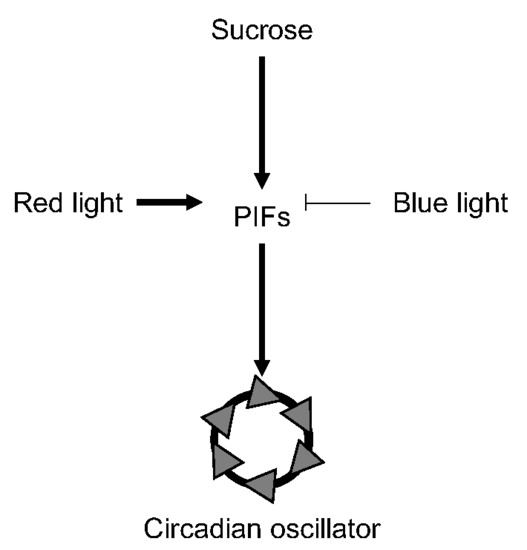
PIF-Mediated Sucrose Regulation of the Circadian Oscillator is Light Quality and Temperature Dependent
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Materials and Growth Conditions
2.2. Bioluminescence Assays
2.3. Leaf Movement Assays
2.4. Sucrose Pulses
2.5. RNA Isolation and Analysis
3. Results
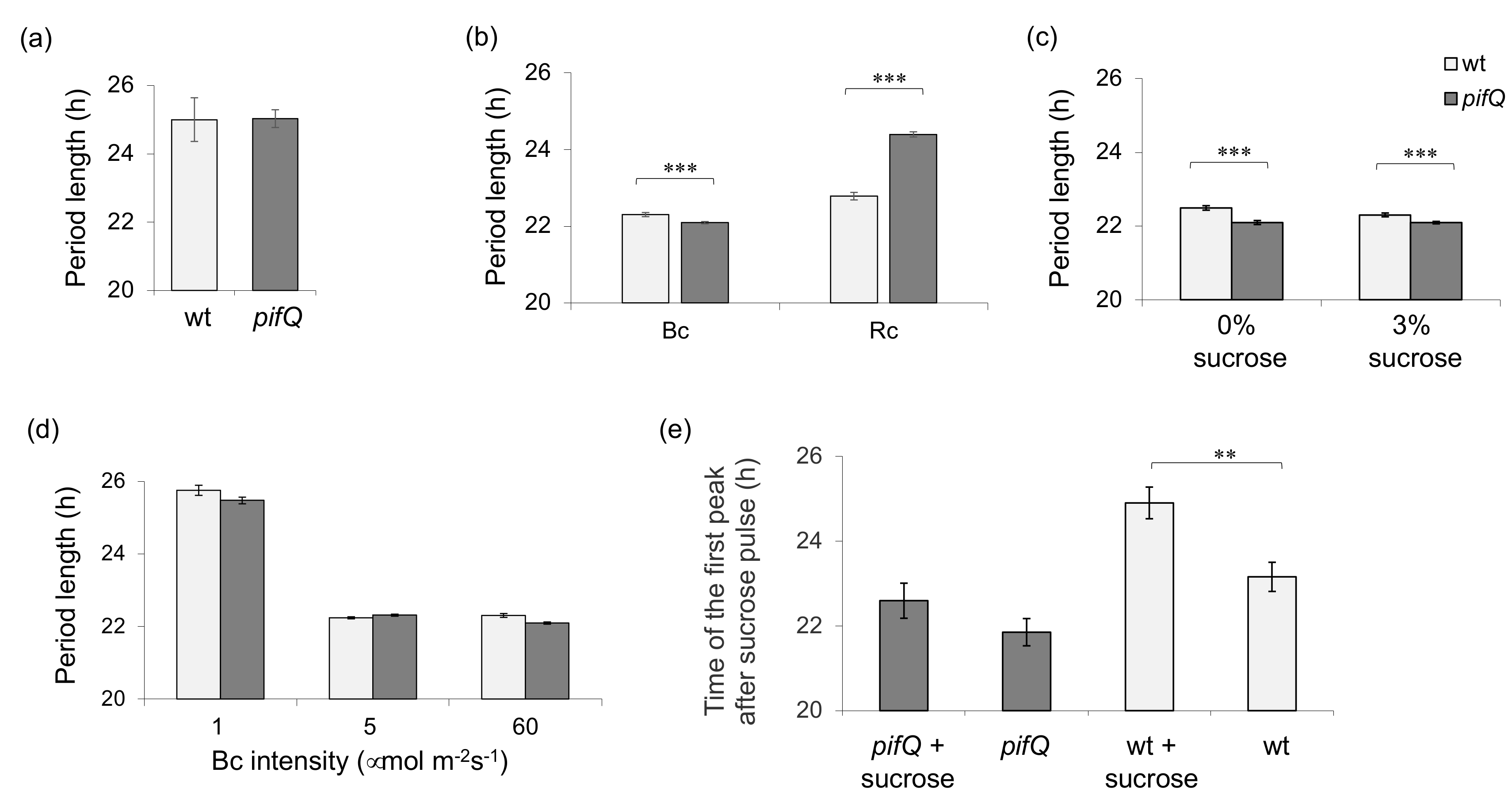
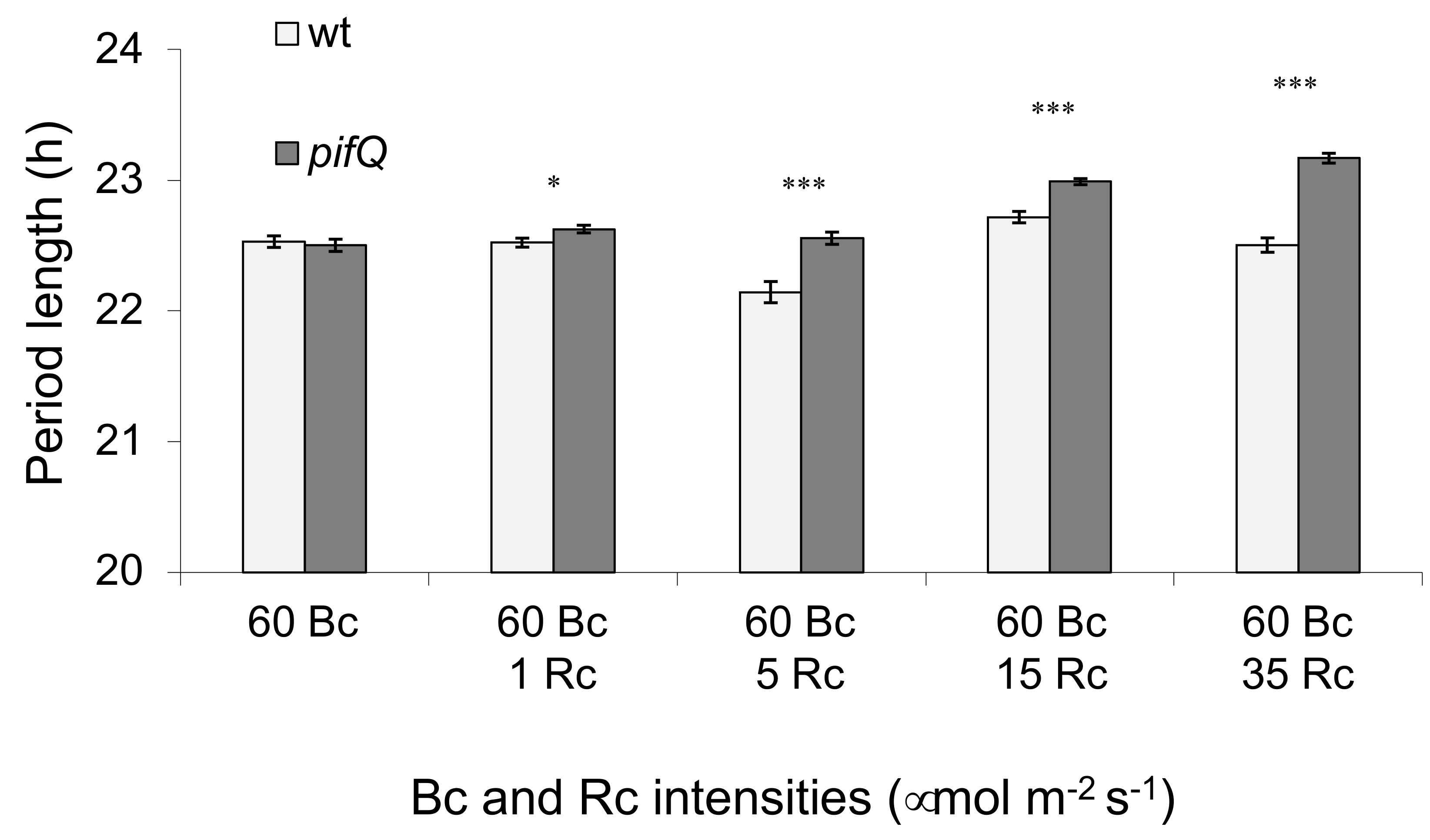
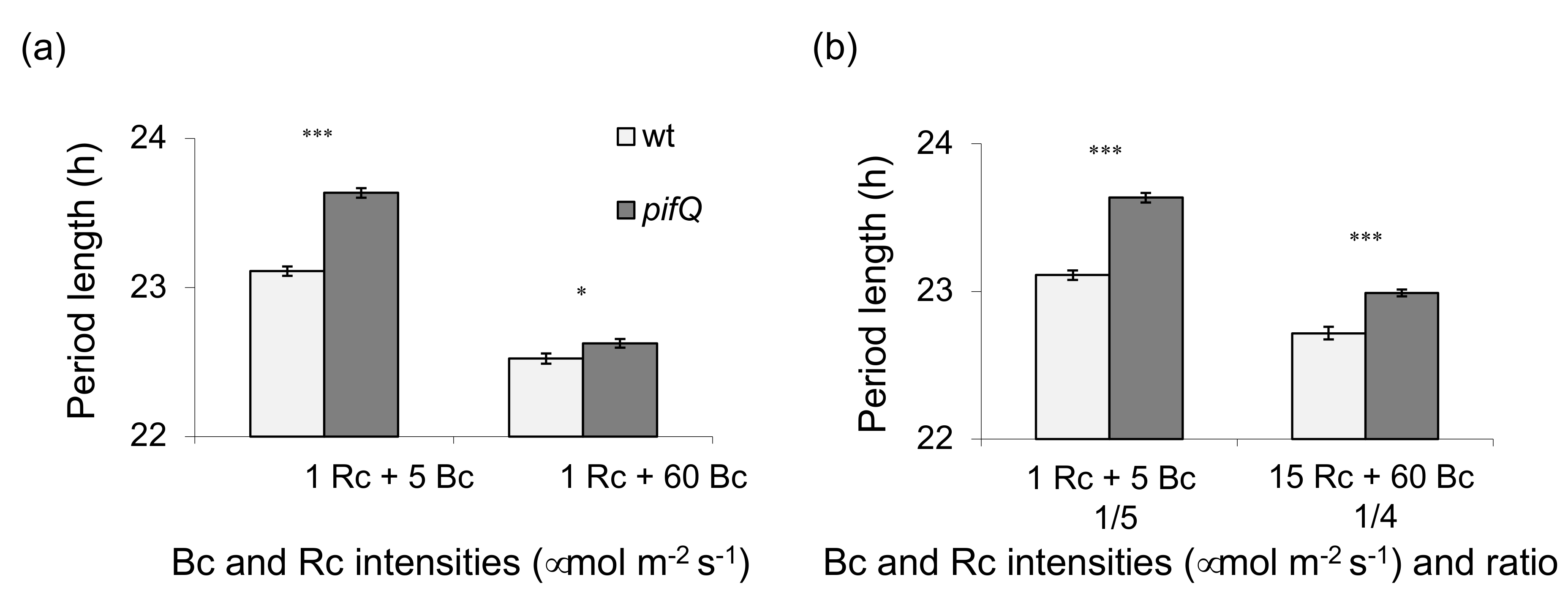
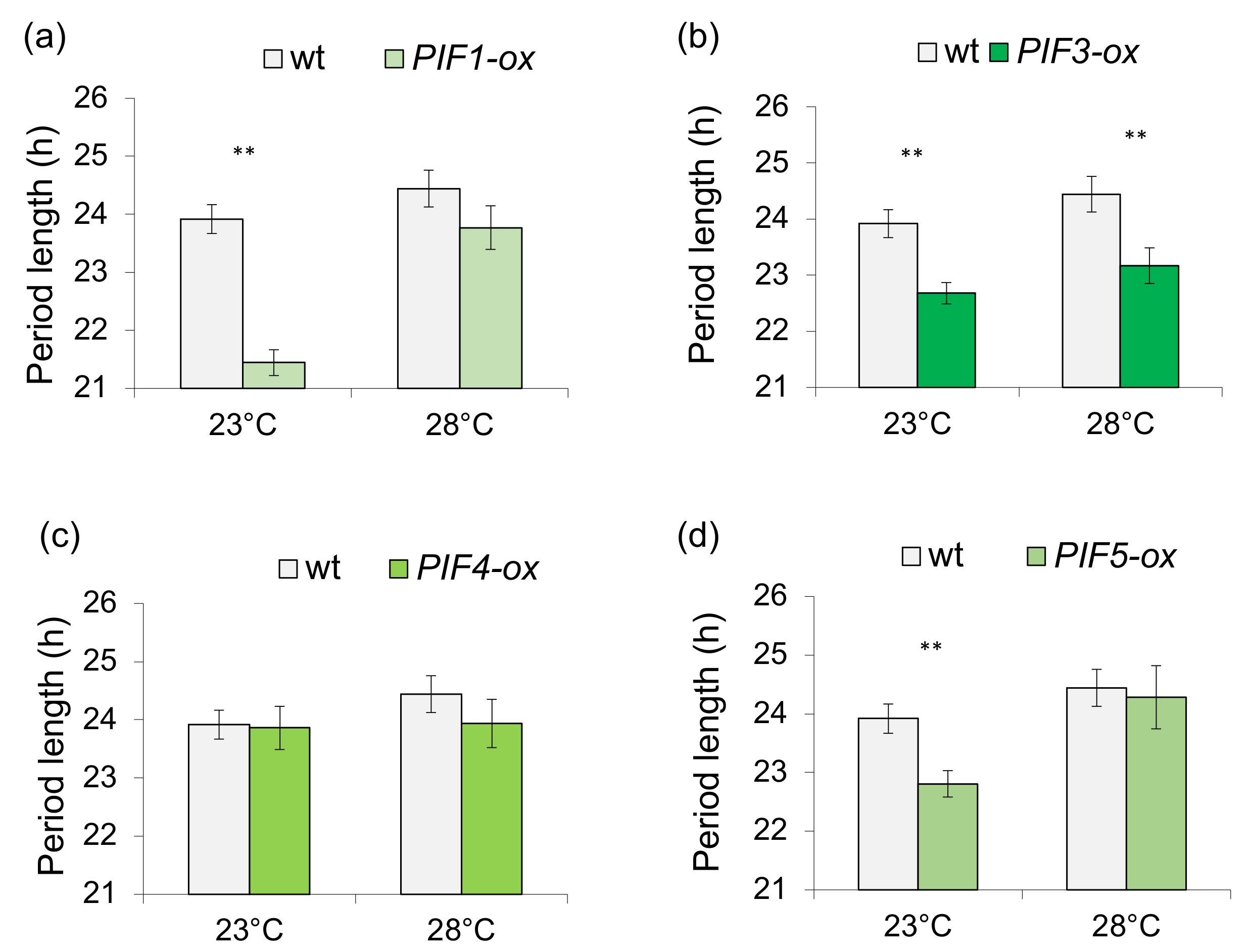
3.1. Light-Dependent Roles for PHYTOCHROME INTERACTING FACTOR Regulation of Sucrose Signaling to the Oscillator
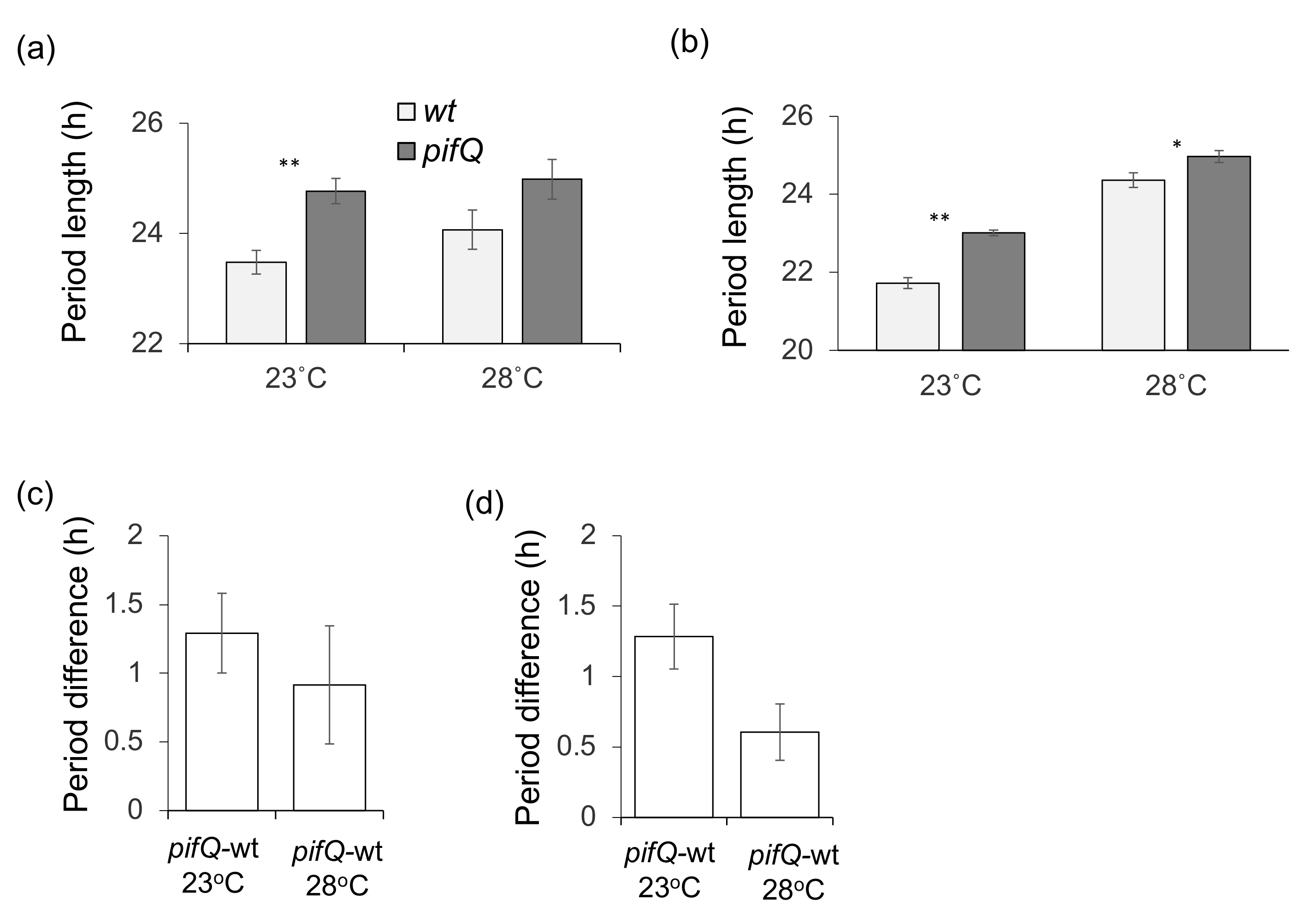
3.2. PHYTOCHROME INTERACTING FACTOR-Mediated Sucrose Signaling Is Temperature Dependent
4. Discussion
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Alabadi, D.; Oyama, T.; Yanovsky, M.J.; Harmon, F.G.; Mas, P.; Kay, S.A. Reciprocal regulation between TOC1 and LHY/CCA1 within the Arabidopsis circadian clock. Science 2001, 293, 880–883. [Google Scholar] [CrossRef] [PubMed]
- Hsu, P.Y.; Harmer, S.L. Wheels within wheels: The plant circadian system. Trends Plant Sci. 2014, 19, 240–249. [Google Scholar] [CrossRef] [PubMed]
- Greenham, K.; McClung, C.R. Integrating circadian dynamics with physiological processes in plants. Nat. Rev. Genet. 2015, 16, 598–610. [Google Scholar] [CrossRef] [PubMed]
- Más, P. Circadian clock function in Arabidopsis thaliana: Time beyond transcription. Trends Cell Biol. 2008, 18, 273–281. [Google Scholar] [CrossRef] [PubMed]
- Gray, J.A.; Shalit-Kaneh, A.; Chu, D.N.; Hsu, P.Y.; Harmer, S.L. The REVEILLE clock genes inhibit growth of juvenile and adult plants by control of cell size. Plant Physiol. 2017, 173, 2308–2322. [Google Scholar] [CrossRef]
- Nolte, C.; Staiger, D. RNA around the clock—Regulation at the RNA level in biological timing. Front. Plant Sci. 2015, 6, 311. [Google Scholar] [CrossRef]
- Romanowski, A.; Yanovsky, M.J. Circadian rhythms and post-transcriptional regulation in higher plants. Front. Plant Sci. 2015, 6, 437. [Google Scholar] [CrossRef]
- Yakir, E.; Hilman, D.; Hassidim, M.; Green, R.M. CIRCADIAN CLOCK ASSOCIATED1 transcript stability and the entrainment of the circadian clock in Arabidopsis. Plant Physiol. 2007, 145, 925–932. [Google Scholar] [CrossRef]
- Más, P.; Kim, W.Y.; Somers, D.E.; Kay, S.A. Targeted degradation of TOC1 by ZTL modulates circadian function in Arabidopsis thaliana. Nature. 2003, 426, 567–570. [Google Scholar] [CrossRef]
- Baudry, A.; Ito, S.; Song, Y.H.; Strait, A.A.; Kiba, T.; Lu, S.; Henriques, R.; Pruneda-Paz, J.L.; Chua, N.H.; Tobin, E.M.; et al. F-box proteins FKF1 and LKP2 act in concert with ZEITLUPE to control Arabidopsis clock progression. Plant Cell. 2010, 22, 606–622. [Google Scholar] [CrossRef]
- Yakir, E.; Hilman, D.; Harir, Y.; Green, R.M. Regulation of output from the plant circadian clock. FEBS J. 2007, 274, 335–345. [Google Scholar] [CrossRef] [PubMed]
- Knight, H.; Thomson, A.J.; McWatters, H.G. SENSITIVE TO FREEZING6 integrates cellular and environmental inputs to the plant circadian clock. Plant Physiol. 2008, 148, 293–303. [Google Scholar] [CrossRef]
- Haydon, M.J.; Mielczarek, O.; Robertson, F.C.; Hubbard, K.E.; Webb, A.A. Photosynthetic entrainment of the Arabidopsis thaliana circadian clock. Nature 2013, 502, 689–692. [Google Scholar] [CrossRef] [PubMed]
- Haydon, M.J.; Mielczarek, O.; Frank, A.; Román, Á.; Webb, A.A. Sucrose and ethylene signaling interact to modulate the circadian clock. Plant Physiol. 2017, 175, 947–958. [Google Scholar] [CrossRef]
- Frank, A.; Matiolli, C.C.; Viana, A.J.C.; Hearn, T.H.; Kusakina, J.; Belbin, F.E.; Newman, D.W.; Yochikawa, A.; Cano-Ramirez, D.L.; Chembath, A.; et al. Circadian entrainment in Arabidopsis by the sugar-responsive transcription factor bZIP63. Curr. Biol. 2018, 28, 2597–2606. [Google Scholar] [CrossRef]
- Shor, E.; Paik, I.; Kangisser, S.; Green, R.; Huq, E. Phytochrome Interacting Factors mediate metabolic control of the circadian system in Arabidopsis. New Phytol. 2017, 215, 217–228. [Google Scholar] [CrossRef] [PubMed]
- Paik, I.; Kathare, P.K.; Kim, J.-I.; Huq, E. Expanding roles of PIFs in signal integration from multiple processes. Mol. Plant 2017, 10, 1035–1046. [Google Scholar] [CrossRef]
- Kumar, S.V.; Lucyshyn, D.; Jaeger, K.E.; Alós, E.; Alvey, E.; Harberd, N.P.; Wigge, P.A. Transcription factor PIF4 controls the thermosensory activation of flowering. Nature 2012, 484, 242–245. [Google Scholar] [CrossRef]
- Leivar, P.; Monte, E.; Oka, Y.; Liu, T.; Carle, C.; Castillon, A.; Huq, E.; Quail, P.H. Multiple phytochrome-interacting bHLH transcription factors repress premature seedling photomorphogenesis in darkness. Curr. Biol. 2008, 18, 1815–1823. [Google Scholar] [CrossRef]
- Zhu, L.; Bu, Q.; Xu, X.; Paik, I.; Huang, X.; Hoecker, U.; Deng, X.W.; Huq, E. CUL4 forms an E3 ligase with COP1 and SPA to promote light-induced degradation of PIF1. Nat. Commun. 2015, 6, 7245. [Google Scholar] [CrossRef]
- Park, E.; Kim, J.; Lee, Y.; Shin, J.; Oh, E.; Chung, W.I.; Liu, J.R.; Choi, G. Degradation of phytochrome interacting factor 3 in phytochrome-mediated light signaling. Plant Cell Physiol. 2004, 45, 968–975. [Google Scholar] [CrossRef] [PubMed]
- Sakuraba, Y.; Jeong, J.; Kang, M.Y.; Kim, J.; Paek, N.C.; Choi, G. Phytochrome-interacting transcription factors PIF4 and PIF5 induce leaf senescence in Arabidopsis. Nat. Commun. 2014, 5, 4636. [Google Scholar] [CrossRef] [PubMed]
- Plautz, J.D.; Straume, M.; Stanewsky, R.; Jamison, C.F.; Brandes, C.; Dowse, H.B.; Hall, J.C.; Kay, S.A. Quantitative analysis of Drosophila period gene transcription in living animals. J. Biol. Rhythm. 1997, 12, 204–217. [Google Scholar] [CrossRef] [PubMed]
- Green, R.M.; Tobin, E.M. Loss of the circadian clock-associated protein 1 in Arabidopsis results in altered clock-regulated gene expression. Proc. Natl. Acad. Sci. USA 1999, 96, 4176–4179. [Google Scholar] [CrossRef] [PubMed]
- Czechowski, T.; Stitt, M.; Altmann, T.; Udvardi, M.K.; Scheible, W.R. Genome-wide identification and testing of superior reference genes for transcript normalization in Arabidopsis. Plant Physiol. 2005, 139, 5–17. [Google Scholar] [CrossRef] [PubMed]
- Nozue, K.; Covington, M.F.; Duek, P.D.; Lorrain, S.; Fankhauser, C.; Harmer, S.L.; Maloof, J.N. Rhythmic growth explained by coincidence between internal and external cues. Nature 2007, 448, 358–361. [Google Scholar] [CrossRef] [PubMed]
- Koini, M.A.; Alvey, L.; Allen, T.; Tilley, C.A.; Harberd, N.P.; Whitelam, G.C.; Franklin, K.A. High temperature- mediated adaptations in plant architecture require the bHLH transcription factor PIF4. Curr. Biol. 2009, 19, 408–413. [Google Scholar] [CrossRef]
- Zhu, J.Y.; Oh, E.; Wang, T.; Wang, Z.Y. TOC1-PIF4 interaction mediates the circadian gating of thermoresponsive growth in Arabidopsis. Nat Commun. 2016, 7, 13692. [Google Scholar] [CrossRef]
- Spitschan, M.; Aguirre, G.K.; Brainard, D.H.; Sweeney, A.M. Variation of outdoor illumination as a function of solar elevation and light pollution. Sci. Rep. 2016, 6, 26756. [Google Scholar] [CrossRef]
- Smith, A.M.; Zeeman, S.C.; Smith, S.M. Starch degradation. Annu. Rev. Plant Biol. 2005, 56, 73–98. [Google Scholar] [CrossRef]
- Devlin, P.F.; Kay, S.A. Cryptochromes are required for phytochrome signaling to the circadian clock but not for rhythmicity. Plant Cell 2000, 12, 2499–2510. [Google Scholar] [CrossRef] [PubMed]
- Somers, D.E.; Devlin, P.F.; Kay, S.A. Phytochromes and cryptochromes in the entrainment of the Arabidopsis circadian clock. Science 1998, 282, 1488–1490. [Google Scholar] [CrossRef] [PubMed]
- Pedmale, U.V.; Huang, S.S.; Zander, M.; Cole, B.J.; Hetzel, J.; Ljung, K.; Reis, P.A.; Sridevi, P.; Nito, K.; Nery, J.R.; et al. Cryptochromes Interact Directly with PIFs to Control Plant Growth in Limiting Blue Light. Cell 2016, 164, 233–245. [Google Scholar] [CrossRef] [PubMed]
- Ma, D.; Li, X.; Guo, Y.; Chu, J.; Fang, S.; Yan, C.; Noel, J.P.; Liu, H. Cryptochrome 1 interacts with PIF4 to regulate high temperature-mediated hypocotyl elongation in response to blue light. Proc. Natl. Acad. Sci. USA 2016, 113, 224–229. [Google Scholar] [CrossRef] [PubMed]
- Leivar, P.; Monte, E. PIFs: Systems integrators in plant development. Plant Cell 2014, 26, 56–78. [Google Scholar] [CrossRef] [PubMed]
- Sawa, M.; Nusinow, D.A.; Kay, S.A.; Imaizumi, T. FKF1 and GIGANTEA complex formation is required for day-length measurement in Arabidopsis. Science 2007, 318, 261–265. [Google Scholar] [CrossRef] [PubMed]
- Kim, W.Y.; Fujiwara, S.; Suh, S.S.; Kim, J.; Kim, Y.; Han, L.; David, K.; Putterill, J.; Nam, H.G.; Somers, D.E. ZEITLUPE is a circadian photoreceptor stabilized by GIGANTEA in blue light. Nature 2007, 449, 356–360. [Google Scholar] [CrossRef]
- Kim, J.; Geng, R.; Gallenstein, R.A.; Somers, D.E. The F-box protein ZEITLUPE controls stability and nucleocytoplasmic partitioning of GIGANTEA. Development 2013, 140, 4060–4069. [Google Scholar] [CrossRef]
- Stavang, J.A.; Gallego-Bartolome, J.; Gomez, M.D.; Yoshida, S.; Asami, T.; Olsen, J.E.; Garcia-Martinez, J.L.; Alabadi, D.; Blazquez, M.A. Hormonal regulation of temperature-induced growth in Arabidopsis. Plant J. 2009, 60, 589–601. [Google Scholar] [CrossRef]
- Franklin, K.A.; Lee, S.H.; Patel, D.; Kumar, S.V.; Spartz, A.K.; Gu, C.; Ye, S.; Yu, P.; Breen, G.; Cohen, J.D.; et al. PHYTOCHROME-INTERACTING FACTOR 4 (PIF4) regulates auxin biosynthesis at high temperature. Proc. Natl. Acad. Sci. USA 2011, 108, 20231–20235. [Google Scholar] [CrossRef]
- Gould, P.D.; Locke, J.C.; Larue, C.; Southern, M.M.; Davis, S.J.; Hanano, S.; Moyle, R.; Milich, R.; Putterill, J.; Millar, A.J.; et al. The molecular basis of temperature compensation in the Arabidopsis circadian clock. Plant Cell 2006, 18, 1177–1187. [Google Scholar] [CrossRef] [PubMed]
- Salomé, P.A.; McClung, C.R. PSEUDO-RESPONSE REGULATOR 7 and 9 are partially redundant genes essential for the temperature responsiveness of the Arabidopsis circadian clock. Plant Cell 2005, 17, 791–803. [Google Scholar] [CrossRef] [PubMed]
- Arana, M.V.; Tognacca, R.S.; Estravis-Barcalá, M.; Sánchez, R.A.; Botto, J.F. Physiological and molecular mechanisms underlying the integration of light and temperature cues in Arabidopsis thaliana seeds. Plant Cell Environ. 2017, 40, 3113–3121. [Google Scholar] [CrossRef] [PubMed]
- Sun, J.; Qi, L.; Li, Y.; Chu, J.; Li, C. PIF4-mediated activation of YUCCA8 expression integrates temperature into the auxin pathway in regulating Arabidopsis hypocotyl growth. PLoS Genet. 2012, 8, e1002594. [Google Scholar] [CrossRef] [PubMed]
- Viczian, A.; Kircher, S.; Fejes, E.; Millar, A.J.; Schafer, E.; Kozma-Bognar, L.; Nagy, F. Functional characterization of phytochrome interacting factor 3 for the Arabidopsis thaliana circadian clockwork. Plant Cell Physiol. 2005, 46, 1591–1602. [Google Scholar] [CrossRef] [PubMed]
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shor, E.; Potavskaya, R.; Kurtz, A.; Paik, I.; Huq, E.; Green, R. PIF-Mediated Sucrose Regulation of the Circadian Oscillator is Light Quality and Temperature Dependent. Genes 2018, 9, 628. https://doi.org/10.3390/genes9120628
Shor E, Potavskaya R, Kurtz A, Paik I, Huq E, Green R. PIF-Mediated Sucrose Regulation of the Circadian Oscillator is Light Quality and Temperature Dependent. Genes. 2018; 9(12):628. https://doi.org/10.3390/genes9120628
Chicago/Turabian StyleShor, Ekaterina, Raya Potavskaya, Ayelet Kurtz, Inyup Paik, Enamul Huq, and Rachel Green. 2018. "PIF-Mediated Sucrose Regulation of the Circadian Oscillator is Light Quality and Temperature Dependent" Genes 9, no. 12: 628. https://doi.org/10.3390/genes9120628
APA StyleShor, E., Potavskaya, R., Kurtz, A., Paik, I., Huq, E., & Green, R. (2018). PIF-Mediated Sucrose Regulation of the Circadian Oscillator is Light Quality and Temperature Dependent. Genes, 9(12), 628. https://doi.org/10.3390/genes9120628