Comparative Transcriptome Analysis Reveals Hormone Signaling Genes Involved in the Launch of Culm-Shape Differentiation in Dendrocalamus sinicus
Abstract
:1. Introduction
2. Materials and Methods
2.1. Plant Materials
2.2. RNA Isolation, Library Construction, and Sequencing
2.3. De Novo Transcriptome Assembly and Functional Annotation
2.4. Functional Analysis of Differentially Expressed Genes
2.5. Validation of DEGs Using qRT-PCR
3. Results
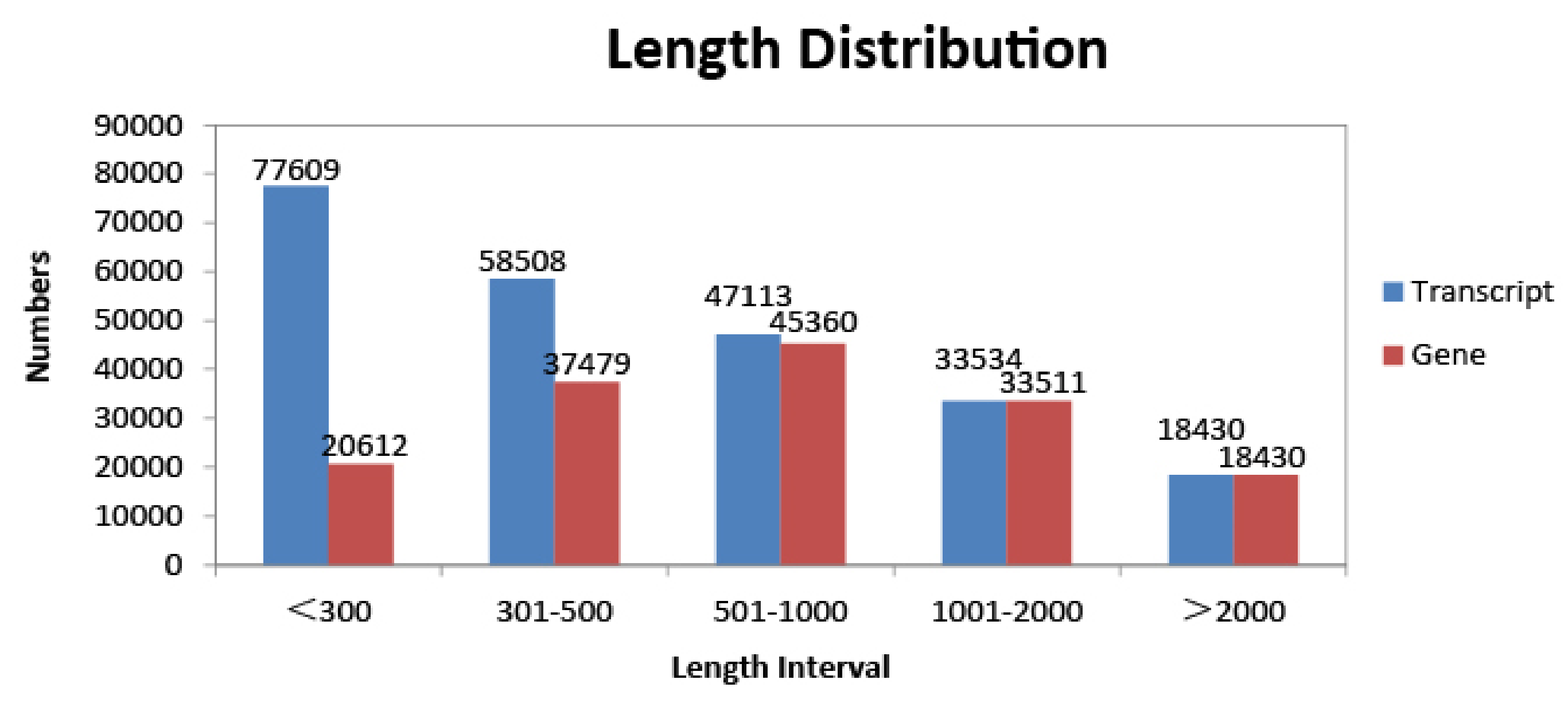
3.1. Transcriptome Sequencing and De Novo Assembly of D. sinicus
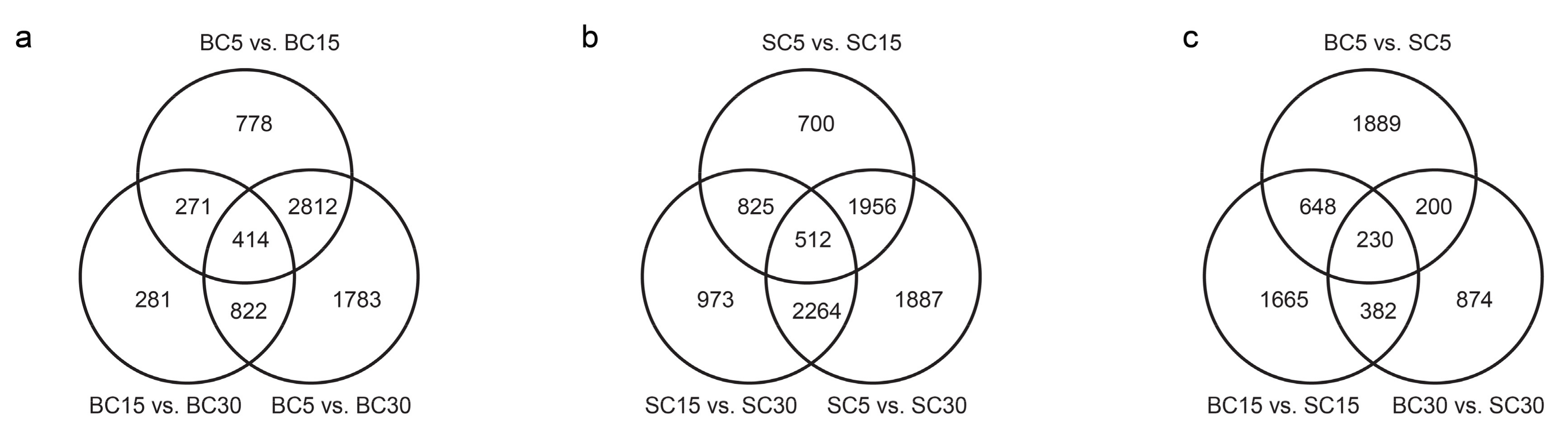
3.2. Differentially Expressed Genes Analysis during Culm Development
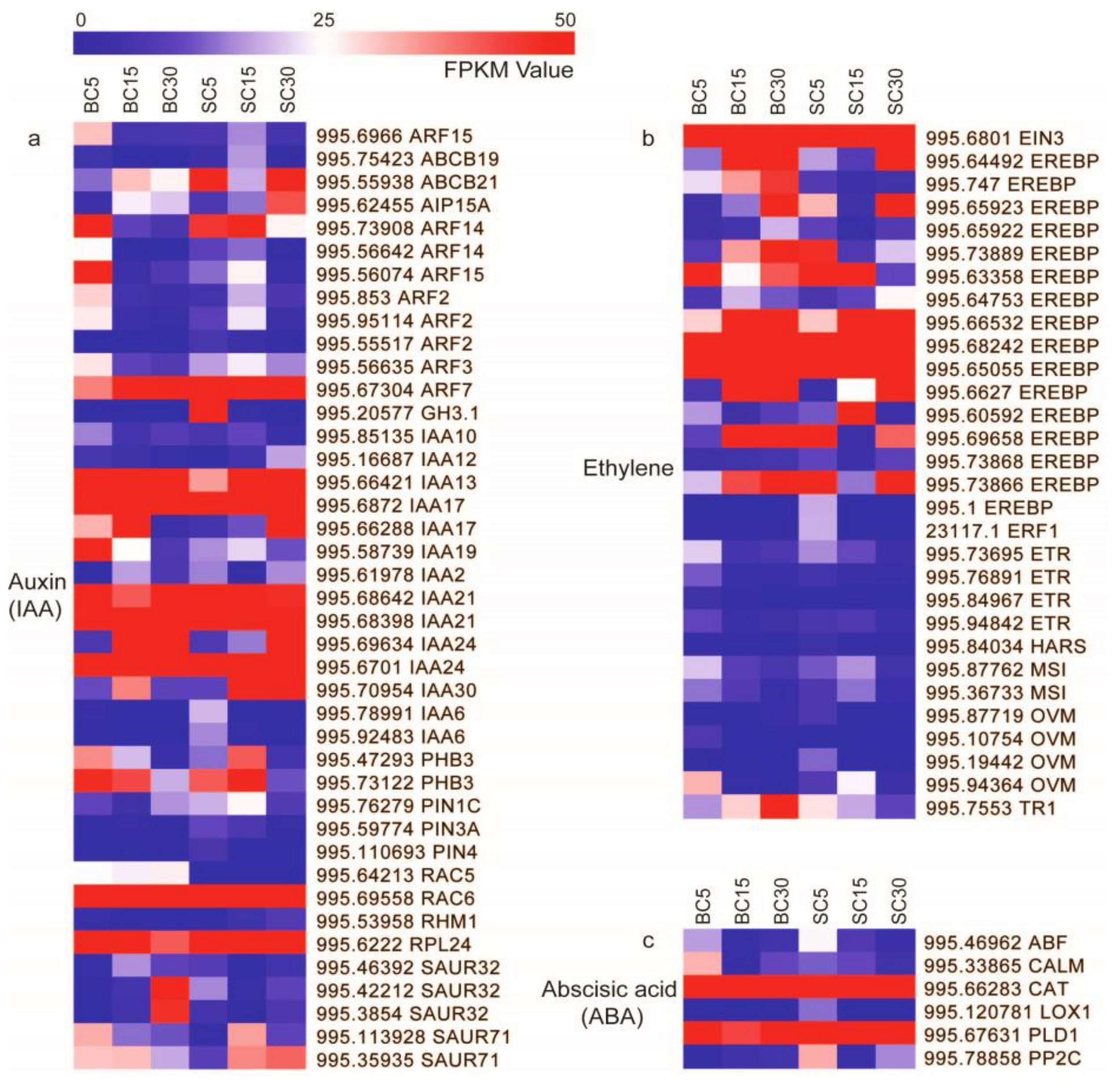
3.3. DEGs Involved in Plant Hormone Signal Transduction
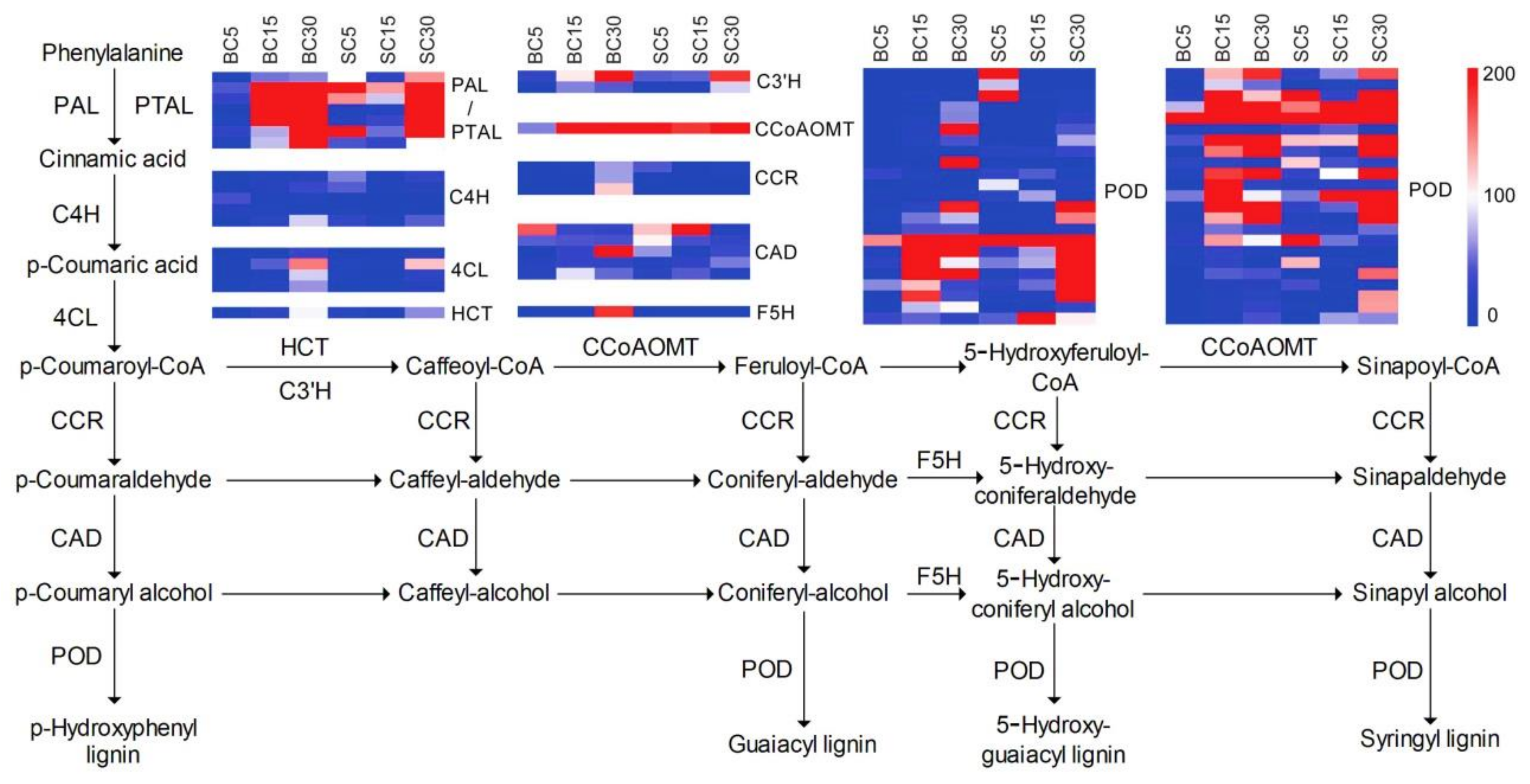
3.4. DEGs Involved in Phenylpropanoid Biosynthesis
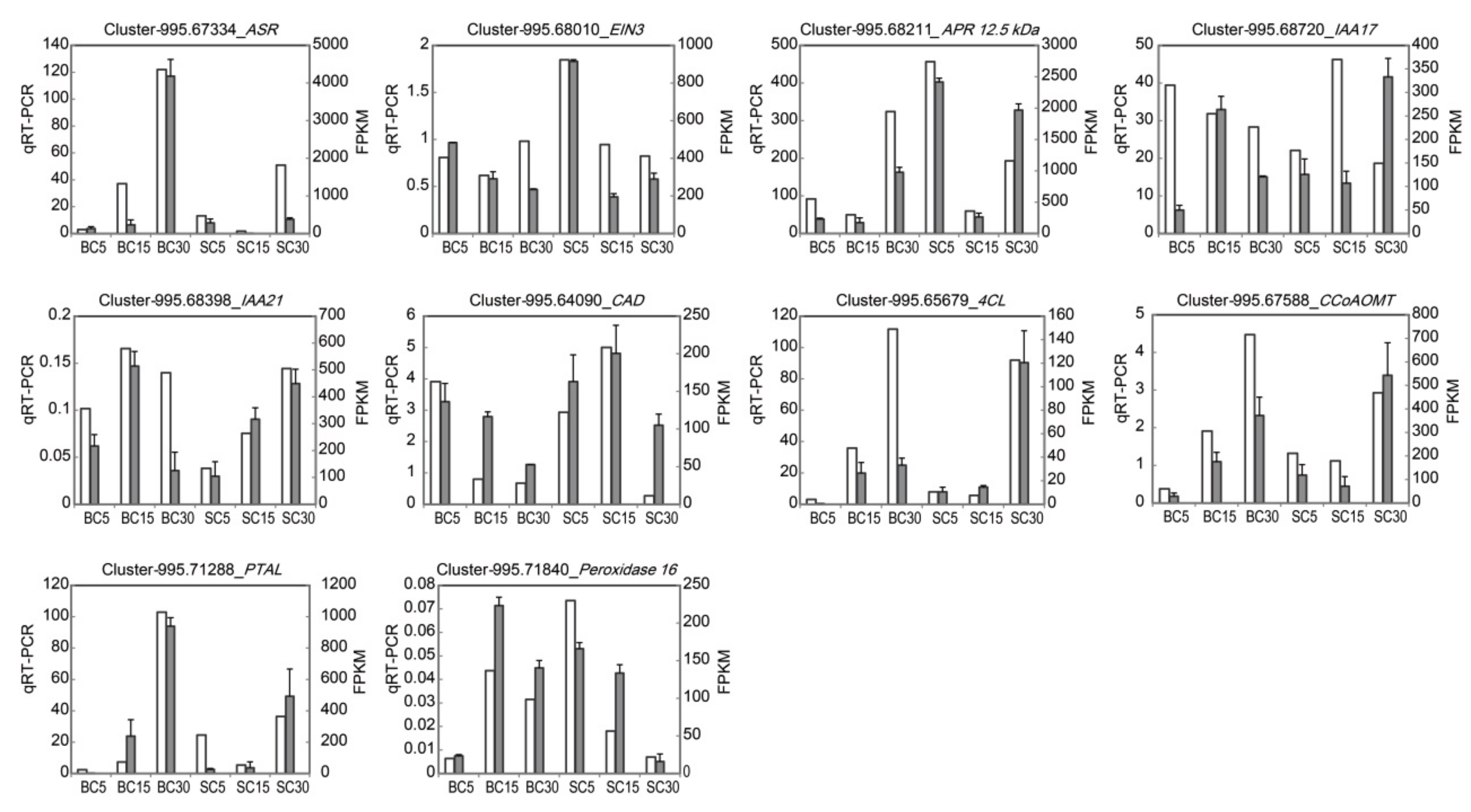
3.5. Verification of the Gene Expression through qRT-PCR
4. Discussion
4.1. Transcriptome Changes in Culm-Shape Differentiation of D. sinicus
4.2. DEGs Involved in Plant Hormone Signal Transduction during Culm-Shape Differentiation
4.3. DEGs Involved in Lignin Biosynthesis during Culm-Shape Differentiation
Supplementary Materials:
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Arteca, R.N.; Arteca, J.M. Characterization of gravitropic inflorescence bending in brassinosteroid biosynthesis and signaling Arabidopsis mutants. J. Plant Physiol. 2011, 168, 1200–1207. [Google Scholar] [CrossRef] [PubMed]
- Kato, T.; Morita, M.T.; Fukaki, H.; Yamauchi, Y.; Uehara, M.; Niihama, M.; Tasaka, M. SGR2, a phospholipase-like protein, and ZIG/SGR4, a SNARE, are involved in the shoot gravitropism of Arabidopsis. Plant Cell 2002, 14, 33–46. [Google Scholar] [CrossRef] [PubMed]
- Multani, D.S.; Briggs, S.P.; Chamberlin, M.A.; Blakeslee, J.J.; Murphy, A.S.; Johal, G.S. Loss of an MDR transporter in compact stalks of maize br2 and sorghum dw3 mutants. Science 2003, 302, 81–84. [Google Scholar] [CrossRef] [PubMed]
- Yang, W.; Ren, S.; Zhang, X.; Gao, M.; Ye, S.; Qi, Y.; Zheng, Y.; Wang, J.; Zeng, L.; Li, Q.; et al. BENT UPPERMOST INTERNODE1 encodes the class II formin FH5 crucial for actin organization and rice development. Plant Cell 2011, 23, 661–680. [Google Scholar] [CrossRef] [PubMed]
- Yamamuro, C.; Ihara, Y.; Wu, X.; Noguchi, T.; Fujioka, S.; Takatsuto, S.; Ashikari, M.; Kitano, H.; Matsuoka, M. Loss of function of a rice brassinosteroid insensitive1 homolog prevents internode elongation and bending of the lamina joint. Plant Cell 2000, 12, 1591–1606. [Google Scholar] [CrossRef] [PubMed]
- Sopian, T.; Feng, J.; Hirata, Y. Characterization of mulberry mutant growth response to gibberellin and abscisic acid application and its molecular analysis. J. Insect Biotechnol. Sericol. 2009, 78, 23–32. [Google Scholar]
- Cui, D.; Neill, S.J.; Tang, Z.; Cai, W. Gibberellin-regulated XET is differentially induced by auxin in rice leaf sheath bases during gravitropic bending. J. Exp. Bot. 2005, 56, 1327–1334. [Google Scholar] [CrossRef] [PubMed]
- Philosoph-Hadas, S.; Friedman, H.; Meir, S. Gravitropic bending and plant hormones. Vitam. Horm. 2005, 72, 31. [Google Scholar] [CrossRef] [PubMed]
- Geng, B.J.; Wang, Z.P. Gramineae Bambusiodeae. In Flora Reipublicae Popularis Sinica; Science Press: Beijing, China, 1996; Volume 9, pp. 3–16. ISBN 978-7-0302-7166-2. [Google Scholar]
- Hong, D.Y.; Blackmore, S. Plants of China. In A Companion to the Flora of China; Science Press: Beijing, China, 2005; pp. 319–320. ISBN 978-7-0303-8574-1. [Google Scholar]
- Hui, C.M.; Yang, X.Y.; Liang, N.; Chen, F. A study on the conservation and development of Dendrocalamus sinicus form Yunnan, China. Appl. Mech. Mater. 2014, 522–524, 1084–1088. [Google Scholar] [CrossRef]
- Hui, C.M.; Yang, Y.M.; Du, F. Basic Research on Eco-Biological Characteristics and Exploitation and Utilization of Dendrocalamus sinicus; Yunnan Science and Technology Press: Kunming, China, 2006; pp. 5–30. ISBN 978-7-5416-2347-9. [Google Scholar]
- Gu, Z.J.; Yang, H.Q.; Sun, M.S.; Yang, Y.M. Distribution characteristics, flowering and seeding of Dendrocalamus sinicus in Yunnan, China. For. Res. 2012, 25, 1–5. [Google Scholar]
- Pu, X.L. Studies on the Biological Characteristics of Dendrocalamus sinicus. J. Nanjing For. Univ. 2004, 28, 93. [Google Scholar]
- Li, P.; Du, F.; Pu, X.L.; Yang, Y.M. RAPD analysis of different variant types of Dendrocalamus sinicus. Acta Bot. Yunnanica 2004, 26, 290–296. [Google Scholar]
- Yang, H.Q.; Ruan, Z.Y.; Tian, B.; Yang, Y.M.; Sun, M.S. Genetic differentiation of four straight-culmed provenances of Dendrocalamus sinicus based on ISSRs. J. Zhejiang For. Coll. 2010, 27, 81–86. [Google Scholar]
- Lin, E.P.; Peng, H.Z.; Jin, Q.Y.; Deng, M.J.; Li, T.; Xiao, X.C.; Hua, X.Q.; Wang, K.H.; Bian, H.W.; Han, N.; et al. Identification and characterization of two Bamboo (Phyllostachys praecox) AP1/SQUA-like MADS-Box genes during floral transition. Planta 2009, 231, 109–120. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.M.; Zhao, L.; Larsonrabin, Z.; Li, D.Z.; Guo, Z.H. De novo sequencing and characterization of the floral transcriptome of Dendrocalamus latiflorus (Poaceae: Bambusoideae). PLoS ONE 2012, 7, e42082. [Google Scholar] [CrossRef] [PubMed]
- Biswas, P.; Chakraborty, S.; Dutta, S.; Pal, A.; Das, M. Bamboo flowering from the perspective of comparative genomics and transcriptomics. Front. Plant Sci. 2016, 7, 1900. [Google Scholar] [CrossRef] [PubMed]
- Wang, K.; Peng, H.; Lin, E.; Jin, Q.; Hua, X.; Yao, S.; Bian, H.; Han, N.; Pan, J.; Wang, J.; et al. Identification of genes related to the development of bamboo rhizome bud. J. Exp. Bot. 2010, 61, 551–561. [Google Scholar] [CrossRef] [PubMed]
- Cui, K.; He, C.Y.; Zhang, J.G.; Duan, A.G.; Zeng, Y.F. Temporal and spatial profiling of internode elongation-associated protein expression in rapidly growing culms of bamboo. J. Proteome Res. 2012, 11, 2492–2507. [Google Scholar] [CrossRef] [PubMed]
- Cui, K.; Wang, H.; Liao, S.; Tang, Q.; Li, L.; Cui, Y.; He, Y. Transcriptome sequencing and analysis for culm elongation of the world’s largest bamboo (Dendrocalamus sinicus). PLoS ONE 2016, 11, e0157362. [Google Scholar] [CrossRef] [PubMed]
- He, C.Y.; Cui, K.; Zhang, J.G.; Duan, A.G.; Zeng, Y.F. Next-generation sequencing-based mRNA and microRNA expression profiling analysis revealed pathways involved in the rapid growth of developing culms in Moso bamboo. BMC Plant Biol. 2013, 13, 119. [Google Scholar] [CrossRef] [PubMed]
- Cui, K.; Wang, H.Y.; He, C.Y.; Sun, D.; Zhang, J.P.; Liao, S.X.; Cui, Y.Z.; Li, L. Analysis of culm elongation in photoheterotrophic status of Dendrocalamus sinicus by comparative proteomics. Trees 2017, 31, 687–704. [Google Scholar] [CrossRef]
- Bhandawat, A.; Singh, G.; Seth, R.; Singh, P.; Sharma, R.K. Genome-wide transcriptional profiling to elucidate key candidates involved in bud burst and rattling growth in a subtropical bamboo (Dendrocalamus hamiltonii). Front. Plant Sci. 2017, 7, 2038. [Google Scholar] [CrossRef] [PubMed]
- Wei, Q.; Jiao, C.; Guo, L.; Ding, Y.; Cao, J.; Feng, J.; Dong, X.; Mao, L.; Sun, H.; Yu, F.; et al. Exploring key cellular processes and candidate genes regulating the primary thickening growth of Moso underground shoots. New Phytol. 2016, 214, 81–96. [Google Scholar] [CrossRef] [PubMed]
- Wu, Q.; Bai, X.; Zhao, W.; Xiang, D.; Wan, Y.; Yan, J.; Zou, L.; Zhao, G. De novo assembly and analysis of tartary buckwheat (fagopyrum tataricum Garetn.) transcriptome discloses key regulators involved in salt-stress response. Genes 2017, 8, 255. [Google Scholar] [CrossRef] [PubMed]
- Danchin, E.G.; Perfus-Barbeoch, L.; Rancurel, C.; Thorpe, P.; Da Rocha, M.; Bajew, S.; Neilson, R.; Guzeeva, E.S.; Da Silva, C.; Guy, J.; et al. The transcriptomes of Xiphinema index and Longidorus elongatus suggest independent acquisition of some plant parasitism genes by horizontal gene transfer in early-branching nematodes. Genes 2017, 8, 287. [Google Scholar] [CrossRef] [PubMed]
- Grabherr, M.G.; Grabherr, M.G.; Haas, B.J.; Yassour, M.; Levin, J.Z.; Thompson, D.A.; Amit, I.; Adiconis, X.; Fan, L.; Raychowdhury, R.; et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 2011, 29, 644–652. [Google Scholar] [CrossRef] [PubMed]
- Li, B.; Dewey, C. RSEM: Accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinform. 2011, 12, 323. [Google Scholar] [CrossRef] [PubMed]
- Anders, S.; Huber, W. Differential expression analysis for sequence count data. Genome Biol. 2010, 11, R106. [Google Scholar] [CrossRef] [PubMed]
- Mao, X.; Cai, T.; Olyarchuk, J.G.; Wei, L. Automated genome annotation and pathway identification using the KEGG Orthology (KO) as a controlled vocabulary. Bioinformatics 2005, 21, 3787–3793. [Google Scholar] [CrossRef] [PubMed]
- Untergasser, A.; Cutcutache, I.; Koressaar, T.; Ye, J.; Faircloth, B.C.; Remm, M.; Rozen, S.G. Primer3—New capabilities and interfaces. Nucleic Acids Res. 2012, 40, e115. [Google Scholar] [CrossRef] [PubMed]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2–ΔΔCT Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Kuhl, J.C.; Cheung, F.; Yuan, Q.; Martin, W.; Zewdie, Y.; McCallum, J.; Catanach, A.; Rutherford, P.; Sink, K.C.; Jenderek, M.; et al. A unique set of 11,008 onion expressed sequence tags reveals expressed sequence and genomic differences between the monocot orders Asparagales and Poales. Plant Cell 2004, 16, 114–125. [Google Scholar] [CrossRef] [PubMed]
- Yang, M.; Zhu, L.; Pan, C.; Xu, L.; Liu, Y.; Ke, W.; Yang, P. Transcriptomic analysis of the regulation of rhizome formation in temperate and tropical lotus (Nelumbo nucifera). Sci. Rep. 2015, 5, 13059. [Google Scholar] [CrossRef] [PubMed]
- Whitford, R.; Fernandez, A.; Tejos, R.; Pérez, A.C.; Kleine-Vehn, J.; Vanneste, S.; Drozdzecki, A.; Leitner, J.; Abas, L.; Aerts, M.; et al. GOLVEN secretory peptides regulate auxin carrier turnover during plant gravitropic responses. Dev. Cell 2012, 22, 678–685. [Google Scholar] [CrossRef] [PubMed]
- Vogt, T.; Dudareva, N.; Pichersky, E.; Werckreichhart, D.; Lewinsohn, E. Phenylpropanoid biosynthesis. Mol. Plant 2010, 3, 2–20. [Google Scholar] [CrossRef] [PubMed]
- Fraser, C.M.; Chapple, C. The phenylpropanoid pathway in Arabidopsis. Arab. Book 2011, 9, e0152. [Google Scholar] [CrossRef] [PubMed]
- Chopra, R.; Burow, G.; Farmer, A.; Mudge, J.; Simpson, C.E.; Burow, M.D. Comparisons of de novo transcriptome assemblers in diploid and polyploid species using peanut (Arachis spp.) RNA-Seq data. PLoS ONE 2014, 9, e115055. [Google Scholar] [CrossRef] [PubMed]
- Peng, Z.; Lu, Y.; Li, L.; Zhao, Q.; Feng, Q.; Gao, Z.; Lu, H.; Hu, T.; Yao, N.; Liu, K.; et al. The draft genome of the fast-growing non-timber forest species moso bamboo (Phyllostachys heterocycla). Nat. Genet. 2013, 45, 456–461. [Google Scholar] [CrossRef] [PubMed]
- Liu, M.; Qiao, G.; Jiang, J.; Yang, H.; Xie, L.; Xie, J.; Zhuo, R. Transcriptome sequencing and de novo analysis for Ma bamboo (Dendrocalamus latiflorus Munro) using the Illumina platform. PLoS ONE 2012, 7, e46766. [Google Scholar] [CrossRef] [PubMed]
- Pnueli, L.; Carmel-Goren, L.; Hareven, D.; Gutfinger, T.; Alvarez, J.; Ganal, M.; Zamir, D.; Lifschitz, E. The SELF-PRUNING gene of tomato regulates vegetative to reproductive switching of sympodial meristems and is the ortholog of CEN and TFL1. Development 1998, 125, 1979–1989. [Google Scholar] [PubMed]
- Li, L.; Cheng, Z.; Ma, Y.; Bai, Q.; Li, X.; Cao, Z.; Wu, Z.; Gao, J. The association of hormone signaling genes, transcription and changes in shoot anatomy during moso bamboo growth. Plant Biotechnol. J. 2017. [Google Scholar] [CrossRef]
- Song, Y.; Xu, Z.F. Ectopic overexpression of an AUXIN/INDOLE-3-ACETIC ACID (Aux/IAA) gene OsIAA4 in rice induces morphological changes and reduces responsiveness to auxin. Int. J. Mol. Sci. 2013, 14, 13645–13656. [Google Scholar] [CrossRef] [PubMed]
- Gamuyao, R.; Nagai, K.; Ayano, M.; Mori, Y.; Minami, A.; Kojima, M.; Suzuki, T.; Sakakibara, H.; Higashiyama, T.; Ashikari, M.; et al. Hormone distribution and transcriptome profiles in bamboo shoots provide insights on bamboo stem emergence and growth. Plant Cell Physiol. 2017, 58, 702–716. [Google Scholar] [CrossRef] [PubMed]
- Bennett, M.J.; Marchant, A.; Green, H.G.; May, S.T.; Ward, S.P.; Millner, P.A.; Walker, A.R.; Schulz, B.; Feldmann, K.A. Arabidopsis AUX1 gene: A permease-like regulator of root gravitropism. Science 1996, 273, 948–950. [Google Scholar] [CrossRef] [PubMed]
- Tatematsu, K.; Kumagai, S.; Muto, H.; Sato, A.; Watahiki, M.K.; Harper, R.M.; Liscum, E.; Yamamoto, K.T. MASSUGU2 encodes Aux/IAA19, an auxin-regulated protein that functions together with the transcriptional activator NPH4/ARF7 to regulate differential growth responses of hypocotyl and formation of lateral roots in Arabidopsis thaliana. Plant Cell 2004, 16, 379–393. [Google Scholar] [CrossRef] [PubMed]
- Hirayama, T.; Umezawa, T. The PP2C-SnRK2 complex: The central regulator of an abscisic acid signaling pathway. Plant Signal. Behav. 2010, 5, 160–163. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.; Zhu, D.; Huang, X.; Li, S.; Gong, Y.; Yao, Q.; Fu, X.; Fan, L.M.; Deng, X.W. Biochemical insights on degradation of Arabidopsis DELLA proteins gained from a cell-free assay system. Plant Cell 2009, 21, 2378–2390. [Google Scholar] [CrossRef] [PubMed]
- Yruela, I. Plant development regulation: Overview and perspectives. J. Plant Physiol. 2015, 182, 62–78. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.Y.; Hsieh, M.H.; Yang, C.C.; Lin, C.S.; Wang, A.Y. Analysis of the cellulose synthase genes associated with primary cell wall synthesis in Bambusa oldhamii. Phytochemistry 2010, 71, 1270–1279. [Google Scholar] [CrossRef] [PubMed]
- Carpita, N.C. Update on mechanisms of plant cell wall biosynthesis: How plants make cellulose and other (1→4)-β-d-glycans. Plant Physiol. 2011, 155, 171–184. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Popko, J.L.; Umezawa, T.; Chiang, V.L. 5-Hydroxyconiferyl aldehyde modulates enzymatic methylation for syringyl monolignol formation, a new view of monolignol biosynthesis in angiosperms. J. Biol. Chem. 2000, 275, 6537–6545. [Google Scholar] [CrossRef] [PubMed]
- Vanholme, R.; Morreel, K.; Ralph, J.; Boerjan, W. Lignin engineering. Curr. Opin. Plant Biol. 2008, 11, 278–285. [Google Scholar] [CrossRef] [PubMed]
- Sewalt, V.; Ni, W.; Blount, J.W.; Jung, H.G.; Masoud, S.A.; Howles, P.A.; Lamb, C.; Dixon, R.A. Reduced lignin content and altered lignin composition in transgenic tobacco down-regulated in expression of l-Phenylalanine Ammonia-lyase or Cinnamate 4-Hydroxylase. Plant Physiol. 1997, 115, 41–50. [Google Scholar] [CrossRef] [PubMed]
- Guo, D.; Chen, F.; Inoue, K.; Blount, J.W.; Dixon, R.A. Downregulation of caffeic acid 3-O-methyltransferase and caffeoyl CoA 3-O-methyltransferase in transgenic alfalfa: Impacts on lignin structure and implications for the biosynthesis of G and S lignin. Plant Cell 2001, 13, 73–88. [Google Scholar] [CrossRef] [PubMed]
- Wagner, A.; Tobimatsu, Y.; Phillips, L.; Flint, H.; Torr, K.; Donaldson, L.; Pears, L.; Ralph, J. CCoAOMT suppression modifies lignin composition in Pinus radiata. Plant J. 2011, 67, 119–129. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Chen, W.; Zhao, Y.; Xiang, Y.; Jiang, H.; Zhu, S.; Cheng, B. Downregulation of caffeoyl-CoA O-methyltransferase (CCoAOMT) by RNA interference leads to reduced lignin production in maize straw. Genet. Mol. Biol. 2013, 36, 540–546. [Google Scholar] [CrossRef] [PubMed]
- Song, J.; Wang, Z. RNAi-mediated suppression of the phenylalanine ammonia-lyase gene in Salvia miltiorrhiza causes abnormal phenotypes and a reduction in rosmarinic acid biosynthesis. J. Plant Res. 2011, 124, 183–192. [Google Scholar] [CrossRef] [PubMed]

| Gene ID | Description | Primer | |
|---|---|---|---|
| Forward (5′-3′) | Reverse (5′-3′) | ||
| Cluster-995.67334 | ASR | GCGAGAACGAGTACGAGAGGT | GATCTCCTCCGCGATCTTGTG |
| Cluster-995.68010 | EIN3 | CGGTCGAAGCGGACCTTCTC | GCAGGACGGCATCCTCAAGTA |
| Cluster-995.68211 | ARP 12.5 kDa | GTACTCGGACAGTGCAGGCTAGA | TGGTGGCGGCCTTGCGGAGCTT |
| Cluster-995.68720 | IAA17 | CTCCGGTTCATGTCCTGCTGCT | AACGCGGCGGTCCGGCGACTCG |
| Cluster-995.68398 | IAA21 | CACAGGAGCGGGACTACATC | CGCCCTTGGGCAGCAG |
| Cluster-995.64090 | CAD | GATTCGTCCGGTGTCCTCTC | GGGAACAACGGGGTACAGAG |
| Cluster-995.65679 | 4CL | GGGGACTTCACCAGCAAGTT | GCACACTGGAGACATCGGTT |
| Cluster-995.67588 | CCoAOMT | TAGTTGTCCTTGTCGGCGTC | TCAACCGCGAGAACTACGAG |
| Cluster-995.71288 | PTAL | AGAGCTCCGTCAAGAACTGC | TCTGCATCAGCGGGTAGTTG |
| Cluster-995.71840 | Peroxidase 16 | TGGTAGTAGCCGTTGTCGAAC | GCGGCTGTACACGTTCAAG |
| BC5 | BC15 | BC30 | SC5 | SC15 | SC30 | |
|---|---|---|---|---|---|---|
| Number of raw reads | 58,731,962 | 65,460,126 | 59,431,136 | 54,367,904 | 56,408,792 | 57,867,298 |
| Number of clean reads (%) | 56,597,392 (96.37%) | 63,275,200 (96.66%) | 57,515,272 (96.78%) | 53,084,380 (97.64%) | 55,287,738 (98.01%) | 54,692,784 (94.51%) |
| Q20 (%) | 97.29 | 97.49 | 97.4 | 97.29 | 97.40 | 97.63 |
| Mapped reads (%) | 39,026,306 (68.95%) | 44,519,052 (70.36%) | 39,786,978 (69.18%) | 36,775,186 (69.28%) | 39,279,750 (71.05%) | 37,541,394 (68.64%) |
| GC (%) | 52.01 | 52.31 | 54.27 | 53.44 | 54.07 | 51.24 |
| Development Stages | KEGG Categories | Rich Factor | Corrected p-Value | No. of Unigenes |
|---|---|---|---|---|
| BC5 vs. SC5 | Plant hormone signal transduction | 0.089 | 0.002 | 60 |
| alpha-Linolenic acid metabolism | 0.158 | 0.002 | 18 | |
| Stilbenoid, diarylheptanoid, and gingerol biosynthesis | 0.222 | 0.002 | 12 | |
| Phenylalanine metabolism | 0.150 | 0.002 | 19 | |
| Phenylpropanoid biosynthesis | 0.099 | 0.003 | 39 | |
| BC15 vs. SC15 | Phenylpropanoid biosynthesis | 0.127 | 5.52 × 10−6 | 50 |
| Cyanoamino acid metabolism | 0.169 | 1.94 × 10−4 | 23 | |
| Steroid biosynthesis | 0.197 | 0.004 | 13 | |
| Citrate cycle (TCA cycle) | 0.119 | 0.021 | 21 | |
| Glycolysis/Gluconeogenesis | 0.087 | 0.038 | 40 | |
| BC30 vs. SC30 | Phenylpropanoid biosynthesis | 0.102 | 1.23 × 10−11 | 40 |
| Flavonoid biosynthesis | 0.137 | 0.001 | 10 | |
| Flavone and flavonol biosynthesis | 0.455 | 0.001 | 5 | |
| Glyoxylate and dicarboxylate metabolism | 0.082 | 0.001 | 16 | |
| Oxidative phosphorylation | 0.058 | 0.001 | 25 |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chen, L.; Guo, X.; Cui, Y.; Zheng, X.; Yang, H. Comparative Transcriptome Analysis Reveals Hormone Signaling Genes Involved in the Launch of Culm-Shape Differentiation in Dendrocalamus sinicus. Genes 2018, 9, 4. https://doi.org/10.3390/genes9010004
Chen L, Guo X, Cui Y, Zheng X, Yang H. Comparative Transcriptome Analysis Reveals Hormone Signaling Genes Involved in the Launch of Culm-Shape Differentiation in Dendrocalamus sinicus. Genes. 2018; 9(1):4. https://doi.org/10.3390/genes9010004
Chicago/Turabian StyleChen, Lingna, Xiaojuan Guo, Yongzhong Cui, Xianggan Zheng, and Hanqi Yang. 2018. "Comparative Transcriptome Analysis Reveals Hormone Signaling Genes Involved in the Launch of Culm-Shape Differentiation in Dendrocalamus sinicus" Genes 9, no. 1: 4. https://doi.org/10.3390/genes9010004
APA StyleChen, L., Guo, X., Cui, Y., Zheng, X., & Yang, H. (2018). Comparative Transcriptome Analysis Reveals Hormone Signaling Genes Involved in the Launch of Culm-Shape Differentiation in Dendrocalamus sinicus. Genes, 9(1), 4. https://doi.org/10.3390/genes9010004





