Dietary Fiber Treatment Corrects the Composition of Gut Microbiota, Promotes SCFA Production, and Suppresses Colon Carcinogenesis
Abstract
1. Introduction
2. Materials and Methods
2.1. Animals
2.2. Histology and Tissue Staining
2.3. Western Blot
2.4. Microbiota Profiling and Bioinformatics Analysis
2.5. SCFA Determination
2.6. Statistical Methods
3. Results
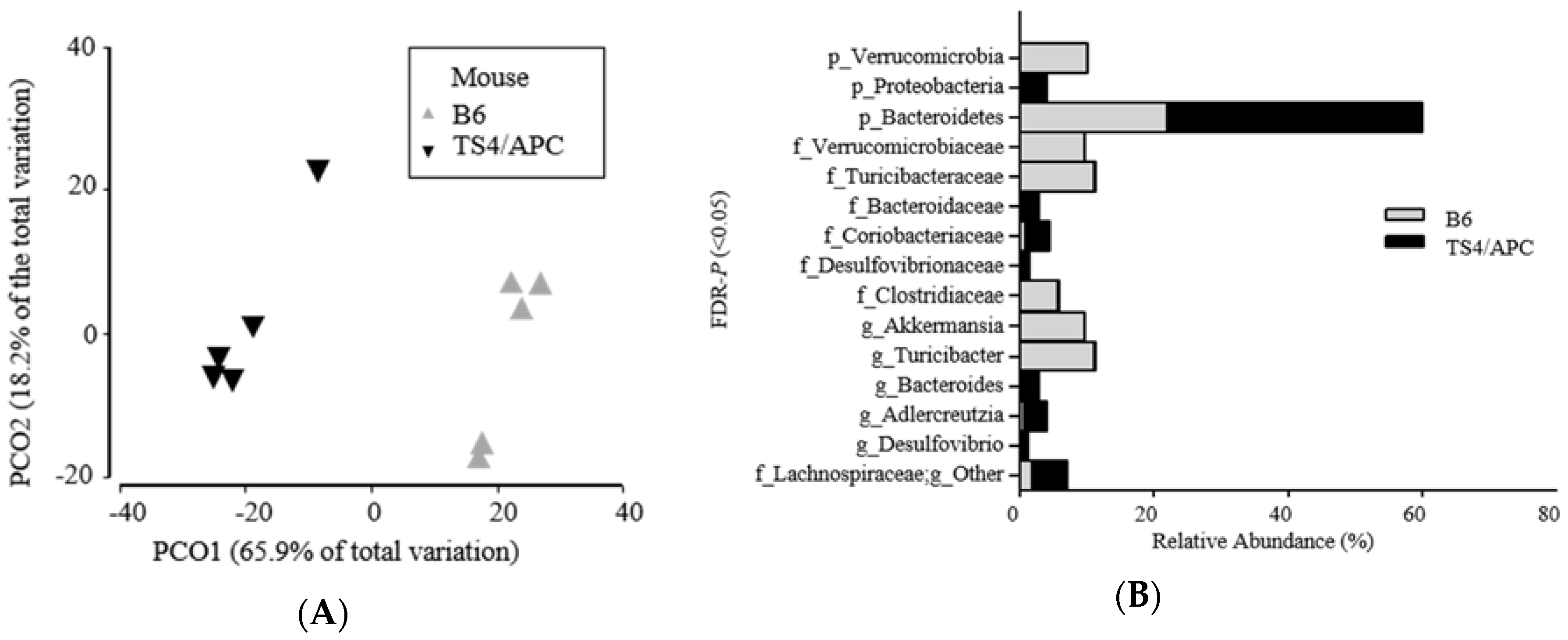
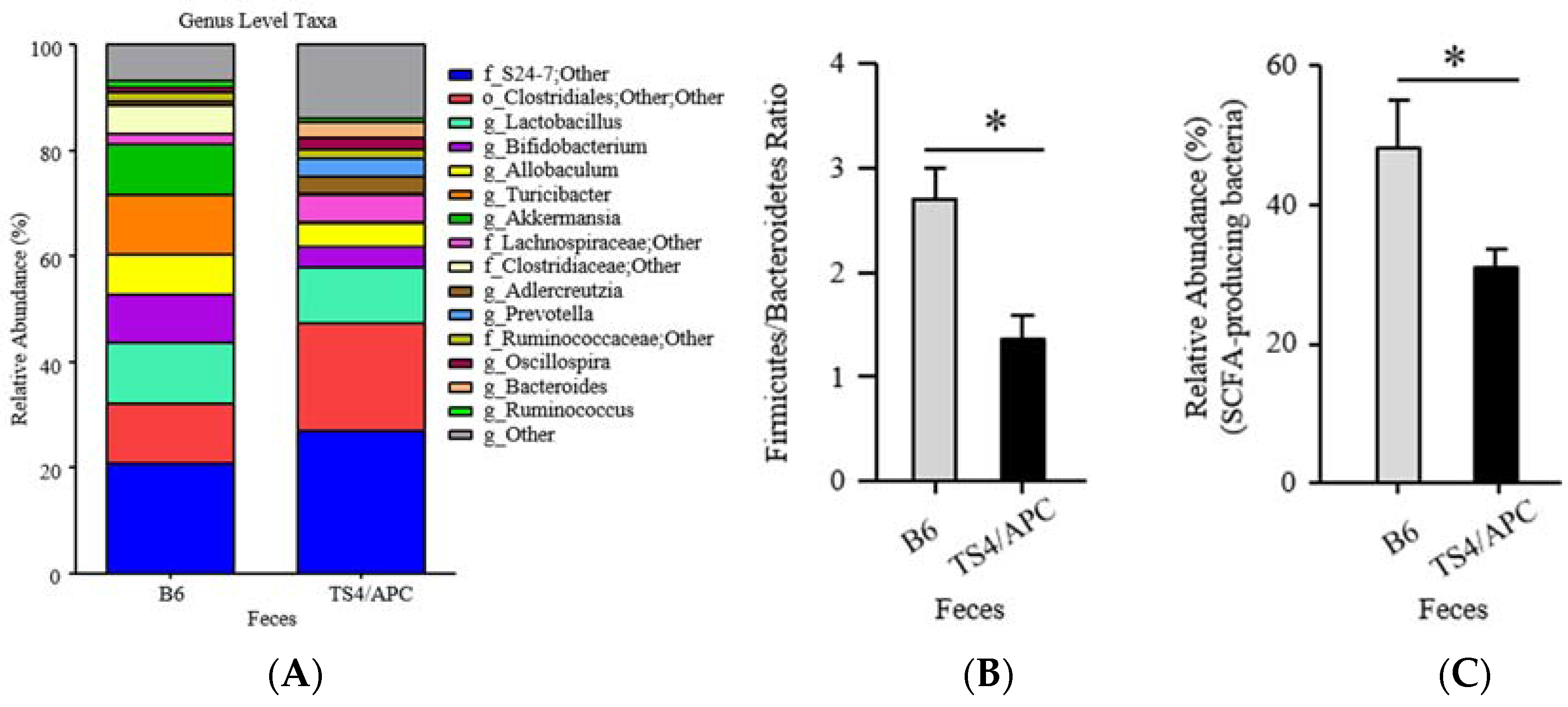
3.1. Polyposis Is Associated with Change in Fecal Microbiota
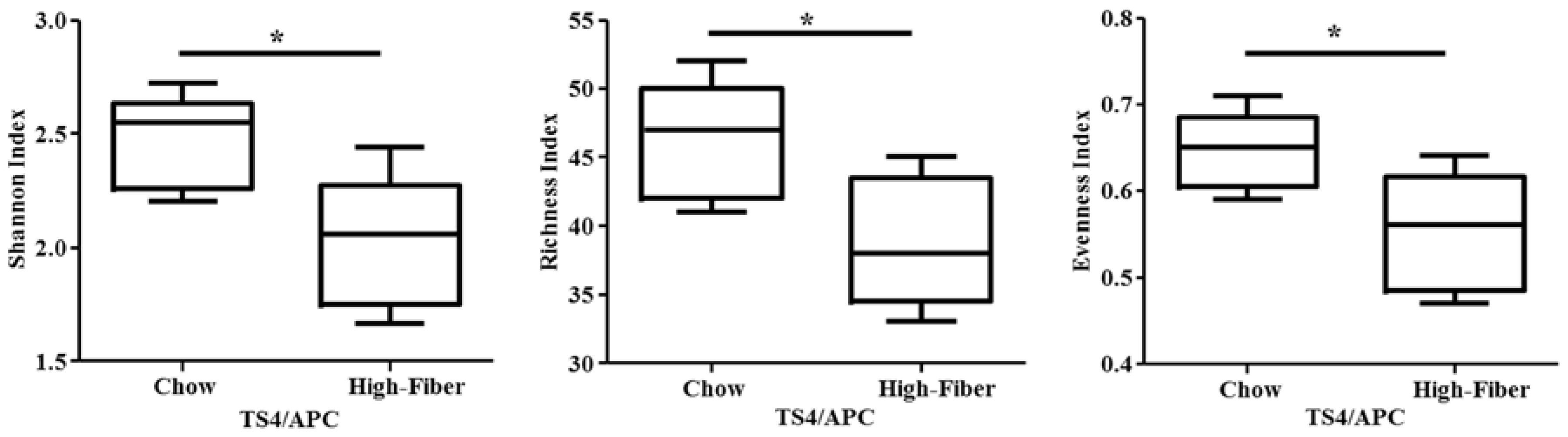
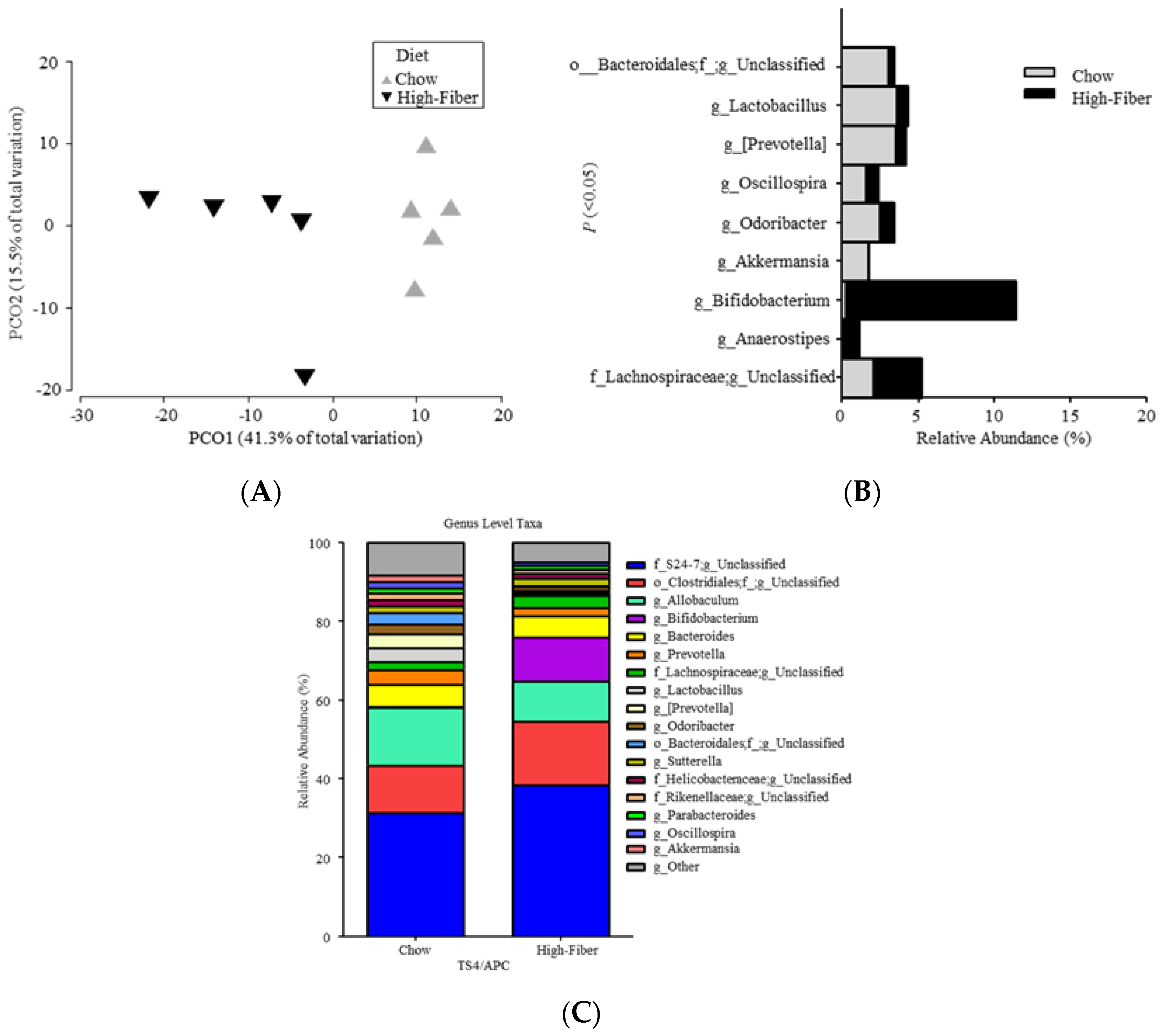
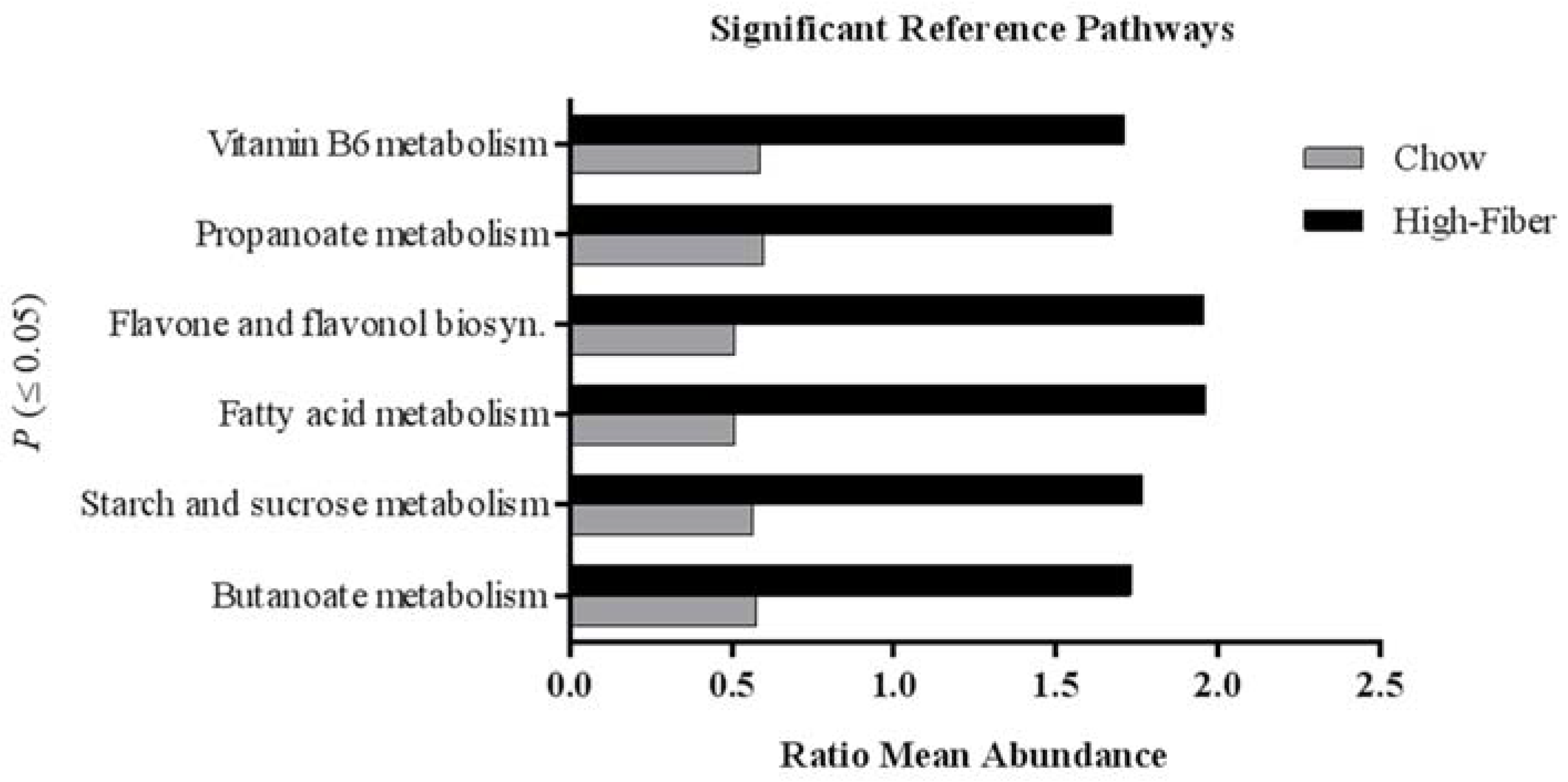
3.2. High Fiber Treatment Alters the Microbiota Composition in Polyposis Mice
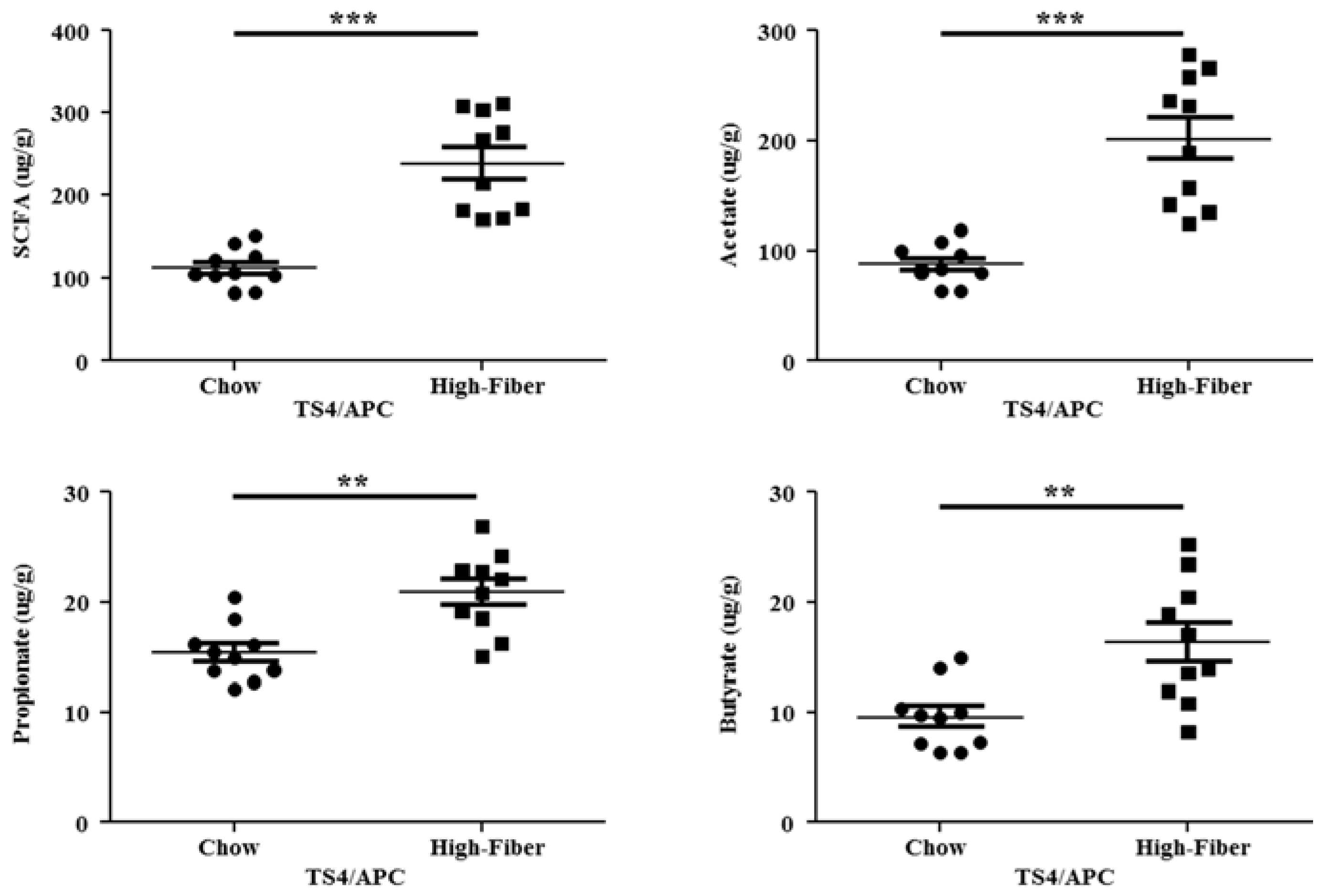
3.3. High Fiber Treatment Increased the Short Chain Fatty Acid Production in Polyposis Mice
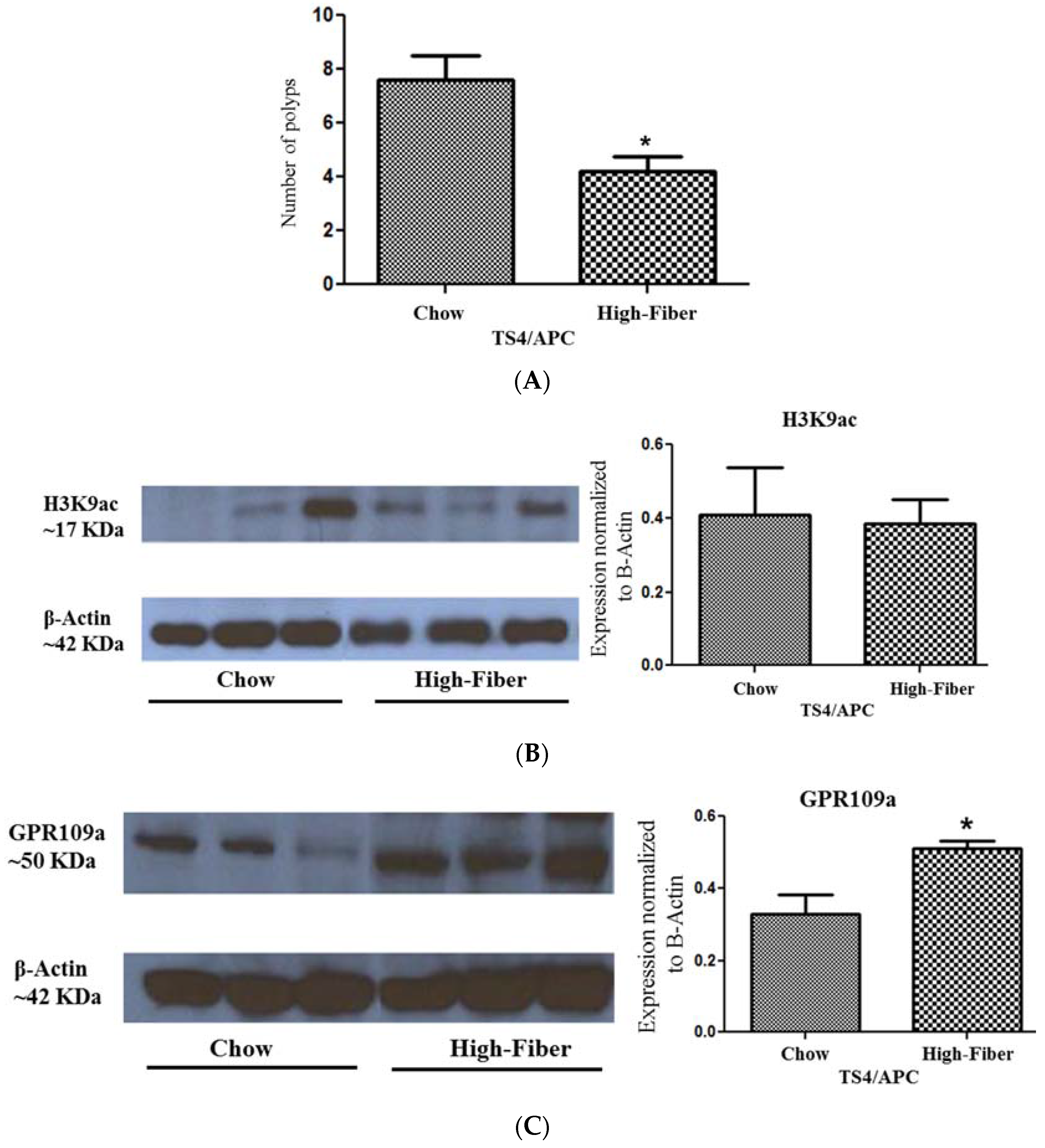
3.4. Increased Short Chain Fatty Acid Is Associated with Improved Polyposis
4. Discussion
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- World Cancer Research Fund/American Institute for Cancer Research. Policy and Action for Cancer Prevention. Food, Nutrition, and Physical Activity: A Global Perspective; AICR: Washington, DC, USA, 2009; p. 201. Available online: http://www.wcrf-hk.org/sites/default/files/Policy_Report.pdf (accessed on 1 December 2017).
- American Cancer Society. Cancer Facts & Figures 2012; ACS: Atlanta, GA, USA, 2012; p. 68. Available online: https://www.cancer.org/content/dam/cancer-org/research/cancer-facts-and-statistics/annual-cancer-facts-and-figures/2012/cancer-facts-and-figures-2012.pdf (accessed on 1 December 2017).
- Kunzmann, A.T.; Coleman, H.G.; Huang, W.Y.; Kitahara, C.M.; Cantwell, M.M.; Berndt, S.I. Dietary fiber intake and risk of colorectal cancer and incident and recurrent adenoma in the prostate, lung, colorectal, and ovarian cancer screening trial. Am. J. Clin. Nutr. 2015, 102, 881–890. [Google Scholar] [CrossRef] [PubMed]
- Murphy, N.; Norat, T.; Ferrari, P.; Jenab, M.; Bueno-de-Mesquita, B.; Skeie, G.; Dahm, C.C.; Overvad, K.; Olsen, A.; Tjonneland, A.; et al. Dietary fibre intake and risks of cancers of the colon and rectum in the european prospective investigation into cancer and nutrition (EPIC). PLoS ONE 2012, 7, e39361. [Google Scholar] [CrossRef] [PubMed]
- O’Keefe, S.J. Diet, microorganisms and their metabolites, and colon cancer. Nat. Rev. Gastroenterol. Hepatol. 2016, 13, 691–706. [Google Scholar] [CrossRef] [PubMed]
- Ganapathy, V.; Thangaraju, M.; Prasad, P.D.; Martin, P.M.; Singh, N. Transporters and receptors for short-chain fatty acids as the molecular link between colonic bacteria and the host. Curr. Opin. Pharmacol. 2013, 13, 869–874. [Google Scholar] [CrossRef] [PubMed]
- Sobhani, I.; Tap, J.; Roudot-Thoraval, F.; Roperch, J.P.; Letulle, S.; Langella, P.; Corthier, G.; Tran Van Nhieu, J.; Furet, J.P. Microbial dysbiosis in colorectal cancer (CRC) patients. PLoS ONE 2011, 6, e16393. [Google Scholar] [CrossRef] [PubMed]
- Gao, R.; Kong, C.; Li, H.; Huang, L.; Qu, X.; Qin, N.; Qin, H. Dysbiosis signature of mycobiota in colon polyp and colorectal cancer. Eur. J. Clin. Microbiol. Infect. Dis. 2017, 36, 2457–2468. [Google Scholar] [CrossRef] [PubMed]
- Yao, Y.; Suo, T.; Andersson, R.; Cao, Y.; Wang, C.; Lu, J.; Chui, E. Dietary fibre for the prevention of recurrent colorectal adenomas and carcinomas. Cochrane Database Syst. Rev. 2017, 1, CD003430. [Google Scholar] [CrossRef] [PubMed]
- Bultman, S.J. Interplay between diet, gut microbiota, epigenetic events, and colorectal cancer. Mol. Nutr. Food Res. 2017, 61. [Google Scholar] [CrossRef] [PubMed]
- Roller, M.; Clune, Y.; Collins, K.; Rechkemmer, G.; Watzl, B. Consumption of prebiotic inulin enriched with oligofructose in combination with the probiotics lactobacillus rhamnosus and bifidobacterium lactis has minor effects on selected immune parameters in polypectomised and colon cancer patients. Br. J. Nutr. 2007, 97, 676–684. [Google Scholar] [CrossRef] [PubMed]
- Corpet, D.E.; Pierre, F. How good are rodent models of carcinogenesis in predicting efficacy in humans? A systematic review and meta-analysis of colon chemoprevention in rats, mice and men. Eur. J. Cancer 2005, 41, 1911–1922. [Google Scholar] [CrossRef] [PubMed]
- Mai, V.; Colbert, L.H.; Perkins, S.N.; Schatzkin, A.; Hursting, S.D. Intestinal microbiota: A potential diet-responsive prevention target in apcmin mice. Mol. Carcinog. 2007, 46, 42–48. [Google Scholar] [CrossRef] [PubMed]
- Zackular, J.P.; Baxter, N.T.; Iverson, K.D.; Sadler, W.D.; Petrosino, J.F.; Chen, G.Y.; Schloss, P.D. The gut microbiome modulates colon tumorigenesis. MBio 2013, 4, e00692-13. [Google Scholar] [CrossRef] [PubMed]
- Hamaker, B.R.; Tuncil, Y.E. A perspective on the complexity of dietary fiber structures and their potential effect on the gut microbiota. J. Mol. Biol. 2014, 426, 3838–3850. [Google Scholar] [CrossRef] [PubMed]
- Saam, J.R.; Gordon, J.I. Inducible gene knockouts in the small intestinal and colonic epithelium. J. Biol. Chem. 1999, 274, 38071–38082. [Google Scholar] [CrossRef] [PubMed]
- Khazaie, K.; Zadeh, M.; Khan, M.W.; Bere, P.; Gounari, F.; Dennis, K.; Blatner, N.R.; Owen, J.L.; Klaenhammer, T.R.; Mohamadzadeh, M. Abating colon cancer polyposis by lactobacillus acidophilus deficient in lipoteichoic acid. Proc. Natl. Acad. Sci. USA 2012, 109, 10462–10467. [Google Scholar] [CrossRef] [PubMed]
- Gounaris, E.; Erdman, S.E.; Restaino, C.; Gurish, M.F.; Friend, D.S.; Gounari, F.; Lee, D.M.; Zhang, G.; Glickman, J.N.; Shin, K.; et al. Mast cells are an essential hematopoietic component for polyp development. Proc. Natl. Acad. Sci. USA 2007, 104, 19977–19982. [Google Scholar] [CrossRef] [PubMed]
- Fischer, A.H.; Jacobson, K.A.; Rose, J.; Zeller, R. Hematoxylin and eosin staining of tissue and cell sections. CSH Protoc. 2008, 2008, pdb-prot4986. [Google Scholar] [CrossRef] [PubMed]
- Forsyth, C.B.; Pulai, J.; Loeser, R.F. Fibronectin fragments and blocking antibodies to alpha2beta1 and alpha5beta1 integrins stimulate mitogen-activated protein kinase signaling and increase collagenase 3 (matrix metalloproteinase 13) production by human articular chondrocytes. Arthritis Rheum. 2002, 46, 2368–2376. [Google Scholar] [CrossRef] [PubMed]
- Banan, A.; Zhang, L.J.; Shaikh, M.; Fields, J.Z.; Choudhary, S.; Forsyth, C.B.; Farhadi, A.; Keshavarzian, A. Theta isoform of protein kinase c alters barrier function in intestinal epithelium through modulation of distinct claudin isotypes: A novel mechanism for regulation of permeability. J. Pharmacol. Exp. Ther. 2005, 313, 962–982. [Google Scholar] [CrossRef] [PubMed]
- Bishehsari, F.; Saadalla, A.; Khazaie, K.; Engen, P.A.; Voigt, R.M.; Shetuni, B.B.; Forsyth, C.; Shaikh, M.; Vitaterna, M.H.; Turek, F.; et al. Light/dark shifting promotes alcohol-induced colon carcinogenesis: Possible role of intestinal inflammatory milieu and microbiota. Int. J. Mol. Sci. 2016, 17, 2017. [Google Scholar] [CrossRef] [PubMed]
- Schmieder, R.; Edwards, R. Quality control and preprocessing of metagenomic datasets. Bioinformatics 2011, 27, 863–864. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R.C. Search and clustering orders of magnitude faster than blast. Bioinformatics 2010, 26, 2460–2461. [Google Scholar] [CrossRef] [PubMed]
- Caporaso, J.G.; Kuczynski, J.; Stombaugh, J.; Bittinger, K.; Bushman, F.D.; Costello, E.K.; Fierer, N.; Pena, A.G.; Goodrich, J.K.; Gordon, J.I.; et al. Qiime allows analysis of high-throughput community sequencing data. Nat. Methods 2010, 7, 335–336. [Google Scholar] [CrossRef] [PubMed]
- McDonald, D.; Clemente, J.C.; Kuczynski, J.; Rideout, J.R.; Stombaugh, J.; Wendel, D.; Wilke, A.; Huse, S.; Hufnagle, J.; Meyer, F.; et al. The biological observation matrix (BIOM) format or: How I learned to stop worrying and love the ome-ome. Gigascience 2012, 1, 7. [Google Scholar] [CrossRef] [PubMed]
- Clarke, K.R. Non-parametric multivariate analyses of changes in community structure. Aust. J. Ecol. 1993, 18, 117–143. [Google Scholar] [CrossRef]
- R Development Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2013. Available online: http://www.R-project.org/ (accessed on 1 December 2017).
- Oksanen, J.; Blanchet, F.G.; Friendly, M.; Kindt, R.; Legendre, P.; McGlinn, D.; Minchin, P.R.; O’Hara, R.B.; Simpson, G.L.; Solymos, P.; et al. Package ‘Vegan’. Community Ecology Package, 2nd ed. 2016. Available online: http://www.pelagicos.net/MARS6910_spring2015/manuals/R_vegan.pdf (accessed on 1 December 2017).
- Langille, M.G.; Zaneveld, J.; Caporaso, J.G.; McDonald, D.; Knights, D.; Reyes, J.A.; Clemente, J.C.; Burkepile, D.E.; Vega Thurber, R.L.; Knight, R.; et al. Predictive functional profiling of microbial communities using 16s rRNA marker gene sequences. Nat. Biotechnol. 2013, 31, 814–821. [Google Scholar] [CrossRef] [PubMed]
- Kanehisa, M.; Goto, S. Kegg: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 2000, 28, 27–30. [Google Scholar] [CrossRef] [PubMed]
- Canani, R.B.; Costanzo, M.D.; Leone, L.; Pedata, M.; Meli, R.; Calignano, A. Potential beneficial effects of butyrate in intestinal and extraintestinal diseases. World J. Gastroenterol. 2011, 17, 1519–1528. [Google Scholar] [CrossRef] [PubMed]
- Louis, P.; Flint, H.J. Diversity, metabolism and microbial ecology of butyrate-producing bacteria from the human large intestine. FEMS Microbiol. Lett. 2009, 294, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Macfarlane, S.; Macfarlane, G.T. Regulation of short-chain fatty acid production. Proc. Nutr. Soc. 2003, 62, 67–72. [Google Scholar] [CrossRef] [PubMed]
- Hakansson, A.; Molin, G. Gut microbiota and inflammation. Nutrients 2011, 3, 637–682. [Google Scholar] [CrossRef] [PubMed]
- Wexler, H.M. Bacteroides: The good, the bad, and the nitty-gritty. Clin. Microbiol. Rev. 2007, 20, 593–621. [Google Scholar] [CrossRef] [PubMed]
- Rajilic-Stojanovic, M.; de Vos, W.M. The first 1000 cultured species of the human gastrointestinal microbiota. FEMS Microbiol. Rev. 2014, 38, 996–1047. [Google Scholar] [CrossRef] [PubMed]
- Louis, P.; Hold, G.L.; Flint, H.J. The gut microbiota, bacterial metabolites and colorectal cancer. Nat. Rev. Microbiol. 2014, 12, 661–672. [Google Scholar] [CrossRef] [PubMed]
- Riviere, A.; Selak, M.; Lantin, D.; Leroy, F.; De Vuyst, L. Bifidobacteria and butyrate-producing colon bacteria: Importance and strategies for their stimulation in the human gut. Front. Microbiol. 2016, 7, 979. [Google Scholar] [CrossRef] [PubMed]
- Reichardt, N.; Duncan, S.H.; Young, P.; Belenguer, A.; McWilliam Leitch, C.; Scott, K.P.; Flint, H.J.; Louis, P. Phylogenetic distribution of three pathways for propionate production within the human gut microbiota. ISME J. 2014, 8, 1323–1335. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Zhao, F.; Wang, Y.; Chen, J.; Tao, J.; Tian, G.; Wu, S.; Liu, W.; Cui, Q.; Geng, B.; et al. Gut microbiota dysbiosis contributes to the development of hypertension. Microbiome 2017, 5, 14. [Google Scholar] [CrossRef] [PubMed]
- Louis, P.; Flint, H.J. Formation of propionate and butyrate by the human colonic microbiota. Environ. Microbiol. 2017, 19, 29–41. [Google Scholar] [CrossRef] [PubMed]
- Antharam, V.C.; Li, E.C.; Ishmael, A.; Sharma, A.; Mai, V.; Rand, K.H.; Wang, G.P. Intestinal dysbiosis and depletion of butyrogenic bacteria in Clostridium difficile infection and nosocomial diarrhea. J. Clin. Microbiol. 2013, 51, 2884–2892. [Google Scholar] [CrossRef] [PubMed]
- Tuncil, Y.E.; Nakatsu, C.H.; Kazem, A.E.; Arioglu-Tuncil, S.; Reuhs, B.; Martens, E.C.; Hamaker, B.R. Delayed utilization of some fast-fermenting soluble dietary fibers by human gut microbiota when presented in a mixture. J. Funct. Foods 2017, 32, 347–357. [Google Scholar] [CrossRef]
- Flemer, B.; Gaci, N.; Borrel, G.; Sanderson, I.R.; Chaudhary, P.P.; Tottey, W.; O’Toole, P.W.; Brugere, J.F. Fecal microbiota variation across the lifespan of the healthy laboratory rat. Gut Microbes 2017, 8, 428–439. [Google Scholar] [CrossRef] [PubMed]
- Earley, H.; Lennon, G.; Balfe, A.; Kilcoyne, M.; Clyne, M.; Joshi, L.; Carrington, S.; Martin, S.T.; Coffey, J.C.; Winter, D.C.; et al. A preliminary study examining the binding capacity of Akkermansia muciniphila and Desulfovibrio spp., to colonic mucin in health and ulcerative colitis. PLoS ONE 2015, 10, e0135280. [Google Scholar] [CrossRef] [PubMed]
- Puppa, M.J.; White, J.P.; Sato, S.; Cairns, M.; Baynes, J.W.; Carson, J.A. Gut barrier dysfunction in the apc(min/+) mouse model of colon cancer cachexia. Biochim. Biophys. Acta 2011, 1812, 1601–1606. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Q.; Jin, Z.; Wu, W.; Gao, R.; Guo, B.; Gao, Z.; Yang, Y.; Qin, H. Analysis of the intestinal lumen microbiota in an animal model of colorectal cancer. PLoS ONE 2014, 9, e90849. [Google Scholar] [CrossRef] [PubMed]
- Mariat, D.; Firmesse, O.; Levenez, F.; Guimaraes, V.; Sokol, H.; Dore, J.; Corthier, G.; Furet, J.P. The Firmicutes/Bacteroidetes ratio of the human microbiota changes with age. BMC Microbiol. 2009, 9, 123. [Google Scholar] [CrossRef] [PubMed]
- Voreades, N.; Kozil, A.; Weir, T.L. Diet and the development of the human intestinal microbiome. Front. Microbiol. 2014, 5, 494. [Google Scholar] [CrossRef] [PubMed]
- Ni, Y.; Wong, V.H.; Tai, W.C.; Li, J.; Wong, W.Y.; Lee, M.M.; Fong, F.L.; El-Nezami, H.; Panagiotou, G. A metagenomic study of the preventive effect of Lactobacillus rhamnosus GG on intestinal polyp formation in ApcMin/+ mice. J. Appl. Microbiol. 2017, 122, 770–784. [Google Scholar] [CrossRef] [PubMed]
- Sugawara, T.; Miyazawa, T. Beneficial effect of dietary wheat glycolipids on cecum short-chain fatty acid and secondary bile acid profiles in mice. J. Nutr. Sci. Vitaminol. 2001, 47, 299–305. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Q.; Gao, R.; Wu, W.; Qin, H. The role of gut microbiota in the pathogenesis of colorectal cancer. Tumour. Biol. 2013, 34, 1285–1300. [Google Scholar] [CrossRef] [PubMed]
- Hold, G.L.; Smith, M.; Grange, C.; Watt, E.R.; El-Omar, E.M.; Mukhopadhya, I. Role of the gut microbiota in inflammatory bowel disease pathogenesis: What have we learnt in the past 10 years? World J. Gastroenterol. 2014, 20, 1192–1210. [Google Scholar] [CrossRef] [PubMed]
- Cammarota, G.; Ianiro, G.; Cianci, R.; Bibbo, S.; Gasbarrini, A.; Curro, D. The involvement of gut microbiota in inflammatory bowel disease pathogenesis: Potential for therapy. Pharmacol. Ther. 2015, 149, 191–212. [Google Scholar] [CrossRef] [PubMed]
- Power, K.A.; Lepp, D.; Zarepoor, L.; Monk, J.M.; Wu, W.; Tsao, R.; Liu, R. Dietary flaxseed modulates the colonic microenvironment in healthy C57BL/6 male mice which may alter susceptibility to gut-associated diseases. J. Nutr. Biochem. 2016, 28, 61–69. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; Wu, J.; Li, J.V.; Zhou, N.Y.; Tang, H.; Wang, Y. Gut microbiota composition modifies fecal metabolic profiles in mice. J. Proteome Res. 2013, 12, 2987–2999. [Google Scholar] [CrossRef] [PubMed]
- Ericsson, A.C.; Akter, S.; Hanson, M.M.; Busi, S.B.; Parker, T.W.; Schehr, R.J.; Hankins, M.A.; Ahner, C.E.; Davis, J.W.; Franklin, C.L.; et al. Differential susceptibility to colorectal cancer due to naturally occurring gut microbiota. Oncotarget 2015, 6, 33689–33704. [Google Scholar] [CrossRef] [PubMed]
- Hinnebusch, B.F.; Meng, S.; Wu, J.T.; Archer, S.Y.; Hodin, R.A. The effects of short-chain fatty acids on human colon cancer cell phenotype are associated with histone hyperacetylation. J. Nutr. 2002, 132, 1012–1017. [Google Scholar] [CrossRef] [PubMed]
- Kiefer, J.; Beyer-Sehlmeyer, G.; Pool-Zobel, B.L. Mixtures of SCFA, composed according to physiologically available concentrations in the gut lumen, modulate histone acetylation in human HT29 colon cancer cells. Br. J. Nutr. 2006, 96, 803–810. [Google Scholar] [CrossRef] [PubMed]
- Forest, V.; Clement, M.; Pierre, F.; Meflah, K.; Menanteau, J. Butyrate restores motile function and actin cytoskeletal network integrity in APC mutated mouse colon epithelial cells. Nutr. Cancer 2003, 45, 84–92. [Google Scholar] [CrossRef] [PubMed]
- Hamer, H.M.; Jonkers, D.M.; Bast, A.; Vanhoutvin, S.A.; Fischer, M.A.; Kodde, A.; Troost, F.J.; Venema, K.; Brummer, R.J. Butyrate modulates oxidative stress in the colonic mucosa of healthy humans. Clin. Nutr. 2009, 28, 88–93. [Google Scholar] [CrossRef] [PubMed]
- Leonel, A.J.; Alvarez-Leite, J.I. Butyrate: Implications for intestinal function. Curr. Opin. Clin. Nutr. Metab. Care 2012, 15, 474–479. [Google Scholar] [CrossRef] [PubMed]
- Thangaraju, M.; Cresci, G.A.; Liu, K.; Ananth, S.; Gnanaprakasam, J.P.; Browning, D.D.; Mellinger, J.D.; Smith, S.B.; Digby, G.J.; Lambert, N.A.; et al. GPR109A is a G-protein-coupled receptor for the bacterial fermentation product butyrate and functions as a tumor suppressor in colon. Cancer Res. 2009, 69, 2826–2832. [Google Scholar] [CrossRef] [PubMed]
- Singh, N.; Gurav, A.; Sivaprakasam, S.; Brady, E.; Padia, R.; Shi, H.; Thangaraju, M.; Prasad, P.D.; Manicassamy, S.; Munn, D.H.; et al. Activation of GPR19A, receptor for niacin and the commensal metabolite butyrate, suppresses colonic inflammation and carcinogenesis. Immunity 2014, 40, 128–139. [Google Scholar] [CrossRef] [PubMed]
- Consortium, H.M.P. Structure, function and diversity of the healthy human microbiome. Nature 2012, 486, 207–214. [Google Scholar] [CrossRef]
- Shen, X.J.; Rawls, J.F.; Randall, T.; Burcal, L.; Mpande, C.N.; Jenkins, N.; Jovov, B.; Abdo, Z.; Sandler, R.S.; Keku, T.O. Molecular characterization of mucosal adherent bacteria and associations with colorectal adenomas. Gut Microbes 2010, 1, 138–147. [Google Scholar] [CrossRef] [PubMed]
- Flemer, B.; Lynch, D.B.; Brown, J.M.; Jeffery, I.B.; Ryan, F.J.; Claesson, M.J.; O’Riordain, M.; Shanahan, F.; O’Toole, P.W. Tumour-associated and non-tumour-associated microbiota in colorectal cancer. Gut 2017, 66, 633–643. [Google Scholar] [CrossRef] [PubMed]
- Purcell, R.V.; Pearson, J.; Aitchison, A.; Dixon, L.; Frizelle, F.A.; Keenan, J.I. Colonization with enterotoxigenic Bacteroides fragilis is associated with early-stage colorectal neoplasia. PLoS ONE 2017, 12, e0171602. [Google Scholar] [CrossRef] [PubMed]
- Moen, B.; Henjum, K.; Mage, I.; Knutsen, S.H.; Rud, I.; Hetland, R.B.; Paulsen, J.E. Effect of dietary fibers on cecal microbiota and intestinal tumorigenesis in azoxymethane treated A/J Min/+ mice. PLoS ONE 2016, 11, e0155402. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Jobin, C. Microbial imbalance and intestinal pathologies: Connections and contributions. Dis. Model. Mech. 2014, 7, 1131–1142. [Google Scholar] [CrossRef] [PubMed]
- Borges-Canha, M.; Portela-Cidade, J.P.; Dinis-Ribeiro, M.; Leite-Moreira, A.F.; Pimentel-Nunes, P. Role of colonic microbiota in colorectal carcinogenesis: A systematic review. Rev. Esp. Enferm. Dig. 2015, 107, 659–671. [Google Scholar] [CrossRef] [PubMed]
- Fukuda, S.; Toh, H.; Hase, K.; Oshima, K.; Nakanishi, Y.; Yoshimura, K.; Tobe, T.; Clarke, J.M.; Topping, D.L.; Suzuki, T.; et al. Bifidobacteria can protect from enteropathogenic infection through production of acetate. Nature 2011, 469, 543–547. [Google Scholar] [CrossRef] [PubMed]
- Belenguer, A.; Holtrop, G.; Duncan, S.H.; Anderson, S.E.; Calder, A.G.; Flint, H.J.; Lobley, G.E. Rates of production and utilization of lactate by microbial communities from the human colon. FEMS Microbiol. Ecol. 2011, 77, 107–119. [Google Scholar] [CrossRef] [PubMed]
- You, H.J.; Oh, D.K.; Ji, G.E. Anticancerogenic effect of a novel chiroinositol-containing polysaccharide from Bifidobacterium bifidum BGN4. FEMS Microbiol. Lett. 2004, 240, 131–136. [Google Scholar] [CrossRef] [PubMed]
- Tirandaz, H.; Mohammadi, E. Efficient tumor targeting by anaerobic butyrate-producing bacteria. Med. Hypotheses 2013, 80, 675–678. [Google Scholar] [CrossRef] [PubMed]
- Velasquez-Manoff, M. Gut microbiome: The peacekeepers. Nature 2015, 518, S3–11. [Google Scholar] [CrossRef] [PubMed]
- O’Keefe, S.J.; Ou, J.; Aufreiter, S.; O’Connor, D.; Sharma, S.; Sepulveda, J.; Fukuwatari, T.; Shibata, K.; Mawhinney, T. Products of the colonic microbiota mediate the effects of diet on colon cancer risk. J. Nutr. 2009, 139, 2044–2048. [Google Scholar] [CrossRef] [PubMed]
- O’Keefe, S.J.; Li, J.V.; Lahti, L.; Ou, J.; Carbonero, F.; Mohammed, K.; Posma, J.M.; Kinross, J.; Wahl, E.; Ruder, E.; et al. Fat, fibre and cancer risk in African Americans and rural Africans. Nat. Commun. 2015, 6, 6342. [Google Scholar] [CrossRef] [PubMed]
- Ohkawara, S.; Furuya, H.; Nagashima, K.; Asanuma, N.; Hino, T. Oral administration of butyrivibrio fibrisolvens, a butyrate-producing bacterium, decreases the formation of aberrant crypt foci in the colon and rectum of mice. J. Nutr. 2005, 135, 2878–2883. [Google Scholar] [CrossRef] [PubMed]
- Davie, J.R. Inhibition of histone deacetylase activity by butyrate. J. Nutr. 2003, 133, 2485S–2493S. [Google Scholar] [CrossRef] [PubMed]
- Bardhan, K.; Paschall, A.V.; Yang, D.; Chen, M.R.; Simon, P.S.; Bhutia, Y.D.; Martin, P.M.; Thangaraju, M.; Browning, D.D.; Ganapathy, V.; et al. IFNγ induces DNA methylation-silenced GPR109A expression via pSTAT1/p300 and H3K18 acetylation in colon cancer. Cancer Immunol. Res. 2015, 3, 795–805. [Google Scholar] [CrossRef] [PubMed]
- Higashimura, Y.; Naito, Y.; Takagi, T.; Uchiyama, K.; Mizushima, K.; Ushiroda, C.; Ohnogi, H.; Kudo, Y.; Yasui, M.; Inui, S.; et al. Protective effect of agaro-oligosaccharides on gut dysbiosis and colon tumorigenesis in high-fat diet-fed mice. Am. J. Physiol. Gastrointest. Liver Physiol. 2016, 310, G367–375. [Google Scholar] [CrossRef] [PubMed]
- Donohoe, D.R.; Holley, D.; Collins, L.B.; Montgomery, S.A.; Whitmore, A.C.; Hillhouse, A.; Curry, K.P.; Renner, S.W.; Greenwalt, A.; Ryan, E.P.; et al. A gnotobiotic mouse model demonstrates that dietary fiber protects against colorectal tumorigenesis in a microbiota- and butyrate-dependent manner. Cancer Discov. 2014, 4, 1387–1397. [Google Scholar] [CrossRef] [PubMed]
- Mehta, R.S.; Nishihara, R.; Cao, Y.; Song, M.; Mima, K.; Qian, Z.R.; Nowak, J.A.; Kosumi, K.; Hamada, T.; Masugi, Y.; et al. Association of dietary patterns with risk of colorectal cancer subtypes classified by fusobacterium nucleatum in tumor tissue. JAMA Oncol. 2017, 3, 921–927. [Google Scholar] [CrossRef] [PubMed]
| Taxa | Total SCFA v. High Fiber | Acetate v. High Fiber | Propionate v. High Fiber | Butyrate v. High Fiber |
|---|---|---|---|---|
| R, p-Value | R, p-Value | R, p-Value | R, p-Value | |
| Phylum | ||||
| Actinobacteria | +0.88, ˂0.00 | +0.91, ˂0.00 | - | - |
| Family | ||||
| Bifidobacteriaceae | +0.89, ˂0.00 | +0.91, ˂0.00 | - | - |
| Prevotellaceae | −0.80, ˂0.00 | −0.78, ˂0.00 | - | −0.73, 0.01 |
| [Paraprevotellaceae] | −0.71, 0.02 | −0.67, 0.03 | - | −0.65, 0.04 |
| Lactobacillaceae | −0.73, 0.02 | −0.71, 0.02 | - | - |
| o_RF32;f_unclassified | −0.65, 0.04 | −0.65, 0.04 | - | - |
| Mycoplasmataceae | −0.65, 0.04 | - | - | - |
| S24-7 | - | - | +0.66, 0.03 | - |
| o_Streptophyta; f_unclassified | - | - | −0.77, 0.01 | −0.82, ˂0.00 |
| Genus | ||||
| Bifidobacterium | +0.89, ˂0.00 | +0.91, ˂0.00 | - | - |
| Prevotella | −0.80, ˂0.00 | −0.78, ˂0.01 | - | −0.66, 0.03 |
| [Prevotella] | −0.73, 0.02 | −0.70, 0.02 | - | - |
| Lactobacillus | −0.73, 0.02 | −0.71, 0.02 | - | - |
| f_Lachnospiraceae; g_unclassified | +0.66, 0.03 | +0.65, 0.04 | - | +0.63, 0.04 |
| Coprococcus | −0.66, 0.03 | −0.68, 0.02 | - | - |
| f_Erysipelotrichaceae; g_unclassified | −0.74, 0.01 | −0.71, 0.02 | - | - |
| o_RF32; f_unclassified; g_unclassified | −0.65, 0.04 | −0.65, 0.04 | - | - |
| f_Mycoplasmataceae; g_unclassified | −0.65, 0.04 | - | - | - |
| S24-7;g_unclassified | - | - | +0.66, 0.03 | - |
| o_Streptophyta; f_unclassifed; g_unclassified | - | - | −0.77, 0.01 | −0.82, ˂0.00 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bishehsari, F.; Engen, P.A.; Preite, N.Z.; Tuncil, Y.E.; Naqib, A.; Shaikh, M.; Rossi, M.; Wilber, S.; Green, S.J.; Hamaker, B.R.; et al. Dietary Fiber Treatment Corrects the Composition of Gut Microbiota, Promotes SCFA Production, and Suppresses Colon Carcinogenesis. Genes 2018, 9, 102. https://doi.org/10.3390/genes9020102
Bishehsari F, Engen PA, Preite NZ, Tuncil YE, Naqib A, Shaikh M, Rossi M, Wilber S, Green SJ, Hamaker BR, et al. Dietary Fiber Treatment Corrects the Composition of Gut Microbiota, Promotes SCFA Production, and Suppresses Colon Carcinogenesis. Genes. 2018; 9(2):102. https://doi.org/10.3390/genes9020102
Chicago/Turabian StyleBishehsari, Faraz, Phillip A. Engen, Nailliw Z. Preite, Yunus E. Tuncil, Ankur Naqib, Maliha Shaikh, Marco Rossi, Sherry Wilber, Stefan J. Green, Bruce R. Hamaker, and et al. 2018. "Dietary Fiber Treatment Corrects the Composition of Gut Microbiota, Promotes SCFA Production, and Suppresses Colon Carcinogenesis" Genes 9, no. 2: 102. https://doi.org/10.3390/genes9020102
APA StyleBishehsari, F., Engen, P. A., Preite, N. Z., Tuncil, Y. E., Naqib, A., Shaikh, M., Rossi, M., Wilber, S., Green, S. J., Hamaker, B. R., Khazaie, K., Voigt, R. M., Forsyth, C. B., & Keshavarzian, A. (2018). Dietary Fiber Treatment Corrects the Composition of Gut Microbiota, Promotes SCFA Production, and Suppresses Colon Carcinogenesis. Genes, 9(2), 102. https://doi.org/10.3390/genes9020102





