Identification of the Ovine Keratin-Associated Protein 26-1 Gene and Its Association with Variation in Wool Traits
Abstract
1. Introduction
2. Materials and Methods
2.1. Sheep Blood and Wool Samples
2.2. Search for an Ovine Homologue of Human KRTAP26-1 in the Sheep Genome
2.3. PCR Primers and Amplification of Sheep Genomic DNA
2.4. Screening for Variation in KRTAP26-1
2.5. Sequencing of Allelic Variants and Sequence Analysis
2.6. Statistical Analyses
3. Results
3.1. Correlations between Wool Traits
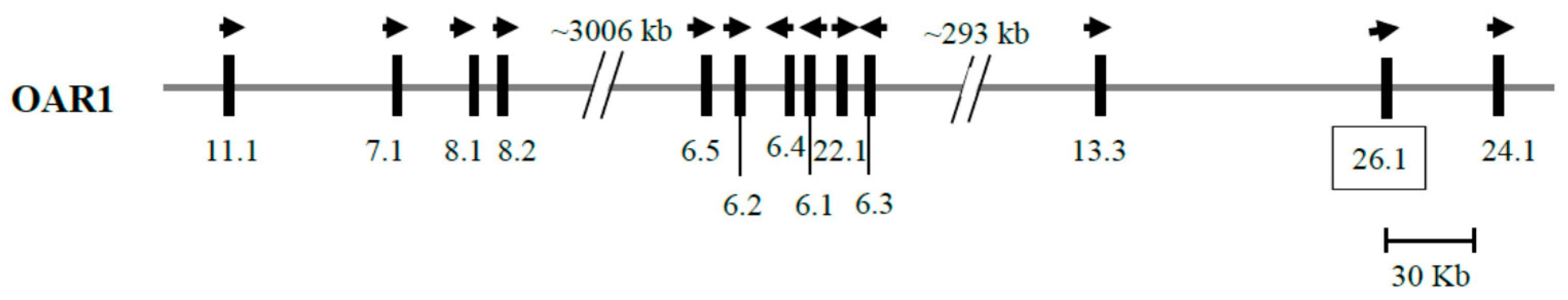
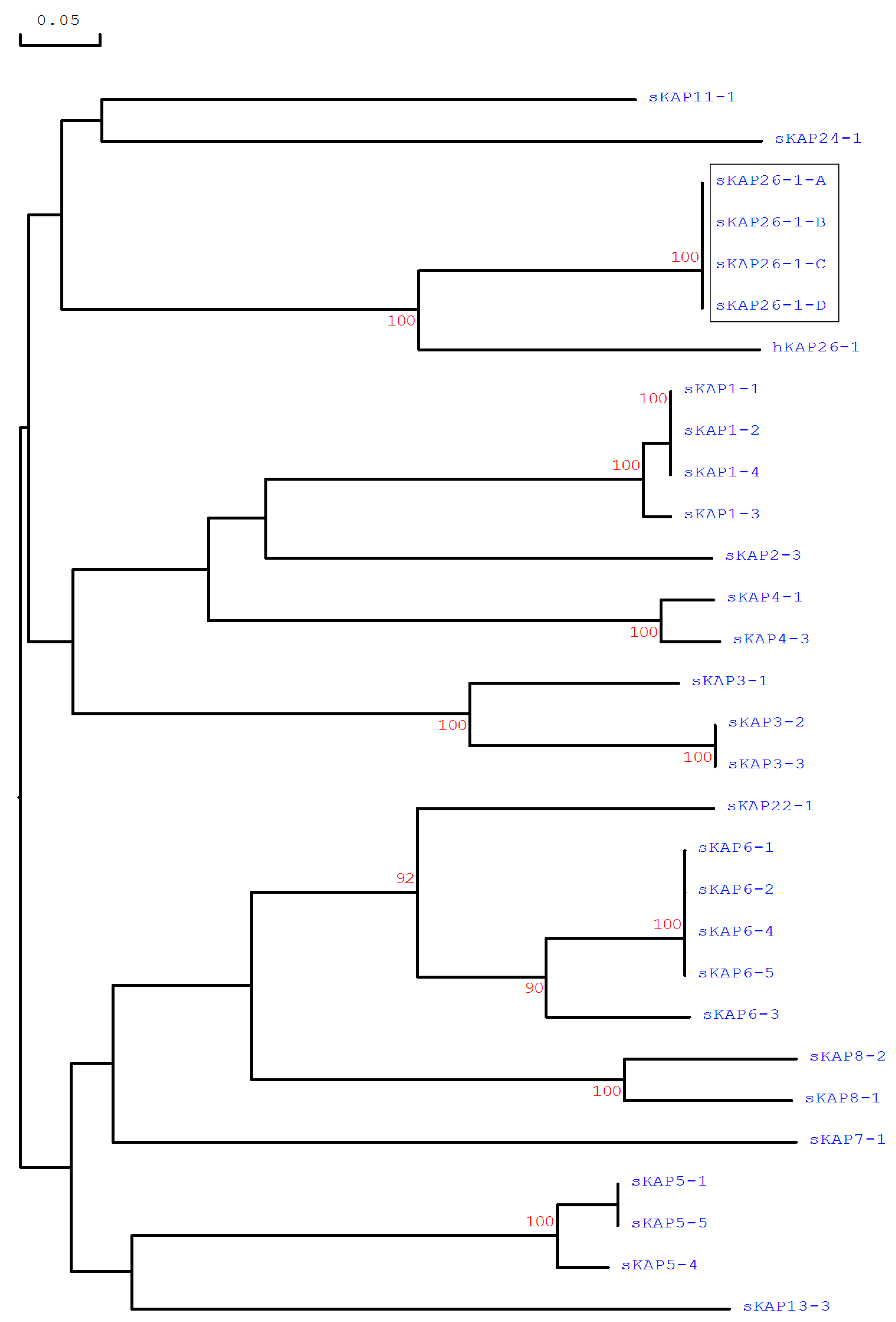
3.2. Identification of KRTAP26-1 in the Sheep Genome
3.3. Variation in Ovine KRTAP26-1
3.4. Comparison of Variant and Genotype Frequencies between NZ Romney and Merino × Southdown-Cross Sheep
3.5. Effect of Variation in KRTAP26-1 on Wool Traits
3.6. Effect of Common Genotypes on Wool Traits
4. Discussion
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Powell, B.C.; Rogers, G.E. The role of keratin proteins and their genes in the growth, structure and properties of hair. In Formation and Structure of Human Hair; Jollès, P., Zahn, H., Höcker, H., Eds.; Birkhäuser Verlag: Basel, Switzerland, 1997; pp. 59–148. [Google Scholar]
- Gong, H.; Zhou, H.; McKenzie, G.W.; Yu, Z.; Clerens, S.; Dyer, J.M.; Plowman, J.E.; Wright, M.W.; Arora, R.; Bawden, C.S.; et al. An updated nomenclature for keratin-associated proteins (KAPs). Int. J. Biol. Sci. 2012, 8, 258–264. [Google Scholar] [CrossRef] [PubMed]
- Gong, H.; Zhou, H.; Forrest, R.H.; Li, S.; Wang, J.; Dyer, J.M.; Luo, Y.; Hickford, J.G. Wool keratin-associated protein genes in sheep—a review. Genes 2016, 7, 24. [Google Scholar] [CrossRef] [PubMed]
- Rogers, M.A.; Schweizer, J. Human KAP genes, only the half of it? Extensive size polymorphisms in hair keratin-associated protein genes. J. Investig. Dermatol. 2005, 124, vii–ix. [Google Scholar] [CrossRef] [PubMed]
- Rogers, M.A.; Winter, H.; Langbein, L.; Wollschläger, A.; Praetzel-Wunder, S.; Jave-Suarez, L.F.; Schweizer, J. Characterization of human KAP24.1, a cuticular hair keratin-associated protein with unusual amino-acid composition and repeat structure. J. Investig. Dermatol. 2007, 127, 1197–1204. [Google Scholar] [CrossRef] [PubMed]
- Rogers, M.A.; Langbein, L.; Praetzel Wunder, S.; Giehl, K. Characterization and expression analysis of the hair keratin associated protein KAP26.1. Brit. J. Dermatol. 2008, 159, 725–729. [Google Scholar] [CrossRef] [PubMed]
- Zhou, H.; Hickford, J.G.H.; Fang, Q. A two-step procedure for extracting genomic DNA from dried blood spots on filter paper for polymerase chain reaction amplification. Anal. Biochem. 2006, 354, 159–161. [Google Scholar] [CrossRef] [PubMed]
- Byun, S.O.; Fang, Q.; Zhou, H.; Hickford, J.G.H. An effective method for silver-staining DNA in large numbers of polyacrylamide gels. Anal. Biochem. 2009, 385, 174–175. [Google Scholar] [CrossRef] [PubMed]
- Gong, H.; Zhou, H.; Hickford, J.G.H. Diversity of the glycine/tyrosine-rich keratin-associated protein 6 gene (KAP6) family in sheep. Mol. Biol. Rep. 2011, 38, 31–35. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Zhou, H.; Gong, H.; Zhao, F.; Wang, J.; Liu, X.; Luo, Y.; Hickford, J.G. Identification of the ovine keratin-associated protein 22-1 (KAP22-1) gene and its effect on wool traits. Genes 2017, 8, 27. [Google Scholar] [CrossRef] [PubMed]
- Gong, H.; Zhou, H.; Hodge, S.; Dyer, J.M.; Hickford, J.G. Association of wool traits with variation in the ovine KAP1-2 gene in Merino cross lambs. Small Rum. Res. 2015, 124, 24–29. [Google Scholar] [CrossRef]
- Safari, E.; Fogarty, N.M.; Gilmour, A.R.; Atkins, K.D.; Mortimer, S.I.; Swan, A.A.; Brien, F.D.; Greeff, J.C.; van der Werf, J.H. Genetic correlations among and between wool, growth and reproduction traits in Merino sheep. J. Anim. Breed. Genet. 2007, 124, 65–72. [Google Scholar] [CrossRef] [PubMed]
- Huisman, A.; Brown, D. Genetic parameters for bodyweight, wool, and disease resistance and reproduction traits in Merino sheep. 4. Genetic relationships between and within wool traits. Anim. Prod. Sci. 2009, 49, 289–296. [Google Scholar] [CrossRef]
- Zhou, H.; Gong, H.; Yan, W.; Luo, Y.Z.; Hickford, J.G.H. Identification and sequence analysis of the keratin-associated protein 24-1 (KAP24-1) gene homologue in sheep. Gene 2012, 511, 62–65. [Google Scholar] [CrossRef] [PubMed]
- Gong, H.; Zhou, H.; Dyer, J.M.; Hickford, J.G. Identification of the ovine KAP11-1 gene (KRTAP11-1) and genetic variation in its coding sequence. Mol. Biol. Rep. 2011, 38, 5429–5433. [Google Scholar] [CrossRef] [PubMed]
- Gong, H.; Zhou, H.; Dyer, J.M.; Plowman, J.E.; Hickford, J.G. Identification of the keratin-associated protein 13-3 (KAP13-3) gene in sheep. Open J. Gen. 2011, 1, 60–64. [Google Scholar] [CrossRef]
- Nakamura, A.; Arimoto, M.; Takeuchi, K.; Fujii, T. A rapid extraction procedure of human hair proteins and identification of phosphorylated species. Biol. Pharm. Bull. 2002, 25, 569–572. [Google Scholar] [CrossRef] [PubMed]
- Zhou, H.; Hickford, J.G.; Fang, Q.; Lin, Y.S. Allelic variation of the ovine Toll-like receptor 4 gene. Dev. Comp. Immunol. 2007, 1, 105–108. [Google Scholar] [CrossRef] [PubMed]
- Gong, H.; Zhou, H.; Hickford, J.G. Polymorphism of the ovine keratin-associated protein 1-4 gene (KRTAP1-4). Mol. Biol. Rep. 2010, 37, 3377–3380. [Google Scholar] [CrossRef] [PubMed]
- Gong, H.; Zhou, H.; Yu, Z.; Dyer, J.; Plowman, J.E.; Hickford, J. Identification of the ovine keratin-associated protein KAP1-2 gene (KRTAP1-2). Exp. Dermatol. 2011, 20, 815–819. [Google Scholar] [CrossRef] [PubMed]
- Nataraj, A.J.; OlivosBGlander, I.; Kusukawa, N.; Highsmith, W.E., Jr. Single-strand conformation polymorphism and heteroduplex analysis for gel-based mutation detection. Electrophoresis 1999, 20, 1177–1185. [Google Scholar] [CrossRef]
- Holman, B.W.B.; Malau-Aduli, A.E.O. A review of sheep wool quality traits. Annu. Rev. Res. Biol. 2012, 2, 1–14. [Google Scholar]
- Rippon, J.A. The structure of wool. In The Coloration of Wool and Other Keratin Fibres; Lewis, D.M., Rippon, J.A., Eds.; John Wiley & Sons in association with the Society of Dyers and Colourists; Plenum Press: Bradford, West Yorkshire, UK, 2013; pp. 1–42. [Google Scholar]

| GFW | CFW | Yield | MSL | MFD | FDSD | CVFD | MSS | MFC | |
|---|---|---|---|---|---|---|---|---|---|
| CFW | 0.916 *** | ||||||||
| Yield | 0.165 ** | 0.528 *** | |||||||
| MSL | 0.463 *** | 0.525 *** | 0.341 *** | ||||||
| MFD | 0.053 | −0.091 | −0.341 *** | −0.270 *** | |||||
| FDSD | 0.160 ** | 0.263 *** | 0.309 *** | 0.350 *** | 0.796 *** | ||||
| CVFD | 0.304 *** | 0.334 *** | 0.174 ** | 0.306 *** | 0.306 *** | 0.815 *** | |||
| MSS | 0.188 *** | 0.220 *** | 0.143 *** | 0.067 | 0.069 | −0.143 ** | 0.290 *** | ||
| MFC | 0.329 *** | 0.492 *** | 0.550 *** | 0.442 *** | 0.425 *** | 0.397 *** | 0.238 *** | 0.079 | |
| PF | 0.022 | 0.085 | 0.251 *** | 0.244 *** | 0.819 *** | 0.794 *** | 0.440 *** | 0.061 | 0.224 *** |
| Nucleotide Position | Variant | Amino Acid Change | |||
|---|---|---|---|---|---|
| KRTAP26-1*A | KRTAP26-1*B | KRTAP26-1*C | KRTAP26-1*D | ||
| c.72 | C | T | C | C | No change |
| c.111 | A | C | T | A | No change |
| c.123 | C | G | C | C | No change |
| c.215 | G | A | G | G | p.Cys72Tyr |
| c.277 | A | A | G | A | p.Ser93Gly |
| c.336 | G | G | G | A | No change |
| c.549 | C | T | C | C | No change |
| Trait | Variant Assessed | n | Single-Variant Model | Multi-Variant Model | ||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Absent | Present | Absent | Present | p | Variants Fitted | Absent | Present | p | ||
| GFW | A | 71 | 312 | 2.37 ± 0.06 | 2.37 ± 0.03 | 0.958 | ||||
| (kg) | B | 207 | 176 | 2.37 ± 0.04 | 2.37 ± 0.04 | 0.902 | ||||
| C | 212 | 171 | 2.39 ± 0.03 | 2.33 ± 0.05 | 0.361 | |||||
| CFW | A | 71 | 312 | 1.72 ± 0.05 | 1.71 ± 0.02 | 0.764 | ||||
| (kg) | B | 207 | 176 | 1.72 ± 0.03 | 1.70 ± 0.03 | 0.678 | ||||
| C | 212 | 171 | 1.71 ± 0.03 | 1.72 ± 0.04 | 0.877 | |||||
| Yield | A | 71 | 312 | 72.2 ± 0.74 | 71.7 ± 0.35 | 0.471 | B, C | 73.0 ± 0.82 | 71.8 ± 0.41 | 0.201 |
| (%) | B | 207 | 176 | 72.4 ± 0.47 | 71.2 ± 0.44 | 0.070 | C | 72.4 ± 0.47 | 71.7 ± 0.52 | 0.291 |
| C | 212 | 171 | 71.2 ± 0.40 | 73.0 ± 0.65 | 0.027 | B | 71.3 ± 0.41 | 72.8 ± 0.68 | 0.082 | |
| MSL (mm) | A | 71 | 312 | 84.7 ± 1.66 | 82.6 ± 0.87 | 0.253 | C | 85.6 ± 1.64 | 84.0 ± 0.93 | 0.377 |
| B | 207 | 176 | 83.9 ± 1.12 | 82.2 ± 1.04 | 0.236 | C | 84.3 ± 1.10 | 84.4 ± 1.18 | 0.950 | |
| C | 212 | 171 | 80.7 ± 0.99 | 87.9 ± 1.51 | <0.001 | None | 80.7 ± 0.99 | 87.9 ± 1.51 | <0.001 | |
| MFD | A | 71 | 312 | 19.8 ± 0.25 | 19.6 ± 0.13 | 0.375 | B, C | 19.6 ± 0.27 | 19.3 ± 0.14 | 0.397 |
| (µm) | B | 207 | 176 | 19.3 ± 0.17 | 19.9 ± 0.15 | 0.001 | C | 19.2 ± 0.16 | 19.6 ± 0.17 | 0.081 |
| C | 212 | 171 | 20.1 ± 0.15 | 18.6 ± 0.22 | <0.001 | B | 20.0 ± 0.15 | 18.7 ± 0.23 | <0.001 | |
| FDSD | A | 71 | 312 | 4.26 ± 0.09 | 4.22 ± 0.04 | 0.680 | B, C | 4.21 ± 0.09 | 4.13 ± 0.04 | 0.450 |
| (µm) | B | 207 | 176 | 4.16 ± 0.06 | 4.28 ± 0.05 | 0.098 | C | 4.13 ± 0.06 | 4.17 ± 0.06 | 0.626 |
| C | 212 | 171 | 4.35 ± 0.05 | 3.96 ± 0.08 | <0.001 | B | 4.34 ± 0.05 | 3.96 ± 0.08 | <0.001 | |
| CVFD | A | 71 | 312 | 21.4 ± 0.28 | 21.9 ± 0.14 | 0.166 | C | 21.4 ± 0.28 | 21.7 ± 0.15 | 0.248 |
| (%) | B | 207 | 176 | 21.9 ± 0.18 | 21.7 ± 0.17 | 0.471 | A, C | 21.7 ± 0.26 | 21.4 ± 0.20 | 0.330 |
| C | 212 | 171 | 22.0 ± 0.16 | 21.4 ± 0.25 | 0.067 | A | 21.8 ± 0.19 | 21.3 ± 0.26 | 0.093 | |
| MSS | A | 71 | 312 | 23.9 ± 1.06 | 22.6 ± 0.55 | 0.237 | B | 23.6 ± 1.19 | 22.7 ± 0.55 | 0.496 |
| (N/ktex) | B | 207 | 176 | 22.1 ± 0.71 | 23.4 ± 0.65 | 0.145 | None | 22.1 ± 0.71 | 23.4 ± 0.65 | 0.145 |
| C | 212 | 171 | 23.1 ± 0.64 | 22.2 ± 0.98 | 0.434 | B | 22.9 ± 0.66 | 22.6 ± 1.01 | 0.774 | |
| MFC | A | 71 | 312 | 91.2 ± 1.80 | 89.5 ± 0.85 | 0.367 | B, C | 89.4 ± 2.00 | 89.1 ± 1.01 | 0.862 |
| (°/mm) | B | 207 | 176 | 87.7 ± 1.15 | 91.4 ± 1.06 | 0.016 | C | 87.6 ± 1.16 | 90.7 ± 1.26 | 0.056 |
| C | 212 | 171 | 90.8 ± 0.98 | 87.2 ± 1.60 | 0.070 | B | 90.3 ± 1.02 | 88.0 ± 1.65 | 0.278 | |
| PF | A | 71 | 312 | 3.00 ± 0.45 | 2.79 ± 0.23 | 0.654 | B, C | 2.55 ± 0.49 | 2.44 ± 0.26 | 0.845 |
| (%) | B | 207 | 176 | 2.33 ± 0.30 | 3.24 ± 0.27 | 0.017 | C | 2.23 ± 0.29 | 2.69 ± 0.31 | 0.237 |
| C | 212 | 171 | 3.47 ± 0.26 | 1.41 ± 0.40 | <0.001 | B | 3.39 ± 0.27 | 1.53 ± 0.41 | <0.001 | |
| Trait | AA (n = 68) | AB (n = 116) | AC (n = 123) | BB (n = 22) | BC (n = 37) | p |
|---|---|---|---|---|---|---|
| GFW (kg) | 2.39 ± 0.05 | 2.36 ± 0.04 | 2.36 ± 0.05 | 2.39 ± 0.09 | 2.40 ± 0.08 | 0.971 |
| CFW (kg) | 1.72 ± 0.04 | 1.68 ± 0.03 | 1.72 ± 0.04 | 1.69 ± 0.07 | 1.77 ± 0.06 | 0.814 |
| Yield (%) | 73.5 ± 1.32 | 72.7 ± 1.22 | 74.6 ± 1.28 | 72.7 ± 1.65 | 74.5 ± 1.58 | 0.310 |
| MSL (mm) | 82.6 ± 1.41 a,b | 81.2 ± 1.07 b | 86.5 ± 1.45 a | 78.2 ± 2.48 b | 88.2 ± 2.17 a | 0.002 |
| MFD (µm) | 19.7 ± 0.22 a,b | 20.2 ± 0.17 a | 18.7 ± 0.23 c | 20.8 ± 0.39 a | 19.1 ± 0.34 b,c | <0.001 |
| FDSD (µm) | 4.41 ± 0.08 a | 4.46 ± 0.06 a | 4.14 ± 0.08 b | 4.67 ± 0.13 a | 4.15 ± 0.12 b | 0.001 |
| CVFD (%) | 22.3 ± 0.26 | 22.0 ± 0.20 | 22.0 ± 0.27 | 22.4 ± 0.47 | 21.7 ± 0.41 | 0.624 |
| MSS (N/ktex) | 22.1 ± 0.89 | 22.7 ± 0.68 | 21.4 ± 0.92 | 25.6 ± 1.58 | 22.7 ± 1.38 | 0.202 |
| MFC (o/mm) | 89.1 ± 1.76 | 91.8 ± 1.31 | 87.0 ± 1.69 | 92.1 ± 3.01 | 88.8 ± 2.61 | 0.260 |
| PF (%) | 3.21 ± 0.40 a,b | 3.50 ± 0.32 a | 1.36 ± 0.15 b | 5.04 ± 1.08 a | 1.39 ± 0.24 b | <0.001 |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Li, S.; Zhou, H.; Gong, H.; Zhao, F.; Hu, J.; Luo, Y.; Hickford, J.G.H. Identification of the Ovine Keratin-Associated Protein 26-1 Gene and Its Association with Variation in Wool Traits. Genes 2017, 8, 225. https://doi.org/10.3390/genes8090225
Li S, Zhou H, Gong H, Zhao F, Hu J, Luo Y, Hickford JGH. Identification of the Ovine Keratin-Associated Protein 26-1 Gene and Its Association with Variation in Wool Traits. Genes. 2017; 8(9):225. https://doi.org/10.3390/genes8090225
Chicago/Turabian StyleLi, Shaobin, Huitong Zhou, Hua Gong, Fangfang Zhao, Jiang Hu, Yuzhu Luo, and Jon G. H. Hickford. 2017. "Identification of the Ovine Keratin-Associated Protein 26-1 Gene and Its Association with Variation in Wool Traits" Genes 8, no. 9: 225. https://doi.org/10.3390/genes8090225
APA StyleLi, S., Zhou, H., Gong, H., Zhao, F., Hu, J., Luo, Y., & Hickford, J. G. H. (2017). Identification of the Ovine Keratin-Associated Protein 26-1 Gene and Its Association with Variation in Wool Traits. Genes, 8(9), 225. https://doi.org/10.3390/genes8090225






