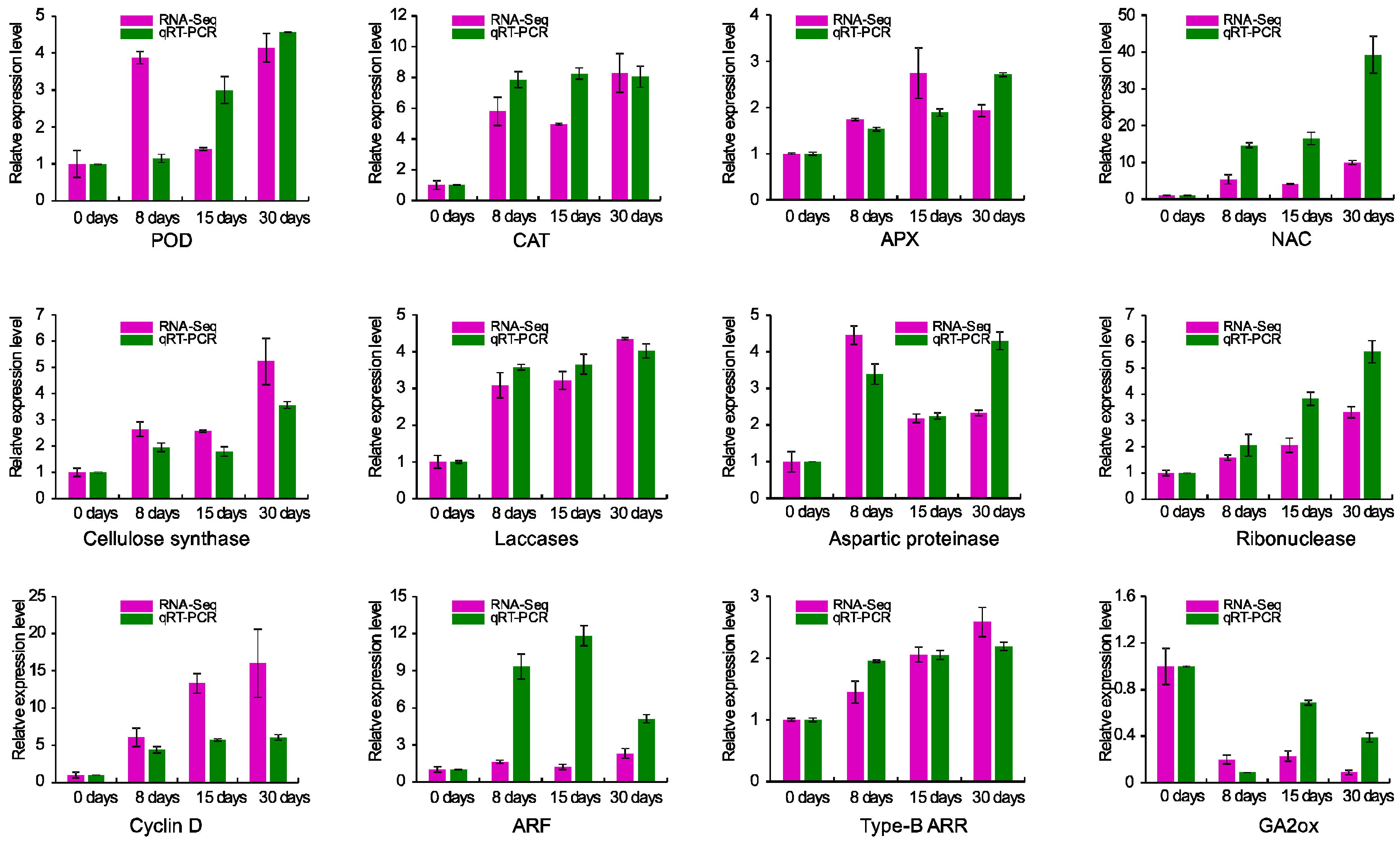
3.8. Validation of RNA-Seq Data by Real-Time RT-PCR (qRT-PCR)
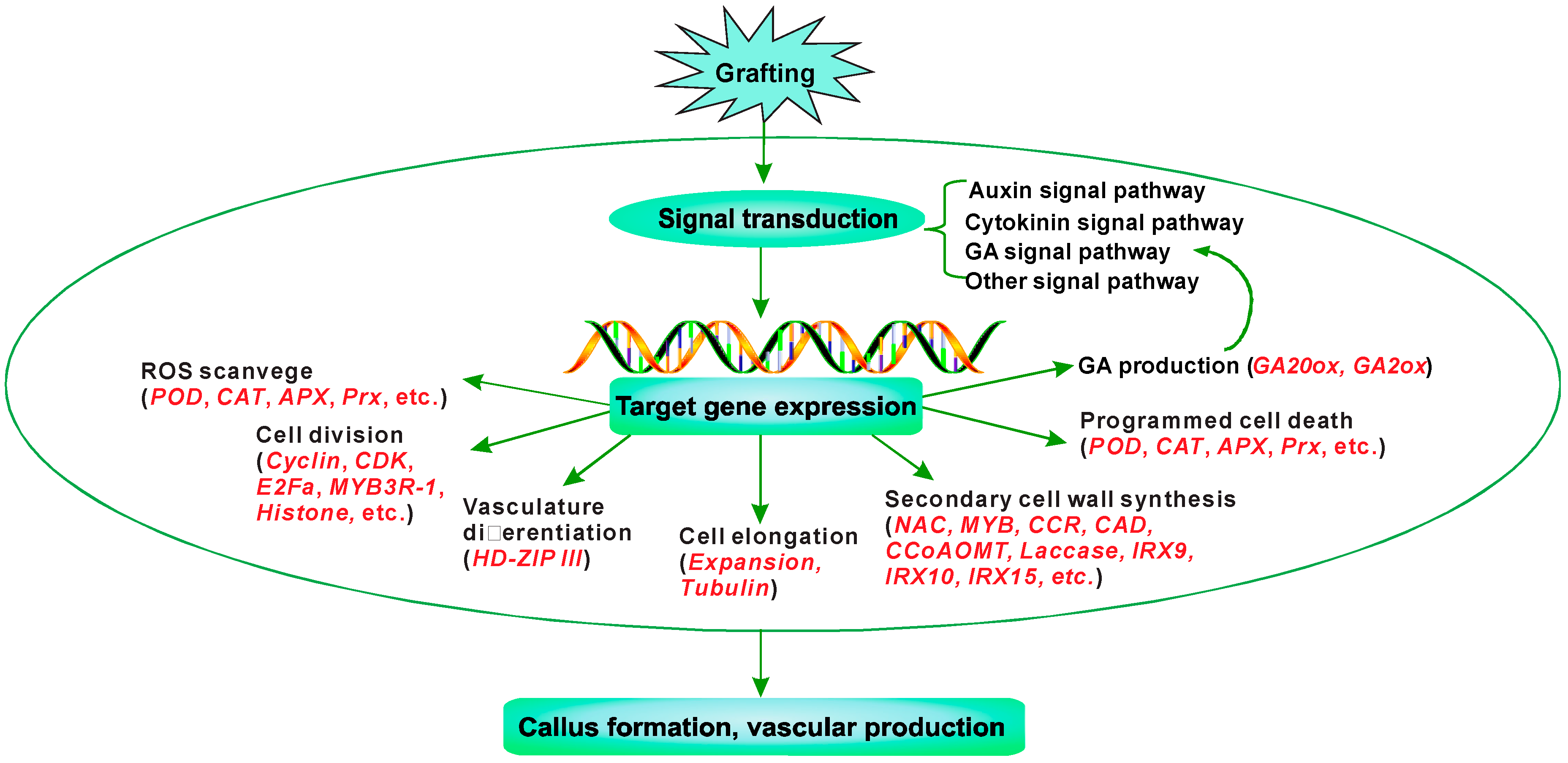
4. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Zhang, R.; Peng, F.; Li, Y. Pecan production in China. Sci. Hortic. 2015, 197, 719–727. [Google Scholar] [CrossRef]
- Asante, A.; Barnett, J. Graft union formation in mango (Mangifera indica L.). J. Hortic. Sci. 1997, 72, 781–790. [Google Scholar] [CrossRef]
- Yang, Z.J.; Feng, J.L.; Chen, H. Study on the anatomical structures in development of the nurse seed grafted union of Camellia oleifera. Plant Sci. J. 2013, 31, 313–320. [Google Scholar] [CrossRef]
- Zheng, B.S.; Chu, H.L.; Jin, S.H.; Huang, Y.J.; Wang, Z.J.; Chen, M.; Huang, J.Q. cDNA-AFLP analysis of gene expression in hickory (Carya carthayensis) during graft process. Tree Physiol. 2009, 30, 297–303. [Google Scholar] [CrossRef] [PubMed]
- Yin, H.; Yan, B.; Sun, J.; Jia, P.; Zhang, Z.; Yan, X.; Chai, J.; Ren, Z.; Zheng, G.; Liu, H. Graft-union development: A delicate process that involves cell–cell communication between scion and stock for local auxin accumulation. J. Exp. Bot. 2012, 63, 4219–4232. [Google Scholar] [CrossRef] [PubMed]
- Cookson, S.J.; Clemente Moreno, M.J.; Hevin, C.; Nyamba Mendome, L.Z.; Delrot, S.; Trossatmagnin, C.; Ollat, N. Graft union formation in grapevine induces transcriptional changes related to cell wall modification, wounding, hormone signalling, and secondary metabolism. J. Exp. Bot. 2013, 64, 2997–3008. [Google Scholar] [CrossRef] [PubMed]
- Cookson, S.J.; Moreno, M.J.C.; Hevin, C.; Mendome, L.Z.N.; Delrot, S.; Magnin, N.; Trossat-Magnin, C.; Ollat, N. Heterografting with nonself rootstocks induces genes involved in stress responses at the graft interface when compared with autografted controls. J. Exp. Bot. 2014, 65, 2473–2481. [Google Scholar] [CrossRef] [PubMed]
- Qiu, L.; Jiang, B.; Fang, J.; Shen, Y.; Fang, Z.; RM, S.K.; Yi, K.; Shen, C.; Yan, D.; Zheng, B. Analysis of transcriptome in hickory (Carya carthayensis), and uncover the dynamics in the hormonal signaling pathway during graft process. BMC Genom. 2016, 17, 935. [Google Scholar] [CrossRef] [PubMed]
- Sima, X.; Jiang, B.; Fang, J.; He, Y.; Fang, Z.; Saravana Kumar, K.M.; Ren, W.; Qiu, L.; Chen, X.; Zheng, B. Identification by deep sequencing and profiling of conserved and novel hickory micrornas involved in the graft process. Plant Biotechnol. Rep. 2015, 9, 115–124. [Google Scholar] [CrossRef]
- Xu, D.; Yuan, H.; Tong, Y.; Zhao, L.; Qiu, L.; Guo, W.; Shen, C.; Liu, H.; Yan, D.; Zheng, B. Comparative proteomic analysis of the graft unions in hickory (Carya carthayensis) provides insights into response mechanisms to grafting process. Front. Plant Sci. 2017, 8. [Google Scholar] [CrossRef] [PubMed]
- Mo, Z.; He, H.; Su, W.; Peng, F. Analysis of differentially accumulated proteins associated with graft union formation in pecan (Carya illinoensis). Sci. Hortic. 2017, 224, 126–134. [Google Scholar] [CrossRef]
- Yang, Y.M.; Xu, C.N.; Wang, B.M.; Jia, J.Z. Effects of plant growth regulators on secondary wall thickening of cotton fibres. Plant Growth Regul. 2001, 35, 233–237. [Google Scholar]
- Pina, A.; Errea, P. Differential induction of phenylalanine ammonia-lyase gene expression in response to in vitro callus unions of Prunus spp. J. Plant Physiol. 2008, 165, 705–714. [Google Scholar] [CrossRef] [PubMed]
- Melnyk, C.W.; Schuster, C.; Leyser, O.; Meyerowitz, E.M. A developmental framework for graft formation and vascular reconnection in Arabidopsis thaliana. Curr. Biol. 2015, 25, 1306–1318. [Google Scholar] [CrossRef] [PubMed]
- Pitaksaringkarn, W.; Ishiguro, S.; Asahina, M.; Satoh, S. ARF6 and ARF8 contribute to tissue reunion in incised Arabidopsis inflorescence stems. Plant Biotechnol. 2014, 31, 49–53. [Google Scholar] [CrossRef]
- Li, S.B.; Xie, Z.Z.; Hu, C.G.; Zhang, J.Z. A review of auxin response factors (ARFs) in plants. Front. Plant Sci. 2016, 7. [Google Scholar] [CrossRef] [PubMed]
- Mattsson, J.; Ckurshumova, W.; Berleth, T. Auxin signaling in Arabidopsis leaf vascular development. Plant Physiol. 2003, 131, 1327–1339. [Google Scholar] [CrossRef] [PubMed]
- Hardtke, C.S.; Berleth, T. The Arabidopsis gene MONOPTEROS encodes a transcription factor mediating embryo axis formation and vascular development. EMBO J. 1998, 17, 1405–1411. [Google Scholar] [CrossRef] [PubMed]
- Aloni, R.; Aloni, E.; Langhans, M.; Ullrich, C. Role of cytokinin and auxin in shaping root architecture: Regulating vascular differentiation, lateral root initiation, root apical dominance and root gravitropism. Ann. Bot. 2006, 97, 883–893. [Google Scholar] [CrossRef] [PubMed]
- Ikeuchi, M.; Sugimoto, K.; Iwase, A. Plant callus: Mechanisms of induction and repression. Plant Cell 2013, 25, 3159–3173. [Google Scholar] [CrossRef] [PubMed]
- Nieminen, K.; Immanen, J.; Laxell, M.; Kauppinen, L.; Tarkowski, P.; Dolezal, K.; Tähtiharju, S.; Elo, A.; Decourteix, M.; Ljung, K. Cytokinin signaling regulates cambial development in poplar. Proc. Natl. Acad. Sci. USA 2008, 105, 20032–20037. [Google Scholar] [CrossRef] [PubMed]
- Hejátko, J.; Ryu, H.; Kim, G.T.; Dobešová, R.; Choi, S.; Choi, S.M.; Souček, P.; Horák, J.; Pekárová, B.; Palme, K. The histidine kinases CYTOKININ-INDEPENDENT1 and ARABIDOPSIS HISTIDINE KINASE2 and 3 regulate vascular tissue development in Arabidopsis shoots. Plant Cell 2009, 21, 2008–2021. [Google Scholar] [CrossRef] [PubMed]
- Del Pozo, J.C.; Lopez-Matas, M.; Ramirez-Parra, E.; Gutierrez, C. Hormonal control of the plant cell cycle. Physiol. Plant. 2005, 123, 173–183. [Google Scholar] [CrossRef]
- Perrot-Rechenmann, C. Cellular responses to auxin: Division versus expansion. Cold Spring Harb. Perspect. Biol. 2010, 2. [Google Scholar] [CrossRef] [PubMed]
- Pils, B.; Heyl, A. Unraveling the evolution of cytokinin signaling. Plant Physiol. 2009, 151, 782–791. [Google Scholar] [CrossRef] [PubMed]
- Mason, M.G.; Mathews, D.E.; Argyros, D.A.; Maxwell, B.B.; Kieber, J.J.; Alonso, J.M.; Ecker, J.R.; Schaller, G.E. Multiple type-B response regulators mediate cytokinin signal transduction in Arabidopsis. Plant Cell 2005, 17, 3007–3018. [Google Scholar] [CrossRef] [PubMed]
- Sakai, H.; Honma, T.; Aoyama, T.; Sato, S.; Kato, T.; Tabata, S.; Oka, A. ARR1, a transcription factor for genes immediately responsive to cytokinins. Science 2001, 294, 1519–1521. [Google Scholar] [CrossRef] [PubMed]
- Mauriat, M.; Moritz, T. Analyses of GA20ox- and GID1-over-expressing aspen suggest that gibberellins play two distinct roles in wood formation. Plant J. 2009, 58, 989–1003. [Google Scholar] [CrossRef] [PubMed]
- Ragni, L.; Nieminen, K.; Pacheco-Villalobos, D.; Sibout, R.; Schwechheimer, C.; Hardtke, C.S. Mobile gibberellin directly stimulates Arabidopsis hypocotyl xylem expansion. Plant Cell 2011, 23, 1322. [Google Scholar] [CrossRef] [PubMed]
- Guo, H.; Wang, Y.; Liu, H.; Hu, P.; Jia, Y.; Zhang, C.; Wang, Y.; Gu, S.; Yang, C.; Wang, C. Exogenous GA3 application enhances xylem development and induces the expression of secondary wall biosynthesis related genes in Betula platyphylla. Int. J. Mol. Sci. 2015, 16, 22960–22975. [Google Scholar] [CrossRef] [PubMed]
- Israelsson, M.; Sundberg, B.; Moritz, T. Tissue-specific localization of gibberellins and expression of gibberellin-biosynthetic and signaling genes in wood-forming tissues in aspen. Plant J. 2005, 44, 494–504. [Google Scholar] [CrossRef] [PubMed]
- Pina, A.; Errea, P. A review of new advances in mechanism of graft compatibility–incompatibility. Sci. Hortic. 2005, 106, 1–11. [Google Scholar] [CrossRef]
- Chen, C.C.; Fu, S.F.; Lee, Y.I.; Lin, C.Y.; Lin, W.C.; Huang, H.J. Transcriptome analysis of age-related gain of callus-forming capacity in Arabidopsis hypocotyls. Plant Cell Physiol. 2012, 53, 1457–1469. [Google Scholar] [CrossRef] [PubMed]
- Xu, K.; Liu, J.; Fan, M.; Xin, W.; Hu, Y.; Xu, C. A genome-wide transcriptome profiling reveals the early molecular events during callus initiation in Arabidopsis multiple organs. Genomics 2012, 100, 116–124. [Google Scholar] [CrossRef] [PubMed]
- Fehér, A.; Magyar, Z. Coordination of cell division and differentiation in plants in comparison to animals. Acta Biol. Szeged. 2015, 59, 275–289. [Google Scholar]
- Riou-Khamlichi, C.; Huntley, R.; Jacqmard, A.; Murray, J.A. Cytokinin activation of Arabidopsis cell division through a D-type cyclin. Science 1999, 283, 1541–1544. [Google Scholar] [CrossRef] [PubMed]
- Cockcroft, C.E.; den Boer, B.G.; Healy, J.S.; Murray, J.A. Cyclin D control of growth rate in plants. Nature 2000, 405, 575–579. [Google Scholar] [PubMed]
- De, V.L.; Beeckman, T.; Beemster, G.T.; De, A.E.J.; Ormenese, S.; Maes, S.; Naudts, M.; Van, D.S.E.; Jacqmard, A.; Engler, G. Control of proliferation, endoreduplication and differentiation by the Arabidopsis E2Fa-DPa transcription factor. EMBO J. 2002, 21, 1360–1368. [Google Scholar]
- Magyar, Z.; Bögre, L.; Ito, M. Dreams make plant cells to cycle or to become quiescent. Curr. Opin. Plant Biol. 2016, 34, 100–106. [Google Scholar] [CrossRef] [PubMed]
- Ye, Z.; Zhong, R. Molecular control of wood formation in trees. J. Exp. Bot. 2015, 66, 4119–4131. [Google Scholar] [CrossRef] [PubMed]
- Du, J.; Miura, E.; Robischon, M.; Martinez, C.; Groover, A. The Populus Class III HD ZIP transcription factor Popcorona affects cell differentiation during secondary growth of woody stems. PLoS ONE 2011, 6, e17458. [Google Scholar] [CrossRef] [PubMed]
- Ilegems, M.; Douet, V.; Meylanbettex, M.; Uyttewaal, M.; Brand, L.; Bowman, J.L.; Stieger, P.A. Interplay of auxin, KANADI and Class III HD-ZIP transcription factors in vascular tissue formation. Development 2010, 137, 975–984. [Google Scholar] [CrossRef] [PubMed]
- De, R.B.; Mähönen, A.P.; Helariutta, Y.; Weijers, D. Plant vascular development: From early specification to differentiation. Nat. Rev. Mol. Cell Biol. 2015, 17, 30–40. [Google Scholar]
- Ye, Z.H. Vascular tissue differentiation and pattern formation in plants. Annu. Rev. Plant Biol. 2002, 53, 183–202. [Google Scholar] [CrossRef] [PubMed]
- Mendu, N.; Silflow, C.D. Elevated levels of tubulin transcripts accompany the GA3-induced elongation of oat internode segments. Plant Cell Physiol. 1993, 34, 973–983. [Google Scholar]
- Suzuki, S.; Li, L.; Sun, Y.H.; Chiang, V.L. The cellulose synthase gene superfamily and biochemical functions of xylem-specific cellulose synthase-like genes in Populus trichocarpa. Plant Physiol. 2006, 142, 1233–1245. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Q.; Nakashima, J.; Chen, F.; Yin, Y.; Fu, C.; Yun, J.; Shao, H.; Wang, X.; Wang, Z.-Y.; Dixon, R.A. Laccase is necessary and nonredundant with peroxidase for lignin polymerization during vascular development in Arabidopsis. Plant Cell 2013, 25, 3976–3987. [Google Scholar] [CrossRef] [PubMed]
- Berthet, S.; Demont-Caulet, N.; Pollet, B.; Bidzinski, P.; Cézard, L.; Le Bris, P.; Borrega, N.; Hervé, J.; Blondet, E.; Balzergue, S. Disruption of Laccase4 and 17 results in tissue-specific alterations to lignification of Arabidopsis thaliana stems. Plant Cell 2011, 23, 1124–1137. [Google Scholar] [CrossRef] [PubMed]
- Zhong, R.; Lee, C.; Zhou, J.; McCarthy, R.L.; Ye, Z.-H. A battery of transcription factors involved in the regulation of secondary cell wall biosynthesis in Arabidopsis. Plant Cell 2008, 20, 2763–2782. [Google Scholar] [CrossRef] [PubMed]
- Mitsuda, N.; Iwase, A.; Yamamoto, H.; Yoshida, M.; Seki, M.; Shinozaki, K.; Ohme-Takagi, M. NAC transcription factors, NST1 and NST3, are key regulators of the formation of secondary walls in woody tissues of Arabidopsis. Plant Cell 2007, 19, 270–280. [Google Scholar] [CrossRef] [PubMed]
- Zhou, J.; Zhong, R.; Ye, Z. Arabidopsis NAC domain proteins, VND1 to VND5, are transcriptional regulators of secondary wall biosynthesis in vessels. PLoS ONE 2014, 9, e105726. [Google Scholar] [CrossRef] [PubMed]
- Stracke, R.; Werber, M.; Weisshaar, B. The R2R3-MYB gene family in Arabidopsis thaliana. Curr. Opin. Plant Biol. 2001, 4, 447–456. [Google Scholar] [CrossRef]
- Yang, J.H.; Wang, H. Molecular mechanisms for vascular development and secondary cell wall formation. Front. Plant Sci. 2016, 7. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Q.; Dixon, R.A. Transcriptional networks for lignin biosynthesis: More complex than we thought? Trends Plant Sci. 2011, 16, 227–233. [Google Scholar] [CrossRef] [PubMed]
- Tang, X.; Zhuang, Y.; Qi, G.; Wang, D.; Liu, H.; Wang, K.; Chai, G.; Zhou, G. Poplar PdMYB221 is involved in the direct and indirect regulation of secondary wall biosynthesis during wood formation. Sci. Rep. 2015, 5. [Google Scholar] [CrossRef] [PubMed]
- Iliev, I.; Savidge, R. Proteolytic activity in relation to seasonal cambial growth and xylogenesis in Pinus banksiana. Phytochemistry 1999, 50, 953–960. [Google Scholar] [CrossRef]
- Courtois-Moreau, C.L.; Pesquet, E.; Sjödin, A.; Muñiz, L.; Bollhöner, B.; Kaneda, M.; Samuels, L.; Jansson, S.; Tuominen, H. A unique program for cell death in xylem fibers of Populus stem. Plant J. 2009, 58, 260–274. [Google Scholar] [CrossRef] [PubMed]
- Iakimova, E.T.; Woltering, E.J. Xylogenesis in zinnia (Zinnia elegans) cell cultures: Unravelling the regulatory steps in a complex developmental programmed cell death event. Planta 2017, 245, 681–705. [Google Scholar] [CrossRef] [PubMed]
- Petzold, H.E.; Zhao, M.; Beers, E.P. Expression and functions of proteases in vascular tissues. Physiol. Plant. 2012, 145, 121–129. [Google Scholar] [CrossRef] [PubMed]
- Ślesak, I.; Ślesak, H.; Libik, M.; Miszalski, Z. Antioxidant response system in the short-term post-wounding effect in Mesembryanthemum crystallinum leaves. J. Plant Physiol. 2008, 165, 127–137. [Google Scholar] [CrossRef] [PubMed]
- Aloni, B.; Karni, L.; Deventurero, G.; Levin, Z.; Cohen, R.; Katzir, N.; Lotan-Pompan, M.; Edelstein, M.; Aktas, H.; Turhan, E. Physiological and biochemical changes at the rootstock-scion interface in graft combinations between Cucurbita rootstocks and a melon scion. J. Hortic. Sci. Biotech. 2008, 83, 777–783. [Google Scholar] [CrossRef]
- Irisarri, P.; Binczycki, P.; Errea, P.; Martens, H.J.; Pina, A. Oxidative stress associated with rootstock–scion interactions in pear/quince combinations during early stages of graft development. J. Plant Physiol. 2015, 176, 25–35. [Google Scholar] [CrossRef] [PubMed]
- Xu, Q.; Guo, S.R.; Li, H.; Du, N.S.; Shu, S.; Sun, J. Physiological aspects of compatibility and incompatibility in grafted cucumber seedlings. J. Am. Soc. Hort. Sci. 2015, 140, 299–307. [Google Scholar]


| Transcript | Unigene | |
|---|---|---|
| Total number | 140,455 | 83,693 |
| Total length | 165,440,800 bp | 74,679,367 np |
| N50 length | 1905 bp | 1350 bp |
| Mean length | 1178 bp | 892 bp |
| 200–300 bp | 18,665 (13.29%) | 15,637 (18.68%) |
| 300–500 bp | 28,705 (20.44%) | 21,661 (25.88%) |
| 500–1000 bp | 36,770 (26.18%) | 24,333 (29.07%) |
| 1000–2000 bp | 31,127 (22.16%) | 13,698 (16.37%) |
| 2000+ bp | 25,188 (17.93%) | 8364 (9.99%) |
| Annotated Databases | Unigene Number | Percentage (%) | 300 nt ≤ Length < 1000 nt | Length ≥ 1000 nt |
|---|---|---|---|---|
| COG | 11,762 | 14.05 | 3747 | 6644 |
| GO | 23,260 | 27.79 | 9028 | 10,879 |
| KEGG | 13,859 | 16.56 | 5567 | 6557 |
| KOG | 21,612 | 25.82 | 8184 | 10,759 |
| Pfam | 25,909 | 30.96 | 8838 | 14,658 |
| Swiss-Prot | 23,243 | 27.77 | 8711 | 12,071 |
| NR | 38,793 | 46.35 | 15,816 | 17,494 |
| Annotated in at least one database | 40,069 | 47.88 | 16,463 | 17,751 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mo, Z.; Feng, G.; Su, W.; Liu, Z.; Peng, F. Transcriptomic Analysis Provides Insights into Grafting Union Development in Pecan (Carya illinoinensis). Genes 2018, 9, 71. https://doi.org/10.3390/genes9020071
Mo Z, Feng G, Su W, Liu Z, Peng F. Transcriptomic Analysis Provides Insights into Grafting Union Development in Pecan (Carya illinoinensis). Genes. 2018; 9(2):71. https://doi.org/10.3390/genes9020071
Chicago/Turabian StyleMo, Zhenghai, Gang Feng, Wenchuan Su, Zhuangzhuang Liu, and Fangren Peng. 2018. "Transcriptomic Analysis Provides Insights into Grafting Union Development in Pecan (Carya illinoinensis)" Genes 9, no. 2: 71. https://doi.org/10.3390/genes9020071
APA StyleMo, Z., Feng, G., Su, W., Liu, Z., & Peng, F. (2018). Transcriptomic Analysis Provides Insights into Grafting Union Development in Pecan (Carya illinoinensis). Genes, 9(2), 71. https://doi.org/10.3390/genes9020071





